Abstract
A considerably small fraction of approximately 60–100 proteins of all chloroplast proteins are encoded by the plastid genome. Many of these proteins are major subunits of complexes with central functions within plastids. In comparison with other subcellular compartments and bacteria, many steps of chloroplast protein biogenesis are not well understood. We report here on the first study of chloroplast-localised trigger factor. In bacteria, this molecular chaperone is known to associate with translating ribosomes to facilitate the folding of newly synthesized proteins. Chloroplast trigger factors of the unicellular green algae Chlamydomonas reinhardtii and the vascular land plant Arabidopsis thaliana were characterized by biophysical and structural methods and compared to the Escherichia coli isoform. We show that chloroplast trigger factor is mainly monomeric and displays only moderate stability against thermal unfolding even under mild heat-stress conditions. The global shape and conformation of these proteins were determined in solution by small-angle X-ray scattering and subsequent ab initio modelling. As observed for bacteria, plastidic trigger factors have a dragon-like structure, albeit with slightly altered domain arrangement and flexibility. This structural conservation despite low amino acid sequence homology illustrates a remarkable evolutionary robustness of chaperone conformations across various kingdoms of life.
Similar content being viewed by others
Introduction
The transformation of the one-dimensional genetic information into complex protein structures is a challenging task for cells. Polypeptides that emerge vectorially from translating ribosomes cannot fold completely to the native conformation unless the polypeptide chain is fully synthesized and released from the ribosome. During this process, nascent polypeptides are highly susceptible to premature folding, misfolding, and aggregation1. In addition, polypeptides synthesized on cytosolic ribosomes and destined for organelles need to be recruited early and kept in an unfolded state to facilitate their translocation across subcellular membranes2. Thus, cells have acquired various factors associating with translating ribosomes to assist protein maturation, to prevent premature folding, and to mediate membrane targeting for translocation3, 4. Many of these factors belong to the abundant and structurally diverse family of molecular chaperones, which act at all stages of a proteins life span to promote a balanced protein homeostasis1. Across all kingdoms of life, specific molecular chaperones associate with translating ribosomes to guide the initial steps of de novo folding of nascent polypeptides5. Further downstream, other chaperones take over nascent polypeptides to assist folding and final maturation. In bacteria, the ATP-independent trigger factor (TF) is the predominant chaperone that transiently binds to ribosomes. This association is mediated by the ribosomal protein Rpl23, which positions TF next to the ribosomal exit tunnel for early binding of nascent polypeptides6. A comprehensive analysis of TF substrates indicated that the chaperone binds most newly translated polypeptides7. In fact, E. coli cells contain a two- to three-fold molar excess of TF relative to ribosomes, thus providing sufficient chaperones to act on all translating ribosomes6, 8,9,10.
High-resolution structural analyses of various bacterial TF molecules showed that TF adopts a unique elongated conformation resembling a “crouching dragon” with three domains11. The N-terminal domain is responsible for ribosome association and contains an essential signature motif for this interaction. This domain further supports the C-terminus to shape the backbone structure of TF. The C-terminal domain is the main module of TF and possesses in vitro chaperone activity on its own12. This domain folds back to interact with the N-terminus and forms the backbone of TF with an open cavity and two protruding arms. Within the amino acid sequence, both termini are separated by the peptidyl–prolyl cis–trans isomerase (PPIase) or so-called head domain, which is situated opposite of the N-terminus in the three-dimensional structure. In contrast with other molecular chaperones, which contain one specific substrate binding site, TF seems to use multiple sites across the entire cavity for both hydrophobic and hydrophilic substrate interactions11. Thus, a bound substrate is accommodated in a protective environment in the interior of TF to prevent misfolding and aggregation. Various structural data further indicated that the protein is rather flexible, which seems essential for binding of the diverse set of substrates and for ribosome association6. Bacterial ribosomes are bound by the monomeric form, while non-ribosome-bound TF shows a fast monomer–dimer equilibrium with a half-life of the dimer of ∼1 s and a KD of ∼1–2 μM13, 14. For the function of the TF dimer, opposing data have been reported. On the one hand, it was hypothesized that this dimer serves as an inactive storage form. On the other hand, it was found that such dimers contribute to the stabilization of unfolded substrate species14,15,16.
In eukaryotes, chloroplasts are the only subcellular compartments that appear to contain molecular chaperones of the trigger factor family. Chloroplasts confer photoautotrophy to plants and algae. These organelles contain their own semi-autonomous genome, which encodes for approximately 60–100 proteins of all ~3000 chloroplast proteins. Most of the chloroplast-encoded proteins are major subunits of central protein complexes involved in gene expression and photosynthesis17. Since chloroplasts are thought to have evolved from a photosynthetically active cyanobacterium more than a billion years ago, many components of the plastidic gene-expression machinery still resemble their cyanobacterial counterparts18. For example, chloroplasts contain 70 S ribosomes each consisting of a small 30 S and a large 50 S subunit19,20,21,22. Consistently, many components of the chloroplast protein folding and quality control machinery are orthologous to the respective system in prokaryotes23. Importantly, the chloroplast gene-expression apparatus also acquired novel mechanisms that are not found in cyanobacteria, which most likely serve the orchestrated expression of the nuclear, mitochondrial, and chloroplast genomes.
While bacterial TF has been studied extensively and might be the best understood molecular chaperone in the literature, no experimental evidence about the function and biophysical properties of plastidic trigger factor exists to date. Here, we describe biophysical and structural properties of plastidic trigger factors from Chlamydomonas (C. reinhardtii) and Arabidopsis (A. thaliana). Despite a high sequence variance among different chloroplast trigger factor species, small-angle X-ray scattering (SAXS) experiments indicated that the eukaryotic chaperones have an architecture strikingly similar to their bacterial counterpart. Yet, chloroplast trigger factors also show distinct molecular features that might have evolved to meet the functional requirements of the chloroplasts.
Results and Discussion
The chloroplast trigger factor family displays low sequence conservation
Genes orthologous to the bacterial TF can be found in all plastid-containing algae and plants but are absent in non-photosynthetic eukaryotes. Thus, it has been postulated that algae and plants encode for a chloroplast isoform24. To gain a first understanding of these chloroplast-localised chaperones, we compared plastidic trigger factor proteins from C. reinhardtii and A. thaliana with the isoform from E. coli. For simplicity and consistency with previous studies, the full-length chloroplast proteins will be abbreviated TIG1, and the E. coli form will be abbreviated EcTF. Excluding the N-terminal chloroplast transit sequence of the TIG1 proteins, chloroplast and bacterial isoforms have a similar overall length and contain the three typical TF domains (i.e., N-terminal ribosome-binding domain, PPI domain, and C-terminal chaperone domain) (Fig. 1a and b). However, the amino acid composition displays only ~18% sequence identity and <30% sequence similarity between the chloroplast TIG proteins and EcTF. Similarly, CrTIG1 and AtTIG1 also share sequences of only 24% identity and 35% similarity (Fig. 1c). Such low sequence conservation between two phyla is remarkable for molecular chaperones where at least some domains tend to be highly conserved among orthologues25, 26. Our phylogenetic analysis of various trigger factor sequences confirmed this high variability among orthologous forms from bacteria, algae, mosses and higher plants. Interestingly, TIG1 orthologues from diatoms and red algae show a closer homology to prokaryotic TFs, while TIG1 proteins from green algae, mosses, and higher plants fall into a separate clade. Within the green lineage, low sequence conservation is observed between chlorophyte and streptophyte TIG1 proteins (Fig. 1d). It might be speculated that the high variety among TIG1s from the green lineage constitutes an evolutionary adaptation of these proteins according to their task during protein biogenesis. Except for the moss P. patens, all investigated green algal and plant genomes encode only one full-length TIG1 species (Supplementary Table S1). Interestingly, mosses and higher plants seem to encode additional, truncated forms of a trigger factor. We propose to term these orthologous forms TIG2, as transit-peptide analyses predict a chloroplast localisation as well. TIG2 proteins seem to contain only the N-terminal ribosome binding domain. Since the N-terminal domain of bacterial TF is the major site for ribosome binding and exhibits some chaperone activity6, an independent or distinct contribution of this truncated variant to protein biogenesis can be envisioned. It has been shown previously by size exclusion chromatography (SEC) and mass spectrometry of A. thaliana chloroplast extracts that TIG2s are expressed and that they co-migrate in high-molecular weight fractions together with TIG1 and plastid ribosomes27. Phylogenetic comparison of TIG1 and TIG2 sequences reveals a clear separation of the two species (Supplementary Figure S1). TIG2 displays a closer relation to the full-length bacterial orthologues compared with TIG1. Thus, it is possible that, during evolution, two trigger factor orthologues were inherited from cyanobacteria and that TIG2 was subsequently lost in algae. Alternatively, TIG2 might have been acquired through horizontal gene transfer after separation of the algal and land-plant lineages.
