Abstract
For somatic point mutations in coding and non-coding regions of the genome, we propose CScape, an integrative classifier for predicting the likelihood that mutations are cancer drivers. Tested on somatic mutations, CScape tends to outperform alternative methods, reaching 91% balanced accuracy in coding regions and 70% in non-coding regions, while even higher accuracy may be achieved using thresholds to isolate high-confidence predictions. Positive predictions tend to cluster in genomic regions, so we apply a statistical approach to isolate coding and non-coding regions of the cancer genome that appear enriched for high-confidence predicted disease-drivers. Predictions and software are available at http://CScape.biocompute.org.uk/.
Similar content being viewed by others
Introduction
Next generation sequencing technologies have accelerated the discovery of single nucleotide variants (SNVs) in the human genome, stimulating the development of predictors for classifying which of these variants are likely functional in disease, and which neutral. Predictors have been developed for variants in both the coding and non-coding regions of the human genome. For example, in Shihab et al.1, we developed such a predictor based on pathogenic disease-driver germline variants from the Human Gene Mutation Database (HGMD2), and assumed neutral variants from the 1,000 Genomes Project Consortium (1000G3). Multiple types of data may be informative, so we used an integrative binary classifier that weighted component data-types according to their relative informativeness1. A variety of similar predictors have been proposed4,5,6,7,8,9.
In this paper we focus on prediction for somatic point mutations in both coding and non-coding regions of the human cancer genome. The development of such predictors will be important for interpreting cancer sequence databases currently being compiled, such as the Cancer Genome Atlas10, the International Cancer Genome Consortium11 and national programmes such as the Genomics England (100,000 genomes) Project12. Mirroring previous methods1, 13, we use an integrative classifier and a wide variety of data sources. Using leave-one-chromosome-out cross-validation (LOCO-CV), the proposed method, which we call CScape, outperforms rival methods, achieving balanced accuracy of 72.3% in coding regions and 62.9% in non-coding regions. Tested on independent data sets, CScape can achieve up to 91% balanced accuracy in coding regions and 70% in non-coding regions. Most methods in this domain attempt to rank mutations to identify the most likely oncogenic examples. One drawback of this approach is that clinical practitioners may find it difficult to interpret scores without provision of specific thresholds for discriminating between benign and oncogenic mutations. Hence in this study we establish two thresholds for each model (a default threshold and a high-confidence threshold) by associating a confidence measure to each predicted label. If we restrict prediction to highest confidence instances only (cautious classification) then balanced accuracy in LOCO-CV rises to 91.7% for coding regions and 76.1% for non-coding regions, with predictions confined to 17.7% (coding) and 14.8% (non-coding) of nucleotide positions across the genome, respectively. These high confidence positive (oncogenic) predictions are typically clustered by genomic location, hence we further introduce a statistical method to find significant contiguous sets of such positive predictions. This latter approach highlights a number of genes as potentially oncogenic via somatic point mutation.
Results
We assembled two distinct datasets: our pathogenic (positive) dataset was constructed using somatic point mutations from the COSMIC database14 (version 75, November 2015) and our control (negative) dataset was constructed using SNVs from the 1,000 Genomes Project3. This control dataset contains many variants which are unannotated and therefore could be true positives. Similarly, we may expect true negatives among the positively labeled COSMIC datapoints, such as passenger mutations co-located with driver mutations. Among the COSMIC database annotations is the recurrence level, or the number of times a mutation has been observed in different cases. To increase the likelihood that positive examples are drivers, not passengers, we use pathogenic examples with a COSMIC recurrence count as high as possible. However, the number of training examples decreases rapidly as we increase the recurrence threshold (Supplementary Figure S1). In addition, an exceedingly high threshold may introduce bias, for example, by restricting the training set to only a subset of relevant genes. Hence for each model we choose as our threshold the highest recurrence level that shows little evidence of bias and provides sufficient training examples to create an accurate classifier (Supplementary Figure S2). Using this approach we thus select recurrence thresholds of r = 5 for coding regions, and r = 3 for non-coding regions.
Bias may also be introduced if negative examples are located in different genomic regions from positive examples. For example, if positives appear predominantly near transcription start sites while negatives are distributed more broadly1, 7, 15. To ensure the locations of putative non-driver mutations approximate those of driver mutations, we select only those putative non-drivers found within a window w of a putative driver mutation. For coding examples we use w = 10,000, and for non-coding examples we use w = 1,000 (Supplementary Figure S3). Hence our final training set, outlined in Supplementary Table S1, consists of 46,420 coding examples and 131,714 non-coding examples.
Feature Groups
All of our data are based on the GRCh37/hg19 version of the human genome and detailed further in the Supplementary. Following our previous work1, 13, we annotated our datasets using more than 30 feature groups that could be predictive of pathogenicity (a detailed description of these feature groups can be found in the Supplementary):
-
Genomic: genomic features included GC content, sequence spectra16, repeat regions and measures of region uniqueness.
-
Evolutionary: as in previous work, evolutionary features included a comprehensive set of conservation-based measures provided by tools such as Phastcons17, PhyloP18 and FATHMM1.
