Abstract
We have recently demonstrated that the rate-limiting enzymes in human glucose metabolism organize into cytoplasmic clusters to form a multienzyme complex, the glucosome, in at least three different sizes. Quantitative high-content imaging data support a hypothesis that the glucosome clusters regulate the direction of glucose flux between energy metabolism and building block biosynthesis in a cluster size-dependent manner. However, direct measurement of their functional contributions to cellular metabolism at subcellular levels has remained challenging. In this work, we develop a mathematical model using a system of ordinary differential equations, in which the association of the rate-limiting enzymes into multienzyme complexes is included as an essential element. We then demonstrate that our mathematical model provides a quantitative principle to simulate glucose flux at both subcellular and population levels in human cancer cells. Lastly, we use the model to simulate 2-deoxyglucose-mediated alteration of glucose flux in a population level based on subcellular high-content imaging data. Collectively, we introduce a new mathematical model for human glucose metabolism, which promotes our understanding of functional roles of differently sized multienzyme complexes in both single-cell and population levels.
Similar content being viewed by others
Introduction
Glucose metabolism consists of glycolysis and gluconeogenesis. Glycolysis converts glucose into pyruvate, whereas gluconeogenesis reverses the sequential reactions to produce glucose1. In normal healthy cells, glucose is converted into pyruvate. Pyruvate then shuttles to mitochondria for oxidative phosphorylation in the presence of oxygen. If the oxygen is limited, however, pyruvate becomes lactate. In addition to its association with energy metabolism, glucose also plays an important role in building block biosynthesis, including the pentose phosphate pathway and serine biosynthesis. This is particularly important in cancer cells because glucose metabolism is dysregulated in proliferating cancer cells in which glucose diverts into building block biosynthesis even in the presence of oxygen. Hence, understanding the mechanism of glucose flux regulation between energy metabolism and building block biosynthesis is important to address altered metabolic phenotypes in human cancer as well as other chronic diseases, including but not limited to diabetes and obesity2.
Meanwhile, multienzyme metabolic complexes catalyzing glycolysis have been identified in various organisms, including protists, plants, yeast, mammalian neurons and erythrocytes, and human cancer cells3. Recently, it has been shown that human liver-type phosphofructokinase 1 (PFKL) forms cytoplasmic clusters in human cancer cells and further colocalizes with other cytoplasmic rate-limiting enzymes of the pathway, including human liver-type fructose-1,6-bisphosphatase (FBPase), pyruvate kinase M2 (PKM2), and phosphoenolpyruvate carboxykinase 1 (PEPCK1), thus indicating the formation of a multienzyme complex, namely the glucosome4. The size of glucosome clusters becomes larger in human breast carcinoma cells (Hs578T), relative to non-cancerous human breast tissue cells (Hs578Bst), demonstrating the spatial alteration of glucose metabolism in cancer cells4. Similarly, tumor-promoting hypoxic conditions also increased the size of glucosome clusters in human hepatocellular carcinoma cells (HepG2)5. Importantly, our quantitative high-content imaging data4 supported a hypothesis that the glucosome clusters regulate the direction of glucose flux between energy metabolism and building block biosynthesis in a cluster size-dependent manner. Unfortunately, however, it has been challenging to prove the hypothesis experimentally by measuring metabolic activities of various sized glucosome clusters at subcellular levels.
In parallel, several mathematical models for glycolysis have been developed using ordinary differential equations (ODEs) for various model organisms: e.g., bacteria, yeast, virus and human (erythrocytes and liver cells)6,7,8,9,10,11. Along with incorporating in vitro kinetic parameters of the enzymes in glycolysis, some ODE-mediated models describe the importance of bistable behavior in glycolysis with strong emphasis on the known allosteric regulation of the rate-limiting enzymes12. Particularly, recent mathematical models for human liver or cancer cells cover a complicated network beyond glycolysis by including the pentose phosphate pathway, serine biosynthesis, glutamine metabolism, and/or mitochondrial metabolism13,14. However, mathematical models for glucose metabolism have barely explained yet how a multienzyme complex influences glucose flux in cancer cells in a cluster size-dependent manner.
In this work, we have constructed a mathematical model to understand how the formation of a multienzyme complex in glucose metabolism affects glucose flux in cancer cells. Since the glucosome is composed of at least four cytoplasmic rate-limiting enzymes of glucose metabolism4, we have formulated a model to accommodate all the members of the glucosome. Our model indeed reveals the impact of their spatial complexation in silico on temporal changes of glycolytic metabolites at a subcellular level in single cells. We further show that our model is capable of predicting metabolic outcomes of a population of cells based on the relative distribution of glucosome clusters in the population. In conclusion, our mathematical model includes not only enzyme kinetics and their allosteric regulation but also spatial compartmentalization of the rate-limiting enzymes of the pathway, thus allowing us to understand the metabolic contributions of glucose flux at both single-cell and population levels.
