Abstract
The toxic metalloid arsenic has been environmentally ubiquitous since life first arose nearly four billion years ago and presents a challenge for the survival of all living organisms. Its bioavailability has varied dramatically over the history of life on Earth. As life spread, biogeochemical and climate changes cyclically increased and decreased bioavailable arsenic. To elucidate the history of arsenic adaptation across the tree of life, we reconstructed the phylogeny of the arsM gene that encodes the As(III) S-adenosylmethionine (SAM) methyltransferase. Our results suggest that life successfully moved into arsenic-rich environments in the late Archean Eon and Proterozoic Eon, respectively, by the spread of arsM genes. The arsM genes of bacterial origin have been transferred to other kingdoms of life on at least six occasions, and the resulting domesticated arsM genes promoted adaptation to environmental arsenic. These results allow us to peer into the history of arsenic adaptation of life on our planet and imply that dissemination of genes encoding diverse adaptive functions to toxic chemicals permit adaptation to changes in concentrations of environmental toxins over evolutionary history.
Similar content being viewed by others
Introduction
Arsenic is a pervasive environmental toxin, and exposure to arsenic was no doubt one of the challenges to the origin of life1. Geologically, there have been many changes in global redox states and climates over the past 4.5 billion years2, 3, which dramatically influenced arsenic concentrations and bioavailability. Specifically, there were dramatic increases in marine arsenic concentrations in the late Archean eon (3–2.5 billon years ago (Bya)) and during interglacial periods after Huronian (~2.4–2.1 Bya) and Cryogenian glaciations (~0.85–0.58 Bya), respectively4. How organisms adapted to arsenic stress in response to the enormous shifts in biogeochemistry of arsenic may have influenced the evolution of modern life1, 4, 5. However, there is sparse evidence with respect to how life coped with toxic arsenic throughout geological time lines.
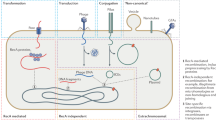
The composition of genomes and proteomes of extant organisms may bear imprints of ancient biogeochemical events6,7,8,9 that can help to reconstruct the evolutionary history of how life adapted to ancient changes in arsenic geochemistry. Multiple pathways for detoxification of both inorganic and organic arsenicals have been evolved in microorganisms. The genes for these defenses are usually encoded in arsenic resistance (ars) operons10. The widely distributed pathway for resistance is efflux of cellular arsenic from bacteria, archaea and fungi catalyzed by either the ArsB or Acr3 efflux permeases (Fig. 1a)11. A parallel pathway widely distributed across various kingdoms of life, for As(III) resistance, is methylation catalyzed by the As(III) S-adenosylmethyltransferase (ArsM, Fig. 1a), which converts As(III) into trivalent methylated species that are non-enzymatically oxidized to the non-toxic pentavalent species methylarsenate [MAs(V)], dimethylarsenate [DMAs(V)] and volatile trimethylarsine [TMAs(III)]12, 13. Metagenomic analysis indicates that the ArsM-catalyzed arsenic methylation strategy may be a major mechanism of resistance in extant marine cyanobacterial populations14. AS3MT, the animal version of ArsM, has been suggested to be the primary route of arsenic detoxification in mammals15. Although potentially significant from an evolutionary standpoint, the evolutionary history of ArsM, which encompasses imprints of how modern life adapts to arsenic stress and how the different kingdoms of life incorporated this machinery, has remained a mystery.
(a) Diagram representing enzymatic reactions catalyzed by arsenite efflux permease (ArsB), arsenite methyltransferase (ArsM), organoarsenical oxidase (ArsH) and methylarsenite efflux permease (ArsP). As(III): arsenite, MAs(III): methylarsenite, MAs(V): methylarsenate; (b) Cluster map of As(III) SAM methyltransferases (ArsM). Protein sequences of ArsM orthologs were clustered based on their pairwise similarity using the CLANS program (Frickey and Lupas, 2004). The clans containing sequences from more than one kingdom of life are indicated by black circles. Clans containing arsM from different phylums of bacteria are indicated by dash circles.
