Abstract
The role of the promoter methylation of O 6-methylguanine-DNA methyltransferase (MGMT) remains controversial for breast and gynecologic cancers. We conducted a meta-analysis to assess the association between hypermethylation of MGMT promoter and the risk of breast and gynecologic cancers. A comprehensive search was conducted in PubMed and Embase electronic databases up to 19th August 2017 for studies about the association between MGMT promoter hypermethylation and breast and gynecologic cancers. A total of 28 articles including 2,171 tumor tissues and 1,191 controls were involved in the meta-analysis. The pooled results showed that MGMT promoter methylation status was significantly associated with an increased risk of breast and gynecologic cancers (OR = 4.37, 95% CI: 2.68–7.13, P < 0.05). The associations were robust in subgroup analysis based on ethnicity, cancer type, methylation detection method, and control source. This meta-analysis indicated that MGMT hypermethylation was significantly associated with the risk of breast and gynecological cancers, and it may be utilized as a valuable biomarker in early diagnostics and prognostication of these cancers. Further efforts are needed to identify and validate this finding in prospective studies, especially in situation with new methylation testing methods and samples from plasma circulating DNA.
Similar content being viewed by others
Introduction
Malignant diseases of the breast and genitals are the most common cancers in women worldwide, and about 2.8 million new cases and 1.0 million cause-specific death each year1. Breast cancer ranks first with 25.5% (1.7 million cases) of all incident cancers, and the genitals (corpus uteri, cervix uteri and ovary) accounts for 16.5% (1.1 million cases) of them. It has generally been accepted that the late diagnosis of breast and gynecologic cancers is a serious global problem, which makes treatment less likely to succeed and reduces their chances of survival2,3. As promoter CpG island hypermethylation is considered to be an early alteration in carcinogenesis and is often present in the precursor lesions of a variety of cancers, DNA hypermethylation might be used as a marker for the early diagnosis of cancer4.
O 6-methylguanine-DNA methyltransferase (MGMT), is a widely expressed DNA repair gene that plays a crucial role in repair of DNA damage caused by alkylating agents5,6. Epigenetic silencing via hypermethylation of specific promoter CpG island is regarded as one of the causes for loss of MGMT activity in tumor tissues7. It has been suggested that loss of MGMT is associated with increased carcinogenic risk and increased sensitivity to therapeutic methylating agents5. Although the exact role of MGMT promoter methylation in malignant transformation and carcinogenesis remains unrevealed completely, it might be a good biomarker candidate for early cancer detection5. The hypermethylation of CpG islands is relatively rare in normal cells, thus the detection of methylated DNA in bodily fluids can be promising8. Several studies have focused on this in other cancers including head and neck squamous cell carcinoma, lung cancer and esophageal cancer9,10,11. Nevertheless, for breast and gynecologic cancers, although many studies have explored the association between their risks and MGMT promoter hypermethylation, the results remain inconsistent12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39. A possible reason to explain the noted discrepancies in results is the inadequate statistical power of the individual studies, especially for relatively rare types (e.g. vaginal cancer and vulvar cancer). Due to breast cancer and gynecologic cancer generally share several common risk factors, such as reproductive history and BRCA1/2 mutations, it is usually adapted to explore or summarize their associated factors together in a few studies or in clinical resource (such as Physician Data Query)40,41,42,43. Therefore, we conducted this systematic review and meta-analysis to assess the association between hypermethylation of MGMT promoter and the risk of breast and gynecologic cancers.
Results
Study selection
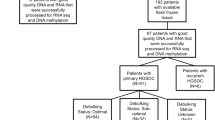
The selection flow of studies was summarized in Fig. 1. The initial search identified 429 studies on breast and gynecological cancers risk and/or clinical outcome assessment for MGMT hypermethylation. According to the inclusion criteria, 28 articles were included in our meta-analysis. One article reported two cancers16 separately and thus was divided to two studies.
The characteristics of included studies
All the eligible studies were issued in English. In total of 2,171 cases and 1,191 controls were involved in the pooled analyses. The publication year of selected studies ranged from 2001 to 2015. All studies focused on Caucasians or Asians except for two studies in the USA14,31 that also included black and other mixed populations. Table 1 presents the primary characteristics and quality assessment of the included studies. The quality of primary studies assessed by NOS showed that most studies (25 out of 29) were rated as “high quality”.