-
Consequences (coding only): using the Variant Effect Predictor19 we used binary vectors to represent allele consequences and the affected amino acids within all transcripts associated with a mutation.
In addition to these feature groups, we developed new features specifically to boost performance for our non-coding predictor, though similar features proved effective in both coding and non-coding regions (see Supplementary Methods for a complete description). Briefly, these new features fall into two categories: sequence features designed to capture disruption that occurs in a sequence as a consequence of a mutation, and genomic context features that encapsulate genomic features within a mutation’s neighborhood, such as known splice sites or start/stop codons. Recent studies have identified regulatory elements, such as transcription factor binding sites (TFBS), as potential targets of oncogenic mutations in non-coding regions20. Consequently at least one method, FunSeq2, already incorporates motif-specific analysis to score mutations in non-coding regions21. The COSMIC database also provides a set of mutational signatures that are specific to oncogenic mutations. These are associated with various distinct forms of mutation, such as DNA replication errors, defective DNA repair, enzymatic DNA modification, and exposure to mutagens22. However, this signature set is relatively new and hence may represent only a subset of potential oncogenic driver signals. Further, the metrics used to develop these signatures are based in part on the COSMIC database and potentially could introduce bias into our models.
CScape models
We evaluated all models using LOCO-CV, omitting mitochondrial and allosomal (X and Y) chromosomes from testing as these tended to yield few examples, and we wanted to focus on performance within autosomal chromosomes. For each fold we leave out one test chromosome while the remaining 21 chromosomes are used to train the model, using the same model parameters for all folds. Except where noted, we trained models using randomly selected, balanced sets of 1,000 positive and 1,000 negative examples. This relatively small subset of examples yields accuracy nearly as high as with complete training sets but takes less time to train, and allows us to estimate the variability of the test results. For testing we used all available examples for the left-out chromosome, resulting in unbalanced test sets for coding but nearly balanced test sets for non-coding (Supplementary Table S1). For this reason we report balanced accuracy for all tests.
We integrated data from the feature groups outlined above and used them to train two distinct classifiers: CS-coding for coding regions, and CS-noncoding for noncoding regions. The simplest kernel method for integrating different data sources is to combine features from all sources into a single kernel. This allows a model to learn how features from one source may interact with features from other sources. Given at least 30 possible data sources, the number of possible combinations of feature groups makes exhaustive testing impractical. Instead, we use an approach similar to previous work in which we found that greedy sequential learning could be an effective means to identify an optimal combination of groups13. To identify the data sources to include in each model, we first build a model based on two top-ranked data sources combined into a single kernel and record its balanced test accuracy in LOCO-CV. We build subsequent models by adding data sources in descending order of their individual balanced accuracy in LOCO-CV. We select as our final combination those data sources associated with a peak or a plateau in LOCO-CV balanced accuracy (Supplementary Figure S6). We evaluate all combinations with and without data normalisation, where we standardise features by subtracting the mean and dividing by the standard deviation. For these models we observed no difference in performance between the raw feature values and standardised data. The final CS-noncoding model includes just five feature groups: 46-way conservation, 100-way conservation, Spectrum, Genomic context and Mappability (Supplementary Table S4, Supplementary Figure S6). For CS-coding the best model also included five feature groups, but a slightly different set: VEP (including amino acid substitutions), 46-way conservation, 100-way conservation, Genomic context and Spectrum.
In previous studies we found that integration at the kernel level, using multiple kernel learning (MKL)23, 24 and sequential learning strategies, tended to outperform integration at the data level13, 25. We thus applied our sequential learning pipeline13 to examples in both coding and non-coding regions: we ranked feature groups by balanced validation accuracy and added them incrementally to aggregate kernels. For non-coding examples, the best MKL aggregates achieved just over 58% balanced accuracy (Supplementary Figure S7, left). For coding examples, the best achieved over 65% balanced accuracy (Supplementary Figure S7, right). However, in both cases, data-level integration yielded greater balanced accuracy than kernel-level integration.
Both of our models output probability scores on [0, 1]: scores below 0.5 are predicted as negative and scores above 0.5, positive (see Methods). These scores (herein called p-scores) lead to a natural interpretation, such as identifying p-scores above 0.95 as high-confidence predictions. The two models exhibit markedly different p-score distributions across the genome (Fig. 1, left): CS-coding yields p-scores in a bi-modal distribution with peaks at 0.18 and 0.91, demonstrating that a substantial portion of its predictions will be highly confident. By contrast, the CS-noncoding model makes few confident predictions, with a majority of p-scores clustered close to 0.6. When we examine the relative distributions of positive and negative examples from the LOCO test sets, the differences between the coding and non-coding models becomes clearer. CS-coding p-score distributions are markedly different between positive and negative examples, exhibiting some separation between the classes (Fig. 1, middle). By contrast, CS-noncoding p-score distributions are highly similar (Fig. 1, right). We have identified feature groups that enable the non-coding model to discriminate between classes with greater balanced accuracy than previous methods (Fig. 2), but achieving clear separation remains challenging. We note that a small proportion of non-coding predictions carry high confidence. Multiplied by the large number of non-coding positions in the human genome, this translates into a large population of high-confidence predictions in non-coding regions.