Horizontal gene transfer (HGT), the movement of genetic information among species, has been recognized as a significant mechanism for prokaryotic adaptation and diversification16. Recently, recognition of rapid expansion of newly sequenced eukaryotic genomes allows for the identification of additional examples of cross-kingdom HGT17,18,19, which contribute to the adaptive potential of the recipients. One report describes the discovery that an extremophilic eukaryote exploits horizontally-acquired genes for environmental adaptation from other kingdoms, e.g., bacteria and archaea20. The widespread occurrence of arsM genes among various kingdoms of life raised the question whether cross-kingdom HGT played a role throughout the evolutionary history of the ArsM As(III) SAM methyltransferase.
To understand the natural history of arsenic adaptation, we analyzed more than 3600 annotated genomes for putative arsM genes. The phylogenetic distribution, geological timing and evolutionary history for arsM were determined using phylogenomic methods. By examining these data, we infer the timing and complex genetic events that have marked the history of arsM, representing the genomic fossils that illustrate how life adapts to toxic arsenic throughout evolutionary history.
Results
Distinct phylogenetic groups
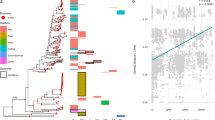
In this study the sequences of 231 bacterial, 19 archaeal and 93 eukaryotic ArsM orthologs were identified in the eggNOG database. Among all the retrieved ArsM sequences, cysteine residues integral to the binding of arsenite to ArsM protein at positions 72, 174 and 224 in CmArsM21 are conserved. In addition, SAM-binding motifs are also conserved among all ArsM sequences. The similarity in these key catalytic residues and domain structure indicate the homology of all ArsMs. The phylogenetic distribution of arsM genes is extensive, including bacteria and archaea, animals and fungi, illustrating the widespread occurrence of the potential for methylation of inorganic arsenic22. Cluster mapping of ArsMs indicates that they segregated into at least 6 distinct groups (Fig. 1b), each of which is comprised of sequences from quite different organisms. Group I consist of ArsMs from cyanobacteria and bacteroidetes phyla. Group II are found in members of cyanobacteria, firmicutes, verrucomicrobia and proteobacteria. Group III are predominantly composed of members of the metazoan, protist, red algae and proteobacteria. Group IV are from fungi and planctomycetes. Group V composed of ArsMs from anaerobes, including chlorobia and anaerobic proteobacteria. Group VI contains ArsMs from archaea and various phyla of bacteria. The observation that sequences from remotely related organisms are closely related, and that they cluster together in a way that is vastly different from the species phylogeny, suggested that evolution of ArsM might not be vertical.
To further explore the evolutionary history of ArsM, phylogenetic analysis was performed for all ArsM homologs. Consistent with the results of cluster mapping, an extraordinary incongruence between our ArsM tree (Fig. 2) and the published phylogeny23 of eukaryotic species was detected, e.g., ArsM homologs in eukaryotes showed a non-monophyletic distribution in three distinct groups with well-bootstrapped support (Group III, IV and VI, Fig. 2). By examining the entire Bayesian posterior ArsM tree samples, we found that none of the trees resolved ArsM from the eukaryotes as being monophyletic. In addition, AU test implemented in CONSEL package rejected (P < 0.01) the alternative tree with monophyletic eukaryotic sequences (Table S1). Here we investigate and discuss the possible causes of the conflict pattern between ArsM tree and species tree.
Phylogeny of ArsM As(III) SAM methyltransferases. The phylogenetic tree was constructed using the maximum likelihood program RAxML. The statistical significance of the branch pattern was estimated by conducting 100 bootstrap replications of the original amino acid alignment; bootstraps >50 were shown as blue circle. The detailed trees of each group are provided in Fig. 3, Figure 4–6 The related SAM-dependent mycolic acid cyclopropane synthetase (cmaS) genes were used as outgroup14.