Meta-analysis
The combining result of the association of MGMT promoter hypermethylation with risk of breast and gynecological cancers was shown in Fig. 2. The random effect model was employed due to the significant heterogeneity among the included studies (I2 = 54.3%, P < 0.05). The pooled results showed that MGMT promoter methylation status was significantly associated with an increased risk of breast and gynecological cancers in women (OR = 4.37, 95% CI: 2.68–7.13, P < 0.05).
Subgroup analysis
We performed subgroup analysis to evaluate the source of the heterogeneity according to ethnicity, cancer type, methylation detection method, and control source (Table 2). No significant differences were observed in subgroup analysis based on neither ethnicity nor cancer type. Most studies used MSP to detect the frequency of MGMT promoter methylation, other methods including pyrosequencing, QMSP, MS-MLPA, MS-HRM and MethyLight were classified as non-MSP group. The ORs were 4.56 (95% CI: 2.62–7.95, P < 0.05) in the MSP group under random effects model, and 4.60 (95% CI: 1.78–11.85, P < 0.05) in the non-MSP group under the fixed-effects model. With regard to the control source, one study20 had both autologous and heterogeneous samples as control and was divided into two studies. Three studies were excluded since they included blood sample as controls. The pooled ORs in heterogeneous and autologous tissue group were overlapped under the fixed effects model, and with the value of 3.33 (95% CI: 2.16–5.14, P < 0.05) and 11.37 (95% CI: 5.11–25.31, P < 0.05), respectively. While in heterogeneous exfoliated cells group, the MGMT promoter methylation status was not significantly associated with cancer risk with a pooled OR of 1.83 (95% CI: 0.83–4.06, P = 0.136).
Sensitivity analysis
Sensitivity analysis performed by excluding the “low quality” study21,23,30,38 which got an NOS score < 6. The pooled results were not significant changed for random effects model (OR = 3.76, 95% CI: 2.30–6.15, P < 0.05), indicating that patients with hypermethylated MGMT may have an increased risk in breast and gynecological cancers.
We also took another sensitivity analysis by excluding the study23 with the biggest OR outlier in the random effects model with statistical significant finding. The overall OR was changed from 4.37 (95% CI: 2.68–7.13, P < 0.05) to 3.97 (95% CI, 2.49–6.35, P < 0.05), which demonstrated that the pooled OR was reliable and stable.
Publication bias
Visual inspection of funnel plots and the Egger’s test were used to evaluate the publication bias in our meta-analysis. The funnel plot displayed in Fig. 3 appeared asymmetrical and the statistical test showed significant result (Egger’s test P < 0.05), suggesting that there might be publication bias due to small-study effects in our study.
Discussion
To the best of our knowledge, this meta-analysis is the first to comprehensively evaluate the association between MGMT promoter methylation status and risk of breast and gynecological cancers in women. A total of 29 studies including 2,171 tumor tissues and 1,191 controls were involved in the meta-analysis. The proportion of MGMT promoter hypermethylation ranged from 3.0% to 70.1% (median: 24.8%) in tumor tissues and 0.0% to 36.9% (median: 0.3%) in non-cancerous controls, respectively. Our major finding suggested that MGMT promoter hypermethylation had a significantly increased risk in tumor tissues (OR = 4.37, 95% CI: 2.68–7.13) compared with non-cancerous tissues and exfoliated cells.
About 8 of 29 included studies presented significant association between hypermethylation of MGMT promoter and risk of breast and gynecological cancers in women16,18,21,23,28,29,31,32, whereas all of the remaining suggested no significant relationship12,13,14,15,16,17,19,20,22,24,25,26,27,30,33,34,35,36,37,38,39. When all studies were pooled into the meta-analysis, cancer risk associated with MGMT promoter hypermethylation was significant in breast and gynecological cancers. The result of sensitivity analysis revealed that this association was quite reliable and stable after excluding the study with the largest OR outlier23, or excluding four studies with lower quality21,23,30,38. Power analysis was also conducted according to our own data. Assuming OR as 4.0 and proportion of MGMT promoter hypermethylation among controls as 0.3%, the powers before and after excluding above studies were both vigorous with a value always larger than 80% in corresponding sample size.