Distributions of CScape p-scores reveal distinct differences in the behaviours of the CS-coding and CS-noncoding models. Left: CS-coding tends to make more confident predictions, as its p-scores exhibit a bi-modal distribution with peaks at 0.18 and 0.91. By contrast CS-noncoding makes few confident predictions, attesting to the difficulty in discriminating between oncogenic and benign mutations in these regions. Middle: p-score distributions for positive and negative examples in the CS-coding LOCO test set are substantially different, further demonstrating that the classes are separable to some degree. Right: p-score distributions for CS-noncoding are highly similar; despite careful selection of informative feature groups, clear differences between the classes remain elusive.
ROC curves for COSMIC data show CScape’s performance relative to other state-of-the-art models in coding regions (left & middle) and non-coding regions (right). Left: CS-coding compares favourably with other methods specifically designed to predict SNVs in the coding regions of the cancer genome. Middle: CS-coding also exhibits better performance than popular general-purpose classifiers when tested on putative oncogenic examples. Right: In non-coding regions we also see performance gains using CS-noncoding, here compared with three general purpose predictors, FATHMM-MKL1, CADD7, DANN8 and cancer-specific FunSeq221.
Applying LOCO-CV to the COSMIC training data, we obtained the performance presented in Table 1 and Fig. 2. Performance at the default threshold is good relative to alternative methods, though overall balanced accuracy (62% for non-coding mutations and 73% for coding mutations) leaves room for improvement, also reflected in the corresponding Matthews correlation coefficients (MCC) of 0.25 and 0.46, respectively. Cautious classification (see below) improves these measurements substantially, raising balanced accuracy by 14 percentage points for the non-coding model and by 19 percentage points for the coding model, and doubling (non-coding) or nearly doubling (coding) the MCC.
Comparison with other Methods
Cancer specific predictors have been proposed for prediction in coding regions of the cancer genome4, 5, 26. General purpose predictors have also been proposed for prediction across the entire genome (coding and non-coding regions) using catalogued disease-drivers across a variety of disease traits (e.g. HGMD2). However, there is currently a dearth of predictors specifically trained for the non-coding regions of the cancer genome. In Fig. 2 we present ROC curves demonstrating that CS-coding and CS-noncoding both outperform rival prediction tools on the COSMIC data. Figure 2 (left) shows CS-coding compared with cancer-specific classifiers (e.g., PolyPhen24 and SIFT5 scores adjusted for somatic mutations via TransFIC27). In Fig. 2 (middle) we compare CS-coding with well established general-purpose pathogenic mutation classifiers (e.g., CADD7, DANN8). In Fig. 2 (right) we compare CS-noncoding with both general-purpose classifiers (FATHMM-MKL1, CADD7, DANN8) and the cancer-specific method FunSeq221. With an area-under-curve measure (AUC) of 0.67 against its nearest competitor with 0.59 (our FATHMM-MKL1, 9), CS-noncoding markedly outperforms these existing methods. The CScape tool at the accompanying website incorporates both CS-coding and CS-noncoding.
Cautious Classification
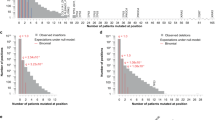
By associating a confidence measure with each prediction, we can evaluate balanced test accuracy against possible confidence measure cutoffs. For each cutoff threshold τ ∈ [0.5, 1] we accept a subset of predictions where p ≥ τ are positive (drivers) and p ≤ (1 − τ) are negative (passengers). We compare these refined sets of predictions to other models in the Supplementary. Figure 3 shows balanced accuracy as a function of p-score cutoff for the coding (left) and non-coding (right) predictors in LOCO-CV, along with the proportion of examples whose p-scores exceed the cutoff. For CS-coding (left), the peak balanced test accuracy is 91.74% which is achieved for a cutoff on the confidence measure at τ = 0.89. With the data used in cross-validation, 15.62% of test examples (5,038 out of 32,253) had a high enough confidence for prediction. Taken across the entire genome, 17.7% of locations in coding regions have a predicted label at this level of confidence. For CS-noncoding (right) the curve has two peaks, as balanced accuracy declines for thresholds between τ = 0.70 and τ = 0.81. In this region small sample size effects begin to influence evaluation of the balanced test accuracy. We choose the lower of the two peaks for the threshold as higher p-scores are of decreasing reliability. At τ = 0.70 the balanced test accuracy is 76.14% for 18.38% of test examples (24,209 out of 131,714). Taken across the entire genome 14.8% of p-scores in non-coding locations had values greater than 0.70.
Cautious classification curves are used to select thresholds that maximise each model’s balanced accuracy. Solid curves show balanced test accuracy (y-axis) as a function of the p-score cutoff (confidence, x-axis). Dashed curves show the proportion of examples in the test datasets where such predictions could be made. Left: For CS-coding, the peak test accuracy is 91.7%, at a p-score of 0.89, with predictions for 15.6% of examples. Right: For non-coding prediction (intergenic and intragenic) the peak test accuracy is 76.1% at a p-score of 0.70, with 18.4% of examples predicted. Beyond the peak, the curve is of decreased reliability since the test set becomes very small.
