Abstract
Engineered nanomaterials hold great promise for the future development of innovative products but their adverse health effects are a major concern. Recent studies have indicated that certain nanomaterials, including carbon nanotubes (CNTs), may be carcinogenic. However, the underlying mechanisms behind their potential malignant properties remain unclear. In this study, we linked SOX9, a stem cell associated transcription factor, to the neoplastic-like properties of human lung epithelial cells chronically exposed to a low-dose of single-walled carbon nanotubes (SWCNTs). We found that SOX9 is upregulated in SWCNT-exposed cells, which is consistent with their abilities to induce tumor formation and metastasis in vivo. We therefore hypothesized that SOX9 overexpression may be responsible for the neoplastic-like phenotype observed in our model. Indeed, SOX9 knockdown inhibited anchorage-independent cell growth in vitro and lung colonization in vivo in a mouse xenograft model. SOX9 depletion also suppressed the formation of cancer stem-like cells (CSCs), as determined by tumor sphere formation and aldehyde dehydrogenase (ALDH) activity (Aldefluor) assays. Furthermore, SOX9 knockdown suppressed tumor metastasis and the expression of the stem cell marker ALDH1A1. Taken together, our findings provide a mechanistic insight into SWCNT-induced carcinogenesis and the role of SOX9 in CSC regulation and metastasis.
Similar content being viewed by others
Introduction
Engineered nanomaterials have increasingly been used for various applications, but their long-term health effects are largely unknown. Carbon nanotubes (CNTs) are one of the most commonly used engineered nanomaterials due to their unique properties such as light weight, high tensile strength, and electrical conductivity1, 2. However, CNTs have some negative properties as well, such as a high aspect ratio and biopersistence; therefore, questions about their potential carcinogenicity have been raised3, 4. Previous animal studies have shown that pulmonary exposure to single-walled carbon nanotubes (SWCNTs) induces inflammation, granulomas, and fibrosis5, 6, conditions that have been associated with an increased risk of lung cancer7, 8. In fact, some CNTs can induce or promote tumor formation in animals3, 9,10,11,12. Furthermore, one type of CNTs, multi-walled carbon nanotubes (MWCNTs) Mitsui-7, was classified as possibly carcinogenic to humans by the International Agency for Research on Cancer (IARC)13, while in vitro data on other CNT types were concluded insufficient to be extrapolated to humans.
We previously reported that long-term, low-dose exposure of human lung epithelial cells to SWCNTs and MWCNTs results in neoplastic-like transformation14, 15. Long-term treatment with CNTs was applied to mimic gradual cellular transformation during cancer development, a process that may require a prolonged exposure to carcinogens16,17,18. We also reported that chronically SWCNT-exposed cells contain a highly invasive and tumorigenic stem-like cell subpopulation19, 20. However, detailed information about the underlying mechanisms remains unknown.
Increasing amounts of evidence suggest that cancer stem cells or stem-like cells (CSCs), also called tumor initiating cells, are the main driving force behind tumor formation and metastasis21, 22. CSCs and regular stem cells share many properties, including self-renewal capacity, potency for differentiation, and resistance to apoptosis. More importantly, CSCs are typically resistant to chemotherapy and eventually give rise to recurrent tumors22, 23.
Many stem cell regulatory proteins are now being recognized as oncogenes because of their ability to regulate CSCs. SOX9 (SRY (sex determining region Y)-box 9) is a member of the SOX family of transcription factors, which play critical roles in embryonic development, lineage commitment, and stem cell maintenance24. Notably, SOX9 is involved in lung branching morphogenesis25, and its expression is elevated in many types of cancer, including lung, skin, brain, and pancreatic cancers26. In non-small cell lung cancer (NSCLC), the most common type of lung cancer, SOX9 expression highly correlates with the disease progression and poor patient survival27, 28. Accumulating evidence also suggests that SOX9 may regulate CSCs29,30,31,32. However, detailed mechanisms have yet to be elucidated. Furthermore, it is not known whether SOX9 plays a role in SWCNT-induced carcinogenesis and CSC formation.
In this study, we demonstrated that chronically SWCNT-exposed human lung cells display high levels of SOX9 expression and contain a distinct CSC subpopulation. We hypothesized that SOX9 overexpression may be responsible for the malignant phenotype observed in these cells. Consequently, we evaluated the effects of SOX9 expression on the tumorigenicity, invasiveness, and stemness of SWCNT-transformed cells in vitro and in vivo.
Results
Exposure to carbon nanotubes induces cell transformation
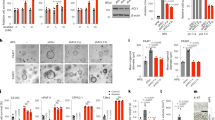
To test the potential role of SOX9 in SWCNT-induced oncogenesis, human bronchial epithelial Beas-2B cells were continuously exposed to occupationally relevant concentrations (0.02 μg/cm2) of SWCNTs for a period of 6 months, as previously described14, 15, 19. Physicochemical properties of SWCNTs used in this study are summarized in Table 1. Potential carcinogenic properties and SOX9 expression were then evaluated in vitro and in vivo. Analysis of anchorage-independent cell growth, a hallmark of cancer33, 34, revealed that the SWCNT-treated cells (termed BSW) formed 5-fold more colonies when compared to passage-matched control (B2B) cells (Fig. 1a). To evaluate the tumorigenic potential of BSW cells in vivo, these cells and the control B2B cells were genetically labeled with luciferase and injected subcutaneously into the flanks of NOD/SCID gamma mice. Tumor formation was examined over a period of 4 weeks by external caliper measurements and by bioluminescence imaging. Unlike control cells, BSW cells formed rapidly growing tumors (Fig. 1b,c). Moreover, ex vivo analysis at the end of the experiments showed spontaneous metastasis of the BSW cells to the mouse lungs and liver (Fig. 1d,e,f and Supplementary Fig. S1). These results indicate that SWCNT-transformed cells possess tumorigenic and metastatic properties.