Since heterogeneity obviously existed among studies, stratified analyses were also performed based on ethnicity, cancer types, methylation detection methods, and control source. The subgroup analysis suggested that hypermethylation of the MGMT gene was associated with the risk of breast and gynecological cancers in almost all these subgroups, except for endometrial cancer and vulvar cancer due to limited samples (<50)16,18,28. Although MSP has some defects which prompt researchers to develop novel test methods, such as pyrosequencing, QMSP,
MS-MLPA, MS-HRM and MethyLight44,45, it is still generally accepted as the best way to evaluate the methylation status of the MGMT promoter46. About 4/5 of included studies have used MSP, and no discrepant results between MSP and non-MSP were showed in our study. We acknowledge that we could not refine the non-MSP in further detail due to the limited related studies, which may need further evaluation in future. In addition, the ORs with autologous tissues as control, were not significantly different from that with heterogeneous tissues, but were significantly larger than that compared with heterogeneous exfoliated cells. It might be explained by the known higher methylation proportion of exfoliated cells in normal or intraepithelial lesions (LSIL, HSIL)21. In our pooled result, the MGMT methylation rate was more than 30% in exfoliated cells but only ranged from 0% to 14% for the adjacent tissues, which also further supported our explanation.
The MGMT gene is ubiquitously expressed in different organs and different tumors and MGMT is responsible for removing the alkyl adducts from the DNA molecules47,48. If repair of the alkylating lesions does not complete entirely, a G → A transition mutation or a strand break can occur, resulting in oncogene mutations in pre-malignant lesions (e.g. KRAS point mutations), or futile cycles of repair that triggers apoptosis (outcome of therapeutic treatment such as Temozolomide in glioblastoma), respectively47,48. In addition, it has been reported that MGMT gene expression in normal and neoplatic tissues varies and MGMT promoter methylation was associated with better suvivial in some cancer types but not all47. In the present study, we showed that MGMT promoter methylation status was significantly associated with an increased risk of breast and gynecologic cancers, which is consistent with previous studies in head and neck squamous cell carcinoma, lung cancer, glioblastoma, and esophageal cancer9,10,11. These works including ours highlighted the possibility of using MGMT promoter methylation status as a biomarker49, based on the facts that MGMT promoter hypermethylation could occur early in the neoplastic process before the clinical manifestation29,50,51, or turn up in normal appearing tissues close to tumors52,53. Currently, it has been indicated that hypermethylation of MGMT in circulating DNA might serve as a surrogate marker for tumor methylation in invasive ductal breast carcinomas26. Therefore, along with the development of different assays for CpGs methylation44,54, our finding provided supporting evidence for diagnosis and prognosis of breast and gynecological cancers with obtaining blood samples instead of biopsies.
We believe that this is the first quantitative study to assess the association between hypermethylation of MGMT promoter and the risk of breast and gynecologic cancers. Our results are reliable according to the stability and consistency in all subgroup analysis and sensitivity analyses. Neither specific factor nor single study could significantly affect the summarized OR. However, the presented information still should be interpreted with caution because some limitations existed. Firstly, funnel plots and results of Egger’s test in our study showed significant result. The small-study effect presented clearly base on visual assessment, but it’s hard to attribute this effect entirely to publication bias55. Nevertheless, publication bias may still exist considering that some studies were excluded due to unavailable information and that studies with negative results often have less chance for publication. Secondly, the lack of the original data limited the further subgroup analysis based on patients’ comorbidity, BMI, lifestyle and other environmental factors, thus, it is still not sure whether MGMT promoter hypermethylation is an independent predictive factor. Thirdly, all of the included studies were retrospective, and prospective cohort studies should be required to confirm our conclusion of its predictive value. Fourth, as an association study, it should be noted that although our results indicated the similar positive associations of MGMT promoter hypermethylation with different types of cancer, the exact underlying mechanisms might be still diverse in different types of cancer.
To sum up, this meta-analysis indicated that MGMT hypermethylation was significantly associated with the risk of breast and gynecological cancers. Consequently, detection of MGMT promoter hypermethylation may be utilized as a valuable biomarker in early diagnostics and prognostication of these cancers. However, further efforts are needed to identify and validate this finding in prospective studies, especially in situation with new methylation testing methods and samples from plasma circulating DNA.