Model and Methods
A Mathematical Model
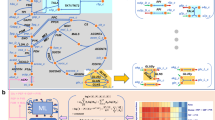
We develop a model to investigate how the size of glucosome clusters affects the direction of glucose flux in cancer environment as an indication of the metabolic activity of glucosome clusters in various sizes. Our model involves 7 metabolic intermediates (S i ), 9 enzyme-associated species (E i ) with various forms by which enzyme activities of the rate limiting steps are regulated, 3 metabolic products (P i ), and 28 reactions (Fig. 1 and Tables 1 and 2). Among the fourteen enzymes participating in glycolysis and/or gluconeogenesis, we only focus on the enzyme activities of the glucosome members (i.e., PFKL, FBPase, PKM2, and PEPCK1), which are hypothesized to have different activity levels in a cluster size-dependent manner. In Fig. 1, we present our simplified metabolic pathway with the enzymes involved in the glucosome. All chemical species and reaction rate constants in our pathway are introduced in Tables 1 and 2. An ordinary differential equations (ODE) model was used to describe temporal dynamics of the metabolic network involving enzymatic reactions associated with glucose metabolism. Temporal concentration changes in each metabolic intermediate are determined by propensity functions of the reactions where the intermediate is produced or consumed. Table 3 summarizes the propensity function R i for the ith reaction. In Table 3, most of the reactions are modeled by the law of mass action, in which the reaction rate is proportional to the substrate concentrations. The allosteric regulations of metabolic enzymes by metabolites are modeled as Michaelis-Menten kinetics, and their propensity functions include rational functions of the substrate concentrations to describe allosteric effects of the corresponding enzymes. Table 4 summarizes a set of all ODEs used in our model.
Simplified glucose metabolism with multienzyme complexes. Seven metabolic intermediates are involved in the pathway: S1 represents glucose, S2 is fructose-6-phosphate, S3 is fructose-1,6-bisphosphate, S4 is 3-phosphoglycerate, S5 is phosphoenolpyruvate, S6 is pyruvate, and S7 is oxaloacetate. Four rate-limiting enzymes are E1 (phosphofructokinase 1, PFK), E2 (fructose-1,6-bisphosphatase, FBPase), E3 (pyruvate kinase M2 dimer, PKM2), and E4 (phosphoenolpyruvate carboxykinase 1, PEPCK1). Pyruvate kinase M2 catalyzes conversion from S5 to S6 when it becomes a tetramer (\({E}_{3}^{\ast }\)). On the other hand, phosphofructokinase 1 is inactivated after post-translational glycosylation (E1gly). PFK forms three differently sized clusters: E S , E M , and E L represent small-, medium- and large-sized clusters, where E M and E L are multienzyme complexes. To measure the direction of glucose flux, we denote three metabolic products as P1, P2, and P3, which represent metabolic outcomes of the pentose phosphate pathway, serine biosynthesis and the downstream of glycolysis. All the used parameters are summarized in Tables 1 and 2.
We also provide reaction rate constants and initial values for the intermediate concentrations used in our model (Tables 1 and 2). We determined the parameters in Tables 1 and 2 using the numerical simulation of the model to be consistent with the qualitative changes in glucose flux in the literature15. As shown in Table 1, we assumed that initial concentrations of metabolic intermediates (S i ), products (P i ), PKM2 tetramers (E3*) and glycosylated PFKL (E1gly) are at the basal levels and set them as 0.01. We also assumed that no multienzyme complex (E S/M/L ) is formed at the beginning of simulation. An enzyme concentration of each glucosome member is assumed to be conserved, setting as 100. Assuming that all reaction rate constants for the enzymes are in the same order of magnitude, the reaction rate constants of metabolic enzymes are set to 10 as shown in Table 2. However, we slightly modified the values for k2, k−2, k4, and k−4 so that glycolysis becomes dominant relative to the metabolic shunt to the pentose phosphate pathway and/or serine biosynthesis in the absence of enzyme clusters (i.e., P3 ≫ P1, P2). Lastly, we assumed that all the metabolic products are produced (k p/s/f ) and degraded (δ p/s/f ) in the same rates, thus setting them to 5 and 0.5, respectively. Note that we provided both forward and backward reaction rate constants in Table 2, which are separated by comma.
Next, different levels of the enzyme activity depending on the size of glucosome clusters are expressed in terms of the parameters c i (for medium-sized enzyme clusters) and e i (for large-sized enzyme clusters) in Table 2. Based on our hypothesis that the medium-sized enzyme clusters promote glucose shunt to the pentose phosphate pathway4, we decelerated glycolysis (c2, c−d ≪ 1) and accelerated gluconeogenesis (c−2, c−6 ≫ 1). Similarly, based on our hypothesis that the large-sized enzyme clusters will shunt glucose flux to serine biosynthesis4, we changed parameters to direct glucose flux toward serine production (e2, e−6 ≫ 1 , and e−2, e−d ≪ 1). An enzyme activity in the small-sized clusters is assumed similar to the activity of a freestanding enzyme because the small-sized clusters formed by PFKL do not represent a multienzyme complex4. Using the initial values and the reaction rate constants shown in Tables 1 and 2, our mathematical model was simulated using MATLAB to compute temporal changes of glucose flux in different scenarios at the subcellular level. Note that our MATLAB simulation codes are provided in the supplementary document.
