Abstract
Breast cancer is a serious problem causing the death of women across the world. At present, one of the major challenges is to design drugs to target breast cancer specific gene(s). RNA interference (RNAi) is an important technique for targeted gene silencing that may lead to promising novel therapeutic strategies for breast cancer. Therefore, identification of such molecules having high oncogene specificity is the need of the hour. Here, we have developed a database named as Breast Oncogenic Specific siRNAs (BOSS, http://bioinformatics.cimap.res.in/sharma/boss/) on the basis of the current research status on siRNA-mediated repression of oncogenes in different breast cancer cell lines. BOSS is a resource of experimentally validated breast oncogenic siRNAs, collected from research articles and patents published yet. The present database contains information on 865 breast oncogenic siRNA entries. Each entry provides comprehensive information of an siRNA that includes its name, sequence, target gene, type of cells, and inhibition value, etc. Additionally, some useful tools like siRNAMAP and BOSS BLAST were also developed and linked with the database. siRNAMAP can be used for the selection of best siRNA against a target gene while BOSS BLAST tool helps to locate the siRNA sequences in deferent oncogenes.
Similar content being viewed by others
Introduction
Breast cancer is one of the major cause of women death in India as well as throughout the world1. Breast cancer is a result of mutation in the genes involved in regulation of cell growth and proliferation2,3,4,5,6,7,8,9. In the process of breast cancer, genes are mutated which may result in gain- or loss-of-function that contribute to the malignant phenotype8. These mutations may be the consequence of spontaneous mutations, environmental factors, viral infection, etc. Anti-cancer drugs target the proteins encoded by these mutated oncogenes3,4,5,6,7,8,9,10,11. Amplification and overexpression of breast oncogenes are the major mechanisms through which these genes participate in the oncogenesis4. RNA interference (RNAi) was first reported by Fire et al. in C. elegans and has been used as a noble technique for cancer gene therapy5. Several studies have revealed the importance of short interfering RNAs (siRNAs) and short hairpin RNAs (shRNAs) in RNAi-mediated silencing of oncogenes as a potential therapeutic strategy for cancer6, 7. siRNAs are generally 19–25 nucleotides in length and have sequence specific gene knockdown capability. Reports on the basis of transfection of synthetic 21- and 22-nucleotide siRNAs with overhanging 3′ ends indicate that siRNA may be a powerful tool to suppress the target-specific gene expression7. In the last decade, several studies reported the use of numerous siRNAs and shRNAs for cancer gene therapy. There are few databases like DSTHO8, siRecords9, siRNAdb10, HuSiDa11, which focus on siRNAs targeting genes of human and other mammals. However, to the best of our knowledge, there is no comprehensive database of siRNAs/shRNAs targeting breast oncogenes which is required to search and analyse the data from the literature. With this in mind, a manually curated specialised databank, “BOSS”, has been developed on the basis of information from experimentally validated and published siRNAs and shRNAs targeting various breast oncogenes to facilitate research on RNAi-based cancer therapy. BOSS database maintains the comprehensive information of breast oncogenic specific siRNAs and comprises information about siRNAs name, target genes, inhibition value, cell line, siRNA sequence, NCBI Accession No., transfection reagent, test method, test objective and Pubmed ID etc. which are directly linked to essential databases. This database is an organised database where breast oncogenic specific siRNAs information is collected from literature and other existing databases to make it an informative tool for the researchers working in this field.
Construction and Content
To develop a comprehensive database for breast oncogenic specific siRNAs, an extensive search was carried out to collect information on siRNA, shRNA and breast cancer. For this, first research articles and patents providing information related to breast oncogenic specific siRNAs were collected from various search engines like PubMed12 and Patent lens13. Specific searches were carried out using a combination of keywords like ‘siRNA’, ‘shRNA’, ‘Breast cancer’, ‘mammary cancer’, ‘cancer’, ‘gene therapy’, ‘gene silencing’ and ‘RNAi’. This exhaustive search yielded around 5613 research articles and 10 patents. From these articles and patents, only experimentally verified breast oncogenic specific siRNAs were retrieved manually. After careful reading of these articles, we scrutinised 88 research articles and 2 patents. Articles were carefully screened and about 865 siRNAs/shRNAs entries targeting breast oncogene with experimental studies were selected to be included in the database. Although most of the siRNA or shRNA entries have been provided with the inhibition value, qualitative representations were considered and included in the database if the quantitative values for siRNAs or shRNA were not given in the corresponding reports. These reports have evaluated the expression level of different test targets (i.e. mRNA, protein, etc) in breast cancer samples14,15,16,17,18,19,20,21,22,23,24,25. The most common method used to evaluate the efficacy was MTT assay. Different experimental methods like WST-8 Assay, RT-PCR, Western blotting etc. were used to validate breast oncogenic specifics siRNAs. Furthermore, 195 siRNAs derived from different assays have also been incorporated with information regarding their alternative efficacies.
