Abstract
Knowledge about genetic drivers of disease increases the efficiency of interpreting patient DNA sequence and helps to identify and prioritize biological points of intervention. Discoveries of genes with single mutations exerting substantial phenotypic impact reliably provide new biological insight, although such approaches tend to generate knowledge that is disjointed from the complexity of biological systems governed by elaborate networks. Here we sought to facilitate diagnostic sequencing for hair disorders and assess the underlying biology by compiling an archive of 684 genes discovered in studies of monogenic disorders and identifying molecular annotations enriched by them. To demonstrate utility for this dataset, we performed two data driven analyses. First, we extracted and analyzed data implicating enriched signaling pathways and identified previously unrecognized contributions from Hippo signaling. Second, we performed hierarchical clustering on the entire dataset to investigate the underlying causal structure of hair disorders. We identified 35 gene clusters representing genetically derived biological modules that provide a foundation for the development of a new disease taxonomy grounded in biology, rather than clinical presentations alone. This Resource will be useful for diagnostic sequencing in patients with diseases affecting the hair follicle, improved characterization of hair follicle biology, and methods development in precision medicine.
Similar content being viewed by others
Introduction
In an age of precision medicine, faced with interpreting DNA sequence in the genomes of patients, it becomes critical to understand both the spectrum of genes that could be contributing to a particular clinical presentation, and the pathways that are mediating genetic effects. An archive of disease genes facilitates diagnostic sequencing1. Rigorous analysis of the functional relationships across a set of genes linked to a particular disease state has the potential to provide robust molecular characterization of both disease pathogenesis and human physiology, and could help illuminate a causal structure that underpins health and tissue homeostasis. Such work can have a profound impact on patient care by prioritizing pathways to therapeutically target, guiding drug development, suggesting drug repurposing opportunities, and improving the efficiency of clinical trials2. Additionally, efforts to functionally organize disease genes would provide a foundation for the development of a new disease taxonomy that is grounded in biology, rather than clinical observations of symptoms alone. The need to develop an improved disease taxonomy by incorporating mechanistic information from molecular data has been identified as a critical challenge in the advancement of precision medicine3. However, such efforts have yet to be rigorously pursued in clinical areas outside of oncology.
Our knowledge of genes that influence human health and disease is largely derived from two complementary gene mapping approaches. Linkage studies and exome sequencing in families segregating rare Mendelian (i.e. monogenic) diseases have identified mutations that are rare in the population and exert strong biological effects that tend to be easy to interpret, thereby facilitating identification of a definitive causal gene and providing insight into disease mechanism. On the other hand, genome-wide association studies (GWAS) are performed in large cohorts of unrelated patients and controls and identify genetic variants with greater population frequencies. Variants identified through GWAS tend to be intergenic and have obscure biological effects, thereby hampering the definitive identification of individual causal genes. Therefore, Mendelian disease genes offer a clear advantage over GWAS loci in gaining biological insight.
Although Mendelian diseases are infrequent within the population, evidence continues to emerge from human genetic studies that etiological information derived from rare diseases caused by single mutations is sometimes generalizable to diseases that are common in the population and have a polygenic architecture1,4,5,6,7. Conceptually, there are a finite number of physiological processes that can drive a particular disease manifestation. If we consider a single biological pathway that contributes to homeostasis in a particular tissue (or set of tissues), there may be genes for which a single mutation exerts an extreme effect, or genes for which an accumulation of variants shifts the tissues towards a disease state. This suggests that diseases across the full spectrum of etiological heterogeneity and population prevalence could share an underlying causal structure. Examples in which gene identification in Mendelian diseases have led to new therapeutic approaches to common diseases provide the most direct evidence for a shared underlying biological architecture1. In further support of this theory, there is evidence that Mendelian disease genes make direct contributions to common disorders, for example when GWAS identify loci that harbor genes that cause Mendelian disorders1,5,6. Finally, it has been shown that deleterious variants in Mendelian loci can contribute non-additively to the risk of developing certain complex diseases affecting similar systems5. Therefore, we propose that constructing a molecular taxonomy from genes implicated in rare disorders could provide valuable insight into the underlying causal structure of common disorders that have clinical presentations overlapping partially with more extreme Mendelian phenotypes.
Dermatological disorders provide salient opportunities for developing methods in precision medicine. Direct visual assessment of diagnostic cues and histological findings allows for a relatively high precision in diagnoses and nuanced phenotypic subtyping. Hair disorders in particular represent a unique opportunity to develop disease taxonomies from genetic data, as gene mapping in humans and animal models has identified hundreds of genes that affect multiple aspects of hair follicle biology, including hair follicle size, density and cycling, as well as hair fiber length, shape, texture and pigmentation. Despite the tremendous amount of data generated from genetic studies of hair, and from molecular and functional studies of genes, there has yet to be a large-scale analysis to integrate all of the available information and generate new biological knowledge about genetic modulators of the hair follicle.