Human bronchial epithelial cells, chronically exposed to carbon nanotubes, undergo malignant transformation. (a) Chronically SWCNT-exposed bronchial epithelial Beas-2B (BSW) cells were cultured in soft agar for 2 weeks, and colonies over 50 μm in diameter were counted, n = 15, p < 0.0001, two-tailed t-test. Scale bar = 500 μm. (b) Luciferase-labeled cells were injected subcutaneously into flanks of NOD/SCID gamma mice (n = 5 per group), and tumor volume was measured weekly by an external caliper. Means are different according to Bonferroni post hoc analysis following 2-way ANOVA, p < 0.05. (c) Weight of isolated subcutaneous tumors at week 4, p = 0.0014, two-tailed t-test. (d) Top – representative bioluminescent signal from tumors at week 4, Bottom – representative luciferase signal from isolated internal organs. (e) Luciferase signal from lungs imaged ex vivo. (f) Representative images of H&E stained lungs from mice bearing BSW cells. Arrows point to metastatic modules. Scale bar, 50 µm. Data are mean ± SEM.
SOX9 overexpression regulates malignant properties of BSW cells
Cancer stem cells (CSCs) are considered to be the main source of cancer metastasis, chemoresistance, and tumor recurrence21. Previous studies have shown that SWCNT-exposed cells contain a CSC-like subpopulation identical to CSCs reported in lung cancer19, 35. This cell population is characterized by the expression of stem cell markers, self-renewal ability, and more importantly by chemoresistance and high tumorigenic potential19, 20. In this study, we examined the molecular mechanisms of CSC regulation to identify possible biomarkers and drug targets for CNT-related malignancies. SOX9, a stem cell transcription factor, has recently been implicated in CSC regulation26, 29 and is overexpressed in non-small cell lung cancer (NSCLC)27, 32. We hypothesized that SOX9 may regulate the tumorigenic and metastatic properties of SWCNT-exposed cells by controlling CSCs. Indeed, BSW cells express a high level of SOX9 protein when compared to passage-matched control cells (Fig. 2a). Furthermore, immunostaining analysis revealed that lungs of all mice in the BSW group contain SOX9 overexpressing micrometastases (Fig. 2b). Immunofluorescent staining for CSC markers in control and SWCNT-exposed cells demonstrated that CSC markers are overexpressed in BSW cells and that the CSCs are the cells that have elevated SOX9 expression (Supplementary Fig. S2).
Human bronchial epithelial cells, chronically exposed to carbon nanotubes, overexpress SOX9. (a) Immunoblot of BSW and control cells, showing significant SOX9 overexpression in BSW cells. Right – relative SOX9 expression presented as fold change over control cells (n = 6). Data are mean ± SEM, *p = 0.0347, paired two-tailed t-test. (b) Representative image of lung metastases from mice bearing BSW cells subcutaneously; sections were stained with anti-SOX9 antibody and antibody against human mitochondria, nuclei stained with DAPI. Scale bar: 20 µm.
To test our hypothesis, SOX9 expression in BSW cells was stably knocked down by shRNAs (Fig. 3a). The knockdown cells (shSOX9) exhibited a slower rate of proliferation (Fig. 3b) and formed substantially fewer colonies in the soft agar assay when compared to vector control cells (Fig. 3c,d). These results suggest that SOX9 depletion alleviates the survival of cells in matrix depleted conditions and potentially decreases their metastatic potential. In vitro migration and invasion assays further demonstrated a marked reduction in cell motility following SOX9 knockdown (Fig. 4). We also used an established NSCLC cell line H460 to compare the results of SOX9 downregulation in BSW cells to those in lung cancer cells. SOX9 knockdown also attenuated colony formation and decreased the proliferation rate of H460 cells (Supplementary Fig. S3), which is consistent with previous reports28, 32. Taken together, our results support the critical role of SOX9 overexpression in the malignant phenotype of SWCNT-exposed cells. Specifically, SOX9 knockdown reduced cell proliferation, colony formation, migration, and invasion - properties that all are considered as hallmarks of cancer33.
SOX9 knockdown inhibits proliferative and colony-forming properties of BSW cells. (a) Immunoblot showing levels of SOX9 protein after knockdown generated by 3 different shRNAs (sh-1, sh-2, sh-3), empty vector was used as a control in BSW cells. (b) Rate of cell proliferation, p < 0.05, Bonferroni post hoc analysis following 2-way ANOVA. (c) Soft agar colony formation assay. Cells were cultured in soft agar for 2 weeks, scale bar −500 μm. (d) Quantification of the soft agar assay, colonies larger than 50 μm in diameter were counted, n = 3. Data presented as mean values ± SEM. Means are different according to Turkey post hoc analysis following ANOVA, p < 0.05.
SOX9 knockdown attenuates migration and invasion of BSW cells in vitro. SOX9 knockdown was generated by 3 different shRNAs (sh-1, sh-2, sh-3), empty vector used as a control in BSW cells. BSW cells were added to control inserts (migration) or inserts coated with Matrigel (invasion) and incubated for 15 h. After removal of non-migrated and non-invaded cells, cells were fixed and stained with crystal violet and counted under a microscope. (a) Transwell migration assay. Bottom – quantification. (b) Transwell invasion assay. Bottom - quantification. Data presented as mean ± SEM, n = 10. Means are different according to Turkey post hoc analysis following ANOVA, p < 0.05.
SOX9 regulates SWCNT-induced cancer stem cells
The ability for self-renewal is a key characteristic of stem cells36, 37. The tumor sphere formation assay has frequently been used as a functional in vitro test to evaluate the self-renewal ability of adult stem cells and cancer stem cells38, 39. To evaluate the sphere forming capability of SWCNT-transformed cells and its regulation by SOX9, 1,000 control and shSOX9 cells were cultured under non-adherent conditions in serum-free medium. After 2 weeks in culture, the number of tumor spheres exceeding 50 µm in diameter was quantified. We found that SOX9 knockdown substantially inhibited sphere formation (Fig. 5a), suggesting that SOX9 positively regulates stem cells in our model.
