Abstract
Ciliates are a suitable microbial model to investigate trait-dependent diversification because of their comparatively complex morphology and high diversity. We examined the impact of seven intrinsic traits on speciation, extinction, and net-diversification of rhynchostomatians, a group of comparatively large, predatory ciliates with proboscis carrying a dorsal brush (sensoric structure) and toxicysts (organelles used to kill the prey). Bayesian estimates under the binary-state speciation and extinction model indicate that two types of extrusomes and two-rowed dorsal brush raise diversification through decreasing extinction. On the other hand, the higher number of contractile vacuoles and their dorsal location likely increase diversification via elevating speciation rate. Particular nuclear characteristics, however, do not significantly differ in their diversification rates and hence lineages with various macronuclear patterns and number of micronuclei have similar probabilities to generate new species. Likelihood-based quantitative state diversification analyses suggest that rhynchostomatians conform to Cope’s rule in that their diversity linearly grows with increasing body length and relative length of the proboscis. Comparison with other litostomatean ciliates indicates that rhynchostomatians are not among the cladogenically most successful lineages and their survival over several hundred million years could be associated with their comparatively large and complex bodies that reduce the risk of extinction.
Similar content being viewed by others
Introduction
Diversity is a phenomenon of life. The asymmetry in the ability of lineages to diversify has become one of the central objectives in trait-dependent diversification studies1,2,3,4. It has been shown that large differences in species numbers between lineages might have originated in two ways, through increasing the speciation rate or decreasing the extinction rate. The high speciation rates cause a direct rise in the number of lineages, while the low extinction rates result in a gradual accumulation of lineages4,5,6. The cladogenic success is also connected with evolution of uniquely beneficial character states, so-called key innovations. Their detection is often associated with a quest for pronounced asymmetry in character state transition rates, i.e., convergent evolution when traits have appeared many times but have been rarely lost7, 8.
Diversification dynamics is an understudied field in microbial groups. Ciliates seem to be a suitable microbial model to explore trait-dependent diversification, because they are both species-rich and comparatively morphologically complex9, 10. These eukaryotic microbes are also diverse functionally, acting in a variety of roles such as fine to coarse filter feeders, hunters, scrapers or sucking feeders11, 12. Within the twelve recognised ciliate classes, the predatory way of life is mostly confined to the Litostomatea and the Prostomatea10. A morphologically very distinct group, the Rhynchostomatia, arose within the Litostomatea during the Palaeozoic period13. Rhynchostomatians are large to gigantic predatory ciliates, being from 100 μm up to 1,500 μm long14! Their raptorial lifestyle is uniquely associated with a conspicuous proboscis that carries batteries of toxicysts, which are extrusive organelles used to kill the prey, and a dorsal brush, which is a sensoric structure possibly involved in prey detection. Interestingly, rhynchostomatians are divided into two fundamental monophyletic groups with highly uneven distribution of taxa: the Tracheliida with just two species and the Dileptida with 64 species and subspecies14.
Using a range of statistical phylogenetic methods, we assessed clade-specific rates and searched for shifts in diversification rates during the evolutionary history of rhynchostomatians. Further, we investigated the impact of key quantitative and qualitative morphological features on speciation, extinction, and net-diversification rates of rhynchostomatian lineages. This complex approach enabled us to recognise which intrinsic traits increase diversification directly through elevating speciation rate, which indirectly via decreasing extinction rate, and which intrinsic traits do not cause asymmetries in cladogenesis.
Results
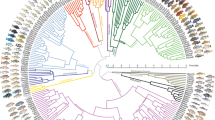
Concatenated and coalescent-based Bayesian phylogenetic analyses of the 18 S rRNA gene, ITS region, and first two domains of the 28 S rRNA gene brought similar results, consistent with previous studies15,16,17,18,19. The subclass Rhynchostomatia was divided into two fundamental lineages, the order Tracheliida and the order Dileptida, whose monophylies were strongly statistically supported by both analyses. Likewise, monophylies of the families Dileptidae and Dimacrocaryonidae were fully statistically supported. As concerns dimacrocaryonids, the subfamily Dimacrocaryoninae consistently received full support, while the subfamily Rimaleptinae was either strongly supported or was depicted as paraphyletic containing the former subfamily (Fig. 1; Supplementary Fig. S1). Since the internal relationships within the three main dileptid lineages were rather poorly resolved, the majority of analyses was performed on a set of 100 randomly selected post-burn-in trees from the coalescent inference.
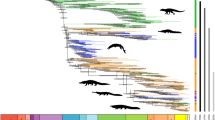
Maximum clade credibility species tree and corresponding cloudogram of the subclass Rhynchostomatia. The cloudogram is shown in green colour and consists of 7,500 superimposed post burn-in trees from the posterior distribution of the BEAST coalescent analysis. Branch colours designate tree partitions inferred to evolve at three significantly different rates by MEDUSA, with maximum-likelihood estimates of net-diversification rate (r) and extinction fraction (ε) for each tree partition. The tracheliid lineage diversified at a net-diversification rate of 4.88 (violet colour), while the dileptid clade at a net-diversification rate of 22.95 with an extinction fraction of 0.85 (red colour). However, there was a statistically significant shift in net-diversification rate to 14.91 within dileptids at the node uniting members with predominantly scattered macronuclear nodules (blue colour). The scale bar indicates number of substitutions. Only posterior probabilities > 0.95 are shown.