Here, we have curated a database of genes for which a single mutation influences hair follicle phenotypes. We identified 684 genes from publicly available resources and from literature describing single gene hair disorders in humans and mammalian models. We annotated these genes across multiple molecular and functional domains and identified 4,937 terms significantly enriched by these genes. In order to demonstrate utility for such a data set, we performed two sets of analyses. First, we extracted data pertaining to cellular signaling pathways to construct and analyze a hair follicle signaling network. Second, we performed hierarchical clustering analysis and natural language processing (NLP) to identify functional clusters of genes and describe relationships within and among these sets of genes. This work provides a valuable resource for advancing the implementation of precision medicine and may be used for diagnostic sequencing, genetic characterization of the hair follicle at an unprecedented scale, and methods development in disease taxonomy.
Results
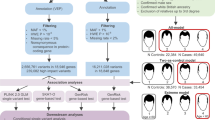
We identified 684 protein-coding genes that influence the integrity of the hair follicle via an inherited genetic mutation in human patients and/or mammalian models and could be mapped to unique Human Genome Organization (HUGO) gene nomenclature committee (HGNC) approved gene symbols (Supplementary Table 1). We characterized the biology implicated by these genes by performing annotation enrichment analysis, which identified 4,937 significantly enriched annotations (Supplementary Table 2), including terms descriptive of gene ontology, biological pathways, protein domains and interactions, gene expression patterns in tissues, and disease connections (Supplementary Table 3).
Several biological themes emerge from a review of the significantly enriched annotations. For example, 962 gene ontology (GO) terms are significantly enriched by 678 genes; 91 of these genes are involved in various cellular metabolic processes, including glucose and lipid metabolism; 222 influence development of organs and tissues outside of the integumentary system including heart, kidney, brain and other tissues of the nervous system, and digestive system including pancreas. Pathway analysis identified 300 pathways significantly enriched by 384 of the 684 genes, including 153 genes that enrich one or more cancer pathways, including not only melanoma and basal cell carcinoma, but also brain, pancreatic, thyroid, lung, endometrial, and colorectal cancers, among others; 134 genes enrich pathways that are annotated to be implicated in response to a viral or bacterial pathogen.
There are 57 cellular signaling pathways significantly enriched by 220 genes, including Wnt, Hippo, TGFβ, Hedgehog, Notch, PI3K-Akt, MAPK, ErbB, Ras, and JAK-STAT pathways, among others (Supplementary Table 4). An analysis of gene distributions across these signaling pathways reveals a complex network in which all pathways are linked by various subsets of genes (Fig. 1). Genes display differing levels of connectivity within this signaling network, participating in as few as one pathway (n = 70) and as many as 40 pathways (MAP2K1). The most highly connected genes in the network (n = 11), participating in 19 or more pathways each, connect 49 of the 57 pathways. Of the remaining eight pathways, which do not contain any of these highly connected genes, seven are connected by a set of 59 genes linking the Hippo pathway to Hedgehog, Wnt, Notch, and p53 signaling pathways (Fig. 1, red edges). This subnetwork is additionally identified by gene community detection (Fig. 1, red gene nodes), which identified a total of four gene groupings (modularity = 0.2346) on the basis of pathway membership (Supplementary Table 4).
Hair follicle signaling network revealed by genes underlying monogenic disorders. Annotations significantly enriched by the 684 genes we identified include 57 cellular signaling pathways (diamond nodes) that are connected by a network of 220 genes (rectangular nodes). Edges represent gene-pathway memberships. The most highly connected genes (black outlines) connect 49 pathways (black outlines). Of the eight pathways that do not contain any of the highly connected genes (red outlines), seven are connected by a set of 59 genes (indicated by red edges). This subnetwork was also identified by the Louvain method for gene community detection (red nodes) as one of four gene communities, and includes all 29 genes of the Hippo pathway. The other three gene communities are color-coded, indicating a consistency of results across both analytic methods.
We next performed a hierarchical clustering analysis to characterize the functional relationships among these 684 genes that are captured by significantly enriched annotations. We utilized an unsupervised agglomerative clustering algorithm to generate a dendrogram that can be used to estimate relative pairwise molecular and functional similarity between any two genes by tracing branches between them (Fig. 2; Supplementary Table 5). For example, a longer distance along branches between two genes indicates fewer similarities, and close neighbors are expected to have more annotations in common.