Evaluation on previously unseen non-coding examples
To evaluate our models on previously unseen examples in non-coding regions, we downloaded putative somatic mutations from two curated databases: The Cancer Genome Atlas (TCGA)10, and the International Cancer Genome Consortium (ICGC)11. Many of the competing methods evaluated here have been trained in part using 1000 Genomes data, so we use that resource to provide negative examples for our tests. For non-coding regions, we compare CScape with scores from FunSeq2 that was designed specifically for predicting oncogenic mutations in non-coding regions21. In addition, we report results for methods capable of predicting pathogenic mutations in both coding and non-coding regions: FATHMM-MKL1, CADD7 and DANN8. We evaluate these methods on a population of randomly-selected test sets and compute the average balanced accuracy, area under ROC curve (AUC), sensitivity, specificity, Matthews correlation coefficient (MCC) and positive predictive value (PPV, see Supplementary for definitions). Testing on randomly selected sets allows us to estimate the variation in performance for each method and to compare methods in a disciplined manner. As a control to establish baseline statistics, we use the CScape scores for each test set and shuffle the labels. To evaluate the other methods, we apply different thresholds to select positive or negative predictions (see Score thresholds in the Supplementary for full details).
Few oncogenic single-point mutations have been verified in non-coding regions. The most prominent to date are three mutations in the TERT promoter region20, 28, 29. These have been characterised as disruptions to putative ETS (E26 transformation specific) family transcription factor binding sites, along with five additional mutations in SDHD and PLEKHS1 20. The three mutations specific to the TERT promoter region are part of the CScape training set, and hence one would expect them to be correctly predicted as oncogenic, but the remaining five mutations were previously unseen. For this small number of documented driver mutations, the CScape non-coding predictor yielded positive predictions for all but one, (Table 2). This one exception is notable as the only one of the eight whose mutation falls outside the core response element20.
The other methods tested also predict a majority of these mutations correctly. For CADD scores we associate negative (−) and positive (+) predictions with negative and positive scores, respectively. For the other methods, we associate negative predictions with scores below 0.5 and positive predictions for the rest. DANN performs as well as CScape on these examples, mis-classifying just one of the mutations. CADD and FunSeq2 mis-classify two mutations, both from the PLEKHS1 gene. FATHMM-MKL (FMKL) mis-classifies the PLEKHS1 mutations as well as two of the SDHD mutations.
International Cancer Genome Consortium (ICGC) data
Within the ICGC data, we found 1,633,449 examples for testing in non-coding regions after we applied our strict filtering criteria and removed examples found in our non-coding training data. For our negative examples from the 1000 Genomes dataset we used examples within 1,000 positions of a positive example and removed the remainder from our training set. This yielded more than 3 million examples, so to achieve a balanced test set, we sampled 1,633,449 of these for our test set. Hence the final ICGC test set contains 3,266,898 examples. CScape yields significantly higher balanced accuracy than competing methods on these data (p < 10−7, Mann-Whitney) even at the default threshold (Fig. 4 (top), Supplementary Table S7). Accuracy at the default threshold is 65%, while FATHMM-MKL yields 54%. Other metrics are similarly strong, with the exception of specificity. At the cautious classification threshold, all statistics improve considerably, with balanced accuracy and AUC scores well over 80%, whilst yielding predictions for 640,312 test examples (19.6%).
CScape yields state-of-the-art performance on unseen examples in non-coding regions. Left: On the ICGC test set, CScape significantly outperforms competitors for all performance measurements except specificity. Right: On the BRCA test set CScape yields balanced accuracy (70%) and AUC (0.84) that are similar to its nearest competitor (FATHMM-MKL, 71% and 0.80, respectively). Error bars indicate a 99% confidence interval for statistics measured across 20 test sets. †For FunSeq2 we report results at its peak balanced accuracy threshold, as described in the Supplementary.
The Cancer Genome Atlas (TCGA) breast cancer dataset
Within the TCGA database we found a small set of 3,368 examples in non-coding regions having a Valid status. The vast majority of these appear in our training set, but 360 examples, all associated with breast cancer (BRCA), were previously unseen. For non-coding tests, we select neutral examples from the 1000 Genomes database in the same way as we did for the ICGC data, by using examples that fall within 1,000 positions of some positive example. After removing examples found in our training set, we had a test set with 3,206 examples (360 positive, 2,846 negative). We compared the CScape non-coding predictions with those of four other methods relevant to non-coding examples: our FATHMM-MKL, CADD, DANN and FunSeq2. On these data CScape at the default threshold is competitive with current state-of-the-art, with average balanced accuracy comparable to FATHMM-MKL and higher than other methods (Fig. 4 (bottom), Supplementary Table S8). At the cautious classification threshold CScape yields higher scores than competitors in all categories except specificity, whilst producing predictions for 795 examples (24.8%).