Evolutionary diversity of chloroplast trigger factor. (a) Alignment of trigger factor amino acid sequences from Escherichia coli (E.c.), Arabidopsis thaliana (A.t.), and Chlamydomonas reinhardtii (C.r.). Sequences were aligned by ClustalOmega and shaded using BoxShade (http://www.ch.embnet.org). Amino acids highlighted in black are perfectly conserved, similar residues are indicated by a grey background. Boxes indicate chloroplast transit-peptide cleavage sites as determined experimentally (green) and as predicted by TargetP and ChloroP (light red for AtTIG1)50, 51. Dark red box indicates putative ribosome-binding site and lines underneath the sequences indicate the three trigger factor domains according to the E. coli structure11. (b) CrTIG1 and AtTIG1 are predicted to comprise the typical three-domain organization of the N-terminal ribosome-binding domain (red), the PPIase domain (turquois), and the C-terminal domain (blue). (c) Sequence homology comparison of EcTF, CrTIG1, and AtTIG1 according to http://imed.med.ucm.es/Tools/sias.html. (d) Phylogram based on amino acid sequence alignments of trigger factor from gram-negative bacteria (Escherichia coli, Vibrio cholerae, Thermotoga maritima), cyanobacteria (Nostoc sp. strain PCC 7120, Synechocystis sp. strain PCC 6803, Prochlorococcus marinus, Synechococcus sp. strain WH8102), the diatom (Phaeodactylum tricornutum), and the mature TIG1 sequences from red algae (Galdieria sulphuraria, Cyanidioschyzon merolae), green algae (Volvox carteri, Chlamydomonas reinhardtii, Coccomyxa subellipsoidea, Micromonas sp. strain RCC299, Ostreococcus lucimarinus), moss (Physcomitrella patens), and land plants (Zea mays, Setaria italica, Oryza sativa, Brachypodium distachyon, Populus trichocarpa, Brassica rapa, Arabidopsis thaliana, Capsella grandiflora). Bootstrap values are given next to the nodes. For sequence information see Supplementary Table S1; for phylogenetic comparison including truncated TIG2 variants see Supplementary Figure S1. The bar indicates branch length.
Most TIG1 is soluble but also thylakoid-associated
For a biophysical comparison of chloroplast CrTIG1 and AtTIG1 with EcTF, mature proteins lacking the N-terminal chloroplast transit peptide were cloned for overexpression and affinity purification followed by subsequent SEC. This yielded highly pure and fully soluble proteins migrating with respective apparent molecular weights of 56 kDa (EcTIG1), 58 kDa (CrTIG1), and 57 kDa (AtTIG1), which is slightly larger than their theoretical molecular weights (i.e., 48.2 kDa (EcTF), 53.7 kDa (CrTIG1), and 53.2 kDa (AtTIG1)) (Fig. 2a). To test if the transit-peptide cleavage sites of CrTIG1 and AtTIG1 proteins were correctly predicted, purified proteins and cell lysates of C. reinhardtii and A. thaliana were separated by SDS-PAGE and analysed by immunoblot (Fig. 2b). Both endogenous and purified TIG1 migrated with the same velocity supporting the correct assumptions of transit-peptide cleavage sites.
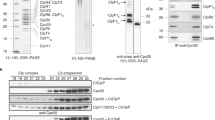
Intracellular localisation of TIG1. (a) EcTF, CrTIG1, and AtTIG1 were heterologously expressed in E. coli, purified via chitin affinity resins and size exclusion chromatography. 0.5 µg or 1 µg of each protein was separated by SDS-PAGE and stained with Coomassie. (b) 7.5 µg or 15 µg of soluble extracts from C. reinhardtii and A. thaliana was separated by SDS-PAGE next to 7.5 ng or 15 ng of purified TIG1 protein, transferred to nitrocellulose, and immunoblotted with antibodies against TIG1 (c). C. reinhardtii chloroplasts (CP) were isolated, lysed by hypo-osmotic shock, and separated into stroma (Str), low-density membranes (LM), and thylakoid membranes (Thyl). Mitochondria (Mito) were separated from the same strain. Whole cells (WC) and 7 µg of each fraction were separated by a 7.5–15% SDS-PAGE, transferred to nitrocellulose, and immunoblotted with antibodies against CrTIG1, chloroplast ribosomal protein PRPL1, HSP70B (stroma and membrane control), CF1β (thylakoid membrane associated), stromal CGE1, and mitochondrial carbonic anhydrase (CA). Note that LM might be over-represented compared to thylakoids. (d) Images of C. reinhardtii cells, stained with antibodies directed against CrTIG1, RbcL, and PRPL1 (FITC, green) and a FISH probe against 70 S ribosomes (red). The numbers of cells with the localisation patterns seen was 15 of 20 (75%) for CrTIG1, 16 of 20 (80%) for RbcL, 13 of 20 (65%) for PRPL1, and 20 of 20 (100%) for FISH. Antibody specificity is shown in Supplementary Figure S2. Scale bars represent 2 µm (e) 20 µg of soluble A. thaliana leaf extract or 20 µg of isolated chloroplasts was separated next to 7.5 ng purified AtTIG1 by SDS-PAGE, transferred to nitrocellulose, and immunoblotted with antibodies against AtTIG1, Actin, and RbcL. Immunoblots in (b, c) and (e) were cropped to the respective size of the displayed bands.
To confirm that CrTIG1 is indeed a chloroplast-localised protein, we performed subcellular fractionation experiments. C. reinhardtii cells were fractionated into mitochondria, chloroplasts, stroma, thylakoids, and thylakoid-associated low-density membranes (LMs)28. The purity and degree of enrichment were tested with antibodies against chloroplast HSP70B, stromal CGE1, thylakoid membrane-associated CF1β, and mitochondrial carbonic anhydrase (CA). CrTIG1 displayed a distribution similar to the chaperone HSP70B26 and was found mainly in the soluble stroma fraction of chloroplasts but is also associated to some extent with LMs and thylakoid membranes (Fig. 2c). Since a major fraction of bacterial TF is ribosome-bound6, the distribution of chloroplast ribosomes was visualised with an antibody directed against a protein of the 50 S subunit, PRPL1. Similar to CrTIG1 and HSP70B, ribosomes were enriched mostly in the stromal fraction but were also detectable in the thylakoid fraction. It has previously been reported that in diurnally growing C. reinhardtii cells, 20–30% of the chloroplast ribosome population is associated with thylakoids29. Here, less than 20% of the ribosomes were found in the thylakoid fractions. However, ribosomes may have dissociated from thylakoids during the long preparation process. For an independent assay of CrTIG1 and PRPL1 distribution, C. reinhardtii cells were examined by immunofluorescence. A hallmark of C. reinhardtii cells is the single cup-shaped chloroplast, which consumes most of the volume within the cell. The chloroplast contains a globular basal region with the pyrenoid and lobes extending apically from the basal region. Chloroplast localisation of CrTIG1 was confirmed by the typical cup-shape staining (Fig. 2d). Plastidic ribosomes were detected by fluorescence in situ hybridization (FISH) against ribosomal RNA and a specific PRPL1 antibody. Comparison of the suborganellar localisations of chloroplast ribosomes and CrTIG1 showed surprisingly different patterns. Ribosomes were mainly found within the globular basal region with highest concentrations proximal to the pyrenoid (highlighted by RbcL staining, Fig. 2d). CrTIG1 staining represented a patchy pattern distributed throughout the chloroplast. A stronger signal was detected in a region which showed little intensity of chloroplast ribosomes and may be outside or at the border of the chloroplast. While it cannot be ruled out that the antibody cross-reacts with cytosolic components, an alternative explanation is that some trigger factor protein may accumulate in a region which was previously shown to contain the cytosolic protein synthesis - and chloroplast import machinery30. The enrichment of trigger factor in chloroplasts was independently confirmed in A. thaliana lysates. Here, immunoblots clearly detect AtTIG1 in plastid fractions (Fig. 2e).