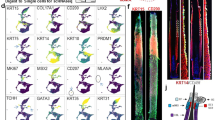
Molecular taxonomy of hair disorder genes revealed by functional hierarchical clustering analysis of 684 genes and 4,937 annotations. Unsupervised agglomerative hierarchical clustering was performed to group 684 genes based on the degree of similarity among their functional annotations. Color-coding distinguishes 35 clusters created by using an arbitrary threshold of height (h) = 1.15, indicated by a black horizontal line. Genes with similar functional annotations are grouped within the same or neighboring clusters. We propose that each cluster represents a biological module, a set of genes that converge on a shared biological feature whose diagnostic and clinical utility remain to be established.
Finally, we sought to investigate the causal structure of hair disorders under the hypothesis that this comprehensive set of genes would converge upon a discrete number of biological processes that are critical for governing hair follicle biology. The dendrogram allows us to optimize the number of gene clusters by varying an arbitrary height threshold. Through an iterative process, we found that setting the height to h = 1.15 generated 35 semantically meaningful gene clusters. We validated this threshold with two sequential procedures. First, principal component analysis (PCA) was performed and the number of components was set to 100, which was indicated to be reasonable on the basis of variance analysis. Second, subsequent t-SNE visualization of the PCA output with 35 clusters labeled was performed, revealing adequate boundaries among clusters, reproduced at various levels of perplexity. NLP identified defining features of each cluster (Supplementary Table 6), which were used to develop semantic descriptions based on functional annotation content for each of the clusters (Table 1).
Discussion
Perturbations in hair follicle biology are manifested in multiple ways, for example interrupting a developmental process leading to hair loss, or affecting the integrity of the hair fiber leading to a change in length, texture or pigmentation. Since one of our goals was to construct a resource that could be used to provide a comprehensive assessment of the biology of this organ, we set out to identify monogenic hair genes regardless of their specific phenotypic consequences. We identified 684 protein-coding genes that alter the hair follicle in human patients and/or mammalian models, representing the most comprehensive archive of hair follicle genes identified in genetic mapping experiments to date (Supplementary Table 1). This resource will be useful for constructing filtering algorithms for human genome sequence data generated for diagnostic or investigative purposes. While this set of genes is larger than has previously been reported in reviews of hair follicle genetics in humans or mouse models (e.g.8,9), as a reference point, gene expression experiments in human and murine models report differential expression of thousands of genes in the hair follicle (e.g.10,11). In order to be comprehensive, we included genes identified in either human or animal studies. We included genes that have only been characterized in animal models because of the possibility that they do contribute to human traits, but have not yet been identified in patients. For example, animal studies had identified fibroblast growth factor 5 (FGF5) as a crucial regulator of hair growth two decades before it was found to underlie a human condition12,13. If this gene list is incorporated into algorithms designed for filtering human DNA sequence data, we recommend including animal model genes and down-weighting evidence scores.
The genes we identified significantly enrich a set of 4,937 annotations, which provide insight into genetic regulators of hair follicle biology and relationships among disease genes (Supplementary Table 2). For example, the metabolic demands of the hair follicle dramatically increase during the growth stage of the hair cycle (anagen) to support the extensive cell proliferation and differentiation that occurs as the organ regenerates, transitioning from a quiescent state. It has been shown previously that glucose is a substantial source of energy in the growing hair follicle, which utilizes aerobic glycolysis14,15. We identified 40 genes annotated by glucose regulation. Likewise, lipid homeostasis is known to be important for maintaining a healthy hair follicle through the identification of several genes in various hair loss disorders16. Our analysis identified a total of 36 genes annotated to be involved with lipid metabolism. Other metabolic pathways implicated in hair follicle biology by our analysis include proteoglycan metabolism (four genes), cellular amino acid metabolism (six genes), and vitamin metabolism (three genes).
Our analysis identified multiple significantly enriched cancer pathways. Limitations of our analytic approach prevent us from inferring a relationship between hair disorders and cancers. Rather, the presence of cancer annotations could simply reflect the critical role that regulation of cell proliferation plays in hair follicle biology, given that this organ undergoes cycling between regeneration and regression throughout the lifespan. Likewise, annotations that implicate tissues outside of the integumentary system may be capturing shared biology or developmental lineage, and/or could be indicative of multisystem disease, but further studies are needed to understand these relationships as well.
