Abstract
Long non-coding RNAs (LncRNAs) have emerged as a relevant class of genome regulators involved in a broad range of biological processes and with important roles in tumor initiation and malignant progression. We have previously identified a p53-regulated tumor suppressor signature of LncRNAs (PR-LncRNAs) in colorectal cancer. Our aim was to identify the expression and function of this signature in gliomas. We found that the expression of the four PR-LncRNAs tested was high in human low-grade glioma samples and diminished with increasing grade of disease, being the lowest in glioblastoma samples. Functional assays demonstrated that PR-LncRNA silencing increased glioma cell proliferation and oncosphere formation. Mechanistically, we found an inverse correlation between PR-LncRNA expression and SOX1, SOX2 and SOX9 stem cell factors in human glioma biopsies and in glioma cells in vitro. Moreover, knock-down of SOX activity abolished the effect of PR-LncRNA silencing in glioma cell activity. In conclusion, our results demonstrate that the expression and function of PR-LncRNAs are significantly altered in gliomagenesis and that their activity is mediated by SOX factors. These results may provide important insights into the mechanisms responsible for glioblastoma pathogenesis.
Similar content being viewed by others
Introduction
Gliomas are relatively rare, around 350,000 people being diagnosed per year worldwide, but they are the most common primary brain tumors and, importantly, account for over 80% of malignant primary central nervous system tumors1. They have been classified into different grades of malignancy based on histopathological and clinical criteria, glioblastomas corresponding to the highest grade2. Glioblastoma is the most common and malignant primary intracranial tumor in adults, with an incidence ranging from 1 to 5 cases per 100,000 people per year. Current therapy consists of maximal surgical tumor resection followed by concomitant radiotherapy and chemotherapy with temozolomide2. This therapy is only partially effective, however, and aggressive growth and recurrence frequently follows even after optimal treatment. In line with this, patients have an associated median survival of 12–15 months and only around 5% of patients survive to 3 years3,4. This dismal patient survival identifies glioblastoma as one of the most aggressive and fatal cancers overall. It is therefore necessary to identify strategies and targets for the early diagnosis and therapeutic treatment of gliomas.
Glioblastoma shows significant variability and heterogeneity at clinical, morphological, histopathological, molecular and cellular levels, and this heterogeneity goes a long way to explaining the poor prognosis2. Several studies have explored the genetic and molecular characteristics of this type of cancer, providing a high-resolution picture of the glioblastoma landscape, in turn, allowing the identification of different subtypes based on the molecular knowledge of the genome and transcriptome5,6,7. Nevertheless, high-throughput sequencing of whole genomes and transcriptomes found that less than 2% of the genome encodes proteins, whilst 75% is actively transcribed into noncoding RNAs8. Increasing evidence is demonstrating that frequent major genomic mutations in cancer reside inside this vast majority of regions that do not encode proteins9. These genomic loci are often transcribed into long noncoding RNAs (LncRNAs)9.
LncRNAs are transcripts of more than 200 nucleotides that miss functional open reading frames and do not have functional protein-coding ability. Mechanistically, LncRNAs can fold into larger structures to provide higher potential for target recognition, which facilitates chromatin remodeling as well as transcriptional and post-transcriptional regulation10. Increasing evidence shows that the levels of LncRNAs are altered in multiple contexts and their deregulation facilitates the modulation of gene expression during both normal biological and pathological processes. Thus, mutations and dysregulations of lncRNAs contribute to the development of several human complex diseases, including cancer11. In brain tumors, a microarray analysis identified over 1000 LncRNAs differentially expressed in glioblastoma and healthy brain tissue12, with a similar quantity of LncRNAs overexpressed and silenced12. Additional studies confirmed differentially expressed LncRNAs in gliomas using further human tissues and also cell lines13,14. Notably, these altered LncRNA expression patterns have been correlated with malignancy grade, patient survival and histological differentiation in human gliomas13,14. In line with this, the development of effective computational models predicted cancer associated-LncRNAs as biomarkers for glioma diagnosis, treatment and prognosis15,16. Moreover, abnormal LncRNA function plays critical roles in the development and progression of gliomas, controlling processes such as proliferation, apoptosis, self-renewal and migration13,14. These studies support the hypothesis that LncRNAs assume an important role in glioblastoma pathogenesis. Nevertheless, the exact functions in normal biological and disease processes have been reported for only a few LncRNAs in glioblastoma17,18.
