Abstract
Disulfides from Allium stipitatum, commonly known as Persian shallot, were previously reported to possess antibacterial properties. Analogues of these compounds, produced by S-methylthiolation of appropriate thiols using S-methyl methanethiosulfonate, exhibited antimicrobial activity, with one compound inhibiting the growth of Mycobacterium tuberculosis at 17 µM (4 mg L−1) and other compounds inhibiting Escherichia coli and multi-drug-resistant (MDR) Staphylococcus aureus at concentrations ranging between 32–138 µM (8–32 mg L−1). These compounds also displayed moderate inhibitory effects on Klebsiella and Proteus species. Whole-cell phenotypic bioassays such as the spot-culture growth inhibition assay (SPOTi), drug efflux inhibition, biofilm inhibition and cytotoxicity assays were used to evaluate these compounds. Of particular note was their ability to inhibit mycobacterial drug efflux and biofilm formation, while maintaining a high selectivity towards M. tuberculosis H37Rv. These results suggest that methyl disulfides are novel scaffolds which could lead to the development of new drugs against tuberculosis (TB).
Similar content being viewed by others
Introduction
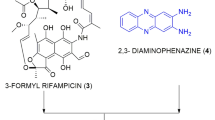
We investigated extracts of bulbs from the plant family Alliaceae for their ability to produce antibacterial compounds, and from Allium neapolitanum, antibacterial canthinone alkaloids and hydroxy acids were characterised1. Of more chemical and pharmacological interest, a study on the Central Asian species Allium stipitatum, led to the isolation of three novel pyridine-N-oxide alkaloids (1–3), displaying outstanding potency towards Mycobacterium tuberculosis (Fig. 1)2. The minimum inhibitory concentrations (MIC) exhibited by these compounds were clinically-relevant and found to range between 2.5–40 µM (0.5–8 mg L−1). Subsequently, a series of structurally-related methyl disulfides were synthesized in an effort to optimize the exceptional antibacterial activity. Structure-activity relationships revealed that the presence of the disulfide moiety was not the only factor responsible for activity, and it is possible that the disulfide is strongly “activated” by the presence of electron-withdrawing functional groups such as pyridine, pyridine-N-oxide, pyrimidine and quinoline, whereas phenyl and thiophene were poorly electron withdrawing and therefore had little effect on the “reactivity” of the disulfide bond (Fig. 1)2. From compounds 4–6, it was clear that the N-oxide was not a prerequisite for antibacterial activity. Based on this rationale, we synthesised a small set of disulphides with proximal electron-withdrawing groups and characterised their antibacterial properties.
Given the continuing issues of multidrug-resistant (MDR) and extensively-drug-resistant (XDR) cases that are increasingly associated with clinically-relevant Gram-positive, Gram-negative and acid-fast human pathogens (such as Staphylococcus aureus, Escherichia coli and Mycobacterium tuberculosis respectively), there is a pressing need to develop new classes of antibacterials3,4,5. Common strategies for effective antimicrobial development are to target novel endogenous effector machinery within a pathogen or to reverse resistance and thereby make the bacteria more susceptible to existing chemotherapy. Increased levels of tolerance towards drugs are observed in bacteria that contain systems to prevent these compounds from reaching their site(s) of action6. Within this paradigm, efflux pump-related multidrug-resistance significantly contributes to a reduction in drug accumulation and often renders antibiotics redundant7. This could be circumvented by molecules that interfere with or inhibit antibiotic efflux8,9. Additionally, multidrug efflux pumps are often transmembrane proteins that secrete metabolites involved in quorum-sensing10. This cross-talk between bacteria is believed to be essential for the formation and dispersion of bacterial biofilms11. Therefore, inhibition of multidrug efflux pumps is also a strategy to inhibit biofilm formation, which is a major contributor to antimicrobial resistance11.
The aim of this study was to synthesise the novel disulphide compounds mentioned earlier and comprehensively evaluate their biological activity to optimise the chemical scaffold as a prospective therapeutic lead.
Results
Synthesis of the antibacterial methyl disulfides
To probe the antibacterial potency, efflux and biofilm inhibitory properties, we chose an initial series of aromatic and heterocyclic thiols on the basis of their commercial availability, namely 4-amino-5-(benzylthio)-4H-1,2,4-triazole-3-thiol (9), 4-aminothieno[2,3-d]pyrimidine-2-thiol (10), 7-fluorobenzo[d]thiazole-2-thiol (11) and 4-ethyl-5-mercapto-4H-1,2,4-triazol-3-ol (12). Each aromatic thiol was treated with S-methyl methanethiosulfonate under alkaline conditions to generate the methyl disulfides, compounds 13–16 (Fig. 1 and Supplementary Information).