While roles for several cellular signaling pathways in hair biology have been previously established, our work provides a comprehensive overview of how these pathways are genetically linked in the hair follicle, revealing important inflection points that await further investigation, including the identification of a new pathway for prioritization in studies focused on modulating hair growth. Previously, the WNT pathway has been shown to be a critical regulator of hair follicle development and cycling, and roles for TGFβ, Notch, Hedgehog and JAK-STAT signaling have also been established17,18. Our network analysis not only demonstrates contributions from these pathways in causing genetic perturbations in the hair follicle, but also implicates Hippo signaling through the identification of 29 genes annotated to participate in this pathway (Fig. 1; Supplementary Table 4). While Hippo signaling has been previously studied in the skin, specifically in mouse epidermal development and human cutaneous squamous cell carcinoma19,20, the pathway has yet to be investigated as a potential modulator of the hair follicle.
Hippo signaling has been extensively studied in the contexts of cancer and development, and has been shown to influence tumor or organ size through the regulation of cell proliferation and apoptosis21,22. In fact, the pathway was originally given its name because genetic perturbations thereof generated “hippopotamus-sized” organs22. Interestingly, the most common form of hair loss, androgenetic alopecia (i.e. male pattern baldness; MPB), has long been characterized as a process of organ miniaturization, whereby hair follicles continue to cycle but undergo a reduction in size, resulting in a transition from thick terminal hair to fine vellus hair23. While Hippo signaling has yet to be specifically implicated in MPB, there is preliminary genetic evidence that is consistent with such a hypothesis. The largest MPB GWAS performed to date included a gene-based analysis that identified 112 autosomal genes with genome-wide significant association (Bonferroni correction of α < 2.769e-06)24, four of which are annotated to participate in Hippo signaling within the Kyoto Encyclopedia of Genes and Genomes (KEGG; pathway hsa04390), including WNT6, WNT10A, WNT3, and CTNNB1. We performed pathway enrichment analysis of these 112 genes and identified hsa04390:Hippo signaling pathway at a significance level of p = 0.049. Three of these genes reside at loci that were also associated with MPB in an independent GWAS (WNT6, WNT10A, WNT3)25. Our analysis of genes that establish a hair follicle cellular signaling network identified the Hippo pathway and a set of 59 genes that link this pathway to Wnt, Notch, Hedgehog and p53 signaling pathways (Fig. 1). A definitive role for Hippo signaling in the pathogenesis of MPB awaits further investigation.
In order to characterize relationships among the 684 genes that influence hair follicle biology through single mutations, we used the set of 4,937 significantly enriched annotations to perform hierarchical clustering. We identified an organizational scheme derived from functional and molecular data, and thus rooted in biology (Fig. 2; Supplementary Table 5). As a preliminary strategy to understand the biological structure suggested by this clustering, we defined 35 gene clusters by optimizing a height threshold (h = 1.15) and using NLP to identify biological themes within clusters (Table 1). We propose that each cluster represents a biological module, a set of genes that converge on a shared biological feature whose diagnostic and clinical utility remain to be established. We found, for example, that Cluster 7 represents a set of genes annotated by terms related to autoimmune disease and pathogen response, and contains a number of genes that mediate tissue interactions with the immune system, including INFG, IL2, IL2RB, and FAS. This supports recent work that has implicated the immune system in hair follicle development26, and suggests further investigation into roles that the immune system may play in hair follicle cycling and homeostasis is warranted.
While our work provides a framework for understanding the biology that influences hair follicle disease, future work linking the biological modules that we identified to disease phenotypes will help to better understand the complex relationship between molecular functions of genes and the disease that arise from mutations in them. There is preliminary evidence that our gene clustering may have diagnostic relevance. For example, cluster 18 is enriched with annotations such as “cell-cell adherens junction” and contains genes that code for components of cellular anchoring junctions. Disruptions in these proteins produce multi-system clinical manifestations that include hypotrichosis and/or woolly hair27. Mutations in Plakophilin 1 (PKP1) cause an inherited disease impacting ectodermal structures, and patients display hypotrichosis, nail dystrophy, and skin fragility9. Mutations in junctional plakoglobin (JUP) and desmoplakin (DSP) cause Naxos disease and Carvajal syndrome respectively, two cardiocutaneous syndromes that include symptoms of woolly hair, cardiomyopathy, and palmoplantar keratoderma9. Our analysis places these three genes adjacent to each other in cluster 18, seemingly capturing biological similarities among disease entities with partially overlapping phenotypes.