Through genome-wide studies, we previously identified a set of LncRNAs, which are differentially expressed upon DNA damage in human colorectal cancer cells in response to active p5319. These LncRNAs, called p53-regulated LncRNAs (PR-LncRNAs), are required for the efficient binding of p53 to some of its well-known target genes and contribute to p53 pro-apoptotic and cell cycle regulatory functions19. Importantly, the expression of the PR-LncRNA signature is lower in colorectal cancer samples than healthy adjacent control tissue, suggesting that PR-LncRNAs might constitute a tumor-suppressor signature19. Since mutations on p53 are common and p53 pathway is frequently deregulated in gliomas20, in this work, we characterized the expression and function of the PR-LncRNA signature, by studying the expression and clinical relevance of four members of this signature in human glioma samples of different grades and assessing the effect of loss of function in glioma cells.
Methods and Materials
Patients and tumor samples
Human glioma patient clinical information were obtained from the Donostia University Hospital. Human glioma samples were provided by the Basque Biobank for Research-OEHUN (http://www.biobancovasco.org). The study included biopsies from 35 patients seen in San Sebastian, and diagnosed as primary glioblastoma grade IV according to the WHO criteria, 4 as a grade III and 4 as a low grade I-II. All study participants signed informed consent form approved by the Institutional Ethical Committee. The study was approved by the ethic committee of Hospital Donostia and all the experiments were performed in accordance with relevant guidelines and regulations.
RNA analysis
Total RNA was extracted with Trizol (Life Technologies). Reverse transcription was performed using random priming and Maxima First Strand cDNA Synthesis Kit (ThermoFisher), according to the manufacturer’s guidelines. Quantitative real-time polymerase chain reaction (PCR) was performed using Absolute SYBR Green mix (Thermo Scientific) in a CFX384 Real-time thermal cycler (BioRad). Variations in input RNA were compensated for by subtracting the PCR threshold cycle values obtained for GAPDH.
Cell lines and cultures
Glioma cell lines U87-MG (U87), U373MG (U373), U251MG (U251) and A172 purchased from the ATCC, were cultured in DMEM (Gibco) supplemented with 10% FBS (Gibco), 100 U/ml penicillin and 100 µg/ml streptomycin for traditional monolayer cultures. Patient-derived GNS166 and GNS179 cell lines, kindly provided by Dr. Steven Pollard21, GB1 established by our group22, and oncospheres from glioma cell lines were cultured in DMEM/F-12 (Sigma) supplemented with N2, B27 supplements (Fisher) and growth factors (20 ng/ml basic fibroblast growth factor, and 20 ng/ml epidermal growth factor; Sigma) for oncosphere cultures22. Cells were maintained under standard conditions in humidified atmosphere of 5% CO2 at 37 °C.
Transfections with antisense oligonucleotides
The antisense oligonucleotide (ASO) sequences used in this study were designed and provided by ISIS Pharmaceuticals and the methodology has been previously described19. Cells were transfected with 50 nM of each ASO, using lipofectamine 2000 (Invitrogen) and inmunofluorescence and mRNA studies were done 48 hours post-transfection.
Lentiviral infections
Lentiviral infections were performed as previously described23. For SOX knockdown, cells were infected with pLKO.1 shSOX1 and pLKO.1 shSOX9 or empty vector (obtained from Sigma). Infected cells were selected in the presence of 2 μg/ml puromycin and then maintained with 0.2 μg/ml puromycin (Sigma).
Oncosphere assays
For quantification studies, 7000 U87-MG cells/well were seeded in non-treated 6-well flat bottom plates and fresh media was added to the plates every 3 days. After 10 days, oncospheres were counted. Then, these oncospheres were disaggregated with accutase (Gibco), and same amount of cells were seeded for secondary oncospheres and maintained in culture for another 10 days.
Immunofluorescence
Immunofluorescence was performed following standard procedures24. The primary antibody was anti-phospho-histone-3 (PH3, 1:2000; Ab14955, Abcam), and the secondary antibody was anti-mouse Alexa Fluor 555 IgG (Invitrogen). Nuclear DNA was stained with Hoechst 33342 (Sigma). Pictures were taken with an Eclipse 80i microscope and processed with the NIS Elements Advances Research software (Nikon).
Discriminant analysis
The PCR expression of the four PR-LncRNAs were used to estimate the heuristic probability distribution functions of LGG and HGG conditions. The heuristic probability distribution functions of LGG and HGG conditions were predicted by fitting the corresponding log2 transformed signals using the generalized extreme value (GEV) model. The GEV model parameters: shape k, scale, σ and location µ, were estimated using the function fitdis of Matlab (MathWorksTM). To estimate the discrimination capability of each PR-LncRNA, we calculated the overlap of the LGG and HGG probability distribution functions (PDFs). For each x coordinate the overlap is the minumun of between the LGG and HGG PDFs. The PR-LncRNAs with best discrimination capabilities are those with smaller overlap between LGG and HGG PDFs. Next, we calculated the performance of all the possible combinations of selected PR-LncRNA using performance metrics of the Receiver Operating Characteristic (ROC), namely, the distance to the optimal point (0, 1) of the ROC space (D01), the accuracy, the specificity and the distance to the optimal point (0, 1) of the ROC space (D01) The D01 is calculated as D01 = (1 − TPR2 + FPR2)0.5, were TPR and FPR are the true and false positive rates, respectively. Finally, the combination of PR-LncRNAs with smallest D01 and highest accuracy and specificity was selected as optimal predictor. We used the same set of samples for our prospective study to choose the optimal predictor and to make the prediction because of limited number of samples from LGG patients. Data processing and graphics were performed with functions developed in Matlab (MathWorksTM).