SOX9 regulates cancer stem cells in SWCNT-exposed cells. SOX9 knockdown was generated by 3 different shRNAs (sh-1, sh-2, sh-3), empty vector used as a control in BSW cells. (a) Tumor sphere formation assay. Right – quantitative analysis of tumor sphere formation, n = 15. (b) Fold change in the number of ALDHhi cells in SOX9 knockdown over control BSW cells, 4 independent experiments combined (n = 4–9) measured by Aldefluor assay. (c) Representative flow cytometry results for Aldefluor assay. Top row – cells stained with the Aldefluor substrate only, bottom – cells stained with the Aldefluor substrate in the presence of the specific ALDH inhibitor DEAB. (d) SOX9 knockdown leads to a depletion of cancer stem cell marker ALDH1A1, right – quantification of relative ALDH1A1 expression, fold changes over control BSW cells, n = 3 independent experiments. Results are mean ± SEM. Means are different according to Turkey post hoc analysis following ANOVA, p < 0.05. DEAB - N,N-diethylaminobenzaldehyde.
Next, we evaluated the expression and activity of the common cancer stem marker ALDH, which is highly active in cancer stem cells and is frequently associated with poor clinical outcomes23, 40, 41. The Aldefluor assay was used to quantify ALDH activity in BSW and BSW-shSOX9 cells. The activated Aldefluor reagent is a cell-permeable fluorescent substrate for ALDH that accumulates inside cells after interaction with ALDH and can be subsequently detected using flow cytometry. Diethylaminobenzaldehyde (DEAB), a specific ALDH inhibitor, was used to evaluate background fluorescence and set up gates for flow cytometry. We found that ALDH activity was substantially decreased in SOX9 knockdown cells, as indicated by a reduced number of cells exhibiting high ALDH activity (ALDHhi cells) (Fig. 5b,c). Consistent with the pattern of ALDH activity, expression levels of ALDH1A1, one of the main CSC-associated ALDH isoforms42, 43, were dramatically reduced following SOX9 knockdown (Fig. 5d). ALDH1A1 expression was also depleted in H460-shSOX9 cells (Supplementary Fig. S3), indicating that a correlation between SOX9 and ALDH expression is not limited to one specific cell system. These results suggest that SOX9 controls CSCs in BSW cells and possibly other cancer cell types through ALDH1A1.
SOX9 knockdown attenuates BSW metastasis in vivo
We next evaluated whether SOX9 depletion affects the metastatic potential of BSW cells in vivo. Luciferase-labeled BSW and BSW-shSOX9 cells were intravenously injected into mice followed by weekly bioluminescence imaging. At the end of the experiment, mice were euthanized; their internal organs (brain, liver, spleen, pancreas, kidneys and lungs) were removed and imaged to evaluate metastatic lesions. Figure 6a shows representative whole-body bioluminescence images of mice bearing either the control or shSOX9 cells. Importantly, lung colonization by shSOX9 cells was dramatically decreased when compared to control cells (Fig. 6b). This was also confirmed by ex vivo lung imaging (Fig. 6c) and by histological analysis (Fig. 6d). The luminescent signal from distant metastases in the liver and brain also decreased (Fig. 6e,f). In addition, we observed an occasional luciferase signal from the kidneys and pancreas in the control but not in the shSOX9 animals (data not shown). Our findings indicate a significant role of SOX9 in the regulation of the metastatic properties of malignantly transformed BSW cells.
SOX9 knockdown inhibits BSW metastasis in vivo. Luciferase-labeled cells were injected intravenously via tail vein into NOD/SCID gamma mice (n = 4 per group) and mice were imaged right after cell injection and weekly thereafter. SOX9 knockdown was generated by 3 different shRNAs (sh-1, sh-2, sh-3), empty vector used as a control in BSW cells. (a) Representative whole body bioluminescence of lung tumors in mice injected with control vector (BSW) or SOX9 knockdown (shSOX9-3). (b) Time course of lung tumor growth generated by weekly whole body imaging, each time point normalized to the signal from day 0. Means are different according to Bonferroni post hoc analysis following 2-way ANOVA, p < 0.05. (c,e,f) Ex vivo luciferase signal from isolated mouse lungs, brain and liver, respectively. Means are different according to Turkey post hoc analysis following ANOVA. (d) Representative pictures of H&E stained lungs showing a notable decrease in tumor lung colonization by SOX9 knockdown cells comparing to control cells, scale bar −250 µm. Data are shown as mean ± SEM, n = 4.
SOX9 overexpression in normal Beas-2B cells induces stem cell-like properties
We next overexpressed SOX9 in passage-matched control (B2B) cells to test whether this would be sufficient to recapitulate the phenotype of BSW cells. Of note, we achieved a maximum 5-fold increase of SOX9 expression (Fig. 7a), while BSW cells have on average a 15-fold increase in SOX9 expression in comparison to Beas-2B cells (Fig. 2a). Nevertheless, a 5-fold SOX9 overexpression in Beas-2B cells was sufficient to promote tumor sphere formation (Fig. 7b,c). Note that these cells normally form a small number of loose cell aggregates under low-attachment conditions, while SOX9 overexpression triggered the appearance of typical, round tumor spheres. Furthermore, mRNA levels of main NSCLC-associated isoforms ALDH1A1 and ALDH1A344 as well as ALDH activity were upregulated by SOX9 (Fig. 7d,e). We also evaluated changes in the expression levels of stem cell related proteins in response to SOX9 overexpression. We observed an increased expression of CD133, a marker of lung CSCs45, and embryonic stem cell factors Nanog, SOX2 and Oct4, although the increase was not statistically significant in the case of Oct4 (Fig. 7f). Together, these results support the regulatory role of SOX9 in lung CSCs induction and ALDH1A1 expression.
SOX9 overexpression in Beas-2B cells induces stem cell-like properties. (a) Level of SOX9 overexpression. (b) Representative images of tumor sphere formation. Scale bar, 200 µm. (c) Quantitative analysis of tumor sphere formation form (b), n = 3, p = 0.0019, two-tailed unpaired t-test. (d) RT-qPCR analysis of 2 ALDH isoforms, n = 6–12, 2 independent experiments combined, p < 0.05 by two-tailed unpaired t-test. (e) Fold change in the number of ALDHhi cells in SOX9 overexpressing cells over control measured by Aldefluor assay, combined results from 4 independent experiments (n = 17), p = 0.004, two-tailed paired t-test. (f) Expression levels of stem cell markers were examined by RT-qPCR, n = 6–9, p < 0.05 according to Bonferroni post hoc analysis following 2-way ANOVA, GFP control vs SOX9 overexpression. n.s. - not significant. Data are shown as mean ± SEM.
