Abstract
Prototheca zopfii (P. zopfii, class Trebouxiophyceae, order Chlorellales, family Chlorellaceae), a non-photosynthetic predominantly free-living unicellular alga, is one of the few pathogens belonging to the plant kingdom. This alga can affect many vertebrate hosts, sustaining systemic infections and diseases such as mastitis in cows. The aim of our work was to sequence and assemble the P. zopfii genotype 1 and genotype 2 mitochondrial and plastid genomes. Remarkably, the P. zopfii mitochondrial (38 Kb) and plastid (28 Kb) genomes are models of compaction and the smallest known in the Trebouxiophyceae. As expected, the P. zopfii genotype 1 and 2 plastid genomes lack all the genes involved in photosynthesis, but, surprisingly, they also lack those coding for RNA polymerases. Our results showed that plastid genes are actively transcribed in P. zopfii, which suggests that the missing RNA polymerases are substituted by nuclear-encoded paralogs. The simplified architecture and highly-reduced gene complement of the P. zopfii mitochondrial and plastid genomes are closer to those of P. stagnora and the achlorophyllous obligate parasite Helicosporidium than to those of P. wickerhamii or P. cutis. This similarity is also supported by maximum likelihood phylogenetic analyses inferences. Overall, the P. zopfii sequences reported here, which include nuclear genome drafts for both genotypes, will help provide both a deeper understanding of the evolution of Prototheca spp. and insights into the corresponding host/pathogen interactions.
Similar content being viewed by others
Introduction
Organisms belonging to the genus Prototheca are achlorophyllous algae widespread in the environment. The genus is classified within the class of Trebouxiophyceae, order Chlorellales and family Chlorellaceae, and historically encompasses six species: P. stagnora, P. ulmea, P. wickerhamii, P. blaschkeae, P. zopfii and P. cutis1,2,3. A seventh species, P. miyajii, has very recently been isolated in a patient with systemic protothecosis and classified as a separate species due to some genetic and phenotypical differences from P. wickerhamii4. Finally, an eighth species, P. moriformis, is not currently considered a species per se because of its biochemical/genetic resemblance to P. zopfii and because of its high intraspecific heterogeneity2,5.
All Prototheca species have forfeited their photosynthetic capabilities, and, consequently, their ability to harvest energy from light and fix carbon, having undergone an evolutionary transition from autotrophy to heterotrophy3, favoured also by the ability of some species to sustain infectious diseases in both humans and animals6,7. P. wickerhamii, P. cutis and P. blaschkeae, in particular, have been associated with human diseases, especially in the presence of impaired immunological-cellular systems1,7,8. Nevertheless, P. wickerhamii, P. blaschkeae and P. zopfii can also infect animals, especially dogs and cows6,9,10. Among them, P. blaschkeae and P. zopfii, are the most important species in the veterinary field because of their ability to sustain bovine mastitis11,12,13. P. zopfii can be further divided into two genotypes, namely genotype 1 and 2, both reported as pathogenic for humans14, whereas genotype 2 is the most isolated Prototheca in bovine mastitis outbreaks worldwide11,12,15,16,17,18.
The sequences of Prototheca species currently available in public databases are those of the 18S rDNA (small subunit of rDNA, SSU) and 28S rDNA (large subunit of rDNA, LSU)2,19, and those of the Internal Transcribed Spacer regions (ITS), as well as some mitochondrial and plastid genomes. Notably, this information is not available for all species and, full-length sequences are very often missing12,20,21: complete sequences of the organellar DNA are currently only available for P. wickerhamii (both mitochondrion and plastid)22,23, P. cutis and P. stagnora (plastid only)24.
In this paper, we present the complete and manually annotated genomes of mitochondria and plastids of P. zopfii genotypes 1 and 2, and the first draft assembly of the whole nuclear genomes of both P. zopfii subspecies. Our work gives, for the first time, a representative overview of the extreme reduction which occurs within the mitochondrial and plastid genomes of these algae and provides basic information for further investigations.
Materials and Methods
Strains and culture conditions
P. zopfii genotype 1 (SAG 2063) and P. zopfii genotype 2 (SAG 2021) were obtained from the Culture Collection of Algae at Göttingen University (“Sammlung von Algenkulturen der Universität Göttingen”, SAG, Göttingen, Germany). The strains were aerobically sub-cultured on Sabouraud Dextrose Agar plates for 48–72 h at 30 °C until DNA isolation was carried out.
Isolation of DNA and RNA
Genomic DNA and RNA were extracted starting from approximately one g of P. zopfii genotype 1 and genotype 2 resuspended in 20 ml wash solution (0.6 M Sucrose, 20 mM Tris, 20 mM MgCl2 and 1 mM DTT, pH 7.5) and centrifuged at 2200 g for 2 min at 4 °C.