Results
Expression of PR-LncRNA signature decreases with glioma grade
To characterize the expression of the PR-LncRNAs in human glioma samples, we measured the expression of four of the PR-LncRNAs (PR-LncRNAs 1, 5 and 10 and Unassigned 4) in a set of biopsies including 4 low-grade gliomas (grade II), 4 anaplasic gliomas (grade III) and 35 glioblastomas (grade IV) obtained from a cohort of patients from Donostia Hospital. First, we found that the levels of the four tested PR-lncRNAs were significantly higher in low-grade than high-grade gliomas (Fig. 1A). The analysis across the different tumor grades revealed that PR-lncRNA expression gradually decreased with the advance of glioma malignancy with the exception of PR-lncRNA-1 (Fig. 1B). In line with previous studies2, patient survival in the Donostia Hospital cohort decreased with advancing glioma grade (Fig. 1C), with a median survival of 77, 49 and 17 months in grades II, III and IV respectively.
PR-LncRNA expression in increasing glioma grade samples. (A) Comparison of PR-LncRNA 1, 5, 10 and Unassigned 4 expression in low (grade II) and high-grade (III and IV) glioma (LGG and HGG respectively). (B) Expression of PR-LncRNA 1, 5, 10 and Unassigned 4 expression in grade II, III and IV gliomas. (C) Kaplan-Meier curve representing the survival of patients with grade II (n = 4), grade III (n = 4) and grade IV (glioblastoma) tumors (n = 35). (D) List of glioma biopsies with high (1) or low(0) (above or below median) expression of PR-LncRNA 1, 5, 10 and Unassigned 4 (E) Heat map of the prediction of a sample to be HGG or LGG using all different combinations (in columns) of expression of PR-LncRNAs as predictors. The colorbar on the top shows the probability of a sample to be HGG or LGG. The negative values are associated with the probabilities of LGG and the positive with the probabilities of HGG. Each row represents the probabilities for a patient. The column table to the right shows the real status of the sample: red for HGG and green for LGG. The optimal predictor variable combination for the discrimination analysis between LGG and HGG grade glioma is chosen as the one with lowest value of the Receiver Operating Characteristic (ROC) of the performance of the predictions and highest value of Accuracy, Specificity and Sensitivity, namely PR-LncRNA 1. (F) Heat map of the predictive value of a sample to be HGG (High) or LGG (Low) using the expression of PR-LncRNA 1 as the optimal predictor. The colorbar on the top shows the probability of a sample to be HGG. The table to the right of the heatmap shows: S, real status of the sample; P, predicted status of the sample; C, comparison between the real and the predicted status. Red color is used for HGG, green for LGG, and white when there is no coincidence between the real and the predicted status using this optimal predictor, PR-LncRNA 1Asterisks (*) indicate statistical significance (p < 0.05).
Next, we studied whether the expression of PR-LncRNAs could form a signature in glioma samples and analyzed their putative association. Indeed, there were significant positive correlations between the expressions of the different PR-LncRNAs (p < 0.001) (Fig. 1D). Moreover, we developed a discriminant analysis method to study the predictive capabilities of the PR-LncRNA to predict low-grade or high grade gliomas. We found that the optimal combination of PR-LncRNAs is actually formed by a single PR-LncRNA, namely PR-LncRNA 1 (Fig. 1E). Finally, we used the optimal predictor PR-LncRNA 1 to perform the discriminant analysis on our sample set. We found that the optimal discriminator classified correctly 93% of the samples, with 100% for the case of high-grade glioma, and 25% for low grade (Fig. 1F). The low prediction rate of low-grade glioma is due to the lack of higher number of available patients (n = 4). These results show that PR-LncRNA levels do represent a signature, expression diminishing with glioma grade.