Evaluation on previously unseen coding examples
To evaluate our models on unseen examples in coding regions, we downloaded putative somatic mutations from two additional curated databases: the Database of Curated Mutations (DoCM)30, and ClinVar31. The DoCM represents a refined subset of examples from the other databases, hence we remove the DoCM examples from the other test data so that we may treat each data set independently. For coding regions, we compare CScape with scores from TransFIC27, which provides scores from MutationAssessor (MAS)6, PolyPhen24 and SIFT5 that have been optimised for predicting oncogenic mutations. As we did in non-coding regions, we also report results for general-purpose predictors FATHMM-MKL, CADD and DANN.
Database of Curated Mutations (DoCM) data
The Database of Curated Mutations (DoCM) represents a carefully curated set of 1,364 somatic mutation examples, including 1,223 distinct SNVs30. Of these, 674 SNVs appeared in our training set and 551 were previously unseen. To create a test set, we found 289 germline mutations in the 1000 Genomes database within 10,000 positions of some DoCM example, of which 173 were absent from the CScape training set. This comprised an unbalanced test set we used to compare CScape’s predictions with other classifiers for coding-region cancer mutations (Fig. 5(a), Supplementary Table S9). Given the relatively small data set, we used N = 10 test samples to generate statistics. At the default threshold, CScape’s balanced accuracy averaged over the 10 sets is 88%, significantly higher than its closest competitor, FATHMM-MKL, at 83% (p < 0.012, Mann-Whitney). At the cautious classification threshold (p-scores below 0.11 or above 0.89) CScape makes predictions for a total of 275 examples (38% overall), but only eight of these are negatives, four of which yield false-positives. This severe imbalance yields extremely low balanced accuracy (see Fig. 3) and high variance for any combination of test sets, but sensitivity is consistently 100% and the PPV is 99%, providing evidence that p-scores above this threshold correctly identify oncogenic examples.
CScape exhibits strong performance on unseen examples in coding regions. (a) For 724 unseen DoCM examples, CScape tends to outperform other methods, with balanced accuracy over 80% averaged over 10 test sets, compared with 75% for FATHMM-MKL, its nearest competitor. (b) For 5,642 unseen ClinVar examples, CScape yields significantly higher values than other methods for all statistics except AUC and Sensitivity. (c) For 108,874 unseen TCGA examples, CScape yields balanced accuracy that is competitive with its closest competitor, FATHMM-MKL. (d) For 92,252 unseen ICGC examples, CScape yields slightly lower balanced accuracy than its top competitor (65% versus 68% for FATHMM-MKL), while PolyPhen2 yields much higher average PPV. Error bars indicate a 99% confidence interval for statistics measured across 20 test sets.
ClinVar
The ClinVar database contains both germline and somatic variants, hence we used annotations to identify somatic mutations. Within its extensive metadata, ClinVar mutations include a clinical significance (CLNSIG) that signifies the likelihood that a mutation is pathogenic, and a variant disease name (CLNDBN) that we use to identify somatic variants. For our test set, we selected SNVs with a CLNSIG designation of pathogenic or likely pathogenic for our positive examples, and those with a benign or likely benign designation as negatives. From these, we selected examples with a CLNDBN containing tumor or cancer designations and filtered out any examples found in our model’s training set. This yielded an unbalanced set of 5,642 examples (1,186 positive, 4,456 negative) for testing.
Results (Fig. 5(b), Supplementary Table S10) show that CScape at the default threshold yields significantly higher average balanced accuracy, PPV and MCC values than the other methods (p < 10−7, Mann-Whitney), and a significantly higher average AUC (p < 10−6) than all methods except CADD. The cautious classification threshold yields similarly strong results, but with significantly higher MCC and PPV even than CScape (p < 10−7, Mann-Whitney) at the default threshold.
The Cancer Genome Atlas (TCGA) data
TCGA data includes a validation status we use to remove low-confidence examples. For our test set, we retained 66,441 out of 4.2 million TCGA examples with a Valid status. Of these, we removed 1,898 that were in the CScape training set, leaving 64,543 positive examples. As in our other tests, we selected neutral examples from the 1000 Genomes database, using examples that fall within 10,000 positions of some positive example. Of 41,319 examples selected from 1000 Genomes, we removed 9,835 that were also in the CScape training set. This left 31,484 negative examples in our test set, for a total of 109,085.
We compared CScape’s performance on the TCGA data with six other methods relevant to coding-region predictions (Fig. 5(c), Supplementary Table S11). At the default threshold (τ = 0.5), CScape yields higher balanced accuracy, MCC and specificity scores than the other methods on these data. In addition, CScape’s PPV and AUC scores are competitive with the best alternatives. CADD and DANN, modeled from the same training set, yield the highest sensitivity but the lowest specificity. At the peak threshold τ = 0.89 established for cautious classification, CScape makes confident predictions for 17,801 test examples (16.3%) and yields the highest performance statistics in all categories.
International Cancer Genome Consortium (ICGC) data
We downloaded 61 mutation files from ICGC containing 47 million distinct single-nucleotide variants (SNVs). ICGC annotates mutations using four different verification statuses: tested and verified, tested and verified to be false, tested and inconclusive and not tested. For positive test examples we selected those having tested and verified for both verification and their biological validation statuses, and we required that examples were shared by at least two distinct donors (donor id). After removing examples found in our coding-region training data, we had 42,515 positive examples. We found just 106 distinct ICGC SNVs with the annotation tested and verified to be false, but none of them met our filtering criteria. Hence as in other tests, we selected as a negative test set examples from 1000 Genomes within 10,000 positions of some positive example and removed those found in our training set. Our final ICGC test set contains 92,732 examples (42,174 positive, 50,558 negative).