Discussion
In this study, we demonstrated that lung epithelial cells chronically exposed to SWCNTs may undergo malignant transformation and gain metastatic properties. These cells displayed traits typically characteristic of cancer cells, including anchorage-independent growth and in vivo tumor formation, supporting the previously published work14, 19. In addition, the metastatic potential of SWCNT-exposed cells was demonstrated for the first time. We observed that BSW cells can metastasize in both subcutaneous (Fig. 1) and tail vein mouse models (Fig. 6). We showed for the first time that SOX9 is a critical player in SWCNT-induced carcinogenesis, and that SOX9 depletion significantly reduces metastatic potential both in vitro (Fig. 4) and in vivo (Fig. 6).
Cancer stem cells (CSCs) have been considered the main source of cancer initiation, dissemination, and recurrence based on their ability for self-renewal, drug- and apoptosis resistance. Furthermore, multiple studies have illustrated that tumor initiating cells share some characteristics with regular stem cells and express stem cell markers. Thus, it is not surprising that many proteins that are involved in embryonic development, such as the SOX family of proteins, appear to be CSC drivers. Our study demonstrated a high level of SOX9 expression in aggressive SWCNT-exposed cells, consistent with recent reports of SOX9 up-regulation in multiple cancer types26. However, the functional role of SOX9 in transformed cells required further investigation. We hypothesized that the elevated SOX9 expression in cancer cells and SWCNT-exposed cells may induce CSC formation, which in turn could drive tumor formation and metastasis. Indeed, we found that SOX9 knockdown strongly inhibited tumor sphere formation and anchorage-independent growth (Figs 5a and 3c,d). Bioluminescence signals from mouse lungs in the tail vein model clearly demonstrated that SOX9 knockdown decreases lung colonization (Fig. 6). Together, these results suggest that SOX9 regulates cancer cell survival, consistent with our previous work in lung cancer32.
We tested how SOX9 affects CSC markers to further evaluate our hypothesis. Elevated activity of ALDH, an enzyme with a variety of functions including detoxification, is associated with poor clinical outcomes due to metastasis and has been linked to CSCs23, 40, 41. We observed a substantial decrease in ALDH activity following SOX9 knockdown (Fig. 5d,c), suggesting that ALDH may be a potential downstream target of SOX9. Likewise, the expression of ALDH1A1, an ALDH isoform most commonly associated with the CSC activity23, 40, 42, was dramatically reduced in both BSW and H460 knockdown cells (Fig. 5d, Supplementary Fig. S3). Notably, ALDH serves not only as a stem cell marker, but also governs stem cell differentiation46, 47 by producing retinoic acid48. To further explore the potential relationship between ALDH and SOX9, we overexpressed SOX9 in control Beas-2B cells. We observed an increase in tumor sphere formation (Fig. 7b,c), which is consistent with the knockdown experiments. Likewise, mRNA levels of the reported to be overexpressed in NSCLC ALDH isoforms ALDH1A1 and ALDH1A344 were elevated, validating our findings in SOX9 knockdown cells. We also observed an increased expression of stem cell makers CD133, Nanog and SOX2 (Fig. 7f), suggesting that SOX9 overexpression renders cells less differentiated. Collectively, our observations strongly support the positive regulatory role of SOX9 in CSC formation, consistent with other studies in pancreatic, esophageal, and colorectal cancers30, 49, 50. To our knowledge, the present study demonstrates the link between SOX9 and ALDH expression for the first time. This provides a key mechanism underlying CSC induction by SOX9, while the exact mode of ALDH upregulation remains a subject for further investigation.
A limitation of this study is that the results were largely derived from one cell model (Beas-2B cells) and one type of CNTs. However, our group observed a similar neoplastic-like transformation in several other models and with different types of CNTs. For examples, MWCNTs have recently been shown to induce neoplastic transformation of primary human small airway epithelial cells51. Both SWCNTs and MWCNTs similarly induced such transformation in hTERT-immortalized human small airway epithelial cells after a long-term exposure15. The induction of fibroblast stem-like cells and ALDH1A1 expression was also reported in primary human lung fibroblasts exposed to SWCNTs or MWCNTs51. Furthermore, co-culture of these activated fibroblasts with lung cancer cells promoted CSC-related properties and tumor formation by cancer cells52. Of note, animal experiments in the current study were conducted in immunodeficient mice, and therefore these results should be interpreted with caution. Nonetheless, several other studies using immunocompetent mice and rats consistently indicate the tumorigenic potential of SWCNTs and MWCNTs9,10,11,12. While SOX9 is implicated as a key regulator of SWCNT-induced carcinogenesis in this report, other regulatory mechanisms are likely to be involved due to the complexity of carcinogenic process. For example, Shvedova et al. showed that inhalation or aspiration of SWCNTs caused lung inflammation and fibrosis along with K-ras mutation in immunocompetent mice53. Since K-ras is a known oncogene, whose mutational activation is frequently associated with an increased risk of lung cancer, it is quite possible that such mutation may be involved in SWCNT-induced carcinogenesis, although this has not been demonstrated yet. In addition, several studies evaluating CNT carcinogenicity via whole transcriptome arrays have identified cancer signatures and known cancer prognostic markers in exposed mouse lung tissues54, 55. Some of the identified genes such as caveolin-1 and Bcl-2 have already been implicated in CNT-induced carcinogenesis20, 56 and may be responsible for the early neoplastic transformation induced by SWCNTs.
Lung cancer is a progressive disease commonly associated with a long-term exposure to carcinogens. However, prolonged exposure studies to nanomaterials are lacking and are greatly needed for risk assessment and for safe-by-design strategies. Our results suggest that long-term exposure to SWCNTs could transform normal epithelial cells into metastatic tumor cells. Although multiple mechanisms and signaling pathways are likely to be involved in the transformation process, we report here the critical role of SOX9 in the transformation through CSC-related mechanisms. Given that CSCs may be involved in cancer initiation steps, we suggest that SOX9 may be used as an early biomarker for SWCNT-induced carcinogenesis. It remains to be elucidated whether SOX9 up-regulation and its contribution to malignancies are specific to SWCNTs or could be extrapolated to other CNTs. Such information will aid in the design of nanomaterial-specific biomarkers for risk assessment and safe-by-design efforts.