PR-LncRNA levels in glioma samples correlate with clinical characteristics
We performed correlation analysis between the expression of the PR-LncRNAs and clinical data concerning Donostia University Hospital glioma cohort. We first analyzed the clinical data for the cohort. Patients were distributed homogeneously in terms of sex, and overall survival did not vary by sex (Table 1). Most of the patients were under 65 years old and, notably, the risk of death was 3.5-fold higher in those who were ≥65 of years age (HR = 1.046, CI 1.008–1.085) (Table 1). The majority of patients were autonomous (Karnofsky score ≥70) at the time of diagnosis, but a preoperative score <70 correlated with poorer outcome (P < 0.007) (Table 1). Finally, patients who underwent complete surgical resection of the tumor (67%) survived for longer than those with subtotal extirpation (p = 0.022) (Table 1). On the other hand, we did not observe any significant correlation between patient survival and molecular markers such as MGMT, IDH1, ATRX, EGFR and p53 status (Table 2)
Next, we compared the expression of PR-LncRNAs with the aforementioned variables, finding that the means of the expression of the four PR-lncRNAs decreased from grade II to grade IV in female patients, and also in male with the exception of grade III (Fig. 2A). The trend of gradual decrease from lower to higher grade glioma is observed particularly in the expression of PR-LncRNA1 and 10 in female patients (Fig. 2B). Interestingly, for PR-LncRNA1, 5 and 10, the expression for female patients is higher than for male patients in grade III and glioblastoma (Fig. 2B, Table 3). These results indicate that PR-LncRNAs expression is sex related. On the contrary, we did not find significant differences in expression of PR-LncRNAs by age or pre-operative Karnofsky score (Table 3). Regarding tumor location, more than half of the tumors with high PR-LncRNAs were in frontal or temporal locations, the right hemisphere and a non-eloquent area (Table 3). Similar percentages had tumor volumes greater than 50 mm3 and a magnetic resonance imaging perfusion parameter higher than 6 (Table 3). When comparing the expression of PR-LncRNAs with MGMT methylation status, and Ki67, IDH1, ATRX and EGFR expression, we did not detect any significant associations (Fig. 2C). We observed that majority of samples with high levels of PR-LncRNAs had wild type p53, while over 50% of cases with low expression of PR-LncRNAs had mutated p53 genes, although this difference was not statistically significant (Fig. 2C).
Correlation between PR-LncRNA expression and patient clinical characteristics. (A) Violin plot associated to the distribution of the expression of the PR-LncRNAs in gliobastoma (IV), anaplastic (III) and low-grade glioma (II) separated by sex (M - male, F - female). The + denote the mean values of the distributions and the black dots are the expression values. (B) Heat map of the expression of the studied PR-LncRNAs separated by sex. Redder color corresponds to higher expression. The colorbar on the top codifies the expression level. (C) Correlation between expression of PR-LncRNAs and molecular markers frequently altered in glioma, namely, p53, Ki67, EGFR, IDH1, ATRX and MGMT. (D) Median overall survival stratified by level of each of the PR-LncRNAs studied.
Finally, we compared PR-LncRNA expression with overall glioblastoma patient survival and we did not observe significant correlations, although we noticed that patients with high levels had longer overall survival for all PR-LncRNAs (Fig. 2D). The mean survival rates of the subgroup of patients with high levels of PR-LncRNAs 1, 5 and 10, and Unassigned 4 were 20, 19.8, 18.1 and 21.9 months respectively, and notably, the survival rates were lower when biopsies presented low levels of these PR-LncRNAs: 16.3, 17, 17.2 and 16.6 months respectively (Fig. 2D). These data indicate a 2–4 month longer survival in patients with higher PR-LncRNA levels. In summary, these results show that the expression of PR-LncRNAs is altered in glioblastoma samples and low levels of these ncRNAs tend to correlate with characteristics linked to increased malignancy.
Silencing of PR-LncRNAs increases cellular proliferation and stem cell properties
In order to characterize the impact of PR-LncRNAs at the cellular level, we inhibited the activity of PR-LncRNAs by transfecting two specific ASOs for PR-LncRNA1 and 10 respectively. Quantitative reverse transcription PCR showed a significant decrease in the expression of the specific PR-LncRNA (Fig. 3A,B). Importantly, when we compared the proliferative potential of control and PR-LncRNA-silenced glioma cells, we detected that the number of phospho-histone-3 (PH-3) positive cells was markedly higher in cells transfected with the ASOs for PR-LncRNA1 and 10 (Fig. 3C,D). Indeed, 3.05% of controls were positive for PH3 compared to 4.9 and 5.38 for PR-LncRNA1 ASOs and 6.84 and 6.1 for PR-LncRNA10 ASOs. Moreover, the expression of p53 downstream targets such as p21cip, BAX and SerpinB5 was lower in cells transfected with ASOs for PR-LncRNA1 and 10 (Fig. 3E). These results support at molecular level the anti-proliferative activity of PR-LncRNAs and, since U87-MG cells present p53 wild-type status, the link with p53 pathway.