Materials and Methods
Literature research
A comprehensive search was conducted to identify all eligible publications in PubMed and Embase electronic databases up to 19th August 201756. We used both the medical subject headings (MeSH) and free-text words. Search terms mainly included methylation, MGMT and different gynecological cancer including endometrial cancer, ovarian cancer, vulvar cancer, uterine cancer, vaginal cancer, cervical cancer, fallopian tube cancer, as well as breast cancer in women. The references of the retrieved articles and related reviews were also carefully checked to find additional eligible studies. No language or other limits were set during the course of literature search.
Inclusion and exclusion criteria
A study was included if it met the following criteria: (1) case-control or cohort study design; (2) evaluated the association between the methylation of MGMT and risk of gynecological or breast cancer in women; (3) provided sufficient data (the numbers of methylation status in two groups, respectively) for calculating the odds ratio (OR) and it 95% confidence interval (CI). Letters, comments, conference reports, laboratory studies and articles that didn’t present enough data for ORs calculation were excluded.
Data extraction
Two reviewers independently read the eligible studies. The following items were extracted from each eligible study: surname of first author, publication year, country of the investigation, ethnicity, diagnosis, method for detecting the methylation status, sample type in case and control groups, and methylation distribution. A discussion was carried out to achieve consensus when discrepancy noted.
Methodological quality assessment
The Newcastle-Ottawa Scale1 (NOS), one of the most commonly used tools for assessing the quality of observational studies in a meta-analysis setting, was employed to evaluate the quality of eligible studies by two investigators independently57. It contains three parts: case and control selection, comparability, and exposure. Each of them respectively comprises four, two, and three items. Each item is given 1 point, 9 points in total. The cut point of 6 points was used to distinguish “low quality” (<6 points) and “high quality” (≥6 points). Disagreements between investigators regarding data extraction were resolved through discussion.
Data analysis
Crude ORs together with their corresponding 95% CIs were calculated to evaluate the association between MGMT promoter hypermethylation and risk of breast and gynecological cancers. We used I2 statistic and Q test to measure the between-study heterogeneity. If I2 < 50% and P > 0.1, the heterogeneity was considered mild, and the summary ORs were combined under a fixed-effects model, otherwise a random-effects model were used. The Z test was used to assess the statistical significance of pooled ORs, and two-tailed P-values <0.05 were considered significant. Moreover, we performed subgroup analysis based on ethnicity, cancer type, methylation detection method, and control source to explore potential sources of heterogeneity. Sensitivity analysis were also performed by the study with “low quality”, and excluding the study with the OR outlier with statistically significant findings. The Egger’s test and visual inspection of funnel plots were utilized to explore any possible publication bias. All statistical analyses were conducted in STATA 12.0 (Stata Corporation, College Station, Texas, USA).
References
Ferlay, J. et al., GLOBOCAN 2012 v1.0, Cancer Incidence and Mortality Worldwide: IARC CancerBase No. 11 [Internet]. Lyon, France: International Agency for Research on Cancer; 2013. Available from: http://globocan.iarc.fr, accessed on 10/24/2016 (2016).
Caplan, L. Delay in breast cancer: implications for stage at diagnosis and survival. Front Public Health 2, 87, https://doi.org/10.3389/fpubh.2014.00087 (2014).
Clarke-Pearson, D. & Soper, J. Gynecological cancer management: identification, diagnosis and treatment. (John Wiley & Sons, 2011).
Laird, P. W. The power and the promise of DNA methylation markers. Nature Reviews Cancer 3, 253–266 (2003).
Gerson, S. L. MGMT: its role in cancer aetiology and cancer therapeutics. Nature Reviews Cancer 4, 296–307 (2004).
Kaina, B., Christmann, M., Naumann, S. & Roos, W. P. MGMT: key node in the battle against genotoxicity, carcinogenicity and apoptosis induced by alkylating agents. DNA repair 6, 1079–1099 (2007).