Methods
Cell culture and exposure to SWCNTs
Immortalized human bronchial epithelial Beas-2B cells were obtained from the American Type Culture Collection (ATCC). Cells were cultured in advanced Dulbecco’s modified Eagle medium (DMEM) (Life Technologies) supplemented with 1% fetal bovine serum (FBS, Atlanta Biologicals), 2 mM L-glutamine, 20 mM HEPES, 100 units/mL penicillin and 100 μg/mL streptomycin in 5% CO2 at 37 °C. This cell model has been reported to be an appropriate model for in vitro lung carcinogenesis studies57. The cells were exposed to well-characterized single-walled CNTs (SWCNTs), as previously described14, 19. Briefly, SWCNTs (Carbon Nanotechnology (CNI)) were purified by acid treatment to remove metal contaminates. Particle characterization studies were performed at NIOSH research facilities, and the results are summarized in Table 1. Particles were treated with acetone and placed in an ultrasonic bath at room temperature for 24 h. The dispersed CNTs were then filtered from the solution using a 20 μm nylon mesh screen followed by a 0.2 μm polytetrafluoroethylene (PTFE) filter. After filter collection, the dispersed CNTs were washed thoroughly with distilled water to remove acetone. The filter was dried overnight in vacuum and weighed to determine the quantity of SWCNTs. The particles were suspended in phosphate-buffered saline (PBS) by brief sonication and added to the cells every 3–4 days when changing the culture medium. Subconfluent cultures of Beas-2B cells were continuously exposed to a low-dose, occupationally relevant concentration (0.02 μg/cm2 or 0.1 μg/mL) of SWCNTs in culture for 6 months and passaged weekly. Cells were rinsed with PBS prior to culture medium changes and cell passaging to reduce potential SWCNT bioaccumulation over the exposure period.
The dose of SWCNTs used in this study was calculated based on reported effects of in vivo MWCNT exposure, normalized to mouse alveolar surface area. The lowest dose, which induced a biological response in vivo, was 10 μg/mouse lung (0.5 mg/kg body weight)6. Dividing this dose by the average mouse alveolar surface area (~500 cm2) gives the in vitro surface area dose of 0.02 μg/cm2, which is roughly equivalent to a human lung burden for 8 hours/day over a month at 400 μg/m3 (high CNT level reported in a research facility)58 or about 3 years at 10 μg/m3 (average CNT level in U.S. facilities)59. The cells were cultured in normal medium without SWCNTs for at least ten passages prior to further experiments Control cells were cultured for the same period of time in the above described culture medium. H460 (NCI-H460) cells were purchased from the ATCC and were passaged less than 20 times. H460 cells were maintained in RPMI medium (Corning) supplemented with 10% FBS, 100 units/mL penicillin, and 100 μg/mL streptomycin.
Plasmids and generation of stable cell lines
pLKO.1 lentiviral plasmids for SOX9 knockdown generation were purchased from GE Healthcare (#RHS4533-EG6662). An empty vector was used as a control, and the virus was produced in HEK293T cells (ATCC). Briefly, HEK293T cells were co-transfected with the shRNA containing pLKO.1 plasmids or empty pLKO.1 vector, pCMV-VSV-G envelope, and psPAX2 packaging plasmids (Addgene #8454, 12260) in the presence of FuGene 6 transfection reagent (Promega). The medium containing viral particles was collected at 24, 36, and 48 hours post-transfection, pooled, and used to infect SWCNT-exposed (BSW) cells in the presence of hexadimethrine bromide (Sigma-Aldrich) at a final concentration of 5 μg/mL. After infection, cells were selected with 1 μg/mL of puromycin (Life Technologies). For SOX9 overexpression, SOX9 cDNA from pCMV-AC-GFP-SOX9 (Origene) was amplified by PCR introducing BamHI/SalI restriction sites and was subcloned into pLenti CMV GFP Zeo (Addgene #17449), replacing GFP. pLenti CMV GFP Zeo vector was used as a control. For flow cytometry experiments (Aldefluor assay), control vector with RFP instead of GFP was used. Viral particles were produced as described above.
Immunoblotting
Cells were lysed in a non-reducing loading buffer containing 63 mM Tris-HCl pH 6.8, 10% glycerol, and 2% sodium dodecyl sulfate (SDS). Protein concentrations were quantified using the Pierce BCA protein assay kit (Thermo-Scientific). Proteins (40 μg) were resolved under denaturing conditions by 7.5–12% SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto PVDF membranes (EMD Millipore). Membranes were blocked in 5% non-fat milk in TBS-T buffer (25 mM Tris-HCl, pH 7.4, 125 mM NaCl, and 0.1% Tween 20) for 1 hour, washed 3 times for 5 minutes with TBS-T buffer, and incubated with appropriate primary antibodies at 4 °C overnight. Membranes were rinsed 3 times for 5 minutes with TBST-T buffer; membranes were then incubated with appropriate HRP-conjugated secondary antibodies (Jackson Immunoresearch), diluted in TBS-T for 1 hour at room temperature. Immune complexes were detected by enhanced chemiluminescence (Pierce ECL, Thermo-Scientific; Immobilon HRP Substrate, Millipore) and quantified with the Image Studio Lite program (Li-COR Biosciences). Antibodies used in this study are listed in the Supplementary Table 1. Full length unprocessed blot images are provided in the Supplementary Materials.
Flow cytometry
The ALDHhi population (cells with high aldehyde dehydrogenase activity) was identified by staining cells with the Aldefluor kit (Stem Cell Technologies), according to the manufacturer’s instructions. Briefly, for each sample, 3 × 105 cells were incubated in the Aldefluor assay buffer with the activated Aldefluor substrate for 45 minutes at 37 °C in the presence or absence of the specific ALDH inhibitor diethylaminobenzaldehyde (DEAB). Tubes stained with both the Aldefluor substrate and DEAB served as a negative control for each sample. ALDH converts Aldefluor substrate into a fluorescent product, and cells with high ALDH activity (ALDHhi) were detected by BD Fortessa cell analyzer (BD Biosciences). The flow cytometry gates were set to obtain 0.1% ALDHhi cells in substrate + inhibitor tubes for each sample. All experiments were performed at least 3 times.