Database Architecture
As shown in Fig. 1, the BOSS database contains the following seventeen fields for each siRNAs entry; (1) BOSS id (2) PubMed id, (3) Sequence, (4) siRNA name, (5) Target gene, (6) GC content, (7) Length of siRNA, (8) Cell types, (9) Year, (10) siRNA source (siRNA/shRNA), (11) Position of siRNA, (12) Test objective, (13) Test method, (14) Gene Bank Accession No., (15) Biological inhibition, (16) Transfection Reagent and (17) Test time.
Database Construction and Maintenance
Database web interface and architecture
BOSS DB is built on Apache HTTP 2.4.9 and MySQL 5.6.17 Servers at the backend, whereas the front-end is built using HTML, PHP, jQuery, and Perl. MySQL is a management system for Open Source Relational SQL database. BOSS web interface and database interfacing scripts have been written in PHP, HTML, PERL, CSS and Java integration programming languages. BOSS database comprises diverse types of information about each siRNA entries, which is collected from different resources. Figure 1 shows the schematic representation of architecture of BOSS database.
Utility
In order to facilitate the users’ search, several web-based tools have been integrated into the database that includes search, advance search and browsing options (Fig. 1).
Search and advance search tool
The search tool is provided for searching all the seventeen fields or selected fields of the database. The advance search provides a refined way to search the database using several combinations of keywords of related fields using logical operators “AND” & “OR” for more specific results.
Browse
A powerful browsing facility has been provided with BOSS database that allows users to browse data using various options. A short description of web interface designed for browsing are as follows:
This option allows the user to browse BOSS database based upon main categories like siRNA Name, Target Gene, Cell Type, Inhibition, GC Content and Test Time of breast oncogenic specific siRNAs. The user can select any of the categories and click the button to find a list of breast oncogenic specific siRNA related to the particular category.
Web-Based Tools
siRNAMAP
The siRNAMAP maps the BOSS database siRNAs on the basis of query nucleotide sequence. It will provide the list of siRNA from the BOSS database which is complementary to the of the user query sequences.
Blast
This tool can be used for similarity-based search of any query sequences with those present in the BOSS database. By using this, user can examine whether a given breast oncogene-specific siRNA sequence or similar siRNA sequence has already been reported or not. The user can submit a query sequence in the Fasta format in the search field and press the submit button. It will display all breast oncogene-specific siRNAs similar to the query sequence. The server provides the option to modify different parameters like scoring matrices, gap penalty, word size etc26.
Data Statistics and Findings
siRNA designing for knock-down target gene expression has been a major challenge. It is obvious from the earlier reports that only a fraction of designed siRNAs is highly effective in gene silencing15,16,17,18,19,20,21,22,23,24,25, 27, 28. Therefore, gene silencing experiments require a cost and labour-intensive optimisation protocol for designing and selection of efficient siRNAs and their delivery into the target cell lines. BOSS database consists of 865 entries for 195 unique breast oncogene-specific siRNAs/shRNAs. It comprises of diverse types of information about each siRNA entry, which is collected from the different resources. The database is an endeavour in the direction of RNA interference (RNAi) for breast cancer gene therapy. While searching for breast oncogene-specific siRNAs in the literature, it was observed that most siRNAs had been tested on various breast cancer cell lines showing different biological inhibition values. The percent efficacy ranges from 0–100 in BOSS database. Negative efficacy values reported in a particular experiment with respect to control was considered as zero, for the sake of simplicity.
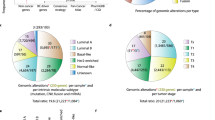
As shown in Fig. 2, number of si/shRNAs with inhibition percentage 81–100, 61–80, 41–60, 21–40 and 0–20, were 26%, 23%, 20%, 18% and 13%, respectively.
The database entries contain siRNA experimentally validated using 23 different cell lines but MCF-7, MDA-MB-231, SKBR-3, MDA-MB-435s and MDA-MB-468 cell lines were mostly used. Different cell lines used to examine siRNA-mediated suppression of different oncogenes are given in supplementary information. There are a number of siRNAs in our database, whose breast oncogenic activity (knockdown efficacy) has already been tested against several types of breast cancer cell lines. Although MCF-7 and MDA-MB-231 were used in most of the cases, this database has siRNA entries targeting a breast oncogene in all the reported breast cancer cell lines. The overall statistics of the different cell lines is depicted in Fig. 3. MCF-7 and MDA-MD-231 lines were used for 47% and 19% siRNAs entries, respectively.
Percentage of different si/shRNAs included in the database suppressing the target gene expression.