Bacterial contaminations during genome sequencing of eukaryotes are not likely to be the explanation for the clustering of eukaryotic and prokaryotic ArsM (Figs 1 and 2), because ArsM homologs in eukaryotes, e.g., fungi24 and animals25, have been well characterized. All eukaryotic arsM genes except that in Hydra magnipapillata carry introns (Table S2). Although arsM genes in H. magnipapillata were found lack introns, they are closely linked to eukaryotic genes with introns on the same genomic scaffolds (Table S2), suggesting that these sequences do not represent bacterial artifacts.
Next, we investigated whether aberrant evolutionary substitution process may have misled the phylogenetic reconstruction. For example, heterogeneous evolutionary rate across lineages can obscure true phylogenetic signal26, thereby causing a gene tree to be incongruent with the species tree. To test whether eukaryotic lineages in three different groups (Group III, IV and VI) evolved at different evolutionary rates, we applied a likelihood ratio test (LRT) between a model in which all eukaryotic lineages evolve at the same rate (homogeneous model) and a model in which different lineage rates are allowed (heterogeneous model). An LRT showed that the homogeneous model cannot be rejected (Table S3), which indicated the difference in evolutionary rates among the eukaryotic lineages is small. Similarly, relation rate test did not show significant difference among evolutionary rates of these eukaryotic lineages (Table S3). Taken together, the evolutionary rate of ArsM among different eukaryotic lineages appear to be similar and are not likely to explain the incongruence between the gene tree and species tree.
Differential loss was suggested to be one of the major reasons for the patchy distributions of genes across eukaryotic lineages27. If this has been the case, all eukaryotic ArsMs should be expected to cluster in a monophyletic group with certain taxonomic lineages missing. However, the converse was observed: ArsMs distributed in non-monophyletic clusters with prokaryotes in ArsM tree (Fig. 2). We therefore hypothesized that the incongruent ArsM tree could be the result of multiple HGT events among prokaryote and/or eukaryote.
Early origin of As(III) methylation
ArsM sequences of cyanobacteria in Group I (Fig. 3a) form a strongly supported monophyletic clade (ML/Bayes = 97/0.99), and its phylogeny is generally congruent with 16 S rRNA phylogeny28. Therefore, the phylogeny of cyanobacterial ArsM in Group I corresponds to that of parent species, indicating linear evolution without HGT. The similarity of topology extends to the details that Gloeobacter is the earliest branch within ArsM phylogeny29, which is in line with the 16S rRNA tree (Fig. S1). This result indicates that the Group I-like ArsM was present before the divergence of Gloeobacter from other cyanobacteria. The evolutionary history of Group I version of ArsM appears to be dated back to an era before the last cyanobacterial common ancestor (LCCA). According to the fossil record and molecular dating, the LCCA is believed to have existed as early as 3 Bya30, considerably before the accumulation of oxygen in the atmosphere (2.45–2.32 Bya) during the Great Oxidation Event (GOE)31. Overall, the finding of an arsM gene in LCCA implies that ArsM arose in an anoxic world before the GOE. This result calls the primordial role of ArsM as a detoxification pathway into question: methylation of arsenite increases toxicity by producing highly toxic MAs(III) and DMAs(III), which are thermodynamically stable in the oxygen-free environment that existed before the GOE32. Thus, ArsM would not be a detoxification enzyme unless there were oxygen to convert the products to less toxic MAs(V) and DMAs(V). We postulate that the enzyme originally evolved to produce extremely toxic trivalent methylarsenicals that functioned as antibiotics to kill competitors in the primordial environment; after the GOE the trivalent species were nonenzymatically oxidized to pentavalent forms, transforming ArsM into a detoxifying enzyme33. Under this model, the original antibiotic producer (ArsM hosts) protected itself by pumping out its own toxic products (MAs(III)) with the methylarsenite efflux permease (ArsP)34, which simultaneously kill off its neighbors. This hypothesis is exemplified by the observation that arsP co-exists with arsM, primarily in anaerobes (Fig. S2b). In addition, clustering of arsP in anaerobic hosts in a monophyletic clade (Group V) in ArsM phylogeny (Fig. S2a) suggests that clustering of ArsP and ArsM might be an ancestral feature in lineages of anaerobes, supporting our hypothesis that ArsP evolved for detoxification of MAs(III) in ArsM hosts.