As concerns diversification of the subclass Rhynchostomatia, a birth-death model was favoured over a pure-birth model (BayesFactor BF = 28.24). Analyses of clade specific rates under the birth-death model indicated that the three main dileptid lineages (Dileptidae, Dimacrocaryoninae, and Rimaleptinae) diversified at a similar rate that was higher than in the tracheliid lineage (Supplementary Table S1). A similar pattern was also recognised by MEDUSA in that the tracheliid lineage diversified at a lower rate (r = 4.8780, yule model, AICc = 41.3196) than the dileptid clade (r = 22.9523, ε = 0.84624, birth-death model, AICc = 34.7045). MEDUSA also indicated that there was a statistically significant shift in diversification rate within the family Dileptidae, i.e., at the node uniting members with predominantly scattered macronuclear nodules (r = 14.9072, yule model, AICc = 37.1112) (Fig. 1).
According to reconstruction analyses, the most recent common ancestor (MRCA) of the order Tracheliida had an ordinary dorsal brush, i.e., all brush rows started at the top of the proboscis. In contrast, the MRCA of the order Dileptida possessed a staggered dorsal brush, i.e., brush rows were arranged in a step-like fashion (Fig. 2; Supplementary Table S2). Since there is a dramatic asymmetry in the number of taxa between these two orders (2 vs. 64), we speculate that transformation of this sensoric structure into a staggered pattern could be associated with the cladogenic success of the dileptid lineage. A further change in the dorsal brush structure happened in the MRCA of the subfamily Dimacrocaryoninae, when a two-rowed brush evolved (Fig. 2; Supplementary Table S2). The number of brush rows is also connected with a rather pronounced asymmetry, i.e., only about 35% of rhynchostomatian taxa have two-rowed brush while about 65% of rhynchostomatians still display more than two rows. Bayesian estimates under the binary-state speciation and extinction (BiSSE) model indicate that two-rowed dorsal brush is associated with reduced extinction rate (Table 1; Supplementary Table S3). Differences in net-diversification rates between two-rowed and multi-rowed lineages in MCMC simulations are distinctly shifted to the positive side of the x-axis (Fig. 3), indicating that lineages with two-rowed brush have a lower risk of extinction.
Maximum-parsimony character state reconstruction of five binary traits. The maximum clade credibility tree shown in Fig. 1 served as a scaffold for reconstruction of character state evolution. Squares at tips of branches show the character state of each taxon. For further details, see Supplementary Table S2. Explanations: CG, cortical granules; CV, contractile vacuoles; DB, dorsal brush; EX, extrusomes; P, proboscis.
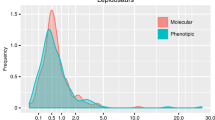
Posterior densities of differences in net-diversification rates between lineages with two-rowed and multi-rowed dorsal brush, between lineages with two types and one type of extrusomes, between lineages with dorsally and dorsally as well as ventrally located contractile vacuoles, and between lineages with more then two and two vacuoles. The x-axis represents the difference in net-diversification rates (∆r) between lineages with particular character states (∆r = r 1 − r 0; subscripts indicate different character states as specified in Supplementary Table S8). The ∆r distribution was obtained from MCMC simulations on a sample of 100 trees from the posterior distribution of the BEAST coalescent analysis. The y-axis represents posterior densities of differences in net-diversification rates, as calculated on the basis of the MCMC simulations. Arrows denote median of the difference in net-diversification rates (∆r). For interquartile ranges and further details, see Table 1. Ninety-five per cent intervals of highest posterior density are indicated below densities.
As concerns the number of extrusome types, reconstruction analyses suggest that the MRCAs of the order Tracheliida and the subfamily Dimacrocaryoninae had a single type of extrusomes anchored in the proboscis oral bulge, while the MRCAs of the family Dileptidae and the subfamily Rimaleptinae displayed two extrusome types (Fig. 2; Supplementary Table S2). The Bayesian BiSSE analyses indicate that two types of extrusomes are, similarly as the two-rowed dorsal brush, associated with reduced extinction rate (Table 1; Supplementary Table S3). This is also indirectly corroborated in that 76% of extant rhynchostomatian taxa possess two types of extrusomes. Differences in net-diversification rates between lineages with one type and two types of extrusomes in MCMC simulations are also conspicuously shifted towards the positive side of the x-axis (Fig. 3), suggesting significant difference in their rates.