For DNA, washed pellets were resuspended in 500 μl of TRIS EDTA lysis buffer (10 mM Tris-HCl, 10 mM EDTA, 250 mM NaCl, pH 8), supplemented with 25 µl of proteinase K (20 mg/ml) (Sigma-Aldrich, St. Louis, MO, USA), 25 µl of SDS 10% and incubated at 56 °C for 2 h. Next, 25 µl of RNaseA (20 mg/ml) (Sigma-Aldrich) were added and the suspensions were incubated at 56 °C for 30 min. DNA was extracted using an equal volume of 1:1 (v/v) phenol:chloroform and precipitated with one volume of cold isopropanol. DNA was rinsed with 70% (v/v) cold ethanol, air dried, resuspended in 30 µl of ultrapure water and stored at −20 °C until use. DNA concentration and quality were estimated by PicoGreen (Thermo Fisher, Waltham, MA, USA) and by agarose gel electrophoresis.
Total RNA was extracted from washed pellets with TRIzol (Invitrogen, Carlsbad, CA, USA) and purified by NucleoSpin miRNA kit (Macherey-Nagel, Duren, Germany), following the manufacturer’s protocol, in combination with TRIzol lysis. RNA concentration (ng/µl) and quality RNA Integrity Number (RIN) were determined by Agilent 2100 Bioanalyzer (Santa Clara, CA, USA). RNA extracts were stored at −80 °C until use.
DNA and RNA library preparation and sequencing
P. zopfii DNA libraries for Illumina sequencing (paired-end and mate-pair sequencing) (Illumina, San Diego, CA, USA) were prepared and sequenced on an Illumina MiSeq instrument, following the manufacturer’s instructions as detailed in Supplementary Table 1. DNA library preparation for GS-FLX sequencing was performed using the GS FLX Titanium Rapid Library Preparation Kit (Roche, Basel, Switzerland) as follows: 1 μg DNA for each P. zopfii strain was used in the preparation of shotgun libraries by genomic DNA fragmentation by nebulization and ligation to specific adapters. According to the manufacturer’s instructions, libraries were subjected to clonal amplification by emulsion PCR reaction, recovered by isopropanol breaking and enriched for positive reaction beads. Each library was separately loaded onto one region of the GS-FLX PicoTiter Plate and sequenced according to the 454 GS-FLX Titanium XL protocol.
One μg of RNA was used for libraries construction using TruSeq® RNA Sample Preparation v2 Kit (Illumina), according to the manufacturer’s instructions using poly(A) enrichment and sequenced on a 2 × 101-cycles Hiseq 2000 run (Illumina).
De novo genome assembly
The whole genomic sequence collections - composed of Illumina paired-end and mate-pair sequences, plus GS-FLX reads - from P. zopfii genotype 1 and 2 were assembled with SPAdes25 in read error correction and assembling mode. Six K values, ranging from K21 to K127, were automatically selected by the algorithm based on the read length and dataset type. CAP326 (-p 96 -o 500) was run on the resulting scaffolds to further assemble contiguous regions. Illumina reads were mapped back to scaffolds by means of bwa mem (v. 0.7.1027), keeping only those having ≤2 mismatches with the reference and Pilon (v 1.22)28 in assembly improvement mode was employed in order to correct substitutions and short indels.
Organelles assembly
Among the assembled scaffolds, sequences of mitochondrion and plastid were detected by homology with a set of 322 mitochondrial and 605 plastid/chloroplast sequences from related species (i.e.: green algae) and circularized, giving rise to full circular sequences for both organelles in both P. zopfii genotypes. Assembly of the mitochondrial and plastid genomes of P. zopfii was independently confirmed by a custom assembly strategy that, starting from “seeds”, implements an iterative procedure aimed at finding reads partially overlapping with the seed and assembling them with the original contig. Then, overlaps among the new contigs were used to generate a “supercontig”. Further alignment-assembly-extension steps were performed on each side of the supercontig until an overlap between the 5′ and 3′ ends of the sequence was found, meaning that the whole circular genome had been covered. A full description of this custom assembly procedure is available in Supplementary Methods.
Nuclear genome annotation
Gene prediction in nuclear genomes was performed with Augustus29 using Chlorella variabilis as reference species. A double annotation procedure was carried out on the predicted protein models, employing both BLAST30 (-evalue 1e−10) versus the UniProtKB database (http://www.uniprot.org/uniprot/), and InterProScan 531 to provide functional analysis. BLAST comparison (-evalue 1e−10) was also performed between the annotated protein datasets of the two P. zopfii genotypes. Annotation of proteins identified as putative DNA-directed RNA polymerases (NEPs) was double-checked by BLASTp against the NCBI’s own non-redundant (“nr”) database. Presence of a chloroplast transit peptide (cTP) in genes annotated as NEPs was assessed by using several prediction tools available on the web: PProwler32, PredAlgo33, TargetP (v1.1)34, and predSL35.