Soft agar colony formation assay
Cells were suspended in a soft agar medium containing 0.33% Difco agar (BD Biosciences) in 2x EMEM medium (Lonza) supplemented with L-glutamine, antibiotics and 15% FBS, and plated onto 6-well plates over a layer of 0.5% soft agar medium. Cells were cultured under standard cell culture conditions and were fed with 0.2 mL of the regular medium twice a week. Plates were observed under a light microscope after 2 weeks, and colonies exceeding 50 μm in diameter were scored. Experiments were performed in triplicates; at least 5 fields of view for each replicate were evaluated.
Migration and invasion assays
Cells were seeded in the upper chamber of Transwell matrigel coated (invasion) or control inserts (migration) with 8 µm pores (Corning) in serum-free medium and were allowed to migrate or invade toward the serum-containing bottom chamber for 15 hours. Cells were subsequently removed from the top part of the chamber using a cotton swab, and cells on the bottom side were stained with crystal violet. Experiments were performed in at least duplicates, 5 random fields were photographed under a light microscope for each replicate.
Cell proliferation assay
The rate of cell proliferation was measured by seeding equal numbers of cells onto 6-well plates in duplicates and counting cells with the Countess automated cell counter (Life Technologies) every 2 days until plates become confluent.
Tumor sphere formation assay
Cells were plated onto ultra-low attachment 24-well plates (Corning) in 0.8% methylcellulose (MC)-based serum-free medium (Stem Cell Technologies, #H4100) supplemented with 20 ng/mL epidermal growth factor (BD Biosciences), 10 ng/mL basic fibroblast growth factor (Sigma-Aldrich), and 5 mg/mL insulin (Sigma-Aldrich). The number of tumor spheres exceeding 50 µm in diameter (20 µm for Beas-2B cells) was quantified under a light microscope after 2 weeks of culture under standard cell culture conditions. All experiments were performed in triplicates and repeated at least twice, with 5 fields of view analyzed for each replicate.
Mouse xenograft experiments
All experiments were performed in accordance with the Guidelines for Animal Experiments at West Virginia University and approved by the Institutional Animal Care and Use Committee. Immunodeficient NOD/SCID gamma mice (NSG), strain NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (Jackson Laboratory), were injected with 1 × 106 of luciferase-labeled cells subcutaneously or via the tail vein. Tumor growth was monitored weekly using IVIS Lumina II in Vivo Imaging system (PerkinElmer), and an external caliper (VWR International) was used for the subcutaneous model. Tumor volume was calculated using the formula: tumor volume [mm3] = 1/2 (length [mm]) × (width [mm]2). At the end of the experiments mice were euthanized, and their organs were removed and imaged ex vivo on a Petri dish to evaluate metastasis. Bioluminescent images were quantified using the Living Image software (PerkinElmer). Tissue processing and H&E staining was carried out by the WVU Pathology Laboratory for Translational Medicine using standard procedures.
Immunohistochemistry
Tissue sections were deparaffinized using standard techniques. Antigen retrieval was carried out by heating slides in a microwave in citrate buffer (10 mM citric acid, 0.05% Tween 20, pH 6.0) for 30 min. Samples were blocked for 30 min with normal goat serum (BioGenex, #HK112–9K) and incubated with primary antibodies overnight at 4 °C (specific antibodies used are listed in Supplementary Table 1). Slides were rinsed with TBS-T buffer 3 times and incubated with fluorescently labeled secondary antibodies (Life Technologies) for 1 hour at room temperature; after which they were rinsed with TBS-T buffer 3 times and mounted with the ProLong DAPI (Life Technologies). Cells were visualized with a Zeiss fluorescent microscope (Carl Zeiss).
Immunofluorescence
Cells were grown overnight on 8-well chamber slides (Nunc), fixed with 4% paraformaldehyde and permeabilized with 100% methanol at −20 °C prior to blocking with 5% BSA in PBS. Cell were incubated with specific primary antibodies (listed in Supplementary Table 1), rinsed with PBS and incubated with secondary antibodies. Secondary antibodies were goat-anti-rabbit Alexa Fluor 546 (Life Technologies) and donkey-anti-mouse Alexa Fluor 488 (Jackson Immunoresearch). Slides were mounted with Fluoroshield with DAPI (Abcam) and visualized with a Zeiss fluorescent microscope (Carl Zeiss).
RT-qPCR
Total RNA was purified using the RNeasy kit (Qiagen). Reverse transcription was performed with SuperScript III Reverse Transcriptase (Life Technologies) using oligo(dT) primers. Quantitative real-time PCR was carried out using the SYBR Green Master Mix (Applied Biosystems) in an ABI 7500 Real-Time PCR Cycler (Applied Biosystems). Results were calculated using the 2−ΔΔCt method; glyceraldehyde-3-phosphate dehydrogenase (GAPDH) served as the internal control. Primer sequences are listed in the Supplementary Table 2.
Statistical analysis
The data represent means ± SEM from two or more independent experiments as indicated. Statistical comparisons were made using two-tailed Student’s t test. When more than two groups were analyzed, ANOVA was used. Statistical analysis was performed with GraphPad Prism software (GraphPad), and p < 0.05 was considered statistically significant.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Disclaimer
The findings and conclusions in this report are those of the authors and do not necessarily represent the views of the National Institute for Occupational Safety and Health.
References
Dai, H. J. Carbon nanotubes: synthesis, integration, and properties. Acc. Chem. Res. 35, 1035–1044 (2002).
Popov, V. N. Carbon nanotubes: properties and application. Mat. Sci. Eng. R. 43, 61–102 (2004).
Poland, C. A. et al. Carbon nanotubes introduced into the abdominal cavity of mice show asbestos-like pathogenicity in a pilot study. Nat. Nanotechnol. 3, 423–428 (2008).
Sanchez, V. C., Pietruska, J. R., Miselis, N. R., Hurt, R. H. & Kane, A. B. Biopersistence and potential adverse health impacts of fibrous nanomaterials: what have we learned from asbestos? WIREs Nanomed. Nanobiotechnol. 1, 511–529 (2009).
Lam, C. W., James, J. T., McCluskey, R. & Hunter, R. L. Pulmonary toxicity of single-wall carbon nanotubes in mice 7 and 90 days after intratracheal instillation. Toxicol. Sci. 77, 126–134 (2004).