Esteller, M., Hamilton, S. R., Burger, P. C., Baylin, S. B. & Herman, J. G. Inactivation of the DNA repair gene O6-methylguanine-DNA methyltransferase by promoter hypermethylation is a common event in primary human neoplasia. Cancer research 59, 793–797 (1999).
Robertson, K. D. DNA methylation and human disease. Nature reviews. Genetics 6, 597–610, https://doi.org/10.1038/nrg1655 (2005).
Cai, F., Xiao, X., Niu, X., Shi, H. & Zhong, Y. Aberrant Methylation of MGMT Promoter in HNSCC: A Meta-Analysis. PloS one 11, e0163534, https://doi.org/10.1371/journal.pone.0163534 (2016).
Gu, C. et al. Association between MGMT promoter methylation and non-small cell lung cancer: a meta-analysis. PloS one 8, e72633, https://doi.org/10.1371/journal.pone.0072633 (2013).
Zhao, J. J., Li, H. Y., Wang, D., Yao, H. & Sun, D. W. Abnormal MGMT promoter methylation may contribute to the risk of esophageal cancer: a meta-analysis of cohort studies. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine 35, 10085–10093, https://doi.org/10.1007/s13277-014-2276-3 (2014).
Alkam, Y. et al. Protein expression and methylation of DNA repair genes hMLH1, hMSH2, MGMT and BRCA1 and their correlation with clinicopathological parameters and prognosis in basal-like breast cancer. Histopathology 63, 713–725, https://doi.org/10.1111/his.12220 (2013).
Banzai, C. et al. Promoter methylation of DAPK1, FHIT, MGMT, and CDKN2A genes in cervical carcinoma. Int J Clin Oncol 19, 127–132, https://doi.org/10.1007/s10147-013-0530-0 (2014).
Brait, M. et al. Association of promoter methylation of VGF and PGP9.5 with ovarian cancer progression. PloS one 8, e70878, https://doi.org/10.1371/journal.pone.0070878 (2013).
Flatley, J. E. et al. Folate status and aberrant DNA methylation are associated with HPV infection and cervical pathogenesis. Cancer epidemiology, biomarkers & prevention: a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology 18, 2782–2789, https://doi.org/10.1158/1055-9965.EPI-09-0493 (2009).
Furlan, D. et al. The high frequency of de novo promoter methylation in synchronous primary endometrial and ovarian carcinomas. Clinical cancer research: an official journal of the American Association for Cancer Research 12, 3329–3336, https://doi.org/10.1158/1078-0432.CCR-05-2679 (2006).
Guerrero, D. et al. Differential hypermethylation of genes in vulvar cancer and lichen sclerosus coexisting or not with vulvar cancer. International journal of cancer. Journal international du cancer 128, 2853–2864, https://doi.org/10.1002/ijc.25629 (2011).
Tsezou. Correlation of promoter hypermethylation in hTERT, DAPK and MGMT genes with cervical oncogenesis progression. Oncology reports 22, https://doi.org/10.3892/or_00000425 (2009).
Kang, S. et al. Polymorphism in folate- and methionine-metabolizing enzyme and aberrant CpG island hypermethylation in uterine cervical cancer. Gynecologic oncology 96, 173–180, https://doi.org/10.1016/j.ygyno.2004.09.031 (2005).
Kekeeva, T. V. et al. Aberrant methylation of tumor suppressor genes and allelic imbalance in cervical intraepithelial neoplasia. Molecular Biology 40, 194–199, https://doi.org/10.1134/s0026893306020038 (2006).
Kim, J. H. et al. Assessment of DNA methylation for the detection of cervical neoplasia in liquid-based cytology specimens. Gynecologic oncology 116, 99–104, https://doi.org/10.1016/j.ygyno.2009.09.032 (2010).
Klajic, J. et al. Quantitative DNA methylation analyses reveal stage dependent DNA methylation and association to clinico-pathological factors in breast tumors. BMC cancer 13, 456, https://doi.org/10.1186/1471-2407-13-456 (2013).
Lin, Z. et al. The hypermethylation and protein expression of p16 INK4A and DNA repair gene O6-methylguanine-DNA methyltransferase in various uterine cervical lesions. Journal of cancer research and clinical oncology 131, 364–370, https://doi.org/10.1007/s00432-004-0657-5 (2005).