Antibacterial Bioassay of the methyl disulfides
The spot culture growth inhibition (SPOTi) assay is a whole-cell phenotypic screen that is routinely used to identify novel antimicrobial molecules with clinical relevance12,13. This rapid but gold-standard assay was applied to evaluate the antimicrobial activity of the synthesized compounds against Gram-positive, Gram-negative and acid-fast bacteria. All of the synthesized methyl disulfides demonstrated antibacterial activity to varying extents (Table 1). Based on the encouraging results when tested against the non-pathogenic model of M. tuberculosis organisms, M. aurum (ATCC23366) and M. bovis BCG (ATCC35734), the compounds were subsequently tested against M. tuberculosis H37Rv and its multidrug-resistant clinical isolates (Mtb-MDR1 and Mtb-MDR2). All four compounds showed anti-mycobacterial activities when tested, with compound 14 having the lowest MIC of 17 µM (4 mg L−1), against the virulent M. tuberculosis H37RV. Additionally, compounds 13–16 exhibited antibacterial activity against the Gram-positive Staphylococcus aureus strains (including effluxing multidrug-resistant strains) and Enterococcus faecalis. In particular, compounds 14 and 16 were active against S. aureus with MIC values ranging between 70–84 µM (16 mg L−1).
Efflux Pump Inhibitory Activity
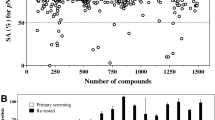
Multi-drug efflux pumps are a key mechanism through which many pathogens, M. tuberculosis in particular, develop intrinsic resistance or tolerance towards xenobiotic compounds14,15. Ethidium bromide (EtBr) is a known substrate for these pumps and its accumulation inside the bacterial cell, when the extrusion mechanism is impaired, can be followed by detecting its fluorescence16. EtBr is usually quenched in an aqueous environment and fluoresces when interacting with the hydrophobic regions within the bacilli17. Verapamil, a calcium channel blocker, is widely used as an inhibitor of efflux in mycobacterial cells and was used as a control in our experiments15. All of the compounds showed inhibition of efflux in the whole-cell model (Fig. 2), with compound 14 and 16 being the most active inhibitors, without affecting the cell viability (a concentration of 25% of the MIC was used for the assay).
Efflux pump inhibition (EPI) of M. aurum under the pressure of methyl disulfides 13–16. Ethidium bromide (EtBr), an efflux pump substrate was used at a final concentration of 1.3 µM (0.5 mg L−1). Its accumulation within the bacterial cells is an indicator of disruption of the efflux mechanism and was detected using fluorescence emissions. Verapamil (VP), a known efflux pump inhibitor, and a drug-free culture were used as positive and negative controls respectively. Low (11–20 rfu) to very high (>50 rfu) inhibition of efflux are represented by the numbers at the side of the graph. The experiments were performed in triplicate (n = 3) and the graph was plotted using the averages. (rfu = relative fluorescence units).
Methyl disulfides as bacterial biofilm inhibitors
As alluded to earlier, efflux mechanisms are involved in quorum-sensing that in turn plays a pivotal role in biofilm formation11. The transcriptional activator LuxR is heavily implicated in quorum sensing and induction of biofilm formation in a variety of bacteria, and is also found in M. tuberculosis and M. leprae18,19. Tubercle bacilli have a natural tendency to form biofilms and other multi-cellular structures, known as cords in liquid culture20. Multi-cellular aggregates resembling biofilms have been detected in the acellular rims of granulomas and necrotic lesions21. Other species belonging to the M. tuberculosis complex (MTBC) such as M. avium are known to form stable biofilms in water reservoirs and can invade lung tissues22. The ability to form cords and biofilms has been correlated with the pathogen’s virulence22. Biofilm-deficient mutants of the pathogen show reduced ability to invade epithelial cells as well as to cause infection in mouse models19.