Alternatively, some clustering results suggest that there may be degenerate mapping between clinical symptoms and molecular or functional characterization of disease genes. For example, uncombable hair syndrome is a nonsyndromic hair disorder with three recently identified causative genes: tricohyalin (TCHH), transglutaminase 3 (TGM3), and peptidylarginine deiminase 3 (PADI3)28. Our hierarchical clustering analysis placed TGM3 and TCHH in cluster 22, whereas PADI3 is in cluster 31. An analysis of annotations that are significantly enriched by these three genes suggests that it is the distribution of transcription factor binding sites that is driving this distinction (Supplementary Table 7). Interestingly, these genes show different patterns of gene expression in The Genotype-Tissue Expression (GTEx) database29, which suggests differences in regulatory elements. While further investigation is required to determine if these results have clinical relevance, this example does suggest possible biological distinctions between diseases that are traditionally grouped together as a single entity on the basis of symptoms, providing motivation for further development of a disease taxonomy that incorporates data from molecular biology experiments. Future work should focus on integrating and evaluating clinical manifestations with molecular annotations.
The goal of clustering methods is to find structure within data, grouping similar elements within the same cluster and dissimilar elements in different clusters. For example, our clustering model separates genes encoding type II (basic) keratins (cluster 13) and type I (acidic) keratins (cluster 14) into adjacent clusters, capturing both their similarities and differences by their relative dendrogram positions. However, as with any unsupervised machine learning method, analytic outcomes may be influenced by the data available for input and the choice of algorithms used to determine similarities among elements (in this case, genes). For example, we annotated genes with functional and molecular data that is currently available in the public domain and integrated into pathway analysis software30. Experimental data that continues to accumulate over time could influence clustering results. Furthermore, there are a number of algorithms available to uncover structure in data. We applied unsupervised agglomerative hierarchical clustering, which required us to empirically determine an optimal number of clusters within the data set. We used an iterative process evaluating and integrating NLP results to partition our dendrogram into 35 clusters, obtained by setting a height threshold of h = 1.15. We believe this to be an optimal operationalization of causal structure because it generated semantically meaningful clusters with NLP. Additionally, while the dimensionality reduction method that we employ has been widely adopted for deriving meaning from high-dimensional data31, interpretation of results has some inherent challenges. For example, the algorithm adapts to the underlying data, performing different translations on different regions of data, which may present a source of confusion in visual interpretation32. The analysis that we report here is presented as an example of the diverse analytic approaches that could be applied to this Resource in future investigations.
Our work in identifying and functionally annotating a comprehensive set of genes that underlie hair disorders provides a valuable resource for both research and clinical communities embarking on precision medicine initiatives for skin and hair disorders, and could be useful for methods development more broadly relevant to the implementation of precision medicine across other clinical areas. Understanding disease causation in patients and devising efficient therapeutic strategies requires knowledge not only of the genes implicated in disease, but also of their interactions through biological pathways, which may reveal a higher order causal structure of disease4. This archive provides a tool for pinpointing loci harboring critical mutations that underlie diseases with clinical manifestations in the hair follicle, and for surveying pathways and biological processes that modulate the hair follicle. We have utilized analytic approaches drawn from the field of machine learning in an initial attempt to functionally link genes based on biological knowledge that is currently available in the public domain, uncovering insight into physiology that is critical to hair biology and disease. Our work invites prioritization of the Hippo signaling pathway in future studies of molecular modulation of hair growth and has identified higher order biological structure among these 684 genes. This work creates an opportunity for future methods development in precision medicine.
Methods
Identification of genes
We compiled a genotype-phenotype database incorporating genes from two publicly available data sources, Online Mendelian Inheritance in Man (OMIM) catalog and Jackson Laboratories Mouse Genome Informatics (MGI) database, as well as human and mammalian model studies from the literature (Supplementary Table 1).
A preliminary list of human genes influencing hair phenotype was created using a series of phenotype searches within OMIM. We defined the following 7 categories to characterize hair phenotype: alopecia, hair cycling, hypertrichosis, hair morphogenesis, hair pigmentation, hair structure, and secondary effects on hair. “Secondary effects” refers to alterations in hair phenotype secondary to a primary alteration in metabolic phenotype. We reduced the risk of false-negative search results by using multiple synonymous descriptors as search terms in OMIM (Supplementary Table 8). We mitigated the risk of false-positive search results by excluding genes that were annotated in OMIM to be without a known gene sequence, and/or with a provisional relationship with the disease, and/or without a gene map locus. Corresponding search terms were used to identify mouse genes with human orthologs linked to hair phenotypes within the MGI database. A list of additional genes known to influence hair phenotype in humans and other mammalian models, including mouse, rat, dog, and horse, was compiled from reports in the literature28,33,34,35,36,37,38,39. We next excluded genes that are not protein-coding and/or do not have human orthologs, removing pseudogenes, heritable phenotypic markers, quantitative trait loci, chromosomal inversions, transgenic mutations that implicated multiple genes, and polygenic mutations. Gene symbols were standardized to HGNC-approved gene symbols for subsequent annotation and analysis.