Muggerud, A. A. et al. Frequent aberrant DNA methylation of ABCB1, FOXC1, PPP2R2B and PTEN in ductal carcinoma in situ and early invasive breast cancer. Breast cancer research: BCR 12, R3, https://doi.org/10.1186/bcr2466 (2010).
Roh, H. J. et al. Inactivation of O(6)-methyguanine-DNA methyltransferase by promoter hypermethylation: association of epithelial ovarian carcinogenesis in specific histological types. J Obstet Gynaecol Res 37, 851–860, https://doi.org/10.1111/j.1447-0756.2010.01452.x (2011).
Sharma, G. et al. Clinical significance of promoter hypermethylation of DNA repair genes in tumor and serum DNA in invasive ductal breast carcinoma patients. Life sciences 87, 83–91, https://doi.org/10.1016/j.lfs.2010.05.001 (2010).
Suehiro, Y. et al. Aneuploidy predicts outcome in patients with endometrial carcinoma and is related to lack of CDH13 hypermethylation. Clinical cancer research: an official journal of the American Association for Cancer Research 14, 3354–3361, https://doi.org/10.1158/1078-0432.CCR-07-4609 (2008).
Sun, L. L. et al. Population-based case-control study on DAPK1, RAR-beta2 and MGMT methylation in liquid-based cytology. Arch Gynecol Obstet 285, 1433–1439, https://doi.org/10.1007/s00404-011-2149-6 (2012).
Yang, H. J. et al. Detection of hypermethylated genes in tumor and plasma of cervical cancer patients. Gynecologic oncology 93, 435–440, https://doi.org/10.1016/j.ygyno.2004.01.039 (2004).
Spitzwieser, M., Holzweber, E., Pfeiler, G., Hacker, S. & Cichna-Markl, M. Applicability of HIN-1, MGMT and RASSF1A promoter methylation as biomarkers for detecting field cancerization in breast cancer. Breast cancer research: BCR 17, 125, https://doi.org/10.1186/s13058-015-0637-5 (2015).
An, J. et al. Messenger RNA expression and methylation of candidate tumor-suppressor genes and risk of ovarian cancer-a case-control analysis. International journal of molecular epidemiology and genetics 1, 1–10 (2010).
Asiaf, A. et al. Protein expression and methylation of MGMT, a DNA repair gene and their correlation with clinicopathological parameters in invasive ductal carcinoma of the breast. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine 36, 6485–6496, https://doi.org/10.1007/s13277-015-3339-9 (2015).
Chmelarova, M. et al. Methylation analysis of tumour suppressor genes in ovarian cancer using MS-MLPA. Folia biologica 58, 246–250 (2012).
de Groot, J. S. et al. Validation of DNA promoter hypermethylation biomarkers in breast cancer–a short report. Cellular oncology (Dordrecht) 37, 297–303, https://doi.org/10.1007/s13402-014-0189-1 (2014).
Dong, S. M., Kim, H. S., Rha, S. H. & Sidransky, D. Promoter hypermethylation of multiple genes in carcinoma of the uterine cervix. Clinical cancer research: an official journal of the American Association for Cancer Research 7, 1982–1986 (2001).
Virmani, A. K. et al. Aberrant methylation during cervical carcinogenesis. Clinical cancer research: an official journal of the American Association for Cancer Research 7, 584–589 (2001).
Shilpa, V. et al. Relationship between promoter methylation & tissue expression of MGMT gene in ovarian cancer. The Indian journal of medical research 140, 616–623 (2014).
Makarla, P. B. et al. Promoter hypermethylation profile of ovarian epithelial neoplasms. Clinical cancer research: an official journal of the American Association for Cancer Research 11, 5365–5369, https://doi.org/10.1158/1078-0432.ccr-04-2455 (2005).
Zemliakova, V. V. et al. Abnormal methylation of several tumor suppressor genes in sporadic breast cancer. Molekuliarnaia biologiia 37, 696–703 (2003).
McTiernan, A., Irwin, M. & Vongruenigen, V. Weight, physical activity, diet, and prognosis in breast and gynecologic cancers. Journal of clinical oncology: official journal of the American Society of Clinical Oncology 28, 4074–4080, https://doi.org/10.1200/JCO.2010.27.9752 (2010).