For these unseen ICGC examples we find that CScape using the default threshold (0.50) is competitive with the other methods tested, though on these data it slightly under-performs its top competitor, FATHMM-MKL (Fig. 5(d), Supplementary Table S12). In particular, the cancer-optimised PolyPhen2 scores yield an exceptionally high average PPV in our tests. With cautious classification threshold, CScape yields substantially higher balanced accuracy, MCC, PPV and AUC scores than all other methods, producing predictions for 13.1% of examples.
Mis-classified test examples
Despite CScape’s strong performance on both coding and non-coding test examples, we investigated mis-classified examples to see if patterns emerged. To this end we used the Variant Effect Predictor (VEP) to identify the regions that may be different between correctly classified and mis-classified test examples (Supplementary Figures S8–S9). We first note that any mutation may impact multiple transcripts, hence the same mutation may have multiple annotations in the VEP database. For our coding classifier, we find the majority of the 627 mis-classified ClinVar examples are annotated as missense mutations for at least one transcript (Supplementary Figure S8). while the majority of correctly classified examples are annotated as synonymous. In our other test sets, there appear to be no strong trends though we do find a slightly higher proportion of correct predictions amongst mis-sense mutations. The non-coding classifier shows little sign of classifying one kind of example better than another (Supplementary Figure S9). On the relatively small BRCA test set (3,206 examples) the model tends to mis-classify examples designated NC transcript and Noncoding exon at a slightly higher rate than others, but on the larger TCGA test set (3.3 million examples) the model shows no preference for any particular gene characteristic.
Cancer-specific thresholds for competitors
Using the default thresholds for all methods, we find that CS-noncoding outperforms competitors on all of our benchmark tests, and that CS-coding either outperforms competitors or yields comparable balanced accuracy. However, several of these other methods (notably CADD, DANN and FATHMM-MKL) have not been optimised for oncogenic mutations. Therefore, we used our training data to identify alternative thresholds for all competing methods across their prediction domains (coding, non-coding or genome-wide) and assessed the potential change in their performance (Supplementary Methods). Overall, results were mixed, with CADD and DANN improving marginally in all regions, while FATHMM-MKL generally performed worse. We also identified region-specific thresholds using either coding or non-coding training sets, and found significant improvements for both CADD and DANN. Hence it may be possible to apply non-cancer-specific methods to cancer variants, provided one can establish appropriate thresholds. However, we believe this requirement would be onerous for most users, and reiterate that CScape achieves similar or higher accuracy at its default threshold.
Regions of Interest
Our Regions of Interest method highlighted multiple segments within the exons of genes as potentially functional as disease-drivers. Since our classifiers were trained on COSMIC data across multiple cancer types, highlighted genes tend to be of broader significance across multiple tumour types, such as the tumour suppressor gene TP53, rather than genes with known associations with a particular type of cancer. We provide files listing these regions of interest, in both coding and non-coding regions, at the CScape website.
Discussion
A stated top balanced test accuracy of 91.7% in coding regions has been made possible through use of a broad set of data sources, with subsequent use of cautious classification to define a more accurate predictor, but with more limited applicability (this accuracy can only be achieved at 17.7% of nucleotide positions). Without cautious classification, CScape yields competitive accuracy in both coding and non-coding regions, but its performance in non-coding regions excels in our tests when compared with other methods. We note that both models, CS-coding and CS-noncoding, benefit from similar but distinct feature groups. In particular, CS-noncoding benefits from the Mappability group that measures the uniqueness of a region, possibly allowing the model to learn the relative oncogenic susceptibility of repeat regions or paralogous sequences. Predictably, CS-coding benefits largely from predicted amino acid substitutions and sequence conservation measures. The positive predictive values (PPV) and numbers of true positives predicted (Table 1) suggest that accurate prediction of positives is achievable, and that the balanced test accuracy is not overwhelmingly achieved through accurate prediction on neutral variants. Currently the paucity of validated oncogenic variants in non-coding regions is a limiting factor in creating accurate predictors for those regions; as more of these examples are confirmed, subsequent models should improve. In addition, it should be possible to improve on these results across the entire cancer genome, via identification of further sources of data relevant to predicting disease-driver status, the use of adaptive recurrences thresholds r in different regions of the coding and non-coding parts of the genome, and the development of cancer-type specific predictors e.g. for breast cancer or prostate cancer (in previous work32 we have established this leads to improved balanced test accuracy). Similarly, we have shown that some competing methods may benefit from using cancer-specific thresholds, given an appropriate data set for evaluating their accuracy at different thresholds. However, it is doubtful that users will be willing to engage in extensive post-processing, and the resulting predictions are unlikely to yield balanced accuracy significantly better than CScape’s.