Organelles annotation
Organelles were annotated using the DOGMA webserver36. Annotation was manually refined, relying on similarities obtained by BLAST versus the organellar genes of the closest known relatives of P. zopfii (i.e.: C. variabilis, Auxenochlorella protothecoides, Helicosporidium sp. and P. wickerhamii). Transfer RNA (tRNA) and ribosomal RNA (rRNA) sequences were determined by BLASTn alignment, whereas coding DNA sequences (CDS) positions were refined on the basis of protein-protein (BLASTp) matches. Circular representations of the mitochondria and plastids were drawn using Gview37. Comparison of CDS among species was performed by BLASTp.
Phylogenetic analysis
The protein sequences inferred from nine conserved plastid ribosomal genes (RPL2, RPL5, RPL14, RPL16, RPL20, RPS8, RPS11, RPS12, RPS14) were retrieved from P. zopfii genotype 1 and 2, from related species (i.e.: C. variabilis, A. protothecoides, Helicosporidium sp., P. wickerhamii, P. cutis and P. stagnora), and from other Trebouxiophyceae species available at NCBI, for a total of 40 species. Only ribosomal proteins whose sequences were available for each considered species were taken into account. Sequences were concatenated and aligned to produce a super-alignment with Clustal-Omega38, which was manually inspected and used to infer a Maximum Likelihood phylogenetic tree with the program PhyML39 using four substitution rate categories, the cpREV substitution model, estimated gamma shape parameter, 1000 bootstraps, and core Trebouxiophyceae as outgroup.
RNA-Seq data analysis
RNA-Seq reads from P. zopfii 1 and 2 were mapped to the respective draft assemblies with STAR aligner (v 2.5.3a)40. Alignments were filtered retaining, for each genotype, only reads aligning for ≥80% of their length and having ≤2 mismatches. The number of reads mapping within the predicted gene models was assessed with BEDtools (v 2.26.0)41, requiring a read to map for at least 90% of its length within a gene to be counted. Counts were then converted to reads-per-kilobase-per-million (RPKM) values.
Results
The genomes of P. zopfii genotype 1 and 2 were sequenced using a combination of different approaches (Supplementary Table 1) resulting in 45,166,626 and 66,488,185 total reads, respectively.
Sequence assembly
Sequence assembly yielded a nuclear genome of 26,448,891 and 24,744,895 bp for P. zopfii genotype 1 and genotype 2, respectively; organelles were very small: mitochondria being 38,164 and 39,222 bp for genotypes 1 and 2, respectively, whereas plastids being 28,698 and 28,686 bp, for genotype 1 and 2 respectively (Table 1).
Mitochondrial structure and annotation
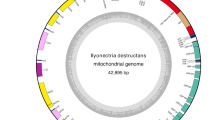
The mitochondrial genomes of both P. zopfii genotype 1 and 2 are sized at about 38–39 Kb (Fig. 1A, Supplementary Fig. 1A). They are extremely compact, with only about 32% of non-coding DNA and are characterized by a substantial loss of any intron-exon structure in their genes. Only P. zopfii genotype 2 shows a single intron (length: 777 bp) in the long ribosomal subunit (lrn) gene, whereas the other species belonging to the Chlorellales (i.e.: C. variabilis, A. protothecoides, Helicosporidium sp., and P. wickerhamii) display a more complex structure, with intron length reaching 4,000–8,000 bp. A putative LAGLIDADG homing endonuclease, a class of restriction enzymes directly involved in the DNA cutting process42, is encoded within the intron of lrn gene.
P. zopfii genotype 2 mitochondrion and plastid circular plot. Circular plots depicting the annotation of P. zopfii genotype 2 mitochondrion (A) and plastid (B). Gene annotation is reported on the outermost circle of the plot; CDS are in blue, tRNA are in green and rRNA are in red. Innermost circles represent gene orientation, GC content and skew. Other rings report the extent and the % identity of the plastid features with those of proximal organisms (C. variabilis, A. protothecoides, Helicosporidium sp., P. wickerhamii plus P. cutis and P. stagnora for plastid only) and with P. zopfii genotype 1. Transparency is proportional to the degree of identity between P. zopfii genotype 2 and each reference genome; no transparency indicates 100% identity. % identity was calculated on the basis of BLASTn (for tRNA and rRNA) and BLASTp (for CDS) matches with the corresponding features on the reference plastid genome.