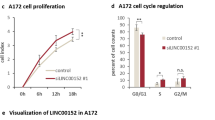
PR-LncRNA1 and 10 silencing leads to increased proliferation and stemness. U87-MG cells were transfected with specific ASOs for the PR-LncRNAs indicated. (A,B) Transfected cells were examined for PR-LncRNA1 and PR-LncRNA10 expression by quantitative reverse transcription polymerase chain reaction (n = 4). (C) Representative immunofluorescence of P-H3 in U87MG cells under the conditions indicated. (D) Quantification of the number of P-H3 positive cells under the conditions indicated (n = 4). (E) Quantification of mRNA levels of p21cip, Bax and SerpinB5 in cells transfected with ASOs for PR-LncRNA1 and 10 and compared to cells with a control ASO (F) Expression of PR-LncRNA 1,5 and 10 in indicated conventional cell lines (U87-MG, U251, U373 and A172) and glioma stem cells (GNS166, GNS179 and GB1) (G) Quantification of primary oncospheres formed in ASO-transfected cells after 10 days in culture (n = 3). (H) Quantification of number of secondary oncospheres generated from disaggregating primary oncospheres in ASO-transfected and control cells. Numbers were assessed after 10 days in culture (n = 3). Asterisks (*,**and ***) indicate statistical significance (p < 0.05, p < 0.01, and p < 0.001, respectively).
Glioma stem cells (GSCs) are a subpopulation of cells linked to glioma malignancy, progression and postulated as therapeutic targets25. We observed that the expression of PR-LncRNAs, particularly PR-LncRNA1 and 5, was lower in patient derived GSC lines (GNS166, GNS179 or GB1) than in conventional glioma cell lines such as U87MG, U251 or A172 (Fig. 3F). Moreover, the ability to form oncospheres was significantly elevated in cells with silencing of PR-LncRNA1 and 10 expression (Fig. 3G). This idea was further corroborated as cells with PR-LncRNA1 and 10 silencing formed significantly more secondary oncospheres (Fig. 3H). In summary, these results demonstrate that PR-LncRNAs regulate the activity of glioma cells.
SOX factors mediate PR-LncRNAs activity
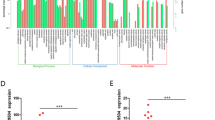
We and others have previously described several regulators of GSCs activity such as members of the SOX family, PML and STAT322,24,25,26,27,28,29. Therefore, we studied whether they were related to PR-LncRNAs activity. Correlation analysis in human biopsies determined that there is a strong inverse correlation between expression of the different PR-LncRNAs and that of SOX1, SOX2 and SOX9 (Fig. 4A,B). The inverse association in human biopsies was especially strong for SOX1 and SOX9 (Fig. 4B). Similar correlation was observed in glioma cells and GSCs, the latter expressing high levels of SOX members (Fig. 4C). To further investigate the link between PR-LncRNAs and SOX members, we studied the expression of the latter in cells transfected with PR-LncRNA1 and 10 ASOs. Interestingly, we observed that specially SOX1 and SOX2 expression was elevated in those cells (Fig. 4D), suggesting that these members of the SOX family may be mediators of PR-LncRNA activity in gliomas. On the contrary, cells with overexpression and silencing of SOX1, SOX2 and SOX9 do not present a clear pattern of decreased and increased levels of PR-LncRNAs respectively (Fig. Suppl 1). These results indicate that PR-LncRNA act upstream SOX family members to regulate glioma cell activity. To further test this idea, SOX1 and SOX9 expression was knocked-down in cells transfected with ASOs for PR-LncRNA1 and 10. Interestingly, silencing of SOX1 or SOX9 abolished the increase in proliferation and oncosphere formation promoted by PR-LncRNA1 and 10 inhibition (Fig. 4E,F), experimentally demonstrating that SOX factors are critical mediators of PR-LncRNA activity.
SOX stem cell factors mediate activity of PR-LncRNAs. (A) Analysis of the association between expression of PR-LncRNAs and stem cell factors in human glioma samples, showing there is a significant inverse correlation between PR-LncRNAs and SOX members. (B) Correlation between PR-LncRNAs and SOX factors in human glioma samples. Spearman Correlation Rho is −0,546 (**), −0,221 and −0,342 (*) for SOX1, SOX2 and SOX9 with PR-LncRNA1. It is −0,670 (***), −0,151 and −0,285 (*) for SOX1, SOX2 and SOX9 with PR-LncRNA5. Finally numbers are −0,521 (**), −0,171 and −0,230 for SOX1, SOX2 and SOX9 with PR-LncRNA10. (C) Violin plots associated to the expression distribution of the PR-LncRNAs (green circles) and SOX factors (red circles). Values in indicated cells are relative to expression of each transcript in U87-MG cells. The + denote the mean values of the distributions and the black dots are the expression values. (D) Quantitative reverse transcription polymerase chain reaction of SOX1, SOX2 and SOX9 in U87-MG control and transfected with ASOs. Data represents the average of 3 independent experiments. (E) Quantification of the number of P-H3 positive cells in U87-MG cells transfected with indicated ASOs and lentivirally infected with shSOX1, shSOX9 or empty vector (pLKO) (n = 2). (F) Quantification of the number of formed oncospheres in U87 cells transfected with indicated ASOs and lentivirally infected with shSOX1, shSOX9 or empty vector (pLKO) (n = 2). P values were determined by Student’s t test. Asterisks (*,**) indicate statistical significance (p < 0.05 and p < 0.01).