The MRCAs of the order Tracheliida and the family Dileptidae possibly had many contractile vacuoles arranged both ventrally and dorsally. In contrast, the MRCAs of the subfamilies Dimacrocaryoninae and Rimaleptinae possessed two dorsally arranged vacuoles (Fig. 2; Supplementary Table S2). BiSSE indicates that the higher number of contractile vacuoles and their dorsal location increase diversification rate on average (Table 1, Fig. 3; Supplementary Table S3). This is also indirectly sustained in that 85% of taxa have more than two contractile vacuoles, and these are arranged dorsally in 72% of taxa. Although intervals of differences in net-diversification rates in MCMC simulations exceed zero towards the negative side of the x-axis (Fig. 3), mean values are significantly positive for both, the contractile vacuole pattern (t = 19.22, df = 9999, p < 2.2e-16, 95% confidence interval of mean 0.26–0.32) and the number of vacuoles (t = 28.90, df = 9999, p < 2.2e-16, 95% confidence interval of mean 0.43–0.49). Thus, Bayesian MCMC analyses suggest a trend that the higher number of contractile vacuoles and their dorsal location are associated with increasing diversification.
The MRCAs of the orders Tracheliida, Dileptida, and the family Dileptidae had multiple micronuclei, while the MRCA of the family Dimacrocaryonidae exhibited only a single micronucleus (Fig. 4; Supplementary Table S2). This shift had no distinct effect on diversification because 52% of rhynchostomatians still possess multiple micronuclei and 48% of taxa have a single micronucleus. Also according to BiSSE analyses, speciation and extinction rates were equal for both states (multiple micronuclei vs. single micronucleus) in the best fitting model (Table 1; Supplementary Table S3). Likewise, the multi-state speciation and extinction analyses (MuSSE) did not recognise any significant difference between models where speciation rates for particular states of the macronuclear pattern were constrained to be equal and where they were allowed to vary (χ2 = 0.1132, df = 3, p = 0.9902) (Supplementary Table S4). Nonetheless, the distribution of macronuclear patterns in extant taxa is rather variable, indicating some asymmetry especially in case of the compact macronucleus. Specifically, 38% of species exhibit two macronuclear nodules, 33% a moniliform strand, 21% many scattered nodules, and only 8% have a compact macronucleus16. In spite of this seeming unbalance among extant species, speciation rates of all four states strongly overlapped in MCMC simulations, where all speciation rates were allowed to vary (Fig. 4). This indicates that lineages with any state have similar probabilities to generate new species.
Maximum-parsimony character state reconstruction of micro- and macronuclear traits as well as posterior densities of speciation rates for four macronuclear patterns in Bayesian MuSSE analyses. The maximum clade credibility tree shown in Fig. 1 served as a scaffold for both reconstruction analyses. Squares at tips of branches show the character state of each taxon. The x-axis represents distribution of speciation rates for mononodulate macronucleus (yellow colour), binodulate macronucleus (blue colour), moniliform macronuclear strand (green colour), and many scattered macronuclear nodules (grey colour). Distributions of speciation rates were obtained from MCMC simulations on the maximum clade credibility tree in the R-package diversitree. The y-axis represents posterior densities of speciation rates calculated on the basis of the MCMC simulations with exponential prior and 10,000 samples. Ninety-five per cent intervals of highest posterior density are indicated below densities.
As concerns quantitative traits, a positive linear relationship between body size and diversification as well as between the relative proboscis length and diversification was preferred over other directional as well as constant and non-directional models analysed (Supplementary Table S5). Likelihood ratio test against the constant rate model supported these positive relationships for body length with χ2 = 5.4 (df = 2, p = 0.067) and for the relative proboscis length with χ2 = 5.1 (df = 2, p = 0.079).
Discussion
Asymmetry in diversification of rhynchostomatians
The class Litostomatea comprises a morphologically and ecologically diverse assemblage of free-living and endosymbiotic ciliates10. Bayesian relaxed molecular clock suggests that litostomateans emerged during the Cryogenian around 680 Mya and all fundamental litostomatean lineages diverged during the Paleozoic and Mesozoic periods13. Distribution of extant taxa is, however, rather uneven between the main free-living litostomatean lineages. According to the classification framework of Vďačný et al.20, there are over 250 species in the order Spathidiida21, more than 100 species in the order Pleurostomatida22,23,24,25,26, about 66 species in the Rhynchostomatia14, and only several dozens of species in the orders Haptorida, Didiniida, and Lacrymariida24, 27, 28. This cursory appraisal indicates that rhynchostomatians are not among the cladogenically most successful litostomateans. And, in addition, the two fundamental rhynchostomatian clades differ drastically in the number of known extant lineages. The order Tracheliida includes only two species, while the order Dileptida collates about 64 taxa14. Such a pronounced asymmetry has not been observed in any other litostomatean lineage, which provokes a question: what caused this conspicuous imbalance between fundamental rhynchostomatian clades?