The annotation revealed a number of coding genes for both P. zopfii species similar to those observed in other Chlorellales species. All the genes encoding for cytochrome units, NADH dehydrogenase, ATP synthases and ribosomal proteins, as well as all the tRNAs and the ribosomal units, have a conserved structure (Table 1, Fig. 2A). As expected, mitochondrial protein similarity between P. zopfii genotype 1 and 2 is above 90%, while similarity with Helicosporidium sp., C. variabilis, A. protothecoides and P. wickerhamii is around 60%.
Plastid structure and annotation
The structure of the plastid genomes of P. zopfii genotype 1 and 2 (Fig. 1B, Supplementary Fig. 1B) is very similar to that of other non-photosynthetic algae belonging to the Trebouxiophyceae class, being extremely compact. The genomes are sized only about 28.7 Kb for both genotypes and are the shortest plastid genomes within their class. Both P. zopfii genotypes possess only 19 CDS, and as a result are simpler than those of Helicosporidium sp. (26 CDS), P. wickerhamii, P. cutis (both possessing 40 CDS) and P. stagnora (28 CDS). On the other hand, the tRNAs and rRNAs are conserved among all these species and other photosynthetic algae (i.e.: C. variabilis and A. protothecoides) (Table 1, Fig. 2B). Plastid protein similarity between genotype 1 and 2 is about 85% while similarity with other organisms is around 41–43%, with the best match being with P. stagnora (i.e.: 49.9% and 49.5% similarity to P. zopfii 1 and 2, respectively) (Fig. 3A). Comparison with other related organisms shows that the plastid genomes of P. zopfii genotype 1 and 2, like those of Helicosporidium sp., P. wickerhamii, P. cutis and P. stagnora, lack the genes associated with photosynthesis (photosystem I and II, chlorophyll biosynthesis and cytochrome components), whereas ATP synthase genes were maintained only in P. wickerhamii and P. cutis. Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (rbcL, or RuBisCO) is absent in P. zopfii, as well as in all other Prototheca spp. and in Helicosporidium sp. Moreover, differently to the other Trebouxiophyceae, P. zopfii also lacks all plastid-encoded RNA polymerases (i.e.: rpoA, rpoB, rpoC1 and rpoC2) (Table 2, Supplementary Table 2). Our transcriptome analysis demonstrated that most plastid genes encoding rRNA and proteins were expressed in P. zopfii (Supplementary Fig. 2A), although three of them (i.e.: rps4, rps7, rpl20) had low RPKM values, suggesting that nuclear-encoded counterparts could compensate for the loss of plastid-encoded RNA polymerases.
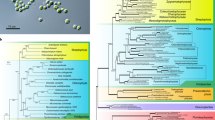
Phylogenetic analysis of P. zopfii. (A) Maximum Likelihood (ML) tree inferred from the super-alignment of 9 plastid ribosomal proteins (i.e.: RPL2, RPL5, RPL14, RPL16, RPL20, RPS8, RPS11, RPS12, and RPS14). Bootstrap values are indicated above the lines. Core Trebouxiophyceae was used as outgroup. Among Chlorellales, (dark green vertical bar) non-photosynthetic species are highlighted in blue; tree lengths for this group are not drawn to scale. (B) Heatmap representing the pairwise average percentage of identity between CDS of 8 organisms belonging to Chlorellales (P. zopfii and its closest relatives).
Both P. zopfii genotypes plastids contain the genes of most of the ribosomal proteins, with the exception of RPL12, RPL19, RPL23, RPL32, RPS2, RPS18 and RPS9. The same proteins, with the exception of RPL19 and RPL32, are also absent in P. stagnora, which, apparently, is the most similar organism. The phylogenetic tree inferred from the super-alignment of 9 shared ribosomal proteins confirms that Helicosporidium sp., P. stagnora and P. zopfii are closely related genera, whereas P. wickerhamii is more distant, closer to A. protothecoides than to other Prototheca species (Fig. 3B).
Nuclear genome annotation
From the genome assembly procedure, 6,956 and 4,555 scaffolds with lengths >1 Kb representing the nuclear genome were obtained for P. zopfii genotype 1 and 2, respectively, indicating a genome size of about 26.5 Mbp and 24.7 Mbp for genotypes 1 and 2, respectively, a size between those of Helicosporidium sp. (12.4 Mb) and C. variabilis (46.2 Mb) and comparable to the genome of A. protothecoides (~23 Mb) and P. wickerhamii (~29 Mb) (Table 1).