Discussion
LncRNAs are a novel group of genome regulators for which there is growing evidence linking their altered expression to a variety of cancers. In this work, we studied the expression and function of a recently described PR-LncRNA signature19 in gliomas. We found that the expression of this set of PR-LncRNAs is altered in samples from different glioma subtypes. Specifically, their levels are higher in low-grade than in high-grade gliomas, and decrease as the grade increases, with the lowest levels observed in glioblastoma samples. These results are in agreement with the previous study in which this set of PR-LncRNAs was found to be downregulated in colorectal cancer samples compared to levels in healthy control tissue19.
Clinically, we observed that patients with low levels of PR-LncRNAs had tumors in frontal or temporal sites, the right hemisphere and areas in contact with ventricles, characteristics, which have been associated with shorter patient survival30,31. In line with this, the subset of biopsies with low levels of the PR-LncRNAs studied overlapped with the subgroup of patients with the lowest median overall survival, although the association was not significant, likely due to the small size of the cohort. Building on the association of downregulated levels of PR-LncRNAs with poor clinical prognosis, in the previous study in colon cancer19, our findings indicate that the signature of PR-LncRNAs is not exclusive to human colorectal cancer and suggest that the pattern of expression of these Lnc-RNAs might constitute a tumor suppressor signature in different types of cancer. This activity might be influenced by the gender since female patients present higher levels of PR-LncRNAs particularly in grade III and glioblastomas. These results further support the link of PR-LncRNAs and poor prognosis because there is evidence that female patients have a survival advantage and live longer than males in glioblastoma32.
There is an increasing interest among scientists to generate tools to predict clinical phenotypes and tumor progression in order to identify biomarkers and to better design patient treatment33. Indeed, there have been generated computational models for lncRNA-cancer association prediction11,34. In our study, we generated computational models in order to determine whether the expression of PR-LncRNAs could predict glioma grade progression. However, the PR-LncRNA signature only discriminated partially between low and high-grade gliomas due to the very low amount of low-grade cases.
Deregulation of the p53 pathway is well known to occur and play a major role in the development and progression of several types of cancer including glioblastoma20. The PR-LncRNA signature was originally identified as p53-regulated LncRNAs19. We did not detect a statistically significant association between p53 status and PR-LncRNA expression, although most of the biopsies with high levels of PR-LncRNAs were positive for wild-type p53 in immunohistochemical analysis, and several p53 downstream targets were decreased in U87-MG cells with PR-LncRNAs silencing. It is important to indicate that the PR-LncRNA signature was identified as p53 regulated upon DNA damage in the HCT116 colorectal cancer cell line19. It is reasonable to surmise that: (i) the response of the PR-LncRNAs might be different in clinical samples and cell lines and also upon stressful conditions in vitro; (ii) the activity and regulation of the signature might be partially different within the different types of cancer; and (iii) some of the PR-LncRNAs might be regulated in a p53-independent manner, given that only around ~3% of the PR-LncRNAs altered by DNA damage in colorectal cells were directly bound by p5319. Further, the association study between p53 protein levels and PR-LncRNAs in human colorectal cancer biopsies has yet to be established. Therefore, further studies need to be conducted with a larger number of samples and in different types of cancer to decipher the link between p53 and PR-LncRNAs in clinical samples. Independently of all these possibilities, our observations confirm the previous results and link expression of the PR-LncRNA signature with poor prognosis and increased malignancy in clinical samples.