Chloroplast TIG proteins are unusually temperature-sensitive
The secondary structure of EcTF is predominantly α-helical for the N- and C-terminal domains, whereas the PPIase mainly contains β-sheets11, 31. To gain insight into the secondary structures of the chloroplast TIG1 isoforms, both mature proteins were analysed by far-UV circular dichroism (CD) spectroscopy and compared with EcTF. Consistent with previous reports31, 32, the far-UV CD spectrum of EcTF contained two pronounced minima at 208 nm and 222 nm, both typical for α-helical secondary structure. The spectra of both plastidic TIG1 proteins were comparable to that of EcTF, indicating that their overall secondary structures are similar (Fig. 3a). Secondary-structure estimation by the K2D2 algorithm33 indicates ~40% α-helix and ~13% β-sheet content for EcTF, close to the 43% α-helical and 16% β-sheet content determined from the respective crystal structure11. Both plastidic TIG1 proteins seem to contain a slightly higher proportion of α-helices (~44%), as indicated by a more intense negative ellipticity at 208 nm as compared with EcTF, and a similar content of β-sheets (12% and 13%, respectively). The data also confirm that all heterologously produced and purified proteins were folded and thus suitable for subsequent in vitro assays.
Secondary structure and thermal stability of TIG1s. (a) Left panels: CD spectra of 0.1 mg/mL of EcTF, CrTIG1, and AtTIG1 in 10 mM KCl, 20 mM Tris pH 7.5 buffer were recorded from 190 nm to 280 nm at 20 °C. Right panels: At the two minima at 208 nm and 222 nm, protein-unfolding curves were recorded from 20 °C to 94 °C at a heating rate of 1°/min. (b) Thermal stability of purified CrTIG1 was measured at 5 mg/mL with indicated buffers by stepwise heating of 1 °C/min over a temperature range of 20–95 °C. Protein-unfolding midpoints (T m) were calculated from the first derivative of intrinsic protein fluorescence emission at 330 nm and 350 nm. (c) Accumulation of insoluble protein aggregates was observed upon cell exposure to various temperatures of heat stress. Protein aggregates and total lysates were separated by 12% SDS-PAGE and visualized by Coomassie staining. Gel slice of total lysates serves as input control. (d) Cropped images of immunoblots of samples shown in (c). show the accumulation of CrTIG1, Rubisco activase (RCA1), HSP90C, CPN20, and Cytochrome F in protein aggregates.
Thermal unfolding was studied by gradual heating of the proteins (Fig. 3a, right panels). The transition curves at 208 nm and 222 nm revealed qualitative differences between the bacterial and the plant proteins. For EcTF, a major unfolding transition was only observed at 222 nm, which started at > 30 °C and had a midpoint (where half of the population is folded and the other half unfolded) between 45 ° and 50 °C. This is consistent with another study34 using differential scanning calorimetry, which determined the unfolding midpoint of EcTF to be at 54 °C. This resistance against thermal unfolding was attributed to the dimeric form of EcTF34. For the chloroplast proteins, unfolding was traceable at both wavelengths, with unfolding starting just above 30 °C and featuring a local maximum in the ellipticity recorded at 222 nm at intermediate temperatures. The midpoints of the thermal unfolding of CrTIG1 and AtTIG1 were between 35 °C and 40 °C, which are far below the values for EcTF (Fig. 3a, right panels). Thus, even mild heat-stress conditions35 seem to cause partial unfolding of at least some domains. Such conformational changes may inactivate the chaperone or reprogram its function for other tasks during heat stress. For a more precise determination of the unfolding midpoint of CrTIG1, shifts of intrinsic tryptophan fluorescence upon thermal unfolding were recorded at wavelengths of 330 nm and 350 nm. We determined the unfolding midpoints T m in buffers having various ionic strengths of 0–300 mM KCl. With no additional salt present, CrTIG1 showed a T m of 35.6 °C, whereas T m was shifted up to 41.6 °C at 300 mM KCl (Fig. 3b). Thus, more physiological salt concentrations render the protein more robust against thermal unfolding.
We next tested if thermal unfolding could also be observed in living cells, which might lead to the accumulation of CrTIG1 in protein aggregates. For this purpose, C. reinhardtii cells were exposed to various temperatures of mild to severe heat stress (i.e., 35–45 °C), and insoluble aggregates were isolated (see Methods). Overall, little aggregation was observed in cells exposed to temperatures between room temperature and 37 °C. At non-lethal heat stress between 39 °C and 42 °C35, increasing protein aggregation was observed with rising temperatures (Fig. 3c). The onset of protein aggregation at 39 °C was confirmed by examining the known heat-labile chloroplast protein rubisco activase (RCA1)36, which started to accumulate in aggregates at temperatures ≥39 °C. Interestingly, CrTIG1 was also detectable in the same fractions as RCA1 (Fig. 3d), while other chloroplast chaperones such as HSP90C or the co-chaperonin CPN20 predominantly accumulated in aggregates at higher temperatures. Thus, in contrast to bacterial TF and other molecular chaperones, chloroplast TIG1 is a remarkably heat-sensitive chaperone, which accumulates in aggregates even under relatively mild heat-stress conditions. Unlike many other chaperones, trigger factor is an ATP-independent chaperone. Thus, the observed heat sensitivity may serve as passive regulator to target this chaperone to protein aggregates which may serve as protective mechanism to rescue unfolded proteins from aggregates upon return to physiological growth conditions.
Chloroplast TIG1 is mainly monomeric
Several structural studies reported the occurrence of dimeric EcTF both in vitro and in vivo 14, 32, 37. To determine if chloroplast TIG1s show similar dimerization behaviours, purified TIG1 protein was separated by SEC. A prominent peak of EcTF protein eluted at a volume of 13.6 mL (Fig. 4, top panel), while CrTIG1 eluted in a minor peak at high molecular weight (12.1 mL) and a major peak at 14.5 mL (Fig. 4, middle panel), and AtTIG1 eluted at 14.1 mL with a small shoulder preceding the actual peak (Fig. 4, bottom panel). Since SEC elution profiles are not precise enough to determine the molecular weights of particles that are clearly nonspherical, as expected at least for EcTF, each fraction of the elution peak was examined online by right-angle static light scattering (RALS). The sloping curve of average molecular weight for EcTF resembles the molecular weight distribution of an earlier study14 with lower apparent masses of the protein at the peak borders and the highest value of 79 kDa in the centre of the peak (Fig. 4, top panel, Table S2). As stated in earlier studies, a heterogeneous population of monomeric and oligomeric EcTF seems to co-elute within this peak, which might represent a fast equilibrium of monomeric and oligomeric states14. By contrast, both CrTIG1 and AtTIG1 were monodisperse, as indicated by the consistent molecular weight values within the major peak area (Fig. 4, centre and bottom panel) with polydispersity values of 1.001 and 1.002 (M w /M n ), respectively. Further, elution profiles resulted in apparent molecular weight values close to their theoretical monomeric masses (Supplementary Table S2). Hence, at the physiological salt conditions used here, chloroplast TIG1s appeared to be present mainly in a monomeric form. In earlier proteomic studies, AtTIG1 migrated in native PAGE as expected for a molecular weight of the dimeric state38. Dimeric species of TIG1 proteins cannot be excluded under certain conditions and in a native context; however, asymmetric and elongated proteins are known to migrate slower than globular proteins in native gels39, which might explain the higher apparent molecular weight of AtTIG1 in the earlier studies. For our subsequent assays, it was essential that SEC fractions of CrTIG1 and AtTIG1 be highly monodisperse, making them ideally suited for three-dimensional bead modelling by SAXS.
Purified chloroplast TIG1 is mainly monomeric. Determination of conformational states of EcTF, CrTIG1, and AtTIG1. 200 µg of protein was separated by SEC in a buffer containing 20 mM Tris pH 7.5 and 150 mM KCl and measured with a right-angle light scattering (RALS) detector combined with a refractive index (RI) detector. Here, the RI detector signal was used to follow the elution. Average molar masses are indicated in blue for each graph (right ordinate). For the full elution profile, see Supplementary Figure S3.