Shvedova, A. A. et al. Unusual inflammatory and fibrogenic pulmonary responses to single-walled carbon nanotubes in mice. Am. J. Physiol. Lung Cell Mol. Physiol. 289, L698–L708 (2005).
Shacter, E. & Weitzman, S. A. Chronic inflammation and cancer. Oncology (Williston Park). 16, 217–226, 229; discussion 230-212 (2002).
Fillon, M. Details linking chronic inflammation and cancer continue to emerge. J. Natl. Cancer Inst. 105, 509–510 (2013).
Nagai, H. et al. Diameter and rigidity of multiwalled carbon nanotubes are critical factors in mesothelial injury and carcinogenesis. Proc. Natl. Acad. Sci. USA 108, E1330–1338 (2011).
Shvedova, A. A. et al. MDSC and TGFbeta are required for racilitation of tumor growth in the lungs of mice exposed to carbon nanotubes. Cancer Res. 75, 1615–1623 (2015).
Suzui, M. et al. Multiwalled carbon nanotubes intratracheally instilled into the rat lung induce development of pleural malignant mesothelioma and lung tumors. Cancer Sci. 107, 924–935 (2016).
Kasai, T. et al. Lung carcinogenicity of inhaled multi-walled carbon nanotube in rats. Part. Fibre Toxicol. 13, 53 (2016).
Grosse, Y. et al. Carcinogenicity of fluoro-edenite, silicon carbide fibres and whiskers, and carbon nanotubes. The Lancet Oncology. 15, 1427–1428 (2014).
Wang, L. et al. Carbon nanotubes induce malignant transformation and tumorigenesis of human lung epithelial cells. Nano Lett. 11, 2796–2803 (2011).
Wang, L. et al. Neoplastic-like transformation effect of single-walled and multi-walled carbon nanotubes compared to asbestos on human lung small airway epithelial cells. Nanotoxicology. 8, 485–507 (2014).
Carel, R. et al. Occupational exposure to asbestos and man-made vitreous fibres and risk of lung cancer: a multicentre case-control study in Europe. Occup. Environ. Med. 64, 502–508 (2007).
Donaldson, K., Murphy, F. A., Duffin, R. & Poland, C. A. Asbestos, carbon nanotubes and the pleural mesothelium: a review of the hypothesis regarding the role of long fibre retention in the parietal pleura, inflammation and mesothelioma. Part. Fibre Toxicol. 7 (2010).
Pintos, J., Parent, M.-E., Rousseau, M.-C., Case, B. W. & Siemiatycki, J. Occupational exposure to asbestos and man-made vitreous fibers, and risk of lung cancer: evidence from two case-control studies in Montreal, Canada. J. Occup. Environ. Med. 50, 1273–1281 (2008).
Luanpitpong, S., Wang, L., Castranova, V. & Rojanasakul, Y. Induction of stem-like cells with malignant properties by chronic exposure of human lung epithelial cells to single-walled carbon nanotubes. Part. Fibre Toxicol. 11, 22 (2014).
Luanpitpong, S. et al. Caveolin-1 regulates lung cancer stem-like cell induction and p53 inactivation in carbon nanotube-driven tumorigenesis. Oncotarget. 5, 3541–3554 (2014).
Kreso, A. & Dick, J. E. Evolution of the Cancer Stem Cell Model. Cell Stem Cell. 14, 275–291 (2014).
Beck, B. & Blanpain, C. Unravelling cancer stem cell potential. Nat. Rev. Cancer. 13, 727–738 (2013).
Charafe-Jauffret, E. et al. Aldehyde dehydrogenase 1-positive cancer stem cells mediate metastasis and poor clinical outcome in inflammatory breast cancer. Clin. Cancer Res. 16, 45–55 (2010).
Jo, A. et al. The versatile functions of Sox9 in development, stem cells, and human diseases. Genes Dis. 1, 149–161 (2014).
Rockich, B. E. et al. Sox9 plays multiple roles in the lung epithelium during branching morphogenesis. Proc. Natl. Acad. Sci. USA 110, E4456–4464 (2013).
Matheu, A. et al. Oncogenicity of the developmental transcription factor Sox9. Cancer Res. 72, 1301–1315 (2012).
Zhou, C. H. et al. Clinical significance of SOX9 in human non-small cell lung cancer progression and overall patient survival. J. Exp. Clin. Cancer Res. 31 (2012).
Jiang, S. S. et al. Upregulation of SOX9 in Lung Adenocarcinoma and Its Involvement in the Regulation of Cell Growth and Tumorigenicity. Clin. Cancer Res. 16, 4363–4373 (2010).
Guo, W. et al. Slug and Sox9 cooperatively determine the mammary stem cell state. Cell. 148, 1015–1028 (2012).
Sun, L. et al. Epigenetic regulation of SOX9 by the NF-kappaB signaling pathway in pancreatic cancer stem cells. Stem Cells. 31, 1454–1466 (2013).
Deng, W. et al. SOX9 inhibits beta-TrCP-mediated protein degradation to promote nuclear GLI1 expression and cancer stem cell properties. J. Cell Sci. 128, 1123–1138 (2015).
Luanpitpong, S. et al. SLUG is required for SOX9 stabilization and functions to promote cancer stem cells and metastasis in human lung carcinoma. Oncogene. 35, 2824–2833 (2016).
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell. 100, 57–70 (2000).
Mori, S. et al. Anchorage-independent cell growth signature identifies tumors with metastatic potential. Oncogene. 28, 2796–2805 (2009).
Singh, S. & Chellappan, S. Lung cancer stem cells: Molecular features and therapeutic targets. Mol. Aspects Med. 39, 50–60 (2014).
Ponti, D. et al. Isolation and in vitro propagation of tumorigenic breast cancer cells with stem/progenitor cell properties. Cancer Res. 65, 5506–5511 (2005).
Falzacappa, M. V. V., Ronchini, C., Reavie, L. B. & Pelicci, P. G. Regulation of self-renewal in normal and cancer stem cells. FEBS J. 279, 3559–3572 (2012).