One of our goals in this study was to exploit the diverse databases emerging from the ENCODE resource. Many of these databases are sparse: some of these feature groups have values for only a tiny proportion of locations across the genome. In this study we explored two data integration methods that work well with whole feature groups: data-level integration, in which we merge groups into a single kernel, and kernel-level integration in which we treat each group as a separate kernel and use MKL to combine them. However the sparsity of many feature groups currently limits their utility in these paradigms. This sparsity could be due to the constraints of the underlying assays, such as expression data obtained for specific conditions, or due to data that are inherently sparse, such as transcription factor binding sites. Regardless of the cause, more sophisticated data integration methods will likely be required to exploit them fully.
As this paper was in preparation, evidence emerged demonstrating that a substantial number of the examples in mutation databases are probably spurious, reflecting damaged DNA rather than mutations33. We mitigate this issue to some extent by using recurrence counts or validation status to filter positive examples, but negative examples could be more severely impacted. Hence we should view the statistics reported here with some scepticism, as the models we have examined have been trained and tested on putative oncogenic driver mutations and, possibly, a mixture of benign mutations and damaged DNA. Only once spurious examples have been largely eliminated from mutation databases will we know whether the added noise renders the discrimination problem more or less difficult than reported here.
Methods
Model selection
All experiments were written in Python (v2.7.6) using classification models from PyML 34 (v0.7.13.3) and scikit-learn 35 (v.0.17). We use LOCO-CV to evaluate a variety of different classification models, to establish the best overall model, using balanced accuracy as our criterion. In each CV fold, we select a training set at random from 21 chromosomes, hence performance will change from one run to the next. For this reason, we ran cross validation 30 times for each model, and use the average balanced accuracy to compare models. Many classifiers have parameters that require tuning, such as a regularisation parameter. To find the best parameter for each model, we selected a range of values (C = 10−4 to 104, in the case of SVM) and ran LOCO-CV 10 times for each value, selecting the parameter that yielded the best average balanced validation accuracy for the model (see Supplementary text and Supplementary Table S5 for more details). We found that Gradient Boosting36 yielded the strongest performance overall, though several models yielded balanced accuracy nearly as high (Supplementary Table S5). In particular, support-vector machines with Gaussian kernels were competitive, facilitating experiments with multiple kernel learning (MKL).
Establishing thresholds
Our classifiers produce scores for every possible mutation, where scores may take on any value in \({\mathbb{R}}\). A common approach for binary classification is to apply a threshold of 0, and predict values below the threshold as negative and those above as positive. An alternative method is to map output scores to a probability distribution on [0, 1] and use a threshold of 0.5. In this work we use Platt scaling37 to associate a confidence measure with each predicted class label, which we interpret as the probability that a particular SNV is a cancer driver. By establishing a suitable confidence threshold we can also restrict prediction to a subset of higher balanced-accuracy predictions (cautious classification). This leads to an alternative, high-confidence threshold for our models that yields improved balanced test accuracy, but at the expense of prediction on a subset of nucleotide positions.
Regions of interest
The resultant positive (oncogenic) predictions are frequently clustered by genomic location. Thus mutations in regions that are highly conserved across species are typically more deleterious than mutations in regions exhibiting high inter-species variability: the motivation for using sequence-conversation features for the classifier. This fact alone should give rise to significant clustering of positive predictions in highly conserved regions. To isolate regions of clustered positive predictions, which we call Regions of Interest (RoIs), we determined the distribution of confidence values and labeled a region as a RoI if the contiguous set of positive confidence values located in this region had a very low expectation of being observed, relative to the overall distribution. Full details on the methods used to identify these regions are given in the Supplementary Methods.
Data availability
CScape predictions are available as an online resource http://CScape.biocompute.org.uk. In addition, a predictions database, software for querying the database, and training and test datasets are all available for download from the website.
References
Shihab, H. et al. An integrative approach to predicting the functional effects of non-coding and coding sequence variation. Bioinformatics 31, 1536–1543 (2015).
Stenson, P. D. et al. The human gene mutation database: building a comprehensive mutation repository for clinical and molecular genetics, diagnostic testing and personalized genomic medicine. Hum. Genet. 133, 1–9 (2014).
The 1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Adzhubei, I. et al. A method and server for predicting damaging missense mutations. Nature Methods 7, 248–249 (2010).
Kumar, P., Henikoff, S. & Ng, P. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–81 (2009).
Reva, B., Antipin, Y. & Sander, C. Predicting the functional impact of protein mutations: application to cancer genomics. Nucleic Acids Research 39, e118 (2011).
Kircher, L. A. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nature Genetics 46, 310–315 (2014).
Quang, D., Chen, Y. & Xie, X. DANN: a deep learning approach for annotating the pathogenicity of genetic variants. Bioinformatics 31, 761–763 (2014).
Liu, X., Li, C. & Boerwinkle, E. The performance of deleteriousness prediction scores for rare non-protein-changing single nucleotide variants in human genes. Journal of Medical Genetics jmedgenet–2016 (2016).
Weinstein, J. N. et al. The cancer genome atlas pan-cancer analysis project. Nature genetics 45, 1113–1120 (2013).
Zhang, J. et al. International cancer genome consortium data portal – a one-stop shop for cancer genomics data. Database 2011 (2011).