SAXS reveals a typical dragon-shape conformation of TIG1
SAXS was used to examine if chloroplast TIG1s possess a similar overall shape as the bacterial isoform or if the eukaryotic chaperones evolved other structural features. Initial static measurements were performed at different protein concentrations in three replicates to examine if the TIG1 proteins remain monodisperse also at higher protein concentrations. Importantly, SAXS parameters of the bacterial EcTF were very comparable to previously published datasets (Supplementary Figure S4)32. At the concentrations used here, the monomer–dimer equilibrium of EcTF was strongly shifted towards the dimeric state, although proteins are known to quickly fluctuate between monomeric and dimeric states even at high concentrations14. Indeed, the values of the Porod volume indicate mainly dimeric proteins in the samples, but the molecular weight values derived from I(0) (i.e., the intensity at zero angle) point to a fast fluctuating population (Supplementary Table S3). Further, it has been reported that the radius of gyration (R g) and the maximal particle size (D max) of EcTF decrease with increasing protein concentration, possibly because of a more densely packed dimer at higher concentrations compared with the monomer at lower concentrations32. Consistently, the R g and D max values determined here also decreased with increasing protein concentration, albeit to a lesser extent (Supplementary Table S3). Chloroplast CrTIG1 and AtTIG1 appeared predominantly as monomers assuming a predicted molecular weight of 53.7 kDa and 53.2 kDa, respectively. In contrast to EcTF, molar masses calculated from the I(0) values slightly increased with rising protein concentrations, most likely because of weak interparticle interference (Supplementary Tables S4 and S5). However, the high linearity of the Guinier plots speaks against aggregation of the samples (Fig. 5a and d, inset)40. To overcome this interference, low-range values at low concentrations and high-range values at high concentrations were merged for further processing. Both plastidic proteins were found to have an R g value of ~38 Å, as determined by the Guinier plot and p(r) function. This very large R g for a 53-kDa protein—as compared to an R g value of 29.9 Å of globular BSA41—indicates an elongated and non-globular shape of both proteins. This notion is supported by the asymmetric peak of the pair distance distribution functions of both TIG1s with a shoulder extending to a maximum particle size of 125 Å for CrTIG1 and 127 Å for AtTIG1, respectively (Fig. 5c and f).
SAXS data from static measurements of trigger factors. Small-angle X-ray scattering data were collected for various concentrations of TIG1 proteins and merged. (a) and (d) Experimental SAXS profiles from merged scattering curves of CrTIG1 and AtTIG1 are indicated by small circles. Respectively, red curves represent the fit by GNOM, blue curves the best CRYSOL fit, grey curves the theoretical scattering of final DAMMIN models. Inset, Guinier plot of ln I(s) versus s −2 obtained from AUTORG. Data points used by AUTORG are labelled in blue. (b) and (e) Kratky plot s 2 I(s) versus s −1 of CrTIG1 and AtTIG, respectively. (c) and (f) Corresponding p(r) function as calculated from the experimental scattering curves using GNOM. “au” is arbitrary units. For SAXS data on EcTIG and the FoXS fit, see Supplementary Figure S4.
All analysed trigger factor proteins showed an overall similar bimodal pattern in Kratky plots with a dominant first peak followed by a distinct small peak indicating well-folded and multi-domain proteins. In addition, pronounced intrinsic flexibilities are indicated by the behaviour of the plots at higher s-range, which do not converge to the abscissa. The Kratky plot of EcTF displays a slight shift compared with both plastidic TIG1 proteins, indicating a more rigid and compact conformation due to the dimeric state32 (Fig. 5b,e and Supplementary Figure S4).
To verify our results regarding the monomeric state obtained from static SAXS measurements, we further performed online SEC coupled to SAXS detection (HPLC-SAXS). Consistent with the static assays, only minor amounts of putative dimer populations were detectable in the CrTIG1 and AtTIG1 samples, as indicated by the slight tailing of the monomeric peak fraction (Fig. 6a and d). To simplify our analysis of the chloroplast proteins, these higher molecular weight fractions were neglected in all subsequent steps. Thus, only frames within the peak section of high quality and constant R g were averaged for further analysis (Fig. 6a,d and Supplementary Figure S5, grey shaded area). The molecular weight was determined using multiple, concentration-independent, methods. These seem to overestimate the mass of the chloroplast TIG1s by up to 17% (Table 1), which might be attributed to traces of dimers or to the influence of capillary fouling on scattering42. However, the masses determined by RALS strongly supported predominantly monomeric states. Importantly, the calculated R g (34.4 Å for CrTIG1, 35.5 Å for AtTIG1) and D max (122 Å for CrTIG1, 124 Å for AtTIG1) values were comparable to the results from static measurement, which allowed us to complementarily process data from both approaches for further modelling.
SAXS data from HPLC-SAXS experiments of trigger factors. Small-angle X-ray scattering data were collected by size-exclusion high-performance liquid chromatography with a ENrich SEC 650 column online with small-angle X-ray scattering (HPLC-SAXS) with 400 µg of the respective protein. (a) and (d) Elution profiles of SEC-SAXS runs, represented by I(0) and R g determined by AUTORG for each frame. Highlighted in grey are the frames used for averaging. (b) and (e) Experimental SAXS profiles from the averaged HPLC-SAXS frames of CrTIG1 and AtTIG1 are indicated by small circles. Respectively, red curves represent the fit by GNOM, blue curve the CRYSOL fit of the RaptorX models, and grey curves the theoretic scattering of final DAMMIN model. Inset: Guinier plot of ln I(s) versus s −2 obtained from AUTORG. Data points used by AUTORG are labelled in blue. (c) and (f) Corresponding p(r) function as calculated from the experimental scattering curves using GNOM. “au” is arbitrary units. For SAXS data on EcTF and the FoXS fit, see Supplementary Figure S5.
All plastidic TIG1 SAXS datasets were applied for ab initio modelling of low-resolution bead envelopes. Modelling of EcTF was not pursued with our dataset since the heterogeneous population of dimeric and monomeric species complicates modelling and different conformations of purified EcTF proteins were comprehensively studied before32. With the static SAXS measurements, more defined scattering at a higher s-range was obtained because of a higher protein concentration in the sample. Hence, ab initio modelling was pursued with the GASBOR algorithm, which allows a chained assembly of dummy residues corresponding to the number of amino acids. The models obtained fitted the experimental data with χ2 values of 0.81 for CrTIG1 and 0.69 for AtTIG1 (Supplementary Table S6). To confirm that the minor traces of putative dimeric species did not significantly influence the models, data were processed in symmetry P1 (no symmetry) and P2 (dimer). The P2 symmetry could be excluded because of the resulting high χ2 values of these models (data not shown) and hence supported the fact that our models were truly based on the monomeric molecules. For the HPLC-SAXS dataset, ab initio modelling was performed with DAMMIF. Consistent with the modelling of the static SAXS data, χ2 values of 0.72 for CrTIG1 and 0.79 for AtTIG1 (Supplementary Table S6) were gained for these low-resolution structures.
The low-resolution bead models of CrTIG1 and AtTIG1 proteins resulting from both datasets clearly resemble the conformation of the bacterial TF with a dragon-like structure containing head, tail and arms (Fig. 7). Interestingly, groove-like structures are visible along the middle domain (Supplementary Figure S6), which could constitute putative binding sites for peptide stretches of chloroplast TIG1 substrates.
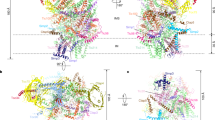
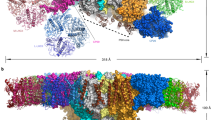
Global shape of the chloroplast trigger factor. Ab initio modelling from SAXS data. Shape topology and low-resolution bead models of CrTIG1 (a,b) and AtTIG1 (c,d) from static SAXS experiments (a,c) and HPLC-SAXS (b,d) were generated using AMBIMETER and DAMMIN based on 10 averaged GASBOR models for static or 20 DAMMIF models for HPLC-SAXS experiments, respectively. Bead models were superimposed by SUPALM on the RaptorX high-resolution model or the improved models by SREFLEX. Secondary structures were coloured in yellow (β-sheets), red (α-helix), and green (random coils). For different side views of the models, see Supplementary Figure S6.