Importantly, we have also performed sensitivity analysis of the parameters to investigate how the concentrations of three metabolic products (P i ) are affected by small changes in each parameter that we have incorporated in our model. In the sensitivity analysis, we have used the method of partial rank correlation coefficient (PRCC)16 to evaluate the level of the linear association between the input values of the parameters and the output concentrations of the metabolic products. To perform the sensitivity analysis, we modified the publically available MATLAB codes which are developed by Kirschner and coworker16. The default parameter values were set as the values used in our model, and we defined an interval for each parameter from half of the default value to twice of the default value. Then, we used Latin hypercube sampling16 to choose each parameter value in the corresponding interval by assuming that each parameter is uniformly distributed in the interval. Afterward, the sampled parameter values were used to calculate the concentrations of the metabolic products at the end time of the simulation. We repeated the sampling process described above 20,000 times and calculated the Spearman correlation coefficients, which correspond to PRCCs.
Experimental Methods
Materials
The plasmid expressing human PFKL with a monomeric form of enhanced green fluorescent protein (mEGFP) was prepared previously (PFKL-mEGFP)4. We have employed PFKL-mEGFP as a marker of the glucosome. 2-Deoxyglucose was purchased from Sigma.
Cell Culture
Human breast carcinoma cells, Hs578T (HTB-126), were obtained from the American Type Culture Collection (ATCC). Hs578T cells were maintained in the Roswell Park Memorial Institute 1640 medium (RPMI1640, Mediatech, Cat# 10-040-CV) supplemented with 10% dialyzed fetal bovine serum (dFBS, Atlanta Biological, Cat# S12850) and 50 µg/mL gentamycin sulfate.
Transfection
To prepare cells for transfection and subsequent imaging, Hs578T cells were gently removed from the culture flask by replacing the culture medium with Trypsin-EDTA solution (Corning, Cat# 25-053-Cl). Fresh, antibiotic-free growth media was subsequently used to harvest and resuspend the cells, followed by plating on glass-bottomed 35 mm Petri dishes (MatTek). When the confluency became ~70–90% on the following day, the cells were transfected with Lipofectamine 2000 (Invitrogen) using the Opti-MEM-I reduced serum media (Opti-MEM-I; Gibco, Cat# 11058). The Opti-MEM-I medium was then exchanged with the fresh antibiotic-free growth medium after a 5 h incubation (37 °C, 5% CO2, and 95% humidity), followed by ~18–24 h incubation in the CO2 incubator at 37 °C.
Fluorescence Live-cell Imaging
On the day of imaging (~18–24 h post-transfection), cells were washed with imaging solution (20 mM HEPES (pH 7.4), 135 mM NaCl, 5 mM KCl, 1 mM MgCl2, 1.8 mM CaCl2, and 5.6 mM glucose) for three 10 min incubations, followed by a ~1–2 h incubation at ambient temperature. All samples were then imaged at ambient temperature (~25 °C) with a 60× 1.45 NA objective (Nikon CFI Plan Apo TIRF) using a Photometrics CoolSnap EZ monochrome CCD camera on a Nikon Eclipse Ti inverted C2 confocal microscope. Epifluorescence imaging was carried out using the following filter set from Chroma Technology; mEGFP detection by a set of Z488/10-HC cleanup, HC TIRF Dichroic and 525/50-HC emission filter.
For cell-based high-content imaging assays, 2-deoxyglucose (25 mM) was added to Hs578T cells after washing three times with the imaging solution. Images showing PFKL-mEGFP clusters were acquired before and after cells were incubated with 2-deoxyglucose at various time points (up to 6 hours). Control experiments were also carried out with 167 μL of vehicle (i.e., water). To ensure reproducibility, our experiments were repeated at least five times over the course of a few months. Statistical analysis was performed using two-sample two-tail t-tests.
Cluster Size Analysis
Cluster size analysis was accomplished using the ImageJ processing software (National Institutes of Health) as we have done before4. Briefly, fluorescent wide-field images were processed through ImageJ using a custom script and macro that automates the counting of fluorescent clusters using its built-in module, so-called robust automatic threshold selection (RATS). In this analysis, the images were scaled according to the pixel size of the microscope (i.e., 0.12 µm/pixel) before the default parameters for RATS (i.e., noise threshold = 25, λ factor = 3) were used in this analysis. Once fluorescent clusters were selected from an image, the particle analysis module was applied to attain both the number and area of fluorescent clusters within an image. This process was repeated for all subsequent cell images. The operator then evaluated the original cell images against the particle mask to eliminate data in which more than one cluster was counted as a single particle.
Results
Modeling Glucose Metabolism with Multienzyme Complexes
Glucose metabolism consists of glycolysis and gluconeogenesis where three irreversible reactions are involved in glycolysis, four irreversible reactions in gluconeogenesis, and seven reversible reactions in both pathways. In our model, we simplify the 14-step pathway by condensing a few reversible steps into one conversion (Fig. 1). Accordingly, we omit some of the metabolic intermediates such as glucose-6-phosphate, dihydroxyacetone phosphate, glyceraldehyde-3-phosphate, 1,3-bisphosphoglycerate, and 2-phosphoglycerate because they are not directly shuttled to other metabolic pathways. In addition, three metabolic products (P i ) represent three metabolic fates of glycolytic metabolites. Importantly, we focus on four cytoplasmic enzymes which spatially organize into multienzyme clusters (i.e., glucosome clusters)4: PFKL, FBPase, PKM2, and PEPCK1. Note that the other ten enzymes are not cytoplasmic or rate-determining in glucose metabolism.