Comparative analyses suggest that rhynchostomatians were caught in a plesiomorphic trap. In contrast to all other litostomateans, they still exhibit several ancient ciliate plesiomorphies, e.g., ventrally located cytostome, preoral kineties (~adoral membranelles) accompanying circumoral kinety (~paroral membrane), and formation of anarchic fields during morphogenesis15, 16, 20, 29, 30. In addition, the morphology of tracheliids has even a more plesiomorphic nature than that of dileptids and, hence, tracheliids are considered to be nearest in morphology to the putative last common progenitor of the Litostomatea20, 29. Specifically, tracheliids have an almost globular body with immobile proboscis whose dorsal side is distinctly shorter than the ventral one, their dorsal brush is not staggered, and their circumoral kinety is composed of dikinetids throughout. On the other hand, the last common ancestor of dileptids very likely possessed a narrow to almost rod-like body with agile proboscis whose ventral and dorsal sides are of a similar length, a staggered dorsal brush, and a hybrid circumoral kinety composed of dikinetids in the proboscis and of monokinetids around the cytostome14, 15, 18. We hypothesise that this suite of apomorphic features might be responsible for the pronounced cladogenic success of dileptids with respect to tracheliids. However, this hypothesis could not be directly tested with the BiSSE approach, because this technique is sensitive to strong tip ratio bias.
Traits with symmetry in net-diversification rates
The present analyses indicated that the dorsal brush structure, contractile vacuole apparatus, and the number of extrusome types have an impact on asymmetry in the ability of particular rhynchostomatian lineages to diversify. On the other hand, particular states of the number of micronuclei and the macronucelar pattern did not differ in their net-diversification rates (Table 1; Supplementary Tables S3 and S4). This is not surprising since the nuclear apparatus is of a paramount significance for survival31, 32. Therefore, we speculate that all nuclear patterns are under similar selection pressure, which in turn causes that lineages with any state have similar probabilities to generate new species. The evolutionary significance of the nuclear apparatus in rhynchostomatians was recognised already by Dragesco33 and Jankowski34, who reflected this in their proposals of classification concepts in dileptids. Later on, most of their suggestions have been corroborated with the onset of molecular phylogenetic techniques15,16,17,18,19.
However, we cannot exclude that various nuclear character states have some indirect effect on asymmetry in net-diversification rates. Thus, different nuclear character states may have different impacts on fitness and may be also correlated with other characters recognised as causing asymmetries in the ability of particular rhynchostomatian lineages to diversify. To cast more light onto this problem, an experimental setting uncovering mutations that negatively or positively influence fitness of one or other of the studied character traits would be needed.
Quantitative traits and diversification of rhynchostomatians
Quantitative traits have been long considered to affect speciation and extinction rates35, although the postulated hypotheses have been conflicting. Species with smaller bodies have been suggested to have higher speciation rates or lower extinction rates than species with larger bodies36,37,38. In contrast, Cope’s rule based on paleontological evidence has indicated that body size increases over evolutionary time, with descendent species tending to be larger than their ancestors39, 40. To overcome these problems, FitzJohn41 developed a likelihood-based method for testing hypotheses about quantitative state speciation and extinction. Using this approach we found support for increasing diversification in rhynchostomatians with growing body length and relative length of the proboscis (Supplementary Table S5). Indeed, rhynchostomatians are much larger than all other litostomateans. They are usually 150–500 μm long, with a maximum of 1,500 μm14. On the other hand, spathidiids are typically 60–150 μm long, with a maximum of 400 μm21 and pleurostomatids are mostly 50–200 μm long, with a maximum of 700 μm22,23,24,25,26. Since spathidiids and pleurostomatids are smaller but more diverse than rhynchostomatians, there must be clade-specific differences in diversification across litostomatean groups. Clade-specific diversification was also detected within primates, whereby elevation of speciation rates along with the increasing body size was concentrated only in the Cercopithecoidea and Hominoidea clades41. Taking into account the diversity and body size patterns of main litostomatean lineages, we hypothesise that large bodies increase the diversification of rhynchostomatians, but very likely through decreasing extinction.
Perspectives
The diversity patterns of ciliates, a major group of microbial eukaryotes, have become a subject of numerous studies9, 42,43,44. On the other hand, intrinsic traits responsible for shaping species richness of particular ciliate groups are largely unknown. Understanding trait-dependent diversification and relationships between evolution of intrinsic and extrinsic traits is principal for sound interpretation of the patterns observed. Therefore, future workers investigating ciliate biodiversity should consider interlinking morphological data with molecular phylogenies. Moreover, our analyses uncovered an interesting trend, survival of some ciliate groups over several hundred million years could be associated with comparatively large and complex bodies. This combination, however, rather lowers extinction than promotes speciation. Our hypothesis could be in the future tested on the ancient, mostly large-bodied karyorelictean ciliates.
Material and Methods
Collection, sample processing and sequencing
During our long-term research we have collected 19 out of ca. 54 comparatively well known rhynchostomatian species from all over the world. There are about 10 further insufficiently described taxa whose identity is obscure14. To minimise bias owing to incomplete taxon sampling, numbers of taxa were held more or less proportional to the size of the two main dileptid lineages. Thus, we sampled six out of the 24 well-described species from the family Dileptidae and 11 out of the 30 well-known species from the family Dimacrocaryonidae. Since we cannot exclude that our sampling is unbiased, potentially type II errors (failing to reject the null hypothesis when the alternate hypothesis is true) can occur in BiSSE analyses. To avoid these errors, we utilised simplified models as recommended by Davis et al.45. Further, we used a Bayesian framework and corrections for missing taxa by specifying the fraction of sampled taxa having particular character states, as recommended by Silvestro et al.46 (for further details, see ‘Diversification analyses’ below).