Next, we attempted to fit high-resolution models into the SAXS-based ab initio shapes. Since, to our knowledge, no structural data exist for any chloroplast trigger factor to date, predicted models were calculated with the RaptorX43 and SAXSTER 44 algorithms on the basis of various structural data of exclusively full-length bacterial isoforms. Different high-resolution structures of TF exist for dimeric apo E. coli TF complexes (X-ray structure, PDB entry 1w2611) and monomeric TF associated with different fragments of the PhoA (NMR structures, PDB entries 2mlx/2mly/2mlz37) or the ribosome (PDB entry 2vrh12). Further, crystal structures of dimeric TF from V. cholerae (PDB entry 1t1145) and T. maritima are available. For the latter, both full-length structures with and without a substrate are monomeric (X-ray structures, PDB entries 3gty/gt046). Different bacterial TF species show various domain orientations47. For example, both the crystal structure and NMR structural analyses reveal different orientations of the head domain among EcTF47, VcTF44, and TmTF46. For VcTF, both domains constituting the arms and the tail show a twist, which is not seen for the other isoforms45. Psi-Blast searches with sequences of CrTIG1 and AtTIG1 resulted in similar matches between the plastidic TIG1 proteins and the bacterial TFs (Supplementary Table S7). The best coverage for CrTIG1 and AtTIG1 was reached with the full-length NMR and cryoEM structures of EcTF (PDB entry 2mlx and 2vrh) (Supplementary Table S7). Our predicted models were validated with CRYSOL and superimposed with the bead models. Again, the models that gave the best χ2 values in CRYSOL were the predicted models from RaptorX based on 2mlx. Importantly, not all high-resolution models fitted with same quality. Superimposing the models based on VcTF and TmTF to the SAXS data was not successful as judged by their high χ2 scores (>25–55, data not shown). To test if we could further improve the domain agreement of our positive models within the SAXS-based bead models, SREFLEX was applied, which examines the flexibility of models. Generally, this refinement improved the χ2 values, albeit with a different outcome. For CrTIG1, χ2 values were slightly improved from 10.4 to 6.2. SREFLEX Refinement of CrTIG1 might have been less than optimal since the orientations of some α-helices within the central body domain were severely altered (Fig. 7a). In contrast, the refinement of AtTIG1 high-resolution models improved χ2 values significantly from 57.2 to 3.3 (Supplementary Table S6) with only mild rearrangements within the domains (Fig. 7c). As shown in Figs 5 and 6, two independent evaluations of our high-resolution models by CRYSOL and a simulation of theoretical scattering and fitting to our experimental data by FoXS resulted in close fits of the models to the SAXS data. Thus, it can be assumed that the E. coli-based models closely describe the overall shape and domain position of the plastidic TIG1 proteins. However, none of the bacteria-derived models tested resulted in a complete match to the SAXS shapes obtained from the static and HPLC- SAXS measurements. This suggests that the intrinsic fold of the chloroplast TIG1 proteins, especially the relative arrangement of the three domains to each other, may be unique for TIG1s (Fig. 7). This is underscored by the relatively high normalized spatial discrepancy values (NSD, a measure of quantitative similarity between sets of three-dimensional points), which are larger than 2.5 for all SREFLEX-refined models of plastidic TIG1s (Supplementary Table S6). These differences seem most pronounced for the arms, which are further away in both chloroplast TIG1 conformations compared with EcTF11, and the head and tail domains, which cannot be described perfectly by our models (Fig. 7 and Supplementary Figure S6). To fully resolve these altered conformations of chloroplast trigger factor, atomic structures are required to determine how chloroplast TIG1 is tailored to perform its function within the chloroplast.
Conclusions
In conclusion, we present here the first description of two eukaryotic chloroplast-localised molecular chaperones of the trigger factor family. Both plastidic TIG1 orthologues are mainly monomeric, as shown independently by RALS and SAXS for the purified proteins. The most surprising finding was the high propensity of plastidic TIG1 for thermal unfolding, which might perform a specific task during heat exposure. Independent static and HPLC SAXS measurements and subsequent ab initio modelling revealed the typical dragon-shape of the molecule as known from bacterial TF containing a head, a tail, and two arms. This trigger-factor-like shape was confirmed by independent high-resolution models, which fitted well to the SAXS data. However, the chloroplast TIG1 models indicate unique domain arrangements and intrinsic conformations, thus suggesting that plastidic TIG1s have structurally diverged from their bacterial orthologues. Given the ubiquitous action of bacterial TF on most newly-synthesized proteins, plastidic TIG1 – and maybe even the truncated version TIG2 – may have specifically adapted to serve the maintenance of protein homeostasis in chloroplasts. It will be interesting to see why chloroplasts are the only compartment of eukaryotic cells that conserved these trigger factor chaperones. Future studies are also required to determine the detailed function and substrate specificity of these chaperones in chloroplasts.
Methods
Cells and Culture Conditions
Most experiments were conducted with the C. reinhardtii strain cw15 CF18548. Microscopy was done with strain cw15–325 (cwd, mt+, arg7−). All cultures were grown photomixotrophically in TAP medium49 on a rotary shaker at 25 °C and an illumination of 70 E m−2 s−1. For chloroplast isolation, cells were grown in TAP medium supplemented with 0.5% (w/v) peptone. For A. thaliana cell extracts, plants were grown under standard conditions.
Cloning, production and purification of the trigger factor species
For CrTIG1 both TargetP and ChloroP50, 51 predicted the transit peptide cleavage site QVC/A. Mature C. reinhardtii TIG1 protein (lacking the putative N-terminal 67 amino-acid transit peptide) was created by site directed mutagenesis of pFW13 resulting in pFW115. For AtTIG1, the predicted cleavage site was RVS/S. Mature A. thaliana TIG1 (lacking the putative N-terminal 27 amino-acid transit peptide), was amplified by PCR from cDNA with the primers (5′-GGGGCATATGTCCCGTCTCTTCCAATCAG-3′) and (5′-CCCCGGATCCTCAACGAGTGATGTATTGAATC-3′) and cloned with NdeI and BamHI into pTyb21 (NEB) resulting pFW141. Since the cleavage site was predicted incorrectly, the cleavage site RLF/A was reassigned based on the alignment of AtTIG1 with CrTIG1. For correction of the mature protein, 49 amino acids were removed from the N-terminal sequence with oligos 5′-GGAAGAGCTCATATGTCCCGTCTCTTCCAATCAGATAGTGT GCTACAGTTTGGTGGGAGGTTGAAGAAACCAATTAGCAGGCCTTTGGACATGTCTTGTGTCTCTAGAAAAATTGGATTTTTCGGA GATTTTATGTCACATGGTGGTAATTTTAGGCTATTC-3′ by site directed mutagenesis resulting in pFW136. The TF sequence from E. coli was amplified by PCR from DH5alpha genomic DNA with primers (5′-GGGGCATATGCAAGTTTCAGTTGAAACCAC-3′) and (5′-CCCCGGATCCTTACGCCTGCTGGTTCATC-3′) and cloned with NdeI and BamHI into pTyb21 (NEB) resulting pFW142. For heterologous protein production pFW115, pFW136 and pFW142 were synthesized in E. coli ER2566 and purified via chitin affinity resins according to the manufacturer’s instructions (NEB). All purified proteins were concentrated with Amicon Ultra-15 concentrator (Merck Millipore, Darmstadt) and purified over a Superdex 200 SEC column (GE healthcare) in 10 mM Tris pH 7.5 and 10 mM KCl buffer. For antisera production, rabbits were immunized with CrTIG1 or AtTIG1.
Phylogenetic analyses and comparison of sequence similarity and identity
Sequences were derived from http://phytozome.jgi.doe.gov, NCBI and http://www.uniprot.org. Phylogenetic analyses were performed with sequences comprising the mature sequences lacking predicted transit peptides using the Phylogeny.fr pipeline52 implementing algorithms T-Coffee53, BioNJ54, and TreeDyn55 setting bootstraps values to 1000. Sequence similarity and identity was determined by http://imed.med.ucm.es/Tools/sias.html.
Immunofluorescence microscopy
Cell fixation and staining was done as published before56. Primary antibodies were against CrTig1, RbcL, and PRPL1 in dilutions of 1:500, 1:2,000, and 1:1,000, respectively. The secondary fluorescein isothiocyanate-labeled antibody (Sigma-Aldrich) was applied in a 1:200 dilution. FISH probes were prepared according to57. After incubation with the secondary antibody, slides were washed in phosphate-buffered saline, and a drop of mounting solution containing DAPI (Vectashield; Vector Laboratories) was applied at the centre of each slide. Samples were analysed with a Leica TCS SP5 II confocal laser-scanning microscope (514-nm excitation and 500-, 592-, or 598–657 nm detection).
Circular dichroism spectroscopy
CD spectra were recorded on a Chirascan-plus (Applied Photophysics) spectropolarimeter. All proteins were measured at a concentration of 0.1 mg/mL in 10 mM KCl, 10 mM Tris, pH 7.5 buffer. At fixed temperatures (20 °C), five scans between 190 nm and 280 nm were recorded with an optical pathlength of 0.1 cm, a step size of 1 nm, and a digital integration time of 1 s. All spectra were baseline-corrected. At the two minima at 222 nm and 208 nm, temperature scans were recorded from 20 °C to 94 °C at a heating rate of 1 °C/min and a digital integration time of 4 s.