M. smegmatis, a non-pathogenic model for M. tuberculosis, forms stable biofilms at the liquid-air interface within 5 days and was used to test whether the impairment of drug efflux could also inhibit the formation of biofilms in mycobacteria15. As compound 14 was found to be the most potent anti-mycobacterial (see Table 1), it was selected for the biofilm inhibition studies. Compound 14 was observed to inhibit the growth of M. smegmatis biofilms in a concentration-dependent manner even at sub-MIC levels (Fig. 3a and b) when compared to controls. This finding was further validated through a quantitative crystal violet staining method23. Scanning electron microscopic24 images (Fig. 3c) of M. smegmatis biofilms revealed a dense lattice-like network of bacterial cells with rough outer coats that are likely to be composed of extracellular polymeric substances (EPS) such as lipids, proteins and extra-cellular DNA. On treatment with compound 14, the outer layer of the bacilli became smoother and they appeared to lose the mesh-like inter-cellular connections within the community.
Inhibition of M. smegmatis biofilm formation in the presence of varying concentrations of compound 14. (a) Dose-dependent inhibition of M. smegmatis biofilm formation, as observed by their thinning in the presence of compound 14. Tubes A and B are ‘no drug’ and solvent (0.1% DMSO) controls respectively. Note that the biofilm formation initiated at the air-liquid interface in M. smegmatis. A newly-formed biofilm becomes stacked on top of the old layer and generates a downwards push. Once a critical biomass was exceeded, the lower part of the mature biofilm was observed to dissociate and settle at the bottom of the stand-culture-tube (see controls in which no inhibitor was added). (b) Crystal violet staining of the biofilms showing a decrease in the intensity of the stain with increasing concentrations of compound 14. (c) SEM images of M. smegmatis planktonic, untreated biofilms and biofilms treated with 50 mg L−1 of compound 14.
Selectivity
The synthesized compounds showed a range of eukaryotic toxicity profiles against murine macrophage RAW264.7 cells (Table 2). Compound 14 demonstrated a promising SI of 16.
Discussion
The multidrug-resistant S. aureus SA-1199B (a strain that overexpresses NorA, a multidrug efflux transporter), proved to be as susceptible to the methyl disulfides as other non-NorA S. aureus isolates (Table 1). This indicated that the methyl disulfides may have a mechanism of action that evades NorA-mediated multi-drug efflux.
Interestingly, compound 16 inhibited the Gram-negative bacteria Klebsiella pneumoniae and Proteus mirabilis at an MIC of 335 µM (64 mg L−1) and whilst this is a moderate activity, it is rare to find compounds demonstrating antibacterial activity toward these organisms. Even the standard antibiotic control used for the assay, norfloxacin, could only inhibit the growth of these organisms at a minimum inhibitory concentration higher than 200 µM (64 mg L−1). The synthesized methyl disulfides exhibited appreciable antibacterial activity against E. coli; particularly compound 16 had good antibacterial activity with an MIC value of 84 µM (16 mg L−1).
Overall, the methyl disulfides exhibited inhibitory effects against Gram-positive bacterial strains and acid-fast Mycobacterium species. However, their moderate activity against the selected Gram-negative bacteria provided further incentive to investigate the endogenous mechanism(s) of action of these compounds.
Compounds 14 and 16 exhibited whole-cell drug efflux pump inhibitory activities higher than 13 and 15. Cells treated with the known efflux pump inhibitor verapamil and inhibitor-free cells were used as positive and negative controls in this assay respectively (see Fig. 2).
In terms of the effects of the compounds on Mycobacterium smegmatis biofilm formation, the ability of compound 14 to concentration-dependently inhibit biofilm formation, even at sub-MIC levels is particularly noteworthy.
The efflux pump and biofilm inhibitory effects indicate the possible mechanisms of action of these compounds. This route of antibacterial activity against Pseudomonas aeruginosa was also noted by Jakobsen et al. (2012) for similar compounds25. In addition, allicin, one of the major volatile compounds present in garlic and also a disulfide has been reported to act through permeabilization of cell membranes and inactivation of metabolic enzymes resulting in depletion of intracellular glutathione pools26,27. Our ongoing genomic and transcriptomic analyses of bacterial cells under pressure of inhibitor compounds, as well as spontaneous resistant mutants followed by molecular and biochemical investigations of the relevant genes and their recombinant products should provide a deeper insight into the molecular mechanism(s) of action of these compounds.