Newly sequenced species were collected from various terrestrial and aquatic habitats (Supplementary Table S6). Terrestrial samples were processed using the non-flooded Petri dish method, while aquatic samples were examined directly after transportation to the laboratory. Taxonomic methods followed Vďačný & Foissner14. After species identification, several cells were picked, washed and transferred into 1.5 ml microtubes, where they were stored for molecular investigations. Genomic DNA of the Spanish population of Rimaleptus binucleatus was extracted as described by Vďačný & Rajter18, while DNA isolation of all other species/populations was according to Jang et al.19. The 18 S rRNA gene, the ITS1–5.8S-ITS2 region, and the D1D2 domain of the 28 S rRNA gene were amplified according to Vďačný et al.16, Vďačný & Rajter18, Jang et al.19, Medlin et al.47, and Sonnenberg et al.48. PCR products were directly sequenced on an ABI 3730 automatic sequencer (Macrogen Inc., Seoul, Korea), using PCR primers and an internal primer (5′-CCG CGG TAA TTC CAG CTC-3′). Sequence fragments from individual samples were checked and assembled into contigs using Geneious ver. 6.1.6 (Biomatters, http://www.geneious.com/).
Tree-building methods
All available 18 S, ITS-region and D1D2 rhynchostomatian sequences were aligned and reliability of alignments was assessed on the GUIDANCE2 Server (http://guidance.tau.ac.il/ver2/), using the MAFFT algorithm and 100 bootstrap repeats49. The most appropriate nucleotide substitution models were chosen for each marker separately (Supplementary Table S7) with the help of the Akaike Information Criterion in jModelTest ver. 2.0.150. Bayesian inference was run under the best fitting evolutionary models in MrBayes ver. 3.2.151, using four independent chains, five million generations and a sample frequency of 1,000 on the CIPRES Portal ver. 1.1552. After reaching stationarity, posterior probabilities of a 50% majority-rule consensus tree were computed. The tree was a posteriori rooted according to our previous studies14,15,16,17,18, i.e. the root was placed on a branch separating the orders Tracheliida and Dileptida.
Apart from the Bayesian gene tree, a species tree was constructed using the Bayesian multispecies coalescent-based approach, as implemented in the program BEAST ver. 2.4.453. This analysis was conducted using five datasets (18 S, ITS1, 5.8 S, ITS2, D1D2). The software BEAUti ver. 2.4.4 was used to create a *BEAST input file with the following settings: (i) GTR + I + Γ evolutionary model for the 18 S dataset, HKY + I + Γ model for the ITS1 dataset, TN93 + I model for the 5.8 S dataset, TN93 + Γ model for the ITS2 dataset, and GTR + Γ model for the D1D2 dataset; (ii) uncorrelated lognormal clock; and (iii) Yule process model for the species tree prior. MCMC analysis was run for 150,000,000 generations, sampling every 15,000th generation. The convergence of all parameters to the stationary distribution in each run was inspected using the program Tracer ver. 1.654. The final tree was summarized in TreeAnnotator ver. 2.4.4 using a 25% burn-in. The coalescent tree was rooted using the molecular clock technique, as implemented in the multispecies coalescent-based approach. The resulting position of the root was congruent with results from our previous phylogenetic analyses of rhynchostomatians, where outgroup taxa were used to root the tree15,16,17.
Reconstruction of character evolution
We selected seven morphological characters that play a key role in classification of rhynchostomatians (Supplementary Table S8). Morphological data were obtained and scored according to our previous studies14, 15, 18. Character history was estimated for key nodes on the maximum clade credibility consensus tree, using the maximum-parsimony method as implemented in MESQUITE55 as well as on 100 trees from the posterior distribution of the BEAST coalescent analysis, using Bayesian and maximum-likelihood methods as implemented in BayesTraits ver. 1.056.
Diversification analyses
Diversification rates were estimated under pure-birth and birth-death models, using a Bayesian framework in the program BayesRate ver. 1.3.4146. This approach accommodates uncertainty in the inferred phylogeny, speciation and extinction-rate parameters, and missing species by specifying the proportion of taxa sampled. Fit of the models with and without extinction was assessed by Bayes factors, for which log marginal likelihoods were estimated over 100 trees from the posterior distribution of the BEAST coalescent analysis, using the thermodynamic integration option. Diversification-rate shifts were identified with maximum-likelihood technique, applied to the maximum clade credibility tree in MEDUSA57. This analysis also employed correction for incomplete taxon sampling and unsampled species were assigned to the extant clades according to Vďačný & Foissner14 and Vďačný & Rajter18.
Character-state associated diversification was estimated using the binary-state (BiSSE), multi-state (MuSSE), and qualitative trait (QuaSSE) methods. Since BiSSE and related methodologies are affected by the sample size and tip ratio bias, we minimised the potential of accepting the null hypothesis when the alternate hypothesis is true (type II error) by a combination of two strategies: (i) specification of the proportion of the taxa sampled for each character state and (ii) reduction of the number of model parameters. Our strategy followed Silvestro et al.46, who implemented a correction for missing taxa by specifying the fraction of sampled taxa having particular character states (Supplementary Table S9), and Davis et al.45, who recognised that simplified models can achieve greater power when tree size is small. Thus, as recommended by Davis et al.45, none of our tested BiSSE models had more than four free parameters.