The BOSS database provides sequences of reported functional siRNAs targeting breast oncogenes and other technical details of the corresponding experiments, including used cell lines, transfection reagents and direct links from the published references. Out of 865 entries, 477 entries were found in Genbank database that lies in 37 different breast genome regions. Users can explore information about the siRNAs/shRNAs sequences, target Homo sapiens genome region, efficacies and the experimental conditions prior to their experiments in user-friendly manner using the search and browsing facility. BOSS database provides experimentally validated siRNAs reported in literature targeting diverse genes of Homo sapiens genome region. The majority of the reported breast oncogenes targeted by RNAi-mediated suppression were NLK (18%), IKKƐ (4%), TTK32 (4%), EGFR (3%) and AurkB (3%) respectively (Fig. 4). Different breast oncogenes targeted by siRNA-mediated suppression are given in supplementary information. The transfection reagents oligofectamine and lipofectamine covers 18% and 35% of the database, respectively. G + C content is the crucial character for functional siRNAs29. The G + C content profile was used for visualisation and analysing the variation of GC content in genomic sequences30. DNA molecules were made up of four nitrogenous bases A, T, G, C. These four nitrogenous bases made different number of hydrogen bonds with each other. Due to three H-bonds between G and C, this base pair is stronger than that of A and T. This makes high G + C containing DNA thermally more stable than AT containing DNA29. It has been reported that sequences of intermediate G + C contents (around 50%) were more effective siRNAs, and our dataset contained 86% of siRNA in the intermediate range (30–65%) G + C content31.
Discussion
Comparison with other databases
Very few databases of siRNAs are available viz., DSTHO8, siRecords9, siRNAdb10, HuSiDa11, which focus on siRNAs targeting genes of both human and other mammals. HuSiDa and siRNAdb are based on published functional siRNA targeting human genes. DSTHO is based on human oncogenes but not for breast oncogenes. The lack of updates and comprehensiveness is another problem with this database8. The present database on breast oncogenic specific siRNAs will be useful in designing and/or evaluating the breast oncogene-specific si/shRNAs, such as VIRsiRNAdb32 database which is specific for experimentally validated viral-specific siRNA/shRNA. In our earlier study, we have also developed, curated and developed a database named HIVsirDB, HIV specific siRNAs against HIV28.
Conclusion
siRNA has been proven to be a valuable tool for knocking down the expression of specific human genes. siRNA exhibit a high degree of specificity and has important medical implementations such as oncogene repression in cancer. The uniqueness of the system, that makes it a powerful tool is the sequence specificity towards a particular gene. It is fast, easy and the most cost-effective processing. This oncogene-specific RNA interference will offer plenty of opportunities for the researchers in exploring the role of breast oncogene specific siRNAs in breast cancer. In addition, siRNA designing algorithms also help to make effective molecules. To the best of the authors’ knowledge, no such database is available for breast oncogene siRNA.
References
Jemal, A. et al. Global cancer statistics. CA. Cancer J. Clin. 61, 69–90 (2011).
Sledge, G. W. & Miller, K. D. Exploiting the hallmarks of cancer: the future conquest of breast cancer. Eur. J. Cancer Oxf. Engl. 1990 39, 1668–1675 (2003).
Croce, C. M. Oncogenes and Cancer. N. Engl. J. Med. 358, 502–511 (2008).
Osborne, C., Wilson, P. & Tripathy, D. Oncogenes and tumor suppressor genes in breast cancer: potential diagnostic and therapeutic applications. The Oncologist 9, 361–377 (2004).
Timmons, L. & Fire, A. Specific interference by ingested dsRNA. Nature 395, 854–854 (1998).
Zheng, Y. et al. Scavenger receptor B1 is a potential biomarker of human nasopharyngeal carcinoma and its growth is inhibited by HDL-mimetic nanoparticles. Theranostics 3, 477–486 (2013).
Elbashir, S. M. et al. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494–498 (2001).
Dash, R., Moharana, S. S., Reddy, A. S., Sastry, G. M. & Sastry, G. N. DSTHO: database of siRNAs targeted at human oncogenes: a statistical analysis. Int. J. Biol. Macromol. 38, 65–69 (2006).
Ren, Y. et al. siRecords: a database of mammalian RNAi experiments and efficacies. Nucleic Acids Res. 37, D146–149 (2009).
Chalk, A. M., Warfinge, R. E., Georgii-Hemming, P. & Sonnhammer, E. L. L. siRNAdb: a database of siRNA sequences. Nucleic Acids Res. 33, D131–134 (2005).
Truss, M. et al. HuSiDa–the human siRNA database: an open-access database for published functional siRNA sequences and technical details of efficient transfer into recipient cells. Nucleic Acids Res. 33, D108–111 (2005).
pubmeddev. Home - PubMed - NCBI. Available at: https://www.ncbi.nlm.nih.gov/pubmed. (Accessed: 21st February 2017).