In agreement with the results obtained in clinical practice and in colorectal cancer cells19, functional studies in glioma cells demonstrated that PR-LncRNA silencing increased the proliferative capacity of cells supporting the potential role of PR-lncRNAs as a tumor suppressor signature in glioma. This effect was regulated, at least in part, by p53 pathway and SOX members. Importantly, we also observed that PR-LncRNAs regulate the activity of glioma stem cells. Indeed, silencing of the PR-LncRNAs1 and 10 significantly increased the oncosphere formation ability of glioma cells and their self-renewal potential. Moreover, the levels of PR-LncRNAs were lower in patient derived glioma stem cell populations than in differentiated conventional glioma cell lines. Of note, there is a small proportion of LncRNAs that have been shown to regulate the activity of the subpopulation of glioma stem cells13,35,36, and little is known concerning the underlying molecular mechanisms regulating this activity37. Therefore, we tried to identify the downstream mechanism of this action revealing that PR-LncRNAs regulate the expression of SOX2 SOX9 and especially SOX1, members of the SOX family transcription factor and stem cell regulators38. It is noteworthy that high levels of these genes have been previously linked to glioma poor prognosis and overall shorter survival in clinical practice and glioma stem cell population at the cellular level22,24,27,39. Thus, we observed that expression of PR-LncRNA was significantly inversely correlated with that of SOX2, SOX9 and SOX1 in human clinical biopsies and in cell cultures in vitro. Moreover, experiments in vitro revealed that their levels, especially of SOX1 and SOX2, were significantly elevated in PR-LncRNA silencing cells and that knock-down of SOX1 and SOX9 expression dramatically reduced the pro-oncogenic activities promoted by PR-LncRNA silencing in glioma cells. Together, these results postulate the SOX family as novel and critical mediator of the PR-LncRNAs activity. In line with these results, gain and loss of function studies demonstrated that SOX members regulate glioma stem cell function as well as glioma cell proliferation and tumorigenic activity27.
It has been postulated that understanding the function and mechanism by which LncRNAs participate in glioma stem cell activity might facilitate development of therapeutic strategies for the treatment of the disease14. In summary, our study identified a novel LncRNA signature associated with glioma progression and malignancy in the clinic and revealed the tumor suppressor role played by these LncRNAs at cellular level. Our data also revealed that members of the SOX family of stem cell regulators, in particular the recently described SOX124, are critical mediators of their activity. These results unravel the expression and function of the PR-LncRNAs in glioma pathobiology, postulating them as novel biomarkers in glioma diagnosis, especially for female patients, as well as potential therapeutic targets.
Change history
20 June 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41598-022-13824-8
References
Ostrom, Q. T. et al. The epidemiology of glioma in adults: a “state of the science” review. Neuro Oncol.16(7), 896–913 (2014 Jul).
Weller, M. et al. European Association for Neuro-Oncology (EANO) guideline on the diagnosis and treatment of adult astrocytic and oligodendroglial gliomas. The lancet oncology (2017).
Stupp, R. et al. Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. The lancet oncology.10(5), 459–66 (2009).
Stupp, R. et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med.352(10), 987–96 (2005).
Noushmehr, H. et al. Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell.17(5), 510–22 (2010).
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell.155(2), 462–77 (2013).
Verhaak, R. G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell.17(1), 98–110 (2010).
Djebali, S. et al. Landscape of transcription in human cells. Nature.489(7414), 101–8 (2012).
Huarte, M. The emerging role of lncRNAs in cancer. Nat Med.21(11), 1253–61 (2015).
Sahu, A., Singhal, U. & Chinnaiyan, A. M. Long noncoding RNAs in cancer: from function to translation. Trends. Cancer.1(2), 93–109 (2015).
Chen, X. & Yan, G. Y. Novel human lncRNA-disease association inference based on lncRNA expression profiles. Bioinformatics.29(20), 2617–24 (2013).
Han, L. et al. LncRNA pro fi le of glioblastoma reveals the potential role of lncRNAs in contributing to glioblastoma pathogenesis. Int J Oncol.40(6), 2004–12 (2012).
Yan, Y., Xu, Z., Li, Z., Sun, L. & Gong, Z. An Insight into the Increasing Role of LncRNAs in the Pathogenesis of Gliomas. Front Mol Neurosci.10, 53 (2017).
Kiang, K. M., Zhang, X. Q. & Leung, G. K. Long Non-Coding RNAs: The Key Players in Glioma Pathogenesis. Cancers (Basel).7(3), 1406–24 (2015).
Chen, X. Predicting lncRNA-disease associations and constructing lncRNA functional similarity network based on the information of miRNA. Sci Rep.5, 13186 (2015).
Wang, W. A., Lai, L. C., Tsai, M. H., Lu, T. P. & Chuang, E. Y. Development of a prediction model for radiosensitivity using the expression values of genes and long non-coding RNAs. Oncotarget.7(18), 26739–50 (2016).
Cao, Y. et al. Identification of prognostic biomarkers in glioblastoma using a long non-coding RNA-mediated, competitive endogenous RNA network. Oncotarget.7(27), 41737–47 (2016).
Li, Y. et al. Identification and characterization of lncRNA mediated transcriptional dysregulation dictates lncRNA roles in glioblastoma. Oncotarget.7(29), 45027–41 (2016).
Sanchez, Y. et al. Genome-wide analysis of the human p53 transcriptional network unveils a lncRNA tumour suppressor signature. Nat Commun.5, 5812 (2014).
Cancer Genome Atlas Research N. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature. 455(7216):1061–8 (2008).
Pollard, S. M. et al. Glioma stem cell lines expanded in adherent culture have tumor-specific phenotypes and are suitable for chemical and genetic screens. Cell Stem Cell.4(6), 568–80 (2009).