For compounds 13, 15 and 16, the bacterial growth inhibition and macrophage cytotoxicity were similar, indicating poor selectivity for their antibacterial action (Table 2). However for compound 14, the SI was 16. The SI provides information on the therapeutic potential of compounds as a function of the concentration range at which they are active against pathogenic mycobacteria while remaining non-toxic to mammalian cells. This provides information on the therapeutic potential of these compound, as a function of the concentration range at which it is active against the growth of pathogenic mycobacteria while remaining non-toxic to mammalian cells (murine macrophages in this case). In conclusion, these synthesized methyl disulfides are new chemical scaffolds that have potential as templates for the discovery of new anti-tubercular leads.
Methods
Materials and synthesis of methyl disulfides
Aromatic thiols 4-amino-5-(benzylthio)-4H-1,2,4-triazole-3-thiol (9), 4-aminothieno[2,3-d]pyrimidine-2-thiol (10), 7-fluorobenzo[d]thiazole-2-thiol (11), 4-ethyl-5-mercapto-4H-1,2,4-triazol-3-ol (12) were purchased from Sigma-Aldrich, Gillingham, U.K. The method of Kitson and Loomes (1985), for the synthesis of methyl 2- and 4-pyridyl disulfide from 2- and 4-thiopyridone and methyl methanethiosulfonate was adapted and modified as follows. The appropriate thiol (2.5 mmol) was dissolved in water (5 mL) containing NaOH (0.10 g, 2.5 mmol, 1 equiv.) and S-methyl methanethiosulfonate (0.315 g, 2.5 mmol, 1 equiv.) added. The solution was stirred for 1 h at room temperature. The cloudy suspension formed was extracted with CH2Cl2 (20 mL). The organic phase was then dried with anhydrous sodium sulfate, filtered, and concentrated under reduced pressure to afford the pure disulfide which was subsequently characterized by spectroscopic techniques – NMR, MS, HRMS, UV and IR (Supplementary Information).
Antibacterial assays (whole-cell phenotypic assays)
Minimum inhibitory concentrations (MIC) of the compounds against Mycobacterium strains were determined using the spot-culture growth inhibition assay (SPOTi)12,13,28,29. The lowest concentration at which mycobacterial growth was completely inhibited by the compound was observed directly. Isoniazid and rifampicin were used as antibiotic controls and the experiments repeated in triplicate.
The antibacterial activity of the compounds was tested against Gram-negative bacteria: Klebsiella pneumoniae, Proteus mirabilis (10830), Escherichia coli (NCTC 10418) and Gram-positive bacteria: Enterococcus faecalis (12697), methicillin-resistant Staphylococcus aureus strains (XU-212 and EMRSA-15) and multidrug-resistant Staphylococcus aureus strain SA-1199B using the microtitre broth dilution assay. Norfloxacin served as a positive control. The assay was performed in 96-well plates and each methyl disulfide was tested in quadruplicate to confirm the reliability and reproducibility of the data. The MIC was determined after the addition of 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) to the 96-well plates. Bacterial growth was indicated by a colour change from yellow to dark blue, which was visually observed. The MIC was recorded as the lowest concentration at which no growth was observed13,30.
Cytotoxicity assay
Eukaryotic cell toxicity assay was carried out using RAW 264.7 macrophage cells, grown in complete RPMI-1640 medium supplemented with 2 mM l-glutamine and 10% heat-inactivated fetal bovine serum and 1% l-glutamine in a 25 cm2 vented, screw-cap cell-culture flask (Flowgen Bioscience Ltd., Hessle, UK) and incubated at 37 °C with a supply of 5% CO2 until confluent growth was observed. Cytotoxicity of the compounds towards the murine macrophages was determined using the resazurin assay29. For quantitative analysis, the fluorescence intensity was measured at λex560/λem590 nm using a FLUOstar OPTIMA micro plate reader. The growth inhibitory concentration (GIC) was reported as the lowest concentration of compound at which no viable eukaryotic cells were detected.
Efflux pump inhibition assay
Efflux pump inhibition assays were performed following previously published protocols and modified using M. aurum cells9,15,31. The effect of the synthesized compounds and verapamil (positive control) on the accumulation of ethidium bromide (EtBr) was determined by measuring fluorescence using a fluorimeter (FLUOstar OPTIMA, BMG Labtech) and fluorescence data was acquired every 60 s for a total period of 60 min. The compounds were used at one quarter of their MICs and EtBr (a known efflux pump substrate) at a concentration of 1.3 µM (0.5 mg L−1).