In addition, we incorporated cancer-relevant mechanisms of the four enzymes forming the glucosome. First, pyruvate kinase catalyzes conversion from phosphoenolpyruvate to pyruvate. The M2 isoform of pyruvate kinase (PKM2) is predominantly expressed in cancer cells while suppressing the expression of pyruvate kinase M1 isoform (PKM1)17. PKM2 plays an important role in aerobic glycolysis as a dimer by impairing pyruvate production, thus redirecting glucose flux into serine biosynthesis18,19. However, a tetrameric form of PKM2 possesses its canonical glycolytic activity in production of pyruvate in cancer cells. We take into account the dimer-tetramer conversion as part of the regulation of glucose metabolism (Fig. 1). Second, PFK catalyzes the conversion of fructose-6-phosphate to fructose-1,6-bisphosphate. Particularly, liver-type PFK (PFKL) is glycosylated in several types of cancer cells under hypoxia conditions, diverting the direction of glucose flux into the pentose phosphate pathway20. Our model includes the impact of glycosylation on PFKL as part of the regulation of its metabolic activity in glycolysis. Third, cytoplasmic PEPCK1 plays an essential role in the first step of gluconeogenesis21. Recently, a new role of PEPCK1 in tumor cell proliferation was discovered in certain types of cancer cells by controlling carbon metabolic flux through the TCA cycle22. However, a molecular level mechanism of PEPCK1 in cancer cells (e.g., post-translational modification-dependent activity or oligomerization-dependent activity) is not firmly established yet to be included in our model. Fourth, FBPase catalyzes the conversion of fructose-1,6-bisphosphate to fructose-6-phosphate. Since FBPase catalyzes the backward reaction of PFK, the activities of PFK and FBPase are reciprocally regulated by various allosteric metabolites. Similarly, an antagonistic role of FBPase for glycolytic flux was validated to play a role in kidney cancer progression23. However, other than the well-established allosteric regulation, cancer-specific alteration of FBPase is not established yet to be included in our model.
Unlike many previous mathematical models for glycolysis, we take into account the recent discovery of the glucosome being formed in various sizes in cancer cells. Briefly, various sizes of PFKL clusters in cancer cells are categorized into three subclasses; small, medium and large-sized cluster4. Small-sized clusters formed by PFKL are single-enzyme assemblies whereas medium- and large-sized clusters represent spatial organizations of multienzyme complexes. When cancer cells display small-sized clusters, the given cells barely include the other sizes of clusters. However, cancer cells showing medium- and large-sized clusters contain smaller sized clusters. Our cluster size analysis has also revealed that HeLa cells with medium-sized clusters show ~86.1 clusters per cell whereas HeLa cells showing large-sized clusters exhibit ~5.6 large-sized clusters and ~25.1 medium-sized clusters per cell4. Accordingly, we assume that 11 medium-sized clusters might form one large cluster in the cytoplasm of cancer cells in our model.
Glucose Flux Analysis at Subcellular Levels
In Fig. 2, metabolic product concentrations are shown in different experimental conditions, which are computed by numerically solving the mathematical model given in Tables 3 and 4. The model results in time-dependent changes of three metabolic products: P1 represent a metabolic product of the pentose phosphate shunt (green line), P2 is a metabolic product of serine biosynthesis (red line), and P3 indicates a metabolic product of the downstream of glycolysis (blue line). In addition, the activity of non-clustering PFKL or clustered PFKL into small sizes are anticipated to be similar in our model. For the simulation of the cases with no cluster or only small-sized PFKL clusters, we set all the parameters related to the formation of medium or large-sized enzyme complexes as zero (i.e., k am , k−am, k al , k−al, c i , and e i ). Please, note that the parameters for small-sized PFKL clusters (i.e., k as and k−as) were also set to zero when we simulated the case showing no cluster. Consequently, our simulation for no cluster or only small-sized PFKL clusters showed the high level of P3 relative to those of P1 and P2 (Fig. 2A and B), indicating that most of glucose flux flows to glycolysis to produce pyruvate and beyond. However, when the rate-limiting enzymes in glucose metabolism are spatially organized into medium-sized glucosome clusters, the enzymes in medium-sized clusters may have different levels of catalytic activity. Because the increased flux of the pentose phosphate shunt is correlated with the high level of medium-sized clusters in cancer cells4, our assumption of c2, c−d ≪ 1 and c−2, c−6 ≫ 1 resulted in a significant increase of P1 but decreased the concentrations of P2 and P3 (Fig. 2C). In this case, we set the values of k al , k−al, and e i as zero. Lastly, we considered all the rate-limiting enzymes being assembled into large-sized glucosome clusters in the presence of smaller clusters. Because the promotion of large-sized clusters in cancer cells is correlated with the increased flux of serine biosynthesis4, our assumption of e2, e−6 ≫ 1 and e−2, e−d ≪ 1 along with the same assumption of c i in the previous case resulted in the substantially increased level of P2 relative to the levels of P1 and P3 (Fig. 2D). Collectively, our mathematical model is developed to understand the cluster size-dependent changes of glucose flux in cancer cells.
Glucose flux analysis at subcellular levels. Time-dependent changes of metabolic product concentrations (P1, P2 and P3) in single cells are graphed for: (A) cells with no spatial organization, (B) cells with small-sized PFKL clusters which indicate single enzyme assemblies, (C) cells with medium-sized multienzyme complexes, and (D) cells with large-sized multienzyme complexes in the presence of smaller clusters. Arbitrary units (a.u.) are used to show relative concentrations or time during our model simulation. P1, P2, and P3 represent metabolic outcomes of the pentose phosphate pathway, serine biosynthesis and the downstream of glycolysis, respectively.