The maximum scaffold length was 97,625 and 57,068 bp, with 1,289 and 1,708 contigs exceeding 5 Kb in length and a N50 value of 6,686 and 7,940 for P. zopfii genotypes 1 and 2, respectively.
Augustus gene prediction led to the individuation of 6,884 and 6,381 gene models for the two genotypes. 56.5% and 62.0% of P. zopfii genotype 1 gene models were annotated versus UniProtKB and InterPro databases, respectively, while the corresponding percentages for genotype 2 were 59.4% and 67.2%. The main features of the nuclear genome, such as gene number and coding density, were consistent for both genotypes, and similar to their counterparts in A. protothecoides genome (i.e.: gene density: 0.26 genes/Kb and 0.32 genes/Kb, for both P. zopfii genotypes and A. protothecoides, respectively). On the contrary, the average exon and intron sizes were higher compared to those previously observed in related species (Supplementary Table 3). BLAST comparison of the predicted proteins of the two P. zopfii genotypes resulted in a set of 6,134 common entities, representing a core set of homologous proteins conserved in the two genotypes. Among the predicted genes and transcripts, we found evidence of the presence and expression of nuclear-encoded polymerases (NEPs) (Supplementary Data 1 and 2). P. zopfii genotype 1 and 2 possess 21 and 19 genes annotated as NEPs, respectively, and both showed a RNA-Seq signal indicating active transcription of many of them (Supplementary Fig. 2B,C). Prediction of target peptides highlighted at least one NEP per genotype as a high-confidence candidate for containing a chloroplast transit peptide (cTP) (genes g5108 and g2780 for genotypes 1 and 2, respectively, for which all the four prediction software employed were concordant); moreover, PredAlgo also suggested two more genes (g3914 and g4216 for genotypes 1 and 2, respectively) to be plastid-directed NEPs. In addition to that, we found evidence of some mitochondrial targeting peptides (mTPs) in more gene models (Supplementary Table 4).
Discussion
In this paper, we describe the complete, manually annotated, circular sequence of both mitochondrial and plastid organellar DNA of P. zopfii genotype 1 and genotype 2, as well as a first draft of the complete genome, by whole genome shotgun sequencing.
Structure of the mitochondrial genomes of both P. zopfii genotypes was revealed to be smaller in size and extremely condensed when compared to that of some related organisms, i.e.: P. wickerhamii22 and Helicosporidium sp.43, but similarly functional, with the size reduction mostly due to the lack of intron-exon structures.
An extremely compact and simplified structure was also observed in P. zopfii plastid genomes, which showed a substantially reduced size (about 28.7 Kb), smaller than those of all other algae belonging to the class of Trebouxiophyceae. As previously observed in the genomes of non-photosynthetic algae belonging to this class, P. zopfii plastids lack all the genes for the synthesis of the proteins involved in the photosynthesis process23,44, and for the RuBisCO large subunit. Fundamental plastid-related functions, however, seem to have been preserved, as indicated by the presence of a RNA-Seq signal on the genes of plastid-encoded ribosomal proteins. The low expression of some of them, however, cannot preclude their presence in the genome as pseudogenes in P. zopfii. However, further experiments should be carried on in order to confirm this observation. More interestingly, the entire set of rpo genes (i.e. rpoA, rpoB, rpoC1 and rpoC2), which codify for the plastid-encoded RNA polymerases (PEPs), was lost in P. zopfii, an unprecedented observation within this class of algae. Loss of PEPs was previously reported for other non-photosynthetic parasitic plants, such as Cuscuta obtusiflora45 and Rhizanthella gardneri46, but not in apicomplexan and algae: plastid genomes of Plasmodium falciparum47, A. protothecoides, P. wickerhamii23, P. cutis, P. stagnora24 and Helicosporidium sp.44 have all retained the complete set of rpo genes. We found no evidence of PEP sequences either in plastid assemblies or in the nuclear draft genomes, whereas nuclear genome contigs contained evidence of 21 and 19 DNA-driven, nuclear-encoded polymerases (NEPs) for P. zopfii genotype 1 and 2, respectively, and at least one of them, per genotype, was predicted to contain plastid-targeting signal peptides. It is therefore possible that P. zopfii codes for other NEPs able to target the plastid, making it possible to transcribe its genetic information.
As previously suggested23,24,48,49, considering the degree of similarity between the few structural genes preserved in its plastid genome and the evidence from the phylogenetic analysis, P. zopfii seems to be more closely related to P. stagnora and to Helicosporidium sp., rather than to P. wickerhamii or P. cutis. Moreover, it is noteworthy that P. wickerhamii appears not to be closely related to P. zopfii, but instead to A. protothecoides, strengthening the evidence that P. wickerhamii is only loosely related to other Prototheca spp, as previously revealed by plastid genome comparison23 and supporting the proposal of either moving P. wickerhamii into Auxenochlorella genus or creating a new genus48.