Biofilm assay (inhibition of biofilm formation)
A late log-phase (OD600 = 3.0) culture of Mycobacterium smegmatis was inoculated into Sauton’s media as 1:100 dilutions. This preparation (2 mL) was transferred into polypropylene tubes and a range of concentrations of compound 14 3–218 µM (0.7–50 mg L−1) was then added to each tube. The cap was tightly closed to avoid evaporation of media and the cultures were incubated at 37 °C in a stationary incubator for 5 days. Tubes containing the diluted cultures without any compounds served as inhibitor-free controls and those with only DMSO served as solvent controls. After 5 days, the biofilm samples were observed using the scanning electron microscope24. The experiments were performed in triplicate.
Once the biofilms were formed, the medium containing planktonic cells was removed carefully using a hypodermic needle. 1% crystal violet was added to the tubes so as to cover the biofilm and was left for 10 min. The crystal violet solution was discarded and the tubes were washed at least three times until no further stain was present in the washings. Ethanol (95% v/v in water) was then added to the tubes and left for 10 min. The solutions were then diluted 1:3 with ethanol and the absorbance of each was measured at 600 nm.
The SEM images were analysed with ImageJ (NIH) software32. Each image was calibrated individually and measurements were recorded for at least 200 cells for each condition from a minimum of five fields with varying magnifications.
References
O’Donnell, G. & Gibbons, S. Antibacterial activity of two canthin-6-one alkaloids from Allium neapolitanum. Phytother. Res. 21, 653–657 (2007).
O’Donnell, G. et al. Bioactive pyridine-N-oxide disulfides from Allium stipitatum. J. Nat. Prod. 72, 360–365 (2009).
Gibbons, S. Phytochemicals for bacterial resistance–strengths, weaknesses and opportunities. Planta Med. 74, 594–602 (2008).
Davies, J. & Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. R. 74, 417–433 (2010).
Fair, R. J. & Tor, Y. Antibiotics and bacterial resistance in the 21st century. Perspect. Medicin. Chem. 6, 25–64 (2014).
Fernandez, L. & Hancock, R. E. Adaptive and mutational resistance: role of porins and efflux pumps in drug resistance. Clin. Microbiol. Rev. 25, 661–681 (2012).
Piddock, L. J. Understanding the basis of antibiotic resistance: a platform for drug discovery. Microbiology 160, 2366–2373 (2014).
Viveiros, M. et al. Inhibitors of mycobacterial efflux pumps as potential boosters for anti-tubercular drugs. Expert Rev. Anti-Infe. 10, 983–998 (2012).
Lechner, D., Gibbons, S. & Bucar, F. Plant phenolic compounds as ethidium bromide efflux inhibitors in Mycobacterium smegmatis. J. Antimicrob. Chemother. 62, 345–348 (2008).
Polkade, A. V., Mantri, S. S., Patwekar, U. J. & Jangid, K. Quorum Sensing: An Under-Explored Phenomenon in the Phylum Actinobacteria. Front. Microbiol. 7, 131 (2016).
Baugh, S., Phillips, C. R., Ekanayaka, A. S., Piddock, L. J. & Webber, M. A. Inhibition of multidrug efflux as a strategy to prevent biofilm formation. J. Antimicrob. Chemother. 69, 673–681 (2014).
Evangelopoulos, D. & Bhakta, S. Rapid methods for testing inhibitors of mycobacterial growth. Methods Mol. Biol. 642, 193–201 (2010).
Guzman, J. D. et al. Antitubercular specific activity of ibuprofen and the other 2-arylpropanoic acids using the HT-SPOTi whole-cell phenotypic assay. BMJ Open 3, https://doi.org/10.1136/bmjopen-2013-002672 (2013).
Blair, J. M., Webber, M. A., Baylay, A. J., Ogbolu, D. O. & Piddock, L. J. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51 (2015).
Rodrigues, L. et al. Thioridazine and chlorpromazine inhibition of ethidium bromide efflux in Mycobacterium avium and Mycobacterium smegmatis. J. Antimicrob. Chemother. 61, 1076–1082 (2008).
Paixao, L. et al. Fluorometric determination of ethidium bromide efflux kinetics in Escherichia coli. J. Biol. Eng. 3, 18 (2009).
Machado, D. et al. Ion Channel Blockers as Antimicrobial Agents, Efflux Inhibitors, and Enhancers of Macrophage Killing Activity against Drug Resistant Mycobacterium tuberculosis. PloS One 11, e0149326, https://doi.org/10.1371/journal.pone.0149326 (2016).