Kauff, N. D. et al. Risk-reducing salpingo-oophorectomy for the prevention of BRCA1- and BRCA2-associated breast and gynecologic cancer: a multicenter, prospective study. Journal of clinical oncology: official journal of the American Society of Clinical Oncology 26, 1331–1337, https://doi.org/10.1200/JCO.2007.13.9626 (2008).
Kurian, A. W., Kingham, K. E. & Ford, J. M. Next-generation sequencing for hereditary breast and gynecologic cancer risk assessment. Curr Opin Obstet Gynecol 27, 23–33, https://doi.org/10.1097/GCO.0000000000000141 (2015).
Board, P. D. Q. C. G. E. In PDQ Cancer Information Summaries (National Cancer Institute (US), 2017).
Kagan, J., Srivastava, S., Barker, P. E., Belinsky, S. A. & Cairns, P. Towards clinical application of methylated DNA sequences as cancer biomarkers: a joint NCI’s EDRN and NIST workshop on standards, methods, assays, reagents and tools. Cancer research 67, 4545–4549 (2007).
Chmelarova, M. & Palicka, V. The most frequent methods used for DNA methylation analysis. Casopis lekaru ceskych 150, 442–445 (2010).
Ammerpohl, O., Martín-Subero, J. I., Richter, J., Vater, I. & Siebert, R. Hunting for the 5th base: Techniques for analyzing DNA methylation. Biochimica et Biophysica Acta (BBA)-General Subjects 1790, 847–862 (2009).
Sharma, S. et al. Role of MGMT in tumor development, progression, diagnosis, treatment and prognosis. Anticancer research 29, 3759–3768 (2009).
Fan, C. H. et al. O6-methylguanine DNA methyltransferase as a promising target for the treatment of temozolomide-resistant gliomas. Cell death & disease 4, e876, https://doi.org/10.1038/cddis.2013.388 (2013).
Laird, P. W. The power and the promise of DNA methylation markers. Nature reviews. Cancer 3, 253–266, https://doi.org/10.1038/nrc1045 (2003).
Nephew, K. P. & Huang, T. H.-M. Epigenetic gene silencing in cancer initiation and progression. Cancer letters 190, 125–133 (2003).
Yan, P. S. et al. Mapping geographic zones of cancer risk with epigenetic biomarkers in normal breast tissue. Clinical Cancer Research 12, 6626–6636 (2006).
Chai, H. & Brown, R. E. Field effect in cancer–an update. Annals of Clinical & Laboratory Science 39, 331–337 (2009).
Dakubo, G. D., Jakupciak, J. P., Birch-Machin, M. A. & Parr, R. L. Clinical implications and utility of field cancerization. Cancer cell international 7, 1 (2007).
Weller, M. et al. MGMT promoter methylation in malignant gliomas: ready for personalized medicine? Nature reviews Neurology 6, 39–51 (2010).
Higgins, J. P. T. & Green, S. (editors). Cochrane Handbook for Systematic Reviews of Interventions Version 5.1.0 [updated March 2011]. The Cochrane Collaboration, 2011. Available from www.handbook.cochrane.org (2017-08-21).
Sagoo, G. S., Little, J. & Higgins, J. P. Systematic reviews of genetic association studies. Human Genome Epidemiology Network. PLoS medicine 6, e28, https://doi.org/10.1371/journal.pmed.1000028 (2009).
Wells, G. et al. The Newcastle-Ottawa Scale (NOS) for assessing the quality of nonrandomised studies in meta-analyses. University of Ottawa, 2009. http://www.ohri.ca/programs/clinical_epidemiology/oxford.asp (2016-09-21)
Acknowledgements
This study was supported by the National Natural Science Foundation of China (Grant No. 81502884)
Author information
Authors and Affiliations
Contributions
S.W. and Y.Z. designed the study. R.C. and L.Z. collected the relevant papers and data, and analyzed the data. R.C. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, R., Zheng, Y., Zhuo, L. et al. Association between MGMT Promoter Methylation and Risk of Breast and Gynecologic Cancers: A Systematic Review and Meta-Analysis. Sci Rep 7, 12783 (2017). https://doi.org/10.1038/s41598-017-13208-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13208-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.