Nuclear genome assemblies of P. zopfii genotype 1 and 2 had a size estimated at about 25–26 Mb for both, consistent with that reported in a previous work50. Although further studies are certainly needed to elucidate the structure of nuclear genomes of P. zopfii, this work adds information to the growing body of genome resources for the plant kingdom, being, although a preliminary draft, the first report of the assembly of nuclear DNA of P. zopfii.
In conclusion, we believe that the information reported herein will be important for the understanding of the evolution and genomic organization of Prototheca spp., with a particular focus on the progressive loss of functions of plastids in the shift from autotrophic, photosynthetic, to obligate, heterotrophic, parasitic algae.
Data Availability
The complete genome sequencing project has been registered in the NCBI BioProject portal (https://www.ncbi.nlm.nih.gov/bioproject/) under the accession number PRJNA388740. Raw DNA sequence reads for P. zopfii genotypes 1 and 2 have been deposited into the NCBI Short-Read Archive (SRA, https://www.ncbi.nlm.nih.gov/sra/) under the accession numbers SRR6319956-SRR6319964. RNA-seq reads are saved under the accession numbers SRR7091517-SRR7091518. Full sequences of mitochondria and plastids are available in GenBank, under accessions MF197533, MF197534, MF197535, and MF197536. The Whole Genome Shotgun project has been deposited (as non-annotated contigs) at DDBJ/ENA/GenBank under the accessions PEIA00000000 and PGFX00000000. The versions described in this paper are PEIA01000000 and PGFX01000000. Sequences annotated as nuclear-encoded polymerases (NEPs) are available as amino acid FASTA files in Supplementary Data 1 and 2.
Change history
11 April 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41598-022-09754-0
References
Satoh, K., Ooe, K., Nagayama, H. & Makimura, K. Prototheca cutis sp. nov., a newly discovered pathogen of protothecosis isolated from inflamed human skin. Int J Syst Evol Microbiol.60, 1236–40, https://doi.org/10.1099/ijs.0.016402-0 (2010).
Roesler, U., Möller, A., Hensel, A., Baumann, D. & Truyen, U. Diversity within the current algal species Prototheca zopfii: a proposal for two Prototheca zopfii genotypes and description of a novel species, Prototheca blaschkeae sp. nov. Int J Syst Evol Microbiol.56, 1419–25 (2006).
Pore, R. S., Barnett, E. A., Barnes, W. C. Jr. & Walker, J. D. Prototheca ecology. Mycopathologia.81, 49–62 (1983).
Masuda, M., Hirose, N., Ishikawa, T., Ikawa, Y. & Nishimura, K. Prototheca miyajii sp. nov., isolated from a patient with systemic protothecosis. Int J Syst Evol Microbiol.66, 1510–1520, https://doi.org/10.1099/ijsem.0.000911 (2016).
Pore, R. S. Selective medium for the isolation of Prototheca. Appl Microbiol.26, 648–9 (1973).
Jánosi, S. et al. Review of the microbiological, pathological, and clinical aspects of bovine mastitis caused by the alga Prototheca zopfii. Vet Q.23, 58–61 (2001).
Lass-Flörl, C. & Mayr, A. Human protothecosis. Clin Microbiol Rev.20, 230–42 (2007).
Nelson, A. M., Neafie, R. C. & Connor, D. H. Cutaneous protothecosis and chlorellosis, extraordinary “aquatic-borne” algal infections. Clin Dermatol.5, 76–87 (1987).
Shank, A. M., Dubielzig, R. D. & Teixeira, L. B. Canine ocular protothecosis: A review of 14 cases. Vet Ophthalmol.18, 437–42, https://doi.org/10.1111/vop.12239 (2015).
Stenner, V. J. et al. Protothecosis in 17 Australian dogs and a review of the canine literature. Med Mycol.45, 249–66 (2007).
Ricchi, M. et al. Molecular characterization of Prototheca strains isolated from Italian dairy herds. J Dairy Sci.93, 4625–31, https://doi.org/10.3168/jds.2010-3178 (2010).
Capra, E. et al. Simultaneous identification by multiplex PCR of major Prototheca spp. isolated from bovine and buffalo intramammary infection and bulk tank. Lett Appl Microbiol.59, 642–7, https://doi.org/10.1111/lam.12326. (2014).
Ricchi, M. et al. First outbreak of bovine mastitis caused by Prototheca blaschkeae. Vet Microbiol.162, 997–9, https://doi.org/10.1016/j.vetmic.2012.11.003 (2013).