Santos, C. L., Correia-Neves, M., Moradas-Ferreira, P. & Mendes, M. V. A walk into the LuxR regulators of Actinobacteria: phylogenomic distribution and functional diversity. PloS One 7, e46758, https://doi.org/10.1371/journal.pone.0046758 (2012).
Yamazaki, Y. et al. The ability to form biofilm influences Mycobacterium avium invasion and translocation of bronchial epithelial cells. Cell. Microbiol. 8, 806–814 (2006).
Middlebrook, G., Dubos, R. J. & Pierce, C. Virulence and Morphological Characteristics of Mammalian Tubercle Bacilli. J. Exp. Med. 86, 175–184 (1947).
Orme, I. M. A new unifying theory of the pathogenesis of tuberculosis. Tuberculosis 94, 8–14 (2014).
Vaerewijck, M. J., Huys, G., Palomino, J. C., Swings, J. & Portaels, F. Mycobacteria in drinking water distribution systems: ecology and significance for human health. FEMS Microbiol. Rev. 29, 911–934 (2005).
O’Toole, G. A. Microtiter dish biofilm formation assay. J. Vis. Exp. https://doi.org/10.3791/2437 (2011).
Kulka, K., Hatfull, G. & Ojha, A. K. Growth of Mycobacterium tuberculosis biofilms. J. Vis. Exp. https://doi.org/10.3791/3820 (2012).
Jakobsen, T. H. et al. Ajoene, a sulfur-rich molecule from garlic, inhibits genes controlled by quorum sensing. Antimicrob. Agents Chemother. 56, 2314–2325 (2012).
Gruhlke, M. C. et al. The defense substance allicin from garlic permeabilizes membranes of Beta vulgaris, Rhoeo discolor, Chara corallina and artificial lipid bilayers. Biochim. Biophys. Acta 1850, 602–611 (2015).
Müller, A. et al. Allicin induces thiol stress in bacteria through S-Allylmercapto modification of protein cysteines. J. Biol. Chem. 291, 11477–11490 (2016).
Danquah, C. A., Maitra, A., Gibbons, S., Faull, J. & Bhakta, S. HT-SPOTi: A Rapid Drug Susceptibility Test (DST) to Evaluate Antibiotic Resistance Profiles and Novel Chemicals for Anti-Infective Drug Discovery. Curr. Protoc. Microbiol. 40, 17.8.1–17.8.12, https://doi.org/10.1002/9780471729259.mc1708s40 (2016).
Guzman, J. D. et al. Anti-tubercular screening of natural products from Colombian plants: 3-methoxynordomesticine, an inhibitor of MurE ligase of Mycobacterium tuberculosis. J. Antimicrob. Chemother. 65, 2101–2107 (2010).
Gibbons, S. & Udo, E. E. The effect of reserpine, a modulator of multidrug efflux pumps, on the in vitro activity of tetracycline against clinical isolates of methicillin resistant Staphylococcus aureus (MRSA) possessing the tet(K) determinant. Phytother. Res. 14, 139–140 (2000).
Groblacher, B., Maier, V., Kunert, O. & Bucar, F. Putative mycobacterial efflux inhibitors from the seeds of Aframomum melegueta. J. Nat. Prod. 75, 1393–1399 (2012).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Acknowledgements
C.A.D. and A.M. thank the Ghanaian Education Trust and Wellcome Trust/Birkbeck for supporting their doctoral research studies respectively.
Author information
Authors and Affiliations
Contributions
S.G., S.B., J.M., P.S. and T.D.M. designed the study. C.A.D., E.K., P.K., A.M., M.R. and D.E. collected and analyzed the data. All authors wrote the manuscript text and S.G., J.M., S.B. and A.M. prepared the figures. All authors reviewed and critically revised the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Danquah, C.A., Kakagianni, E., Khondkar, P. et al. Analogues of Disulfides from Allium stipitatum Demonstrate Potent Anti-tubercular Activities through Drug Efflux Pump and Biofilm Inhibition. Sci Rep 8, 1150 (2018). https://doi.org/10.1038/s41598-017-18948-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-18948-w
This article is cited by
-
In Silico Approach for Phytocompound-Based Drug Designing to Fight Efflux Pump-Mediated Multidrug-Resistant Mycobacterium tuberculosis
Applied Biochemistry and Biotechnology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.