Hirschhaeuser, F. et al. Multicellular tumor spheroids: An underestimated tool is catching up again. J. Biotechnol. 148, 3–15 (2010).
Pastrana, E., Silva-Vargas, V. & Doetsch, F. Eyes wide open: a critical review of sphere-formation as an assay for stem cells. Cell Stem Cell. 8, 486–498 (2011).
Ginestier, C. et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell. 1, 555–567 (2007).
Moreb, J. S. Aldehyde dehydrogenase as a marker for stem cells. Curr. Stem Cell Res. Ther. 3, 237–246 (2008).
Marcato, P., Dean, C. A., Giacomantonio, C. A. & Lee, P. W. K. Aldehyde dehydrogenase Its role as a cancer stem cell marker comes down to the specific isoform. Cell Cycle. 10, 1378–1384 (2011).
Sullivan, J. P. et al. Aldehyde dehydrogenase activity selects for lung adenocarcinoma stem cells dependent on notch signaling. Cancer Res. 70, 9937–9948 (2010).
Patel, M. et al. ALDH1A1 and ALDH3A1 expression in lung cancers: correlation with histologic type and potential precursors. Lung Cancer. 59, 340–349 (2008).
Bertolini, G. et al. Highly tumorigenic lung cancer CD133+ cells display stem-like features and are spared by cisplatin treatment. Proc Natl Acad Sci USA 106, 16281–16286 (2009).
Gudas, L. J. & Wagner, J. A. Retinoids regulate stem cell differentiation. J. Cell. Physiol. 226, 322–330 (2011).
Chute, J. P. et al. Inhibition of aldehyde dehydrogenase and retinoid signaling induces the expansion of human hematopoietic stem cells. Proc Natl Acad Sci USA 103, 11707–11712 (2006).
Ginestier, C. et al. Retinoid signaling regulates breast cancer stem cell differentiation. Cell Cycle. 8, 3297–3302 (2009).
Kopp, J. L. et al. Identification of Sox9-dependent acinar-to-ductal reprogramming as the principal mechanism for initiation of pancreatic ductal adenocarcinoma. Cancer Cell. 22, 737–750 (2012).
Song, S. et al. Hippo coactivator YAP1 upregulates SOX9 and endows esophageal cancer cells with stem-like properties. Cancer Res. 74, 4170–4182 (2014).
Luanpitpong, S. et al. Induction of stemlike cells with fibrogenic properties by carbon nanotubes and its role in fibrogenesis. Nano Lett. 14, 3110–3116 (2014).
Luanpitpong, S. et al. Induction of cancer-associated fibroblast-like cells by carbon nanotubes dictates its tumorigenicity. Sci. Rep. 6, 39558 (2016).
Shvedova, A. A. et al. Inhalation vs. aspiration of single-walled carbon nanotubes in C57BL/6 mice: inflammation, fibrosis, oxidative stress, and mutagenesis. Am. J. Physiol. Lung Cell Mol. Physiol. 295, L552–L565 (2008).
Pacurari, M. et al. Multi-walled carbon nanotube-induced gene expression in the mouse lung: association with lung pathology. Toxicol. Appl. Pharmacol. 255, 18–31 (2011).
Poulsen, S. S. et al. Transcriptomic Analysis Reveals Novel Mechanistic Insight into Murine Biological Responses to Multi-Walled Carbon Nanotubes in Lungs and Cultured Lung Epithelial Cells. PLoS One. 8 (2013).
Chen, D. et al. Gene expression profile of human lung epithelial cells chronically exposed to single-walled carbon nanotubes. Nanoscale Res Lett. 10, 12 (2015).
Park, Y.-H., Kim, D., Dai, J. & Zhang, Z. Human bronchial epithelial BEAS-2B cells, an appropriate in vitro model to study heavy metals induced carcinogenesis. Toxicol. Appl. Pharmacol. 287, 240–245 (2015).
Han, J. H. et al. Monitoring multiwalled carbon nanotube exposure in carbon nanotube research facility. Inhal. Toxicol. 20, 741–749 (2008).
Erdely, A. et al. Carbon nanotube dosimetry: from workplace exposure assessment to inhalation toxicology. Part. Fibre Toxicol. 10, 53 (2013).
Acknowledgements
This work was supported by grants from the NIH (R01-ES022968, R01-EB018857) and NSF (CBET-1434503). Flow Cytometry experiments were performed in the WVU Flow Cytometry & Single Cell Core Facility, which is supported by the NIH equipment grant S10OD016165 and the Institutional Development Award (IDeA) from the National Institute of General Medical Sciences of the NIH - P30GM103488 (CoBRE) and P20GM103434 (INBRE). Small animal imaging and analysis were performed in the WVU Animal Models & Imaging Facility supported by the WVU Cancer Institute and NIH grants P20 RR016440, P30 GM103488 and S10 RR026378. Imaging experiments and analysis were performed in the WVU Microscope Imaging Facility, supported by the WVU Cancer Institute and NIH grants P20 RR016440, P30 GM103488 and P20 GM103434.
Author information
Authors and Affiliations
Contributions
Conception and design: M.A.V., S.L., Y.R. Acquisition of data: M.A.V. Analysis and interpretation of data: M.A.V., L.R., V.C., H.R., Y.R. Administrative, technical, or material support: L.R., V.C., H.R., Y.R., C.Z.D. Drafting or revision of the manuscript: M.A.V., H.R., V.C., Y.R. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Voronkova, M.A., Luanpitpong, S., Rojanasakul, L.W. et al. SOX9 Regulates Cancer Stem-Like Properties and Metastatic Potential of Single-Walled Carbon Nanotube-Exposed Cells. Sci Rep 7, 11653 (2017). https://doi.org/10.1038/s41598-017-12037-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-12037-8
This article is cited by
-
Photobiomodulation Facilitates Rat Cutaneous Wound Healing by Promoting Epidermal Stem Cells and Hair Follicle Stem Cells Proliferation
Tissue Engineering and Regenerative Medicine (2024)
-
3-Nitrobenzanthrone promotes malignant transformation in human lung epithelial cells through the epiregulin-signaling pathway
Cell Biology and Toxicology (2022)
-
Identification of a stemness-related gene panel associated with BET inhibition in triple negative breast cancer
Cellular Oncology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.