Gene annotation
The list of official gene symbols was uploaded to the functional annotation tool on the Database for Annotation, Visualization and Integrated Discovery (DAVID) v.6.8. Species and background were set to “Homo sapiens.” Queried categories of annotations are listed in Supplementary Table 3. Functional annotations for which p < 0.05 were downloaded to an Excel database (Excel 2016, Microsoft Corp, Seattle, WA). Significantly enriched annotations that were molecularly uninformative were removed (“disease mutation”, “polymorphism”, and “sequence variant”). To prepare the data for functional hierarchical clustering, a binary matrix of annotations for the set of genes was created in Excel.
Signaling network construction
Significantly enriched pathways that contain the term “signaling” were extracted from the gene annotation database (Supplementary Table 2). Pathway names and gene names were imported to Cytoscape v.3.4.0 (Supplementary Table 4) to construct a network with the edge-weighted spring embedded layout40. Highly connected genes are defined as being within the 95th percentile of pathway connections, which was empirically determined to be participating in more than 16 pathways (Supplementary Figure 1).
Gene community detection
To identify communities of genes with similar pathway membership we used the Louvain method as implemented in R 3.3.3 using the igraph package, first constructing an adjacency matrix from Supplementary Table 4 in R41.
Hierarchical clustering
Subsequent analyses were performed using Python in Jupyter Notebook, using the numpy, sklearn, pandas, and SciPy packages, as well as matplotlib and Seaborn visualization libraries. Dimensionality reduction was performed using the principal component analysis function from sklearn with the number of components set to 100, followed by visualization using t-distributed stochastic neighbor embedding (t-SNE) via the TSNE function from sklearn, with perplexity values in the range (5–250). A pairwise distance matrix was created using the pdist function in the scipy.spatial.distance module in Python, applying the Jaccard metric. Unsupervised agglomerative hierarchical clustering was performed with the linkage function in the scipy.cluster.hierarchy module, with method set to Ward (i.e. Ward variance minimization). The output was plotted with the Seaborn visualization library in Python. Several iterations of NLP (see below) were performed at various height thresholds partitioning the dendrogram into different numbers of clusters. An arbitrary height threshold of 1.15 was set to partition the dendrogram into a set of 35 clusters, which was found to yield semantically meaningful results from NLP.
Natural language processing
NLP using pandas and numpy packages and matplotlib plotting library in Python identified the most frequent annotations associated with each cluster. All significantly enriched annotations that appeared in fewer than 60% of the clusters (n ≤ 20) were used for the analysis. Weights were assigned based on the relative frequency of a given annotation across clusters, to preferentially down-weight common annotations (Supplementary Table 5). The relative weights were defined as the count of a given annotation in a given cluster, multiplied by the natural log of the inverse quotient of the count of that annotation across all clusters divided by the total count of all annotations across all clusters. This allowed for the rational development of semantic descriptions of clusters, derived from frequent annotations associated with a given cluster (Table 1).
References
Chong, J. X. et al. The Genetic Basis of Mendelian Phenotypes: Discoveries, Challenges, and Opportunities. American journal of human genetics 97, 199–215, https://doi.org/10.1016/j.ajhg.2015.06.009 (2015).
Brooks, P. J., Tagle, D. A. & Groft, S. Expanding rare disease drug trials based on shared molecular etiology. Nat Biotechnol 32, 515–518, https://doi.org/10.1038/nbt.2924 (2014).
In Toward Precision Medicine: Building a Knowledge Network for Biomedical Research and a New Taxonomy of Disease The National Academies Collection: Reports funded by National Institutes of Health (2011).
Bauer-Mehren, A. et al. Gene-disease network analysis reveals functional modules in mendelian, complex and environmental diseases. PloS one 6, e20284, https://doi.org/10.1371/journal.pone.0020284 (2011).
Blair, D. R. et al. A nondegenerate code of deleterious variants in Mendelian loci contributes to complex disease risk. Cell 155, 70–80, https://doi.org/10.1016/j.cell.2013.08.030 (2013).
Lupski, J. R., Belmont, J. W., Boerwinkle, E. & Gibbs, R. A. Clan genomics and the complex architecture of human disease. Cell 147, 32–43, https://doi.org/10.1016/j.cell.2011.09.008 (2011).