Thermal stability/melting point determination using Differential Scanning Fluorimetry
Protein thermal stability was measured at 5 mg/mL protein concentration in different buffer compositions using the Prometheus NT.48 (NanoTemper Technologies). Thermal unfolding was performed in nanoDSF grade high-sensitivity glass capillaries (NanoTemper Technologies) with a heating rate of 1 °C/min over a temperature range of 20–95 °C. Protein-unfolding midpoints (T m) were calculated from the first derivative of the ratio of intrinsic protein fluorescence emission intensities at 330 nm and 350 nm58, 59.
Analysis of protein aggregates
C. reinhardtii cells were grown to logarithmic phase. For each sample, 3 × 108 cells were harvested and resuspended in 50 mL fresh medium at the respective temperatures. After 45 min cells were harvested quickly and chilled on ice. Aggregate preparations were done as published previously60. In brief, cell pellets were resuspended in lysis buffer (20 mM NaPi pH 6.8, 10 mM DTT, 1 mM EDTA, 0.25x proteinase inhibitor (Roche, cOmplete-EDTA-free)) and lysed by sonication (Bandelin Sonopuls UW 2200, Berlin). Cell debris was removed by centrifugation at 500 g, and supernatants were adjusted to same protein concentrations. As input control, “Total lysate” aliquots were removed and supplemented with 1 volume of sample buffer (125 mM Tris-HCl, pH 6.8, 20% glycerol, 4% SDS, 100 mM DTT, and 0.005% bromophenol blue). From remaining lysates, protein aggregates were isolated by repetitive centrifugation at 19,000 g and washes with 20 mM sodium phosphate pH 6.8 and 2% NP-40. The final pellet was dissolved in 6 M urea and supplemented with 1 volume of sample buffer before separation on SDS-PAGE, transfer to nitrocellulose membrane, and immunoblotting.
RALS measurements
SEC coupled to RALS was performed on an OMNISEC system (Malvern Instruments, Worcestershire, UK) using a Superdex 200 Increase 10/300 SEC column (GE Healthcare). Prior to each run, the system was extensively equilibrated in running buffer (20 mM Tris, pH 7.5, 150 mM KCl). Protein concentrations were determined from online refractive index (RI) measurements assuming a dn/dc of 0.185 mL/mg. 50 µL of a calibration standard (BSA, Sigma, at 3 mg/mL) and samples (at 4 mg/mL) were injected twice. All experiments were conducted at 20 °C with a flow rate of 0.5 mL/min for 70 min. Molar masses were calculated as described by the manufacturer using OMNISEC v10 software.
X-ray scattering experiments and data analysis
Small-angle X-ray scattering data were collected on the BM29 beamline61 at the European Synchrotron Radiation Facility (ESRF, Grenoble) with a Pilatus 1 M detector (16.9 cm × 17.9 cm) at a wavelength of 0.9919 Å (12.5 keV) and a sample-detector distance of 2.867 m corresponding to a q-range of 0.025–5 nm−1. For static measurements, five to eight different protein concentrations were measured with 1 s exposure times per frame and 10 frames per concentration (n = 3) at 20 °C. Scattering by proteins samples were measured in 20 mM Tris buffer, pH 7.5 supplemented with 150 mM KCl. Scattering by the corresponding buffers was measured before and after one run, averaged and subtracted from the protein scattering. BSA standards were used to calibrate the I(0) values and the scattering of pure water was used to calibrate the intensity to absolute units62. Prior to measurements, protein concentrations were determined from absorption at 280 nm (A 280) using a NanoDrop One photometer (Thermo Scientific) and the corresponding molar coefficient calculated with ProtParam (www. expasy.ch).
For SEC high-performance liquid chromatography/small-angle X-ray scattering (HPLC-SAXS), an online HPLC system (Shimadzu, Japan) connected to the sample changer of the BM29 was used63. Per run, 400 µg protein (50 µL with 8 mg/mL protein concentration) was injected onto an ENrich SEC 650 10 mm × 300 mm column (BioRad) and separated at a flow rate of 0.75 mL/min at room temperature. 1500 frames with an exposure time of 1.5 s per frame were collected per run.
The data processing was done with the ATSAS 2.7.2 program package64, 65. The forward scattering I(0) and the radius of gyration R g were evaluated with PRIMUS66 using the Guinier approximation assuming that for a spherical particles at very small angles (s < 1.3/R g) the intensity is represented by I(s) = I(0) exp[−(sR g)2/3]). The distance distribution function p(r) and the maximum particle dimension (D max) were obtained using GNOM 67.
Frames collected from static measurements were checked for radiation damage before averaging and buffer subtraction. Scattering curves from the concentration series were investigated individually for indications of aggregation and particle attraction or repulsion. Curves affected were excluded from further analysis. For the characterization of the oligomerization state, scattering curves (from n = 3) with same protein concentration were scaled and merged. For ab initio modelling, frames from different concentrations were merged. Frames of the HPLC-SAXS, which were automatically processed by EDNA, were checked and inspected with HDFview. Frames with a consistent R g from the peak scattering intensity and good quality were scaled manually and averaged to yield a single frame. Buffer subtraction was applied using buffer frames before or after the protein peak avoiding over-subtraction with frames from the aggregate peak.
A first insight into the overall shape of the TIG1 proteins was given by the AMBIMETER tool. These models yielded a crude shape topology, which was further refined and supported by ab initio modelling. For the latter, 10–20 models were generated with GASBOR 68 or DAMMIF 69 and then averaged with DAMAVER 70. As a last refinement step, the damstart model generated with DAMAVER was refined with one cycle in DAMMIN. Protein structure modelling was performed with the SAXSTER server71 on the basis of all available full-length TF structures (i.e. from E. coli, T. maritima and V. cholerae) as templates or RaptorX43 algorithm based on the EcTF structure. Superposition of ab initio models derived from SAXS data and 3D models were calculated with SUPCOMB 72 or SUPALM 73. Fits between the SAXS data and the structures were evaluated with CRYSOL 74 and FoXS 75 and superposed with the bead models using SUPALM 73. The models were improved using SREFLEX in rigid or flexible mode with domain partition76. To this end, sequences regions of the domains were determined from the RaptorX model in accordance with the domain annotation of the available TF structures (Supplementary Table S8). Additionally, theoretical scattering profiles from the models were generated and fitted against the experimental data with the FoXS Server75.
Miscellaneous
Chloroplast and mitochondria were isolated from C. reinhardtii cells as described earlier26. SDS-PAGE was performed as published77. Antibodies described earlier were against HSP70B48, CGE178, HSP90C26, mitochondrial carbonic anhydrase79, CytF80 and CF1β81. Antibody purification was described earlier26.
References
Hartl, F. U., Bracher, A. & Hayer-Hartl, M. Molecular chaperones in protein folding and proteostasis. Nature 475, 324–332, doi:10.1038/nature10317 (2011).
Sjuts, I. & Soll, J. & Bolter, B. Import of Soluble Proteins into Chloroplasts and Potential Regulatory Mechanisms. Front Plant Sci 8, 168, doi:10.3389/fpls.2017.00168 (2017).
Pechmann, S., Willmund, F. & Frydman, J. The ribosome as a hub for protein quality control. Mol Cell 49, 411–421, doi:10.1016/j.molcel.2013.01.020 (2013).
Preissler, S. & Deuerling, E. Ribosome-associated chaperones as key players in proteostasis. Trends Biochem Sci 37, 274–283, doi:10.1016/j.tibs.2012.03.002 (2012).
Frydman, J. Folding of newly translated proteins in vivo: the role of molecular chaperones. Annu Rev Biochem 70, 603–647, doi:10.1146/annurev.biochem.70.1.603 (2001).
Hoffmann, A., Bukau, B. & Kramer, G. Structure and function of the molecular chaperone Trigger Factor. Biochim Biophys Acta 1803, 650–661, doi:10.1016/j.bbamcr.2010.01.017 (2010).
Oh, E. et al. Selective ribosome profiling reveals the cotranslational chaperone action of trigger factor in vivo. Cell 147, 1295–1308, doi:10.1016/j.cell.2011.10.044 (2011).
Teter, S. A. et al. In Cell Vol. 97 755–765 (1999).
Lill, R., Crooke, E., Guthrie, B. & Wickner, W. The “trigger factor cycle” includes ribosomes, presecretory proteins, and the plasma membrane. Cell 54, 1013–1018 (1988).
Teter, S. A. et al. Polypeptide flux through bacterial Hsp70: DnaK cooperates with trigger factor in chaperoning nascent chains. Cell 97, 755–765 (1999).
Ferbitz, L. et al. Trigger factor in complex with the ribosome forms a molecular cradle for nascent proteins. Nature 431, 590–596, doi:10.1038/nature02899 (2004).