Maximum likelihood tree of ArsM orthologs in different groups. (a) Maximum likelihood tree of ArsM orthologs in Group I; (b) Maximum likelihood tree of ArsM orthologs in Group III; (c) Maximum likelihood tree of ArsM orthologs in Group IV; (d) Maximum likelihood tree of ArsM orthologs in Group VI. Maximum likelihood bootstrap support value (>50) are indicated below branches. Bayesian posterior probability values of critical bipartition appear above the branch for comparison.
Alternatively, ArsM could function as arsenite detoxification pathway in the anoxic environment of the primordial earth through co-evolution with ArsH, which enzymatically oxidizes of MAs(III) to MAs(V)35. However, biochemical studies and cluster analysis suggests that this explanation is unlikely. First, ArsH uses molecular oxygen in its catalytic mechanism35, so it most likely evolved after the appearance of atmospheric O2 rather than functioning in the primordial anoxic environment. Second, ArsH and ArsM show a distinct taxonomic distribution, which suggests an independent evolutionary trajectory rather than co-evolution (Fig. S3b). Third, cluster analysis shows that ArsH and ArsM coexist primarily in aerobes, not anaerobes (Fig. S3c). This implies that it is organisms with an aerobic lifestyle that possess both ArsM and ArsH and suggests that ArsH did not evolve to play a role in detoxification of MAs(III) in ArsM hosts in anoxic environments.
HGT of arsM from bacteria to metazoa
HGT of arsM genes may have contributed to the evolution of metazoans by allowing them to adapt to ubiquitous presence of arsenic. The metazoan group forms a strongly supported (ML/Bayes = 80/1.0) sister group to the bacteria/diatom group (Fig. 3b; Fig. S4). Formation of this cluster indicates that the metazoan group originated via the first ancient HGT from bacteria to the ancestor of the metazoan lineages. All metazoan ArsMs, including those from early-branching metazoan lineages (cnidaria), form a well-supported monophyletic clade (ML/Bayes = 100/1.0), so it is likely that the bacterial arsM gene was transferred to the common ancestors of all metazoans (urmetazoan) around 600–700 million years ago (Mya)36,37,38, and subsequent later gene loss might explain for differential presence of arsM gene among metazoan genomes. Urmetazoans possibly acquired arsM from bacteria that formed their food source through the operation of a gene transfer ratchet mechanism39, 40. In addition, a second but more recent HGT from bacteria to metazoan may also have occurred. As shown in Fig. 3d, Hydra magnipapillata are nested within bacterial species in ArsM phylogeny, which indicates a recent HGT from bacteria to this member of the Cnidaria phylum. The acquired arsM may have alleviated arsenic stress by biotransformation of arsenic into the less toxic species such as those commonly found in mammalian urine40. These observations suggest that HGT had a critical role in the adaptation of metazoans to environmental arsenic stress.
HGT of arsM from bacteria to diatoms
We propose that the acquisition of an arsM gene by eukaryotes from bacteria conferred diatoms with the ability to cope with stressful marine arsenic concentrations. ArsM from two diatoms, Thalassiosira pseudonana and Phaeodactylum tricornutum, form a sister group to a bacterial-only clade with high bootstrap support (ML/Bayes = 87/100, Fig. 3b), indicating that an arsM gene was transferred from bacteria to diatoms. Although HGT is considered rare, with limited evolutionary significance in eukaryotes compared to prokaryotes, our results are supported by the observation that HGT between bacteria and diatoms is pervasive and occurs more frequently than that in other eukaryotes41. The occurrence of bacterial arsM genes in diatoms serves as part of a generally metabolic strategy in addition to the role in arsenic detoxification. For example, ArsM not only permits diatoms to produce less toxic MAs(V) or DMAs(V) but also provides the substrates for biosynthesis of organoarsenical compounds such as arsenosugars and arsenolipids42, 43. Arsenolipids, for example, are reported to compensate for the deficiency of phosphorus in diatoms growing in phosphorus-depleted oceans44. Arsenolipids have a phosphate sparing effect in synthesis of membrane phospholipids45.