The 100,000 genomes project protocol v3, genomics england. http://www.genomicsengland.co.uk/ (2017).
Rogers, M. et al. Sequential data selection for predicting the pathogenic effects of sequence variation. In Bioinformatics and Biomedicine (BIBM), 2015 IEEE International Conference on, 639–644 (2015).
Ritchie, G., Dunham, I., Zeggini, E. & Flicek, P. Functional annotation of noncoding sequence variants. Nat. Methods 11, 294–296 (2014).
Leslie, C. S., Eskin, E. & Noble, W. S. The spectrum kernel: A string kernel for SVM protein classification. In Pacific Symposium on Biocomputing, vol. 7, 566–575 (World Scientific, 2002).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Research 15, 1034–1050 (2005).
Pollard, K. S., Hubisz, M., Rosenbloom, K. & Siepel, A. Detection of non-neutral substitution rates on mammalian phylogenies. Genome Research 20, 110–121 (2010).
McLaren, W. et al. The ensembl variant effect predictor. Genome biology 17, 122 (2016).
Weinhold, N., Jacobsen, A., Schultz, N., Sander, C. & Lee, W. Genome-wide analysis of noncoding regulatory mutations in cancer. Nature Genetics 46, 1160–1165 (2014).
Fu, Y., Liu, Z., Lou, S., Bedford, J., Mu, X.J., Yip, K.Y., Khurana, E. and Gerstein, M., FunSeq2: a framework for prioritizing noncoding regulatory variants in cancer. Genome Biology 15 (2014).
Wellcome Trust Sanger Institute. COSMIC: Signatures of Mutational Processes in Human Cancer. http://cancer.sanger.ac.uk/cosmic/signatures.
Campbell, C. & Ying, Y. Learning with Support Vector Machines (Morgan and Claypool, 2011).
Gönen, M. & Alpaydn, E. Multiple kernel learning algorithms. Journal of Machine Learning Research 12, 2211–2268 (2011).
Shihab, H., Rogers, M., Gaunt, T. & Campbell, C. HIPred: an integrative approach for predicting haploinsufficiency in the human genome. in press (2017).
Wong, W. et al. CHASM and SNVBox: toolkit for detecting biologically important single nucleotide mutations in cancer. Bioinformatics 27 (2011).
Gonzalez-Perez, A., Deu-Pons, J. & Lopez-Bigas, N. Improving the prediction of the functional impact of cancer mutations by baseline tolerance transformation. Genome medicine 4, 1 (2012).
Huang, F. W. et al. Highly recurrent tert promoter mutations in human melanoma. Science 339, 957–959 (2013).
Horn, S. et al. Tert promoter mutations in familial and sporadic melanoma. Science 339, 959–961 (2013).
Ainscough, B. J. et al. DoCM: a database of curated mutations in cancer. Nature Methods 13, 806–807, https://doi.org/10.1038/nmeth.4000 (2016).
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Research 42, D980–D985 (2014).
Shihab, H. et al. Ranking non-synonymous single nucleotide polymorphisms based on disease concepts. Human Genomics 8, 11 (2014).
Chen, L., Liu, P., Evans, T. C. & Ettwiller, L. M. DNA damage is a pervasive cause of sequencing errors, directly confounding variant identification. Science 355, 752–756 (2017).
Ben-Hur, A. PyML: PyML - machine learning in Python. http://pyml.sourceforge.net/ (2010).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. Journal of Machine Learning Research 12, 2825–2830 (2011).
Mason, L., Baxter, J., Bartlett, P. & Frean, M. Boosting algorithms as gradient descent in function space (NIPS, 1999).
Platt, J. Probabilistic outputs for support vector machines and comparison to regularised likelihood methods. In Advances in large margin classifiers 61–74 (MIT Press, 1999).
Acknowledgements
This work was supported by Medical Research Council grants MC_UU_12013/8 and G1000427/1 and Cancer Research UK grant C18281/A19169. M.R. was supported by an EPSRC grant (EP/K008250/1).
Author information
Authors and Affiliations
Contributions
M.R. had primary responsibility for developing the predictor and H.S. contributed through resourcing of data for the project. T.G. and C.C. oversaw the project and contributed to preparing the paper along with M.R.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rogers, M.F., Shihab, H.A., Gaunt, T.R. et al. CScape: a tool for predicting oncogenic single-point mutations in the cancer genome. Sci Rep 7, 11597 (2017). https://doi.org/10.1038/s41598-017-11746-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-11746-4
This article is cited by
-
The mutational landscape of a US Midwestern breast cancer cohort reveals subtype-specific cancer drivers and prognostic markers
Human Genomics (2023)
-
Clinically actionable cancer somatic variants (CACSV): a tumor interpreted dataset for analytical workflows
BMC Medical Genomics (2022)
-
A bioinformatics approach to the identification of novel deleterious mutations of human TPMT through validated screening and molecular dynamics
Scientific Reports (2022)
-
Impact of deleterious missense PRKCI variants on structural and functional dynamics of protein
Scientific Reports (2022)
-
GhostKnockoff inference empowers identification of putative causal variants in genome-wide association studies
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.