Garros-Regulez, L. et al. mTOR inhibition decreases SOX2-SOX9 mediated glioma stem cell activity and temozolomide resistance. Expert Opin Ther Targets.20(4), 393–405 (2016).
Etxaniz, U. et al. Neural-competent cells of adult human dermis belong to the Schwann lineage. Stem Cell Reports.3(5), 774–88 (2014).
Garcia, I. et al. Oncogenic activity of SOX1 in glioblastoma. Sci Rep.7, 46575 (2017).
Carrasco-Garcia, E. et al. Therapeutic strategies targeting glioblastoma stem cells. Recent Pat Anticancer Drug Discov.8(3), 216–27 (2013).
Garros-Regulez, L. et al. Targeting SOX2 as a Therapeutic Strategy in Glioblastoma. Front Oncol.6, 222 (2016).
de la Rocha, A. M., Sampron, N., Alonso, M. M. & Matheu, A. Role of SOX family of transcription factors in central nervous system tumors. Am J Cancer Res.4(4), 312–24 (2014).
Iwanami, A. et al. PML mediates glioblastoma resistance to mammalian target of rapamycin (mTOR)-targeted therapies. Proc Natl Acad Sci USA110(11), 4339–44 (2013).
Rahaman, S. O. et al. Inhibition of constitutively active Stat3 suppresses proliferation and induces apoptosis in glioblastoma multiforme cells. Oncogene.21(55), 8404–13 (2002).
Mistry, A. M. et al. Influence of glioblastoma contact with the lateral ventricle on survival: a meta-analysis. Journal of neuro-oncology.131(1), 125–33 (2017).
Awad, A. W. et al. Impact of removed tumor volume and location on patient outcome in glioblastoma. Journal of neuro-oncology.135(1), 161–71 (2017).
Franceschi, E. et al. The Prognostic Roles of Gender and O6-Methylguanine-DNA Methyltransferase Methylation Status in Glioblastoma Patients: The Female Power. World Neurosurg.112, e342–e7 (2018).
Wang, E. et al. Predictive genomics: a cancer hallmark network framework for predicting tumor clinical phenotypes using genome sequencing data. Semin Cancer Biol.30, 4–12 (2015 Feb).
Chen, X., You, Z. H., Yan, G. Y. & Gong, D. W. IRWRLDA: improved random walk with restart for lncRNA-disease association prediction. Oncotarget.7(36), 57919–31 (2016).
Katsushima, K. et al. Targeting the Notch-regulated non-coding RNA TUG1 for glioma treatment. Nat Commun.7, 13616 (2016).
Brodie, S. et al. The novel long non-coding RNA TALNEC2, regulates tumor cell growth and the stemness and radiation response of glioma stem cells. Oncotarget.8(19), 31785–801 (2017).
Shi, J. et al. Long non-coding RNA in glioma: signaling pathways. Oncotarget.8(16), 27582–92 (2017).
Sarkar, A. & Hochedlinger, K. The sox family of transcription factors: versatile regulators of stem and progenitor cell fate. Cell Stem Cell.12(1), 15–30 (2013).
Alonso, M. M. et al. Genetic and epigenetic modifications of Sox2 contribute to the invasive phenotype of malignant gliomas. PLoS One.6(11), e26740 (2011).
Acknowledgements
PA, JA-I and AS-A were recipients of a predoctoral fellowship from the Spanish Association Against Cancer (AECC Gipuzkoa), Basque Government and Instituto Salud Carlos III. This work was supported by grants from the Carlos III Institute of Health and the European Regional Development Fund (PI13/02277, CP16/00039, DTS16/084, and PI16/01580) and Industry and Health Departments of the Basque Country.
Author information
Authors and Affiliations
Contributions
S.-T.B., P.A., J.-A.I., A.-S.A. and I.G. performed experimental research; S.T.-B., M.A., J.U., A.Q., L.E., J.V. I.R., E.U. and N.S. collected patient information and samples, M.H. contributed with production of reagents; C.S. performed bioinformatics and biostatistical analysis. D.G. and M.-A.B. performed computational predictive models. N.S., I.G. and M.H. helped to analyze data, supervised experiments and all of them revised the manuscript; A.M. designed the research, directed the work, analyzed data, provided funding and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained errors in Figure 3. In Figure 3C, the representative image for ASO10A was inadvertently duplicated as ASO1B. Data from the fourth experimental replicate was also omitted from Figure 3D.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Torres-Bayona, S., Aldaz, P., Auzmendi-Iriarte, J. et al. PR-LncRNA signature regulates glioma cell activity through expression of SOX factors. Sci Rep 8, 12746 (2018). https://doi.org/10.1038/s41598-018-30836-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-30836-5
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.