HGT of arsM from bacteria to fungi
Phylogenetic analysis of Group IV indicates a planctomycetes origin for fungal ArsM orthologs (Fig. 3c). ArsM orthologs from fungi form a well-supported clade (ML/Bayes = 94/1.0), and cluster as a sister group to the planctomycetes clade (Fig. 3c), suggesting that an ancestral fungal species acquired an arsM gene from a planctomycetes. The species-specific arsM gene duplications in Aspergillus fumigatus and Thielavia terrestris, respectively (Fig. 3c), indicate arsM genes evolve dynamically by gene duplications after ancestral acquisition. Although fungi represent the most recalcitrant of all organisms to gene transfer due to their robust cell walls and lost phagotrophic capacities, conjugation-like transfer and ecological association with other microorganisms could have promoted HGT between prokaryotes and fungi46. A phylogenomic analysis of 60 sequenced fungal genomes suggests that the gene encoding ArsC, a bacterial arsenite reductase which catalyzes a critical step in arsenate metabolism1, was transferred into lineages of the fungal kingdom47. In addition, the fungal species A. fumigatus has more arsenic resistance genes than any other eukaryote, and they all resemble bacterial arsenic resistance determinants. The original HGT must have been at least one entire bacterial ars operon. This includes ArsH, which gives MAs(III) resistance by oxidation to MAs(V)35, implying that A. fumigatus comes into contact with both inorganic and organic arsenicals. The acquisition of bacterial arsenate reductase genes as well as other arsenic resistance genes by fungi reflects the arsenic dilemma faced by fungi during their evolution trajectory. Thus, HGT of multiple ars genes from bacteria to fungi, including those involved in As(V) reduction, As(III) methylation and MAs(III) oxidation, provides synergistic effect in arsenic tolerance.
HGT of arsM from bacteria to archaea
HGT of arsM also occurred between bacteria and archaea. The archaeal ArsMs are paraphyletic clustered or nested within other bacterial sequences (Fig. 2), indicative of several independent bacteria-to-archaea HGT events. For example, ArsM sequences from 4 halobacteria are nested within firmicutes and proteobacteria (Fig. 3d), indicating that the ArsM of the halobacterium group originated via HGT from bacteria, although the identification of the donor lineage is not straightforward. As(III) methylation represents a useful ecological function for the halobacteria and other extremophilic archaea. The natural habitat of halobacteria is often rich in heavy metals and metalloids, including arsenic48. In addition, other archaeal species are extremophiles, living in extreme environments such as hot springs, where the arsenic concentration is often very high1. Therefore, in addition to utilizing arsenic for bioenergetics purpose49, 50, acquiring an arsenite methylation pathway might be critical for adaptation of archaea to extreme environments51, 52.
Eukaryote-to-eukaryote HGT of arsM
A eukaryote-to-eukaryote HGT of arsM gene may also have occurred between heterolobosea and green algae. The grouping (ML/Bayes = 95/0.87) of Naegleria gruberi (heterolobosea) and green alga in the ArsM phylogeny is surprising (Fig. 3b), as Naegleria and green alga are classified into different supergroups of eukaryotes23. A plausible explanation could be that the arsM gene from a green alga was transferred into N. gruberi through a secondary endosymbiotic event, as suggested by the ‘plastid-early’ hypothesis53. According to this evolutionary scenario, a secondary endosymbiosis of green alga occurred in the common ancestor of heterolobosea and euglenida, and the plastid-lack heterolobosea (N. gruberi) secondarily lost its plastid, after successful integration of an arsM gene into its nuclear genome. The endosymbiotically-transferred arsM gene logically would have allowed N. gruberi to adapt to high arsenic concentrations in its ecological niche. For example, the capacity for As(III) methylation provides Euglena gracilis, a protist related to N. gruberi, with a distinct arsenite metabolism that allows it to cope with arsenic stress in an acid drainage environment54. In addition, fusion of putative arsM genes and selenophosphate synthetase (SelD), as observed in the genome of N. gruberi, is advantageous to a free-living protist faced with potentially threatening environments55, 56.