BiSSE analyses were performed on dorsal brush, contractile vacuole and extrusome features in the program BayesRate. The Bayesian framework allows a more robust hypothesis testing, because it accounts for uncertainty in the inferred phylogeny as well as in the speciation and extinction-rate parameters. We tested four nested diversification models: (i) model 1 allowed both speciation and extinction rates to vary in both states; (ii) model 2 constrained speciation rates to be equal in both states but allowed extinction rates to vary; (iii) model 3 constrained extinction rates to be equal in both states but allowed speciation rates to vary; and (iv) model 4 constrained speciation and extinction rates to be equal in both states. Bayesian analyses were performed for each model over a sample of 100 trees from the posterior distribution of the BEAST coalescent analysis, running the MCMC simulation for 220 iterations of slice sampling per tree, discarding the first 20 samples per tree as burn-in. Corrections for incomplete taxon sampling were employed and diffuse priors were set for the diversification-rate parameters as suggested by Johnson et al.7. Fit of the models was assessed by Bayes factors, for which log marginal likelihoods were estimated over 100 trees from the posterior distribution, using the thermodynamic integration option. Dorsal brush pattern was not subjected to the BiSSE analyses because ordinary brush occurs only in two tracheliid species and staggered brush in all dileptids, which causes a strong tip ratio bias that could not be appropriately handled by BiSSE45.
MuSSE analyses were performed for macronuclear characteristics and QuaSSE analyses for body length and relative length of the proboscis, using the R-package diversitree41, 58. Maximum-likelihoods were estimated for several models, as specified in Supplementary Tables S4 and S5, on the maximum clade credibility consensus tree. We assessed the relative fit of the models to the data by calculating differences in AIC scores and AIC weights as well as by performing likelihood ratio test. Speciation rate of particular macronuclear characteristics was also subjected to Bayesian MCMC analysis with exponential prior and 10,000 samples.
Data availability
Accession numbers of newly obtained DNA sequences are listed in Supplementary Table S6 and are available at https://www.ncbi.nlm.nih.gov/nucleotide/. Results of all analyses are included in this published article and its Supplementary information files. The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Maddison, W. P., Midford, P. E. & Otto, S. P. Estimating a binary character’s effect on speciation and extinction. Syst. Biol. 56, 701–710 (2007).
Fuchs, J., Johnson, J. A. & Mindell, D. P. Rapid diversification of falcons (Aves: Falconidae) due to expansion of open habitats in the Late Miocene. Mol. Phylogenet. Evol. 82, 166–182 (2015).
Sinn, B. T., Kelly, L. M. & Freudenstein, J. V. Putative floral brood-site mimicry, loss of autonomous selfing, and reduced vegetative growth are significantly correlated with increased diversification in Asarum (Aristolochiaceae). Mol. Phylogenet. Evol. 89, 194–204 (2015).
de Vos, J. M., Hughes, C. E., Schneeweiss, G. M., Moore, B. R. & Conti, E. Heterostyly accelerates diversification via reduced extinction in primroses. Proc. R. Soc. B 281, 20140075 (2014).
Nee, S. Birth–death models in macroevolution. Ann. Rev. Ecol. Syst. 31, 1–17 (2006).
Benton, M. J. & Emerson, B. C. How did life become so diverse? The dynamics of diversification according to the fossil record and molecular phylogenetics. Palaeontology 50, 23–40 (2007).
Johnson, M. T. J., FitzJohn, R. G., Smith, S. D., Rausher, M. D. & Otto, S. P. Loss of sexual recombination and segregation is associated with increased diversification in evening primroses. Evolution 65, 3230–3240 (2011).
Armbruster, W. S. Floral specialization and angiosperm diversity: phenotypic divergence, fitness trade-offs and realized pollination accuracy. AoB Plants 6, plu003 (2014).
Foissner, W., Chao, A. & Katz, L. A. Diversity and geographic distribution of ciliates (Protista: Ciliophora). Biodivers. Conserv. 17, 345–363 (2008).
Lynn, D. H. The Ciliated Protozoa. Characterization, Classification, and Guide to the Literature. 3rd edn (Springer Verlag, 2008).
Fenchel, T. Ecology of Protozoa. The Biology of Free-living Phagotrophic Protists. (Springer Verlag, 1987).
Weisse, T. et al. Functional ecology of aquatic phagotrophic protists – concepts, limitations, and perspectives. Eur. J. Protistol. 55, 50–74 (2016).
Vďačný, P. Estimation of divergence times in litostomatean ciliates (Ciliophora: Intramacronucleata), with Bayesian relaxed clock and 18S rRNA gene. Eur. J. Protistol. 51, 321–334 (2015).