Hirose, N. et al. Molecular Characterization of Prototheca strains isolated in China revealed the first cases of protothecosis associated with Prototheca zopfii genotype 1. Med Mycol. https://doi.org/10.1093/mmy/myx039 (2017).
Jagielski, T., Lassa, H., Ahrholdt, J., Malinowski, E. & Roesler, U. Genotyping of bovine Prototheca mastitis isolates from Poland. Vet Microbiol.149, 283–7, https://doi.org/10.1016/j.vetmic.2010.09.034 (2011).
Möller, A., Truyen, U. & Roesler, U. Prototheca zopfii genotype 2: the causative agent of bovine protothecal mastitis? Vet Microbiol.120, 370–4 (2007).
Sobukawa, H. et al. Short communication: Molecular typing of Prototheca zopfii from bovine mastitis in Japan. J Dairy Sci.95, 4442–6, https://doi.org/10.3168/jds.2011-5101 (2012).
Gao, J. et al. Characterization of Prototheca zopfii associated with outbreak of bovine clinical mastitis in herd of Beijing, China. Mycopathologia.173, 275–81, https://doi.org/10.1007/s11046-011-9510-y (2012).
Kishimoto, Y. et al. 26S rDNA-based phylogenetic investigation of Japanese cattle-associated Prototheca zopfii isolates. J Vet Med Sci.72, 123–6 (2010).
Hirose, N., Nishimura, K., Inoue-Sakamoto, M. & Masuda, M. Ribosomal internal transcribed spacer of Prototheca wickerhamii has characteristic structure useful for identification and genotyping. PLoS One. 8, e81223, https://doi.org/10.1371/journal.pone.0081223. eCollection2013 (2013).
Marques, S., Huss, V. A., Pfisterer, K., Grosse, C. & Thompson, G. Internal transcribed spacer sequence-based rapid molecular identification of Prototheca zopfii and Prototheca blaschkeae directly from milk of infected cows. J Dairy Sci.98, 3001–9, https://doi.org/10.3168/jds.2014-9271 (2015).
Wolff, G., Plante, I., Lang, B. F., Kück, U. & Burger, G. Complete sequence of the mitochondrial DNA of the chlorophyte alga Prototheca wickerhamii. Gene content and genome organization. J Mol Biol.237, 75–86 (1994).
Yan, D. et al. Auxenochlorella protothecoides and Prototheca wickerhamii plastid genome sequences give insight into the origins of non-photosynthetic algae. Sci Rep.5, 14465, https://doi.org/10.1038/srep14465 (2015).
Suzuki, S., Endoh, R., Manabe, R. I., Ohkuma, M. & Hirakawa, Y. Multiple losses of photosynthesis and convergent reductive genome evolution in the colourless green algae Prototheca. Sci Rep.8(1), 940 (2018).
Bankevich, A. et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. Journal of Computational Biology.19, 455–477, https://doi.org/10.1089/cmb.2012.0021 (2012).
Huang, X. & Madan, A. CAP3: A DNA sequence assembly program. Genome Res.9, 868–77 (1999).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv1303, 3997v1 (2013).
Walker, B. J. et al. Pilon: An Integrated Tool for Comprehensive Microbial Variant Detection and Genome Assembly Improvement. PLoS ONE.9(11), e112963, https://doi.org/10.1371/journal.pone.0112963 (2014).
Stanke, M. & Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Research.33, W465–W467, https://doi.org/10.1093/nar/gki458 (2005).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J Mol Biol.215, 403–10 (1990).
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Research.33, W116–W120, https://doi.org/10.1093/nar/gki442 (2005).
Bodén, M. & Hawkins, J. Prediction of subcellular localization using sequence-biased recurrent networks. Bioinformatics21, 2279–2286 (2005).
Tardif, M. et al. PredAlgo: a new subcellular localization prediction tool dedicated to green algae. Mol Biol Evol.29(12), 3625–39, https://doi.org/10.1093/molbev/mss178. (2012).
Emanuelsson, O., Nielsen, H., Brunak, S. & von Heijne, G. Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol300, 1005–1016 (2000).
Petsalaki, E. I., Bagos, P. G., Litou, Z. I. & Hamodrakas, S. J. PredSL: a tool for the N-terminal sequence-based prediction of protein subcellular localization. Genomics Proteomics Bioinformatics4, 48–55 (2006).
Wyman, S. K., Jansen, R. K. & Boore, J. L. Automatic annotation of organellar genomes with DOGMA. Bioinformatics.20, 3252–5 (2004).