HGT of arsM between cyanobacteria and other bacteria phyla
Cyanobacteria do not form a monophyletic clade in the ArsM tree. One cyanobacterial group (Group II, Fig. S5), including Synechococcus, Prochlorococcus, Cyanothece, Lyngbya, Trichodesmium erythraeum and Arthrospira, clusters within firmicutes, proteobacteria and verrucomicrobia with high bootstrap support (ML/Bayes = 100/1.0). The remaining cyanobacteria (Group I, Fig. 3a) form a well-supported monophyletic group (ML/Bayes = 97/0.99). The paraphyletic grouping of cyanobacterial ArsM strikingly conflicts with 16 S rRNA-based phylogenies, which tends to place cyanobacteria as one coherent (monophyletic) phyla in bacteria57, indicating HGT occurred between cyanobacteria and other bacterial phyla. The current cyanobacterial ArsM topology is most easily explained by displacement of the original arsM gene in Group II cyanobacteria with an inter-phylum transferred ortholog from other Group II bacteria (xenologous gene displacement). Inter-phylum transfers between cyanobacteria and other phyla should not be surprising, as it has been estimated that individual cyanobacterial genomes acquired between 9.5% and 16.6% of their genes through HGT58. HGT between Group II organisms is further corroborated by the ecology of these species: Group II organisms are frequently found in sea bacterioplankton communities, providing a viable environment for gene sharing59. The alternative hypothesis - that both Group I and Group II-like arsM genes were present in the last common cyanobacterial ancestor and subsequently differentially lost - is less likely for several reasons60. First, all extant cyanobacteria have either Group I or Group II versions of ArsM - none has been found to encode both - which argues against retention of both versions in a single genome over a long evolutionary timescale, as required by an “ancient paralogy and differential loss” scenario59. Second, too many parallel independent losses would have to be postulated if the both Group I and Group II-like ArsMs were ancestral to cyanobacteria, which is not parsimonious18.
ArsMs from marine cyanobacteria (Group II, Fig. S5) and those from freshwater cyanobacteria (Group I, Fig. 3a) form phylogenetically-distinct clusters, suggesting that a HGT that occurred in marine cyanobacteria contributed to niche differentiation of these two cyanobacteria “ecotypes”. Marine cyanobacteria are more likely to suffer arsenic stress than those in freshwater due to higher arsenic concentrations in open seawater61. In addition, because arsenate is taken up by phosphate transport systems, low phosphorus concentrations in the ocean may contribute to higher levels of arsenate uptake, thereby increasing the stress of arsenic to marine organisms62. Consistently, a number of marine cyanobacteria are more efficient in arsenic methylation than freshwater cyanobacteria63. Natural selection has favored human AS3MT haplotypes that associate with more efficient arsenic metabolism in populations with generations of severe arsenic exposure as a result of increased expression of the gene64, and we suggest that HGT followed by selection for higher arsM expression in marine cyanobacteria that resulted in an increase in arsenic methylation rates may contribute to the ability of cyanobacteria to thrive in arsenic-rich marine environments.
HGT of arsM among anaerobes
Inter-phylum HGT of ArsM between distantly related anaerobic bacteria may have played a role in their adaptation to an ecological niche. As shown in Fig. S6, ArsMs from two strictly anaerobic chlorobi, Prosthecochloris aestuarii and Chlorobium phaeobacterioides, are nested within a branch of strictly anaerobic genera of δ-proteobacteria, such as sulfur-reducing bacteria and ferric-iron reducing bacteria, strongly suggesting a transfer event from δ-proteobacteria to chlorobi. Although genetic exchange between distantly related organisms representing different bacterial phyla is thought to be very infrequent, extensive HGT within anaerobic mesophilic environments was previously described between Sphaerochaeta spp and Clostridia 65. Indeed, there is a higher frequency of HGT within anaerobic rather than aerobic bacteria, and a greater niche overlap and/or physical proximity among organisms within anaerobic environments favors HGT66. Inter-phylum HGTs appear to be fundamental for the adaptation of the anaerobes to their perspective ecological niches, although the ecological role of ArsM in anoxic environments is still unknown. These results suggest that the capabilities for As(III) methylation under anaerobic conditions were transferred between distantly related phyla.