Sensitivity Analysis
Next, we performed a sensitivity analysis by which we evaluated without bias how the input values of simulation parameters influence the production of specific metabolic flux (P i ). According to the publically available MATLAB codes developed by Kirschner and coworker16, we modified to calculate PRCCs in our model (Figure S1). If the PRCC values are close to −1 or +1, the corresponding parameters have strong correlations with the formation of the given metabolic product. On the other hand, if the correlation coefficient is close to 0, there is no correlation between the input parameters and the given metabolic product. When the absolute values of the PRCCs are greater than ±0.2 and their p-values are less than 0.01, we defined the corresponding parameters as ‘sensitive’ parameters (Fig. 3) to determine which simulation parameters are critical for our model and how sensitive each parameter is to understand its contribution to the outcomes of glucose flux. Noticeably, the formation of each metabolic product was sensitive not only to the enzyme activities of the glucosome members but also the efficiencies of enzyme clustering into the certain sizes of glucosomes.
The partial rank correlation coefficients (PRCCs) between the ‘essential’ input parameters and the concentrations of metabolic products. The PRCCs at time = 10 are graphed to provide relative strengths of the correlations between the input parameters and the concentrations of metabolic products (P1, P2, and P3). The horizontal lines at ±0.2 indicate the thresholds we used to distinguish sensitive essential parameters from non-essential parameters. The PRCCs of all parameters are shown in supplementary figure S1. P1, P2, and P3 represent metabolic outcomes of the pentose phosphate pathway, serine biosynthesis and the downstream of glycolysis, respectively.
In Fig. 4, we further analyzed a few critical parameters that influence the enzyme activities of the glucosome members (i.e., k2, k −2 , and k−d). Each selected parameter was perturbed in one direction while all other parameter values were fixed as default values as shown in Table 2. The changes in the metabolic product concentrations were then presented under four cases; (i) no spatial organization, (ii) small-sized PFKL clusters, (iii) medium-sized multienzyme complexes, and (iv) large-sized multienzyme complexes. Note that for the cases dealing with the medium-sized or large-sized complexes, the smaller-sized clusters are also formed in the model. When k2 was decreased from 40 to 10 (Fig. 4A), the conversion from S2 to S3 was reduced. We thus observed significant increase in P1 while decrease in P2 and P3. On the other hand, when k−2 was increased from 7 to 10 (Fig. 4B), a similar pattern was observed as in Fig. 4A although the changes were small due to the narrow ranges of the parameter. Finally, when k−d was increased from 1 to 10 (Fig. 4C), glucose flux moved toward the production of P3, which resulted in significant increase in P3 and decrease in P1 or P2. Note that we did not observe much increase of P3 in the cases where no spatial organization or only small-sized PFKL clusters are dominant (Fig. 4C) because the flux was already committed toward the production of P3 in these cases. Collectively, the simulated flux changes reflect the changes of enzymatic activities involved in glucosome clusters.
Simulated concentration changes of metabolic products with varying ranges of the catalytic activities of glucosome members. Time-dependent concentration changes of three metabolic products are expressed as shaded regions, when one ‘essential’ kinetic rate constant is perturbed for k2 ranging from 40 to 10 (A), k−2 ranging from 7 to 10 (B), and k−d from 1 to 10 (C). In each case, time-dependent concentration changes of three metabolic products in single cells are graphed for (i) cells with no spatial organization, (ii) cells with small-sized PFKL clusters, (iii) cells with medium-sized multienzyme complexes, and (iv) cells with large-sized multienzyme complexes.
In Fig. 5, we also analyzed more critical parameters that influence the efficiencies of enzyme clustering into the glucosomes (i.e., c−2, e2, and e−d). Similarly, each selected parameter was perturbed in one direction while fixing the other parameter values. The changes in the metabolic product concentrations were shown under the four cases as well. When c−2 was decreased from 10 to 1 (Fig. 5A), the level of P1 decreased and that of P2 increased significantly due to the decreased contribution of the medium-sized cluster to the conversion from S3 to S2. When e2 was decreased from 2.5 to 1 (Fig. 5B), the resulting effect was quite opposite from the one shown Fig. 5A because the decreased e2 reduces the contribution of the large-sized clusters to the conversion of S2 to S3. When e−d was increased from 0.05 to 1 (Fig. 5C), the level of P2 decreased and that of P3 increased significantly with slight decrease in P1. This is due to the increased efficiency of the large-sized cluster on its contribution to the conversion of E3 to \({E}_{3}^{\ast }\).
Simulated concentration changes of metabolic products with varying ranges of the clustering efficiency of glucosomes. Time-dependent concentration changes of three metabolic products are expressed as shaded regions, when one ‘essential’ clustering efficiency is perturbed for c−2 ranging from 10 to 1 (A), e2 from 2.5 to 1 (B), and e−d from 0.05 to 1 (C). In each case, time-dependent concentration changes of three metabolic products in single cells are graphed for (i) cells with no spatial organization, (ii) cells with small-sized PFKL clusters, (iii) cells with medium-sized multienzyme complexes, and (iv) cells with large-sized multienzyme complexes.