Antonarakis, S. E. & Beckmann, J. S. Mendelian disorders deserve more attention. Nat Rev Genet 7, 277–282, https://doi.org/10.1038/nrg1826 (2006).
Nakamura, M., Schneider, M. R., Schmidt-Ullrich, R. & Paus, R. Mutant laboratory mice with abnormalities in hair follicle morphogenesis, cycling, and/or structure: an update. Journal of dermatological science 69, 6–29, https://doi.org/10.1016/j.jdermsci.2012.10.001 (2013).
Shimomura, Y. Journey toward unraveling the molecular basis of hereditary hair disorders. Journal of dermatological science 84, 232–238, https://doi.org/10.1016/j.jdermsci.2016.08.006 (2016).
Chew, E. G. et al. Differential Expression between Human Dermal Papilla Cells from Balding and Non-Balding Scalps Reveals New Candidate Genes for Androgenetic Alopecia. The Journal of investigative dermatology 136, 1559–1567, https://doi.org/10.1016/j.jid.2016.03.032 (2016).
Rezza, A. et al. Signaling Networks among Stem Cell Precursors, Transit-Amplifying Progenitors, and their Niche in Developing Hair Follicles. Cell Rep 14, 3001–3018, https://doi.org/10.1016/j.celrep.2016.02.078 (2016).
Hebert, J. M., Rosenquist, T., Gotz, J. & Martin, G. R. FGF5 as a regulator of the hair growth cycle: evidence from targeted and spontaneous mutations. Cell 78, 1017–1025 (1994).
Higgins, C. A. et al. FGF5 is a crucial regulator of hair length in humans. Proceedings of the National Academy of Sciences of the United States of America 111, 10648–10653, https://doi.org/10.1073/pnas.1402862111 (2014).
Philpott, M. P. & Kealey, T. Metabolic studies on isolated hair follicles: hair follicles engage in aerobic glycolysis and do not demonstrate the glucose fatty acid cycle. The Journal of investigative dermatology 96, 875–879 (1991).
Adachi, K. & Uno, H. Glucose metabolism of growing and resting human hair follicles. Am J Physiol 215, 1234–1239 (1968).
Stenn, K. S. & Karnik, P. Lipids to the top of hair biology. The Journal of investigative dermatology 130, 1205–1207, https://doi.org/10.1038/jid.2010.52 (2010).
Harel, S. et al. Pharmacologic inhibition of JAK-STAT signaling promotes hair growth. Sci Adv 1, e1500973, https://doi.org/10.1126/sciadv.1500973 (2015).
Paus, R. & Cotsarelis, G. The biology of hair follicles. The New England journal of medicine 341, 491–497, https://doi.org/10.1056/NEJM199908123410706 (1999).
Walko, G. et al. A genome-wide screen identifies YAP/WBP2 interplay conferring growth advantage on human epidermal stem cells. Nat Commun 8, 14744, https://doi.org/10.1038/ncomms14744 (2017).
Zhang, H., Pasolli, H. A. & Fuchs, E. Yes-associated protein (YAP) transcriptional coactivator functions in balancing growth and differentiation in skin. Proceedings of the National Academy of Sciences of the United States of America 108, 2270–2275, https://doi.org/10.1073/pnas.1019603108 (2011).
Attisano, L. & Wrana, J. L. Signal integration in TGF-beta, WNT, and Hippo pathways. F1000Prime Rep 5, 17, https://doi.org/10.12703/P5-17 (2013).
Yu, F. X., Zhao, B. & Guan, K. L. Hippo Pathway in Organ Size Control, Tissue Homeostasis, and Cancer. Cell 163, 811–828, https://doi.org/10.1016/j.cell.2015.10.044 (2015).
Whiting, D. A. Possible mechanisms of miniaturization during androgenetic alopecia or pattern hair loss. Journal of the American Academy of Dermatology 45, S81–86 (2001).
Hagenaars, S. P. et al. Genetic prediction of male pattern baldness. Plos Genet 13, e1006594, https://doi.org/10.1371/journal.pgen.1006594 (2017).
Heilmann-Heimbach, S. et al. Meta-analysis identifies novel risk loci and yields systematic insights into the biology of male-pattern baldness. Nat Commun 8, 14694, https://doi.org/10.1038/ncomms14694 (2017).
Ali, N. et al. Regulatory T Cells in Skin Facilitate Epithelial Stem Cell Differentiation. Cell 169, 1119–1129 e1111, https://doi.org/10.1016/j.cell.2017.05.002 (2017).