Discussion
The concentrations and bioavailability of arsenic varied dramatically during major periods of phylogenetic diversification. As a result, there was early evolution of an arsenic methylation gene that then continually transferred horizontally among kingdoms of life. These HGT events mark the natural history of arsenic adaptation. Our phylogenetic analysis suggests an early cyanobacterial ArsM origin toward the end of the Archean Eon, more than 2.5 Bya (Fig. 3a). In the geological records, the concentration of arsenic in the late Archean Eon (3 Bya-2.5 Bya) increased in oceans67, probably due to significant oxidative weathering of arsenic-rich minerals triggered by low levels of O2 available before the GOE68,69,70,71. Hence, the innovation of arsenic methylation systems in cyanobacteria at the end of Archean could have produced fitness advantages, leading to an increase in cyanobacterial diversity and abundance57, which in turn would supercharge cyanobacteria, the engine of life. Subsequently inter-kingdom HGT of arsM represented a major mechanism of evolutionary adaptation to arsenic stress. Our results suggested that the arsenic methylation pathway found in the common ancestor of metazoa (~600 Mya) was acquired via HGT (Fig. 3b; Fig. S4), quite likely as a result of an ‘arsenic burst’ after the Cryogenian glaciations (850–580 Mya)4. In this scenario, a burst of deglaciating metal/As-rich waters and intensified chemical weathering4 would have led to massive increases in the concentration and bioavailability of arsenic in the ocean, selecting for stable transfer of arsM from prokaryote to metazoa. We proposed that increased fitness of ArsM-containing species led to a higher frequency and wider distribution of metazoa during post-Cryogenian.
Given that ArsM may have played a critical role in promoting the adaptation and diversification of life under arsenic changing environment, our study has implications for the evolutionary significance of inter-kingdom HGT. Conventional belief is that HGT between distantly related organisms from different kingdoms is infrequent, especially when multiple cellular eukaryotes are involved. However, our analysis reveals that at least 6 HGT events (in the case of arsM) occurred among different kingdoms of life (Fig. 4). An extremophilic eukaryote was recently suggested to exploit horizontally-acquired genes in adaptation to toxic metal-rich environments from other kingdoms, e.g., bacteria and archaea20. Therefore, inter-kingdom HGT has been overlooked as a potentially significant factor in facilitating adaptation of organisms to changing environments and/or the development of resistance to not only arsenic but also other environmental toxic substances72,73,74.
Diagram illustrating the inter-kingdom HGT of arsM gene. Horizontal lines and arrows show HGT donors and recipients. LCCA, last cyanobacterial common ancestor. Information about tree of life is based on Yue et al.17.
Two lines of evidence further suggest that the horizontally transferred arsM genes provide an adaptive function to their hosts. Firstly, signatures of purifying selection were observed acting on the arsM genes from HGT hosts, such as metazoa, fungi, diatoms and archaea, as indicated by the overall gene dN/dS ratio (less than 1) of these gene groups (Table S1). Consistently, individual codons with significant (P < 0.05) signatures of purifying selection were identified in different arsM groups (Table S1), and no sites were found to be under positive selection. These results imply that arsM genes were under rigorous functional conservation in response to strong selective pressure after HGT, which highlights the significant role of arsM in allowing their host to adapt to novel ecological niches. Secondly, the majority of ArsM proteins from HGT hosts possess four cysteine residues integral to arsenite binding and flanking motifs of their corresponding bacterial ArsM groups (Fig. S7), indicating the retention of enzymatic function after HGT. The domestication of ArsM function within recipient organisms is further demonstrated by a genome-wide association study among human populations living in a region with elevated arsenic exposure shows that adaptive polymorphisms in the AS3MT gene, the metazoan version of arsM, play a role in human acclimatization to arsenic rich environments64.