Merz, F. et al. The C-terminal domain of Escherichia coli trigger factor represents the central module of its chaperone activity. J Biol Chem 281, 31963–31971, doi:10.1074/jbc.M605164200 (2006).
Kaiser, C. M. et al. Real-time observation of trigger factor function on translating ribosomes. Nature 444, 455–460, doi:10.1038/nature05225 (2006).
Patzelt, H. et al. Three-state equilibrium of Escherichia coli trigger factor. Biol Chem 383, 1611–1619, doi:10.1515/BC.2002.182 (2002).
Lakshmipathy, S. K. et al. Identification of nascent chain interaction sites on trigger factor. J Biol Chem 282, 12186–12193, doi:10.1074/jbc.M609871200 (2007).
Liu, C. P., Perrett, S. & Zhou, J. M. Dimeric trigger factor stably binds folding-competent intermediates and cooperates with the DnaK-DnaJ-GrpE chaperone system to allow refolding. J Biol Chem 280, 13315–13320, doi:10.1074/jbc.M414151200 (2005).
Allen, J. F., de Paula, W. B., Puthiyaveetil, S. & Nield, J. A structural phylogenetic map for chloroplast photosynthesis. Trends Plant Sci 16, 645–655, doi:10.1016/j.tplants.2011.10.004 (2011).
Gray, M. W. Origin and evolution of organelle genomes. Curr Opin Genet Dev 3, 884–890 (1993).
Sharma, M. R. et al. Cryo-EM study of the spinach chloroplast ribosome reveals the structural and functional roles of plastid-specific ribosomal proteins. Proc Natl Acad Sci USA 104, 19315–19320, doi:10.1073/pnas.0709856104 (2007).
Manuell, A. L., Quispe, J. & Mayfield, S. P. In PLoS Biol Vol. 5 e209 (2007).
Bieri, P., Leibundgut, M., Saurer, M., Boehringer, D. & Ban, N. The complete structure of the chloroplast 70S ribosome in complex with translation factor pY. Embo J, doi:10.15252/embj.201695959 (2016).
Graf, M. et al. Cryo-EM structure of the spinach chloroplast ribosome reveals the location of plastid-specific ribosomal proteins and extensions. Nucleic Acids Res, doi:10.1093/nar/gkw1272 (2016).
Trosch, R., Muhlhaus, T., Schroda, M. & Willmund, F. ATP-dependent molecular chaperones in plastids-More complex than expected. Biochim Biophys Acta 1847, 872–888, doi:10.1016/j.bbabio.2015.01.002 (2015).
Breiman, A., Fieulaine, S., Meinnel, T. & Giglione, C. The intriguing realm of protein biogenesis: Facing the green co-translational protein maturation networks. Biochim Biophys Acta 1864, 531–550, doi:10.1016/j.bbapap.2015.11.002 (2016).
Schroda, M. In Photosyn. Res Vol. 82 221–240 (2004).
Willmund, F. & Schroda, M. HEAT SHOCK PROTEIN 90C is a bona fide Hsp90 that interacts with plastidic HSP70B in Chlamydomonas reinhardtii. Plant Physiol 138, 2310–2322, doi:10.1104/pp.105.063578 (2005).
Olinares, P. D., Ponnala, L. & van Wijk, K. J. Megadalton complexes in the chloroplast stroma of Arabidopsis thaliana characterized by size exclusion chromatography, mass spectrometry, and hierarchical clustering. Mol Cell Proteomics 9, 1594–1615, doi:10.1074/mcp.M000038-MCP201 (2010).
Zerges, W. & Rochaix, J. D. In J. Cell Biol. Vol. 140 101–110 (1998).
Chua, N. H., Blobel, G., Siekevitz, P. & Palade, G. E. Attachment of chloroplast polysomes to thylakoid membranes in Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 70, 1554–1558 (1973).
Uniacke, J. & Zerges, W. Chloroplast protein targeting involves localized translation in Chlamydomonas. Proc Natl Acad Sci USA 106, 1439–1444, doi:10.1073/pnas.0811268106 (2009).
Zarnt, T. et al. Modular structure of the trigger factor required for high activity in protein folding. J Mol Biol 271, 827–837, doi:10.1006/jmbi.1997.1206 (1997).
Rathore, Y. S., Dhoke, R. R., Badmalia, M. & Sagar, A. & Ashish. SAXS data based global shape analysis of trigger factor (TF) proteins from E. coli, V. cholerae, and P. frigidicola: resolving the debate on the nature of monomeric and dimeric forms. J Phys Chem B 119, 6101–6112, doi:10.1021/acs.jpcb.5b00759 (2015).
Perez-Iratxeta, C. & Andrade-Navarro, M. A. K2D2: estimation of protein secondary structure from circular dichroism spectra. BMC Struct Biol 8, 25, doi:10.1186/1472-6807-8-25 (2008).
Fan, D. J., Ding, Y. W., Pan, X. M. & Zhou, J. M. Thermal unfolding of Escherichia coli trigger factor studied by ultra-sensitive differential scanning calorimetry. Biochim Biophys Acta 1784, 1728–1734, doi:10.1016/j.bbapap.2008.05.002 (2008).
Schroda, M., Hemme, D. & Muhlhaus, T. The Chlamydomonas heat stress response. Plant J 82, 466–480, doi:10.1111/tpj.12816 (2015).
Feller, U., Crafts-Brandner, S. J. & Salvucci, M. E. Moderately High Temperatures Inhibit Ribulose-1,5-Bisphosphate Carboxylase/Oxygenase (Rubisco) Activase-Mediated Activation of Rubisco. Plant Physiol 116, 539–546 (1998).
Saio, T., Guan, X., Rossi, P., Economou, A. & Kalodimos, C. G. Structural basis for protein antiaggregation activity of the trigger factor chaperone. Science 344, 1250494, doi:10.1126/science.1250494 (2014).
Peltier, J. B. et al. The oligomeric stromal proteome of Arabidopsis thaliana chloroplasts. Mol Cell Proteomics 5, 114–133, doi:10.1074/mcp.M500180-MCP200 (2006).
Willmund, F., Dorn, K. V., Schulz-Raffelt, M. & Schroda, M. The chloroplast DnaJ homolog CDJ1 of Chlamydomonas reinhardtii is part of a multichaperone complex containing HSP70B, CGE1, and HSP90C. Plant Physiol 148, 2070–2082, doi:10.1104/pp.108.127944 (2008).
Jacques, D. A. & Trewhella, J. Small-angle scattering for structural biology–expanding the frontier while avoiding the pitfalls. Protein Sci 19, 642–657, doi:10.1002/pro.351 (2010).
Mylonas, E. & Svergun, D. I. Accuracy of molecular mass determination of proteins in solution by small-angle X-ray scattering. Journal of Applied Crystallography 40, S245–S249, doi:10.1107/S002188980700252x (2007).
Jeffries, C. M., Graewert, M. A., Svergun, D. I. & Blanchet, C. E. Limiting radiation damage for high-brilliance biological solution scattering: practical experience at the EMBL P12 beamline PETRAIII. J Synchrotron Radiat 22, 273–279, doi:10.1107/S1600577515000375 (2015).
Kallberg, M. et al. Template-based protein structure modeling using the RaptorX web server. Nat Protoc 7, 1511–1522, doi:10.1038/nprot.2012.085 (2012).
dos Reis, M., Savva, R. & Wernisch, L. Solving the riddle of codon usage preferences: a test for translational selection. Nucleic Acids Res 32, 5036–5044, doi:10.1093/nar/gkh834 (2004).
Ludlam, A. V., Moore, B. A. & Xu, Z. The crystal structure of ribosomal chaperone trigger factor from Vibrio cholerae. Proc Natl Acad Sci USA 101, 13436–13441, doi:10.1073/pnas.0405868101 (2004).
Martinez-Hackert, E. & Hendrickson, W. A. Promiscuous substrate recognition in folding and assembly activities of the trigger factor chaperone. Cell 138, 923–934, doi:10.1016/j.cell.2009.07.044 (2009).
Thomas, A. S., Mao, S. & Elcock, A. H. Flexibility of the bacterial chaperone trigger factor in microsecond-timescale molecular dynamics simulations. Biophys J 105, 732–744, doi:10.1016/j.bpj.2013.06.028 (2013).
Schroda, M., Vallon, O., Wollman, F. A. & Beck, C. F. In Plant Cell Vol. 11 1165–1178 (1999).
Harris, E. H., Boynton, J. E. & Gillham, N. W. In J. Cell Biol. Vol. 63 160–179 (1974).