Vďačný, P. & Foissner, W. Monograph of the dileptids (Protista, Ciliophora, Rhynchostomatia). Denisia 31, 1–529 (2012).
Vďačný, P. et al. Morphological and molecular phylogeny of dileptids: resolution at the base of the class Litostomatea (Ciliophora, Intramacronucleata). Eur. J. Protistol. 47, 295–313 (2011).
Vďačný, P., Bourland, W. A., Orsi, W., Epstein, S. S. & Foissner, W. Genealogical analyses of multiple loci of litostomatean ciliates (Protista, Ciliophora, Litostomatea). Mol. Phylogenet. Evol. 65, 397–411 (2012).
Vďačný, P. et al. The chaos prevails: molecular phylogeny of the Haptoria (Ciliophora, Litostomatea). Protist 165, 93–111 (2014).
Vďačný, P. & Rajter, Ľ. Reconciling morphological and molecular classification of predatory ciliates: evolutionary taxonomy of dileptids (Ciliophora, Litostomatea, Rhynchostomatia). Mol. Phylogenet. Evol. 90, 112–128 (2015).
Jang, S. W., Vďačný, P., Shazib, S. U. A. & Shin, M. K. Morphological and molecular characterization of the name-bearing type species Rimaleptus binucleatus (Kahl, 1931), with a phylogenetic re-analysis of dileptid evolutionary history (Ciliophora: Litostomatea: Rhynchostomatia). Eur. J. Protistol. 50, 456–471 (2014).
Vďačný, P., Bourland, W. A., Orsi, W., Epstein, S. S. & Foissner, W. Phylogeny and classification of the Litostomatea (Protista, Ciliophora), with emphasis on free-living taxa and the 18S rRNA gene. Mol. Phylogenet. Evol. 59, 510–522 (2011).
Foissner, W. & Xu, K. Monograph of the Spathidiida (Ciliophora, Haptoria). Vol. I: Protospathidiidae, Arcuospathidiidae, Apertospathulidae. Monograph. Biol. 81, 1–485 (2007).
Kahl, A. Urtiere oder Protozoa I: Wimpertiere oder Ciliata (Infusoria) 2. Holotricha außer den im 1. Teil behandelten Prostomata. Tierwelt Dtl. 21, 181–398 (1931).
Song, W. & Wilbert, N. Taxonomische Untersuchungen an Aufwuchsciliaten (Protozoa, Ciliophora) im Poppelsdorfer Weiher, Bonn. Lauterbornia 3, 2–221 (1989).
Foissner, W., Berger, H., Blatterer, H. & Kohmann, F. Taxonomische und ökologische Revision der Ciliaten des Saprobiensystems – Band IV: Gymnostomatea, Loxodes, Suctoria. Informationsberichte des Bayerischen Landesamtes für Wasserwirtschaft 1/95, 1–540 (1995).
Lin, X., Warren, A. & Song, W. Pleurostomatids in Free-living Ciliates in the Bohai and Yellow Seas, China (eds Song, W., Warren, A. & Hu, X.) 93–134 (Science Press, 2009).
Vďačný, P. & Rajter, Ľ. An annotated and revised checklist of pleurostome ciliates (Protista: Ciliophora: Litostomatea) from Slovakia, Central Europe. Zootaxa 3760, 501–521 (2014).
Kahl, A. Urtiere oder Protozoa I: Wimpertiere oder Ciliata (Infusoria) 1. Allgemeiner Teil und Prostomata. Tierwelt Dtl. 18, 1–180 (1930).
Foissner, W., Agatha, S. & Berger, H. Soil ciliates (Protozoa, Ciliophora) from Namibia (Southwest Africa), with emphasis on two contrasting environments, the Etosha Region and the Namib Desert. Denisia 5, 1–1459 (2002).
Vďačný, P., Orsi, W. & Foissner, W. Molecular and morphological evidence for a sister group relationship of the classes Armophorea and Litostomatea (Ciliophora, Intramacronucleata, Lamellicorticata infraphyl. nov.), with an account on basal haptorid litostomateans. Eur. J. Protistol. 46, 298–309 (2010).
Vďačný, P. & Foissner, W. Ontogenesis of Dileptus terrenus and Pseudomonilicaryon brachyproboscis (Ciliophora, Haptoria). J. Eukaryot. Microbiol. 56, 232–243 (2009).
Paulin, J. J. Morphology and cytology of ciliates in Ciliates: Cells as Organisms (eds Hausmann, K. & Bradbury, P. C.) 1–40 (Gustav Fischer Verlag, 1996).
Raikov, I. B. Nuclei of ciliates in Ciliates: Cells as Organisms (eds Hausmann, K. & Bradbury, P. C.) 221–242 (Gustav Fischer Verlag, 1996).
Dragesco, J. Révision du genre Dileptus, Dujardin 1871 (Ciliata Holotricha) (systématique, cytologie, biologie). Bull. Biol. Fr. Belg. 97, 103–145 (1963).
Jankowski, A. W. Novye rody klassov Gymnostomatea i Ciliostomea [New genera of the classes Gymnostomatea and Ciliostomea]. Mater. IV. Konf. Uc. Sekc. Zool. (Kishinev) year 1967, 36 (1967).