Metabolic Flux Analysis at Ensemble Levels
We have further evaluated our model by quantifying the changes of glucose flux in the presence of methylene blue, fructose-1,6-bisphosphate or epidermal growth factor (EGF), which are known to regulate the direction of glucose flux at ensemble levels24,25,26,27,28,29. As shown in Table 5, we have counted the percentages of cells showing no cluster, small, medium, and large clusters in the absence and presence of the listed glucose flux regulators. In combination with our subcellular flux analysis (Fig. 2), we have computed the time-dependent changes of metabolic products (P1, P2, and P3) at an ensemble level in the four published cases4. To predict the levels of metabolic products at an ensemble level, we multiplied the levels of P1, P2, and P3 of the four cases shown in Fig. 2 by the corresponding proportions of the cells showing differently sized clusters. In a control population of Hs578T cells without any exogenous stimulus (Fig. 6A), the level of P3 appears to be slightly greater than those of P1 and P2. However, relative to Fig. 6A, the cancer cells that were treated with methylene blue (Fig. 6B) or fructose-1,6-bisphosphate (Fig. 6C) increase the level of P1 while decreasing the level of P3 at an ensemble level. On the other hand, Fig. 6D shows the case where the cancer cells have been treated with EGF. In this case, the level of P2 increases but the level of P1 decreases relative to the other cases. It appears clear that the simulation is capable of showing that glucose flux shunt from glycolysis to either the pentose phosphate pathway or the serine biosynthesis at ensemble levels, whose trend is indeed consistent with the already known functions of the glucose flux regulators in populations24,25,26,27,28,29. Collectively, our mathematical model with high-content imaging analysis is adequate to predict metabolic consequences of glucose flux regulators in a population of cancer cells.
Metabolic flux analysis at ensemble levels. Time-dependent concentration changes of three metabolic products are simulated by our mathematical model in the four scenarios: cancer cells without a glucose flux regulator as a control (A), cancer cells that are treated with 5 nM methylene blue (B) or 15 mM fructose-1,6-bisphosphate (C), and cancer cells with 30 ng/ml epidermal growth factors (D). Note that human breast cancer cells (Hs578T) were cultured in the medium of RPMI1640 and 10% dialyzed FBS. P1, P2, and P3 represent metabolic outcomes of the pentose phosphate pathway, serine biosynthesis and the downstream of glycolysis, respectively.
Mathematical Prediction of Metabolic Flux with 2-Deoxyglucose at Ensemble Levels
Our model is now applied to assess metabolic consequences of 2-deoxyglucose in Hs578T cells. 2-Deoxyglucose is a competitive inhibitor of phosphoglucose isomerase catalyzing step 2 in glycolysis30. However, due to the lack of 2-hydroxyl group, 2-deoxyglucose cannot further metabolize into fructose-6-phosphate in downstream glycolysis, thereby resulting in the inhibition of glycolysis. Accordingly, 2-deoxyglucose appeared to impair cell growth and thus implemented for anti-tumor therapeutics. However, its pharmacological effects on cancer treatment31,32,33 remain inconclusive, consequently raising a number of questions particularly including potential metabolic alterations by 2-deoxyglucose in cancer cells.
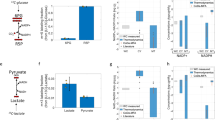
In this work, we have employed our mathematical model to predict whether 2-deoxyglucose can potentially alter the direction of glucose flux in cancer cells beyond glycolysis inhibition. First, we performed cell-based high-content imaging assays to validate the metabolic effect of 2-deoxyglucose at single cell levels (Fig. 7). Briefly, we expressed PFKL-mEGFP as a glucosome marker in Hs578T cells. To be consistent with Table 5, Hs578T cells were cultured in RPMI1640 with 10% dialyzed FBS. When we treated transfected Hs578T cells with 25 mM of 2-deoxyglucose for 6 hours, the percentage of cells showing small-sized clusters was significantly reduced from 58.3 % to 34.7% (Fig. 7A). Meanwhile, the percentages of cells showing medium- and large-sized clusters were significantly increased from 13.4% to 21.2% and from 26.7% to 44.1%, respectively (Fig. 7A), indicating the strong promotion of glucosome formation and thus metabolic shunts to anabolic pathways.
The effect of 2-deoxyglucose on the distribution of Hs578T cells with various sizes of PFKL-mEGFP clusters. The percentage (%) of Hs578T cells displaying each size of PFKL-mEGFP cluster was analyzed in the presence of 2-deoxyglucose. (A) The graph shows the average percentages (%) of cells displaying the given sized clusters along with their standard deviations (±) in the absence (black bars) and presence (red bars) of 2-deoxyglucose. At least five independent imaging sessions were performed and total 1200 transfected cells were analyzed. Statistical analyses were performed using two-sample two-tail t-tests. *p < 0.01. (B) Metabolic flux analysis were performed using our mathematical model with the high-content imaging data of 2-deoxyglucose. Relative to a control flux (Fig. 2A), glycolytic flux (P3) decreased, but the metabolic shunts of glucose to the pentose phosphate pathway (P1) and serine biosynthesis (P2) increased. (C and D) Representative images of Hs578T cells show subcellular localization of PFKL-mEGFP before and after treatment of 2-deoxyglucose (25 mM) for 6 hours. Scale bar, 10 µm.