Petkau, A., Stuart-Edwards, M., Stothard, P. & Van Domselaar, G. Interactive microbial genome visualization with GView. Bioinformatics.26, 3125–6, https://doi.org/10.1093/bioinformatics/btq588 (2010).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol.7, 539, https://doi.org/10.1038/msb.2011.75 (2011).
Guindon, S. et al. New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Systematic Biology59(3), 307–21 (2010).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics29(1), 15–21, https://doi.org/10.1093/bioinformatics/bts635 (2013).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics26(6), 841–842, https://doi.org/10.1093/bioinformatics/btq033 (2010).
Hafez, M. & Hausner, G. Homing endonucleases: DNA scissors on a mission. Genome.55, 553–69, https://doi.org/10.1139/g2012-049 (2012).
Pombert, J. F. & Keeling, P. J. The mitochondrial genome of the entomoparasitic green alga Helicosporidium. PLoS One.5(1), e8954 (2010).
deKoning, A. P. & Keeling, P. J. The complete plastid genome sequence of the parasitic green alga Helicosporidium sp. is highly reduced and structured. BMC Biol.21, 4:12 (2006).
McNeal, J. R., Kuehl, J. V., Boore, J. L. & de Pamphilis, C. W. Complete plastid genome sequences suggest strong selection for retention of photosynthetic genes in the parasitic plant genus Cuscuta. BMC Plant Biol.7, 57 (2007).
Delannoy, E., Fujii, S., Colas des Francs-Small, C., Brundrett, M. & Small, I. Rampant gene loss in the underground orchid Rhizanthella gardneri highlights evolutionary constraints on plastid genomes. Mol Biol Evol.28, 2077–86, https://doi.org/10.1093/molbev/msr028 (2011).
Wilson, R. et al. Complete gene map of the plastid-like DNA of the malaria parasite Plasmodium falciparum. J Mol Biol.261, 155–172 (1996).
Ueno, R., Urano, N. & Suzuki, M. Phylogeny of the non-photosynthetic green micro-algal genus Prototheca (Trebouxiophyceae, Chlorophyta) and related taxa inferred from SSU and LSU ribosomal DNA partial sequence data. FEMS Microbiol Lett.223, 275–80 (2003).
Tartar, A., Boucias, D. G., Becnel, J. J. & Adams, B. J. Comparison of plastid 16S rRNA (rrn16) genes from Helicosporidium spp.: evidence supporting the reclassification of Helicosporidia as green algae (Chlorophyta). Int J Syst Evol Microbiol.53, 1719–23 (2003).
Suzuki, T. Electrophoretic separation of chromosomes in an achlorophyllous microalga, Prototheca zopfii. J Tokyo Univ Nat Sci.50, 13–6 (2006).
Jagielski, T. et al. An optimized method for high quality DNA extraction from microalga Prototheca wickerhamii for genome sequencing. Plant Methods.13, 77, https://doi.org/10.1186/s13007-017-0228-9 (2017).
Green, B. R. Chloroplast genomes of photosynthetic eukaryotes. Plant J.66(1), 34–44, https://doi.org/10.1111/j.1365-313X.2011.04541.x (2011).
Acknowledgements
We thank Dr. Tomasz Jagielski from the University of Warsaw (Poland) for providing P. zopfii genotype 2 strain. We also thank Mr. Bryan Manchi for his valuable revision of the English text.
Author information
Authors and Affiliations
Contributions
M.S. and B.L. assembled and annotated the genome, conducted analyses and wrote the paper. E.C. prepared P. zopfii genome samples, performed genomic analyses and wrote the paper. R.B. prepared and sequenced P. zopfii samples on GS-FLX platform. M.R. helped during the conception of the study, provided P. zopfii and wrote the paper. M.L. provided P. zopfii and helped draft the manuscript. S.C. sequenced P. zopfii samples on Illumina MiSeq instrument and helped draft the manuscript. B.C. conceived the study and helped draft the manuscript. P.C. conceived the study, helped in preparing P. zopfii genome samples, sequenced P. zopfii samples on Illumina MiSeq instrument and wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained errors in the Materials and Methods section where the donation of the P. zopfii genotype 2 (SAG 2021) was incorrectly attributed to Dr. Jagielski, University of Warsaw (Poland).
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Severgnini, M., Lazzari, B., Capra, E. et al. Genome sequencing of Prototheca zopfii genotypes 1 and 2 provides evidence of a severe reduction in organellar genomes. Sci Rep 8, 14637 (2018). https://doi.org/10.1038/s41598-018-32992-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-32992-0
Keywords
This article is cited by
-
A first insight into the genome of Prototheca wickerhamii, a major causative agent of human protothecosis
BMC Genomics (2021)
-
Protothecosis in Dogs and Cats—New Research Directions
Mycopathologia (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.