Gittleman, J. & Purvis, A. Body size and species-richness in carnivores and primates. Proc. R. Soc. Lond. B Biol. Sci. 265, 113–119 (1998).
Cardillo, M. et al. Multiple causes of high extinction risk in large mammals. Science 309, 1239–1241 (2005).
Paradis, E. Statistical analysis of diversification with species traits. Evolution 59, 1–12 (2005).
Clauset, A. & Erwin, D. H. The evolution and distribution of species body size. Science 321, 399–401 (2008).
Alroy, J. Cope’s rule and the dynamics of body mass evolution in North American fossil mammals. Science 280, 731–734 (1988).
Jablonski, D. Body-size evolution in Cretaceous molluscs and the status of Cope’s rule. Nature 385, 250–252 (1997).
FitzJohn, R. G. Quantitative traits and diversification. Syst. Biol. 59, 619–633 (2010).
Filker, S., Sommaruga, R., Vila, I. & Stoeck, T. Microbial eukaryote plankton communities of high-mountain lakes from three continents exhibit strong biogeographic patterns. Mol. Ecol. 25, 2286–2301 (2016).
Zhao, F. & Xu, K. Microbial genetic diversity and ciliate community structure along an environmental gradient in coastal soil. Eur. J. Protistol. 49, 516–525 (2013).
Zhao, F. & Xu, K. Biodiversity patterns of soil ciliates along salinity gradients. Eur. J. Protistol. 53, 1–10 (2016).
Davis, M. P., Midford, P. E. & Maddison, W. Exploring power and parameter estimation of the BiSSE method for analyzing species diversification. BMC Evol. Biol. 13, 38 (2013).
Silvestro, D., Schnitzler, J. & Zizka, G. A Bayesian framework to estimate diversification rates and their variation through time and space. BMC Evol. Biol. 11, 311 (2011).
Medlin, L., Elwood, H. J., Stickel, S. & Sogin, M. L. The characterization of enzymatically amplified eukaryotic l6S-like rRNA-coding regions. Gene 71, 491–499 (1988).
Sonnenberg, R., Nolte, A. W. & Tautz, D. An evaluation of LSU rDNA D1-D2 sequences for their use in species identification. Front. Zool. 4, 6 (2007).
Sela, I., Ashkenazy, H., Katoh, K. & Pupko, T. GUIDANCE2: accurate detection of unreliable alignment regions accounting for the uncertainty of multiple parameters. Nucleic Acids Res. 43, W7–W14 (2015).
Posada, D. jModelTest: phylogenetic model averaging. Mol. Biol. Evol. 25, 1253–1256 (2008).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Miller, M. A., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees in Proceedings of the Gateway Computing Environments Workshop (ed. GCE) 1–8 (New Orleans, 2010).
Bouckaert, R. et al. BEAST 2: A Software Platform for Bayesian Evolutionary Analysis. PLoS Comput. Biol. 10, e1003537 (2014).
Rambaut, A. & Drummond, A. J. Tracer v1.4. http://beast.bio.ed.ac.uk/Tracer (2007).
Maddison, W. P. & Maddison, D. R. Mesquite: A Modular System for Evolutionary Analysis. Version 2.0. http://mesquiteproject.org (2007).
Pagel, M. & Meade, A. Bayesian analysis of correlated evolution of discrete characters by reversible-jump Markov chain Monte Carlo. Am. Nat. 167, 808–825 (2006).
Alfaro, M. E. et al. Nine exceptional radiations plus high turnover explain species diversity in jawed vertebrates. Proc. Natl. Acad. Sci. USA 106, 13410–13414 (2009).
FitzJohn, R. G. diversitree: comparative phylogenetic analyses of diversification in R. Methods Ecol. Evol. 3, 1084–1092 (2012).
Acknowledgements
We are very grateful to Ing. Ivan Rúrik for his help and thoughtful comments. This work was supported by the Slovak Research and Development Agency under the contract No. APVV-15-0147, by the Grant Agency of the Ministry of Education, Science, Research and Sport of the Slovak Republic and Slovak Academy of Sciences under the Grant VEGA 1/0041/17, and by the National Research Foundation of Korea funded by the Korean government of MOE and MSIP under Grants 2015R1D1A1A09058911 and 2015M1A5A1041804. This study was also supported by the Comenius University in Bratislava (Grant Nos UK/40/2016 and UK/400/2017).
Author information
Authors and Affiliations
Contributions
P.V. and M.K.S. designed the research. P.V., S.W.J. and M.K.S. conducted fieldwork. S.U.A.S., L.R. and S.W.J. conducted laboratory work. P.V. prepared the data sets, conducted analyses and wrote a first draft of the article. All authors contributed to revisions of the draft.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vďačný, P., Rajter, Ľ., Shazib, S.U.A. et al. Diversification dynamics of rhynchostomatian ciliates: the impact of seven intrinsic traits on speciation and extinction in a microbial group. Sci Rep 7, 9918 (2017). https://doi.org/10.1038/s41598-017-09472-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-09472-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.