Emanuelsson, O., Nielsen, H., Brunak, S. & von Heijne, G. Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol 300, 1005–1016, doi:10.1006/jmbi.2000.3903 (2000).
Emanuelsson, O., Nielsen, H. & von Heijne, G. ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci 8, 978–984, doi:10.1110/ps.8.5.978 (1999).
Dereeper, A. et al. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 36, W465–469, doi:10.1093/nar/gkn180 (2008).
Notredame, C., Higgins, D. G. & Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302, 205–217, doi:10.1006/jmbi.2000.4042 (2000).
Gascuel, O. BIONJ: an improved version of the NJ algorithm based on a simple model of sequence data. Mol. Biol. Evol. 14, 685–695 (1997).
Chevenet, F., Brun, C., Banuls, A. L., Jacq, B. & Christen, R. TreeDyn: towards dynamic graphics and annotations for analyses of trees. BMC Bioinformatics 7, 439, doi:10.1186/1471-2105-7-439 (2006).
Uniacke, J., Colon-Ramos, D. & Zerges, W. FISH and immunofluorescence staining in Chlamydomonas. Methods Mol Biol 714, 15–29, doi:10.1007/978-1-61779-005-8_2 (2011).
Daims, H., Bruhl, A., Amann, R., Schleifer, K. H. & Wagner, M. The domain-specific probe EUB338 is insufficient for the detection of all Bacteria: development and evaluation of a more comprehensive probe set. Syst Appl Microbiol 22, 434–444, doi:10.1016/S0723-2020(99)80053-8 (1999).
Temel, D. B., Landsman, P. & Brader, M. L. Orthogonal Methods for Characterizing the Unfolding of Therapeutic Monoclonal Antibodies: Differential Scanning Calorimetry, Isothermal Chemical Denaturation, and Intrinsic Fluorescence with Concomitant Static Light Scattering. Methods Enzymol 567, 359–389, doi:10.1016/bs.mie.2015.08.029 (2016).
Alexander, C. G. et al. Novel microscale approaches for easy, rapid determination of protein stability in academic and commercial settings. Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics 1844, 2241–2250, doi:10.1016/j.bbapap.2014.09.016 (2014).
Willmund, F. et al. The cotranslational function of ribosome-associated Hsp70 in eukaryotic protein homeostasis. Cell 152, 196–209, doi:10.1016/j.cell.2012.12.001 (2013).
Pernot, P. et al. Upgraded ESRF BM29 beamline for SAXS on macromolecules in solution. J Synchrotron Radiat 20, 660–664, doi:10.1107/S0909049513010431 (2013).
Orthaber, D., Bergmann, A. & Glatter, O. SAXS experiments on absolute scale with Kratky systems using water as a secondary standard. Journal of Applied Crystallography 33, 218–225, doi:10.1107/S0021889899015216 (2000).
Hutin, S., Brennich, M., Maillot, B. & Round, A. Online ion-exchange chromatography for small-angle X-ray scattering. Acta Crystallogr D 72, 1090–1099, doi:10.1107/S2059798316012833 (2016).
Konarev, P. V., Petoukhov, M. V., Volkov, V. V. & Svergun, D. I. ATSAS 2.1, a program package for small-angle scattering data analysis. Journal of Applied Crystallography 39, 277–286, doi:10.1107/S0021889806004699 (2006).
Petoukhov, M. V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. Journal of Applied Crystallography 45, 342–350, doi:10.1107/S0021889812007662 (2012).
Konarev, P. V., Volkov, V. V., Sokolova, A. V., Koch, M. H. J. & Svergun, D. I. PRIMUS: a Windows PC-based system for small-angle scattering data analysis. Journal of Applied Crystallography 36, 1277–1282 (2003).
Svergun, D. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. Journal of Applied Crystallography 25, 495–503, doi:10.1107/S0021889892001663 (1992).
Svergun, D. I., Petoukhov, M. V. & Koch, M. H. J. Determination of domain structure of proteins from X-ray solution scattering. Biophysical Journal 80, 2946–2953 (2001).
Franke, D. & Svergun, D. I. DAMMIF, a program for rapid ab-initio shape determination in small-angle scattering. J Appl Crystallogr 42, 342–346, doi:10.1107/S0021889809000338 (2009).
Volkov, V. V. & Svergun, D. I. Uniqueness of ab initio shape determination in small-angle scattering. Journal of Applied Crystallography 36, 860–864, doi:10.1107/S0021889803000268 (2003).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40, doi:10.1186/1471-2105-9-40 (2008).
Kozin, M. B. & Svergun, D. I. Automated matching of high- and low-resolution structural models. Journal of Applied Crystallography 34, 33–41, doi:10.1107/S0021889800014126 (2001).
Konarev, P. V., Petoukhov, M. V. & Svergun, D. I. Rapid automated superposition of shapes and macromolecular models using spherical harmonics. J Appl Crystallogr 49, 953–960, doi:10.1107/S1600576716005793 (2016).
Svergun, D. I. B. C. K. M. H. J. CRYSOL - a Program to Evaluate X-ray Solution Scattering of Biological Macromolecules from Atomic Coordinates. J. Appl. Cryst. 28, 768–773 (1995).
Schneidman-Duhovny, D., Hammel, M. & Sali, A. FoXS: a web server for rapid computation and fitting of SAXS profiles. Nucleic Acids Res 38, W540–544, doi:10.1093/nar/gkq461 (2010).
Panjkovich, A. & Svergun, D. I. Deciphering conformational transitions of proteins by small angle X-ray scattering and normal mode analysis. Phys Chem Chem Phys 18, 5707–5719, doi:10.1039/c5cp04540a (2016).
Laemmli, U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685 (1970).
Schroda, M., Vallon, O., Whitelegge, J. P., Beck, C. F. & Wollman, F. A. The chloroplastic GrpE homolog of Chlamydomonas: two isoforms generated by differential splicing. Plant Cell 13, 2823–2839 (2001).
Eriksson, M., Gardestrom, P. & Samuelsson, G. Isolation, Purification, and Characterization of Mitochondria from Chlamydomonas reinhardtii. Plant Physiol 107, 479–483 (1995).
Pierre, Y. & Popot, J. L. Identification of two 4-kDa miniproteins in the cytochrome b6f complex from Chlamydomonas reinhardtii. C R Acad Sci III 316, 1404–1409 (1993).
Lemaire, C. & Wollman, F. A. The chloroplast ATP synthase in Chlamydomonas reinhardtii. II. Biochemical studies on its biogenesis using mutants defective in photophosphorylation. J Biol Chem 264, 10235–10242 (1989).
Acknowledgements
We thank Francis-André Wollman for the antibodies against Cytf and CF1β, Michael Schroda for the antibodies against HSP70B, HSP90C and CGE1 and help with phylogenetic analyses, and Mats Eriksson for the antibody against mitochondrial carbonic anhydrase. For cloning of CrTIG1, we thank the Kazuza DNA Research Institute for providing cDNA clone AV639812. We thank Catherine Birck from the IGBMC Strasbourg for access to the Prometheus NT.48 and Sabine Heit for help in performing measurements. We thank the BM29 staff at the ESRF Grenoble for support with SAXS experiments, which were performed within the framework of the Frankfurt–Homburg Block Allocation Group (BAG, MX-1745 and MX-1836). We thank Erik Frotscher and Anne Grethen for help with CD and SEC-RALS, respectively. For critical reading of this manuscript, we thank Michael Schroda and Karolin Willmund. This work was supported by the Deutsche Forschungsgemeinschaft (TRR175-A05) and the Marie-Curie Career Integration Grant to FW and the Landesforschungsförderungsprogramm des Saarlandes (LFFP 11/02 and 15/04) to CRDL.
Author information
Authors and Affiliations
Contributions
F.R. purified proteins, conducted C.D., RALS, and SAXS. experiments and analysed SAXS data; Y.C. helped with SAXS measurements, performed differential scanning fluorimetry, analysed SAXS data and wrote parts of the manuscript; M.R. contributed immunofluorescence and comparison of purified proteins with cell lysates; K.G. purified the proteins; S.K. provided instruments for C.D. and SEC-RALS measurements and helped with data analysis; C.R.D.L. coordinated SAXS analysis and helped with SAXS data evaluation; F.W. designed experiments, prepared the figures and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ries, F., Carius, Y., Rohr, M. et al. Structural and molecular comparison of bacterial and eukaryotic trigger factors. Sci Rep 7, 10680 (2017). https://doi.org/10.1038/s41598-017-10625-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-10625-2
This article is cited by
-
Chloroplasten-Proteinbiogenese: angepasst an das phototrophe Leben
BIOspektrum (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.