Porter, P. S. The genetics of human hair growth. Birth defects original article series 7, 69–85 (1971).
FB, U. B. et al. Mutations in Three Genes Encoding Proteins Involved in Hair Shaft Formation Cause Uncombable Hair Syndrome. American journal of human genetics 99, 1292–1304, https://doi.org/10.1016/j.ajhg.2016.10.004 (2016).
Carithers, L. J. & Moore, H. M. The Genotype-Tissue Expression (GTEx) Project. Biopreserv Biobank 13, 307–308, https://doi.org/10.1089/bio.2015.29031.hmm (2015).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature protocols 4, 44–57, https://doi.org/10.1038/nprot.2008.211 (2009).
Maaten, L. V. D. & Hinton, G. Visualizing data using t-SNE. Journal of Machine Learning Research 9, 2579–2605 (2008).
Wattenberg, M., Viégas, F. & Johnson, I. How to Use t-SNE Effectively. Distill 1, e2 (2016).
Dierks, C., Momke, S., Philipp, U. & Distl, O. Allelic heterogeneity of FGF5 mutations causes the long-hair phenotype in dogs. Animal genetics 44, 425–431, https://doi.org/10.1111/age.12010 (2013).
Drogemuller, C. et al. A mutation in hairless dogs implicates FOXI3 in ectodermal development. Science 321, 1462, https://doi.org/10.1126/science.1162525 (2008).
Kaelin, C. B. & Barsh, G. S. Genetics of pigmentation in dogs and cats. Annual review of animal biosciences 1, 125–156, https://doi.org/10.1146/annurev-animal-031412-103659 (2013).
Oguro-Okano, M., Honda, M., Yamazaki, K. & Okano, K. Mutations in the melanocortin 1 receptor, beta-defensin103 and agouti signaling protein genes, and their association with coat color phenotypes in Akita-inu dogs. The Journal of veterinary medical science 73, 853–858 (2011).
Parker, H. G., Chase, K., Cadieu, E., Lark, K. G. & Ostrander, E. A. An insertion in the RSPO2 gene correlates with improper coat in the Portuguese water dog. The Journal of heredity 101, 612–617, https://doi.org/10.1093/jhered/esq068 (2010).
Schoenebeck, J. J. & Ostrander, E. A. Insights into morphology and disease from the dog genome project. Annual review of cell and developmental biology 30, 535–560, https://doi.org/10.1146/annurev-cellbio-100913-012927 (2014).
Shirokova, V. et al. Foxi3 Deficiency Compromises Hair Follicle Stem Cell Specification and Activation. Stem cells (Dayton, Ohio) 34, 1896–1908, https://doi.org/10.1002/stem.2363 (2016).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13, 2498–2504, https://doi.org/10.1101/gr.1239303 (2003).
Blondel, V. D., Jean-Loup, G., Renaud, L. & Etienne, L. Fast unfolding of communities in large networks. Journal of Statistical Mechanics: Theory and Experiment 2008, P100008 (2008).
Acknowledgements
We received support from P30AR069632 Columbia University Skin Disease Resource-Based Center (epiCURE) and the National Alopecia Areata Foundation (to L.P.). Funding from Collaboratory@Columbia supported this collaboration between the Columbia University Data Science Institute and the Columbia University College of Physicians and Surgeons. We thank Drs Iuliana Ionita-Laza and Zihuai He for biostatistics advice, Dr. Claire Higgins for critical insights and perspectives on hair follicle biology, and Dr. Annemieke de Jong for help with interpreting lipid data. We are grateful to Drs Katherine A. Fantauzzo, Angela M. Christiano and Richard Mayeux for helpful feedback on this work.
Author information
Authors and Affiliations
Contributions
R.K.S. contributed to the development and execution of the search algorithm and analytic plan. K.C. provided additional analysis. X.L. and A.C.M. contributed to the development of the analytic plan and helped perform data analysis. R.K.S. and L.P. wrote the manuscript and prepared displays. L.P. contributed to data analysis and is responsible for the conception, design, oversight, and execution of this study, the interpretation of data, and the management of collaborations.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Severin, R.K., Li, X., Qian, K. et al. Computational derivation of a molecular framework for hair follicle biology from disease genes. Sci Rep 7, 16303 (2017). https://doi.org/10.1038/s41598-017-16050-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16050-9
This article is cited by
-
Inborn Errors of Immunity in Hidradenitis Suppurativa Pathogenesis and Disease Burden
Journal of Clinical Immunology (2023)
-
Shedding light on cashmere goat hair follicle biology: from morphology analyses to transcriptomic landascape
BMC Genomics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.