The Lens. Available at: https://www.lens.org/lens/. (Accessed: 21st February 2017).
Dressing, G. E., Alyea, R., Pang, Y. & Thomas, P. Membrane progesterone receptors (mPRs) mediate progestin induced antimorbidity in breast cancer cells and are expressed in human breast tumors. Horm. Cancer 3, 101–112 (2012).
Sun, L. et al. Knockdown of S-phase kinase-associated protein-2 expression in MCF-7 inhibits cell growth and enhances the cytotoxic effects of epirubicin. Acta Biochim. Biophys. Sin. 39, 999–1007 (2007).
Abdelrahim, M., Smith, R. & Safe, S. Aryl hydrocarbon receptor gene silencing with small inhibitory RNA differentially modulates Ah-responsiveness in MCF-7 and HepG2 cancer cells. Mol. Pharmacol. 63, 1373–1381 (2003).
Qiao, H. et al. Synergistic suppression of human breast cancer cells by combination of plumbagin and zoledronic acid In vitro. Acta Pharmacol. Sin. 36, 1085–1098 (2015).
Qin, B. & Cheng, K. Silencing of the IKKε gene by siRNA inhibits invasiveness and growth of breast cancer cells. Breast Cancer Res. BCR 12, R74 (2010).
Liang, Y. et al. siRNA-based targeting of cyclin E overexpression inhibits breast cancer cell growth and suppresses tumor development in breast cancer mouse model. PloS One 5, e12860 (2010).
Li, J. et al. Role for ezrin in breast cancer cell chemotaxis to CCL5. Oncol. Rep. 24, 965–971 (2010).
Pillé, J.-Y. et al. Anti-RhoA and anti-RhoC siRNAs inhibit the proliferation and invasiveness of MDA-MB-231 breast cancer cells in vitro and in vivo. Mol. Ther. J. Am. Soc. Gene Ther. 11, 267–274 (2005).
Glondu-Lassis, M. et al. PTPL1/PTPN13 regulates breast cancer cell aggressiveness through direct inactivation of Src kinase. Cancer Res. 70, 5116–5126 (2010).
Howlin, J., Rosenkvist, J. & Andersson, T. TNK2 preserves epidermal growth factor receptor expression on the cell surface and enhances migration and invasion of human breast cancer cells. Breast Cancer Res. BCR 10, R36 (2008).
Groth-Pedersen, L. et al. Identification of cytoskeleton-associated proteins essential for lysosomal stability and survival of human cancer cells. PloS One 7, e45381 (2012).
Luo, X.-G., Zou, J.-N., Wang, S.-Z., Zhang, T.-C. & Xi, T. Novobiocin decreases SMYD3 expression and inhibits the migration of MDA-MB-231 human breast cancer cells. IUBMB Life 62, 194–199 (2010).
Nucleotide BLAST: Search nucleotide databases using a nucleotide query. Available at: https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE_TYPE=BlastSearch (Accessed: 21st February 2017).
Dar, S. A., Thakur, A., Qureshi, A. & Kumar, M. siRNAmod: A database of experimentally validated chemically modified siRNAs. Sci. Rep. 6, 20031 (2016).
Tyagi, A. et al. HIVsirDB: a database of HIV inhibiting siRNAs. PloS One 6, e25917 (2011).
Gong, W. et al. Integrated siRNA design based on surveying of features associated with high RNAi effectiveness. BMC Bioinformatics 7, 516 (2006).
Elbashir, S. M., Harborth, J., Weber, K. & Tuschl, T. Analysis of gene function in somatic mammalian cells using small interfering RNAs. Methods San Diego Calif 26, 199–213 (2002).
Kumar, R., Conklin, D. S. & Mittal, V. High-throughput selection of effective RNAi probes for gene silencing. Genome Res. 13, 2333–2340 (2003).
Thakur, N., Qureshi, A. & Kumar, M. VIRsiRNAdb: a curated database of experimentally validated viral siRNA/shRNA. Nucleic Acids Res. 40, D230–236 (2012).
Acknowledgements
This work was supported by funds from Indian Council of Medical Research (Project No. BIC/11(34)/2014 and CSIR-Central Institute of Medicinal and Aromatic Plants, Lucknow India. We are thankful to Dr. Mukti Nath Mishra, CSIR-CIMAP for his assistance for proof-reading of manuscripts.
Author information
Authors and Affiliations
Contributions
A.T. collected and compiled the data, and developed the website. A.T. and M.S. contributed in manuscript writing. A.S. conceived and coordinated the project, helped in the interpretation of data, refined the drafted manuscript and gave overall supervision to the project.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tyagi, A., Semwal, M. & Sharma, A. A database of breast oncogenic specific siRNAs. Sci Rep 7, 8706 (2017). https://doi.org/10.1038/s41598-017-08948-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-08948-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.