Subsequently, we computed an ensemble-level outcome of glucose flux in the presence 2-deoxyglucose. Similarly to Fig. 6, we calculated the changes of metabolic outcomes at ensemble levels based on our subcellular flux analysis (Fig. 2) and high-content imaging analysis (Fig. 7A, Table 5). Relative to a control flux in Hs578T cells where glycolytic flux reaches to ~7 arbitrary units at t = 10 (Fig. 6A), glycolytic flux was indeed inhibited at ~ 5 arbitrary units in the presence of 2-deoxyglucose (Fig. 7B). Importantly, our model provided that metabolic shunts from glycolysis to anabolic biosynthetic pathways were promoted and relatively became dominant in the population of Hs578T cells. Although more experimental validation may be necessary, it is clear that 2-deoxyglucose promoted the formation of glucosome clusters in both single-cell (Fig. 7C,D) and ensemble levels (Fig. 7B). Therefore, we propose that along with the inhibitory role of 2-deoxyglucose in glycolysis, its commitment diverting glucose flux into anabolic pathways may explain in part cancer progression observed during its clinical trials31.
Discussion
We constructed a simple mathematical model to understand the cluster size-dependent functional contributions of metabolic enzymes and their multienzyme complexes to cancer cell metabolism. Briefly, glycolysis is interconnected with energy metabolism and anabolic biosynthetic pathways, including the pentose phosphate pathway and serine biosynthesis. Three metabolic products (P i ) thus represent downstream pathways of glycolysis, allowing us to investigate how glucose flux changes its direction at metabolic nodes between energy metabolism and anabolic pathways in cancer cells. Based on our experimental data4, we developed a model to predict that medium-sized clusters of glucosomes shunt glucose flux into the pentose phosphate pathway whereas large-sized clusters of glucosome divert glucose flux into serine biosynthesis at subcellular levels. Importantly, our model supports that the changes of relative ratios of cancer cells displaying small-, medium- and large-sized clusters in a population appear to be significant enough to influence overall net metabolic outcomes of cancer cells at ensemble levels. Moreover, our mathematical model is further evaluated to predict the effect of 2-deoxyglucose on the fate of glucose, thus providing new quantitative insights of how 2-deoxyglucose alters glucose flux in cancer cells. Collectively, we conclude that our mathematical model supports the hypothesis that glucosomes divert glucose flux into the pentose phosphate pathway and serine biosynthesis in a cluster size-dependent manner.
It is important to emphasize here that our mathematical model accounting for size-dependent metabolic functions of glucosome clusters is significantly different from other models that mostly rely on in vitro enzyme kinetics. Our model explains altered glycolysis in cancer cells that redirects glucose flux into the pentose phosphate pathway and serine biosynthesis in a cluster size-dependent manner. Importantly, we can now predict the direction of glucose flux in the presence of small-molecule drug candidates as long as quantitative high-content imaging data is obtained in single-cell levels. Additionally, we can quantify relative partition ratios of glucose flux between glycolysis, the pentose phosphate pathway and serine biosynthesis in various conditions. Collectively, our mathematical model fully integrates various cancer-associated mechanisms discovered in recent years, thus advancing our understanding of cancer cell metabolism in single cells.
References
Hers, H. G. & Hue, L. Gluconeogenesis and related aspects of glycolysis. Annu Rev Biochem 52, 617–653 (1983).
Scrutton, M. C. & Utter, M. F. Regulation of Glycolysis and Gluconeogenesis in Animal Tissues. Annu Rev Biochem 37, 249 (1968).
Schmitt, D. L. & An, S. Spatial Organization of Metabolic Enzyme Complexes in Cells. Biochemistry 56, 3184–3196 (2017).
Kohnhorst, C. L. et al. Identification of a multienzyme complex for glucose metabolism in living cells. J Biol Chem 292, 9191–9203 (2017).
Jin, M. et al. Glycolytic Enzymes Coalesce in G Bodies under Hypoxic Stress. Cell Rep 20, 895–908 (2017).
Holzhutter, H. G., Jacobasch, G. & Bisdorff, A. Mathematical modelling of metabolic pathways affected by an enzyme deficiency. A mathematical model of glycolysis in normal and pyruvate-kinase-deficient red blood cells. Eur J Biochem 149, 101–111 (1985).
Joshi, A. & Palsson, B. O. Metabolic dynamics in the human red cell. Part I—A comprehensive kinetic model. J Theor Biol 141, 515–528 (1989).
Munger, J. et al. Systems-level metabolic flux profiling identifies fatty acid synthesis as a target for antiviral therapy. Nat Biotechnol 26, 1179–1186 (2008).
Rizzi, M., Baltes, M., Theobald, U. & Reuss, M. In vivo analysis of metabolic dynamics in Saccharomyces cerevisiae: II. Mathematical model. Biotechnol Bioeng 55, 592–608 (1997).
Teusink, B. et al. Can yeast glycolysis be understood in terms of in vitro kinetics of the constituent enzymes? Testing biochemistry. Eur J Biochem 267, 5313–5329 (2000).
Voit, E. O. et al. Regulation of glycolysis in Lactococcus lactis: an unfinished systems biological case study. Syst Biol (Stevenage) 153, 286–298 (2006).
Mulukutla, B. C., Yongky, A., Daoutidis, P. & Hu, W. S. Bistability in glycolysis pathway as a physiological switch in energy metabolism. PLoS One 9, e98756 (2014).
Khazaei, T., McGuigan, A. & Mahadevan, R. Ensemble modeling of cancer metabolism. Front Physiol 3, 135 (2012).
Marin-Hernandez, A. et al. Modeling cancer glycolysis. Biochim Biophys Acta 1807, 755–767 (2011).
Benjamin, D. I., Cravatt, B. F. & Nomura, D. K. Global profiling strategies for mapping dysregulated metabolic pathways in cancer. Cell Metab 16, 565–577 (2012).
Marino, S., Hogue, I. B., Ray, C. J. & Kirschner, D. E. A methodology for performing global uncertainty and sensitivity analysis in systems biology. J Theor Biol 254, 178–196 (2008).
Mazurek, S., Boschek, C. B., Hugo, F. & Eigenbrodt, E. Pyruvate kinase type M2 and its role in tumor growth and spreading. Semin Cancer Biol 15, 300–308 (2005).
Christofk, H. R. et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 452, 230–233 (2008).
Wong, N., De Melo, J. & Tang, D. PKM2, a Central Point of Regulation in Cancer Metabolism. Int J Cell Biol 2013, 242513 (2013).
Yi, W. et al. Phosphofructokinase 1 glycosylation regulates cell growth and metabolism. Science 337, 975–980 (2012).
Xiong, Y., Lei, Q. Y., Zhao, S. & Guan, K. L. Regulation of Glycolysis and Gluconeogenesis by Acetylation of PKM and PEPCK. Cold Spring Harb. Symp. Quant. Biol. 76, 285–289 (2011).
Montal, E. D. et al. PEPCK Coordinates the Regulation of Central Carbon Metabolism to Promote Cancer Cell Growth. Mol Cell 60, 571–583 (2015).
Li, B. et al. Fructose-1,6-bisphosphatase opposes renal carcinoma progression. Nature 513, 251–255 (2014).
Bolanos, J. P., Delgado-Esteban, M., Herrero-Mendez, A., Fernandez-Fernandez, S. & Almeida, A. Regulation of glycolysis and pentose-phosphate pathway by nitric oxide: impact on neuronal survival. Biochim. Biophys. Acta 1777, 789–793 (2008).
Delgado, T. C., Castro, M. M., Geraldes, C. F. & Jones, J. G. Quantitation of erythrocyte pentose pathway flux with [2-13C]glucose and 1H NMR analysis of the lactate methyl signal. Magn. Reson. Med. 51, 1283–1286 (2004).
Kelleher, J. A., Chan, P. H., Chan, T. Y. & Gregory, G. A. Energy metabolism in hypoxic astrocytes: protective mechanism of fructose-1,6-bisphosphate. Neurochem. Res. 20, 785–792 (1995).
Lewis, I. A., Campanella, M. E., Markley, J. L. & Low, P. S. Role of band 3 in regulating metabolic flux of red blood cells. Proc. Natl. Acad. Sci. USA 106, 18515–18520 (2009).
Martino, G. et al. The fate of human peripheral blood lymphocytes after transplantation into SCID mice. Eur. J. Immunol. 23, 1023–1028 (1993).
Possemato, R. et al. Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature 476, 346–350 (2011).
Wick, A. N., Drury, D. R., Nakada, H. I. & Wolfe, J. B. Localization of the primary metabolic block produced by 2-deoxyglucose. J Biol Chem 224, 963–969 (1957).
Raez, L. E. et al. A phase I dose-escalation trial of 2-deoxy-D-glucose alone or combined with docetaxel in patients with advanced solid tumors. Cancer Chemother Pharmacol 71, 523–530 (2013).
Ralser, M. et al. A catabolic block does not sufficiently explain how 2-deoxy-D-glucose inhibits cell growth. Proc Natl Acad Sci USA 105, 17807–17811 (2008).
Urakami, K., Zangiacomi, V., Yamaguchi, K. & Kusuhara, M. Impact of 2-deoxy-D-glucose on the target metabolome profile of a human endometrial cancer cell line. Biomed Res 34, 221–229 (2013).
Acknowledgements
We would like to thank Dr. Bradford Peercy (UMBC) for critically reading our manuscript. We thank UMBC for providing the Start-Up funds (S.A. and H.-W.K., respectively). This work was also supported in part by the 2016 AACR-Bayer Innovation and Discovery Grant (#16-80-44-ANSO; S.A.), NIH/NIGMS (R01GM125981; S.A.), and the National Science Foundation (DMS-1620403; H.-W.K.). M.J. was also funded in part by NIH/NIGMS Training Grant (T32GM066706).
Author information
Authors and Affiliations
Contributions
S.A. and H.-W.K. designed research and developed a model; M.J. performed high-content imaging experiments; H.-W.K. performed numerical simulation of the models; and S.A. and H.-W.K. wrote the paper.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jeon, M., Kang, HW. & An, S. A Mathematical Model for Enzyme Clustering in Glucose Metabolism. Sci Rep 8, 2696 (2018). https://doi.org/10.1038/s41598-018-20348-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-20348-7
This article is cited by
-
Expression of fructose-1,6-bisphosphatase 1 is associated with [18F]FDG uptake and prognosis in patients with mesial temporal lobe epilepsy
European Radiology (2023)
-
Temperature effect on water dynamics in tetramer phosphofructokinase matrix and the super-arrhenius respiration rate
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.