An emerging pattern is that most inter-kingdom HGT of the arsM gene involve acquisition from bacteria. For example, eukaryotic ArsM, including those in metazoa, fungi, diatoms, can be traced back to multiple acquisitions from independent bacterial groups (Fig. 4). Although we also propose eukaryote-to-eukaryote HGT (Fig. 3b and 5), current evidence indicates that bacteria appear to be the common candidate donors of new physiological functions for eukaryotes20. Indeed, bacteria-to-eukaryote HGT underlies the adaptation of many microbial eukaryotes20, 59. Bacteria are abundant in nearly every ecological niche, and many eukaryotes survive by eating and digesting them in large numbers75, so bacterial genes, including arsM, are a source of foreign genetic material for eukaryotes. Secondly, bacteria are metabolically more diverse than eukaryotes, and, as such, have more to offer in the way of genetic information for new and beneficial functions such as arsenic methylation. In addition, many bacteria have mechanisms such as the type IV secretion system to transfer DNA to other organisms76. These ecological links between donor and recipient lineages further imply that bacteria provide a rich source of genetic material that allows rapid adaptations to new ecological niches, in this case, arsenic stress.
The finding of recurrent HGT events during the evolution of ArsM raises the question of why ArsM was so frequently transferred horizontally among different kingdoms of life. Genes are logical candidates for domestication after HGT if 1) they independently provide functionality and 2) are immediately beneficial to a recipient organism18. arsM genes, which were transferred from various bacteria to other kingdoms of life, meet these qualifications. Firstly, ArsMs contain both SAM binding and catalytic domains, and require no other proteins for their activity21; secondly, the ability to methylate arsenic into innocuous organic forms is of immediate advantages for organisms suffering from environmental arsenic stress. As such, ArsM could easily be domesticated as a single functional unit in other kingdoms of life such as metazoans, diatoms, archaea and fungi (Fig. 4). In the case of recurrent transfer and retention of a gene encoding a specific protein in diverse species, a model that allows for immediate selection seems more likely18, although many genes transferred between prokaryotes are pre-adaptive traits with neutral value77.
In summary, we demonstrate here that life largely avoided arsenic stress in the late Archean Eon and Proterozoic Eon, respectively, by the propagation of arsM genes and bacterial genes encoding As(III) S-adenosylmethyltransferase were horizontally transferred in at least six distinct events to diverse kingdoms of life. The recurrent and independent transfer of arsM genes to distinct kingdoms of life suggests that these genes confer immediate fitness benefits by supplying a new dominant function to the host organisms. Diverse genetic and biochemical mechanisms have been demonstrated to allow organisms to physiologically adapt to the stress of toxic environmental substances78. We propose that the spread of genes that underpinning the ability to tolerate toxic substances - beyond the arsM genes – has occurred multiple times among different kingdoms in response to selective pressures imposed by changing environmental conditions through geological history.
Methods
Identification of arsM genes and cluster mapping
To obtain sequences of arsM genes encoding candidate orthologs, the Clusters of Orthologous Groups (COG) entry associate with ArsM function, NOG07857, was used to extract additional ArsM sequences in the eggNOG79 database. Conserved domains of the extracted protein sequences were identified with the Batch Web CD-Search Tool in the NCBI Conserved Domain Database (CDD) (http://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi). To identify the As(III) binding site of ArsMs, each extracted protein sequence was aligned to the reference alignment80 of known ArsM sequences with hmmalign program from the HMMER3 package (http://hmmer.janelia.org). Only those sequences containing SAM-binding domains, as well as at least 3 of the four conserved cysteine residues corresponding to residues 44, 72, 174 and 224 in the ArsM sequence of the eukaryotic red algae Cyanidioschyzon merolae sp. 5508 (CmArsM)21, were used for further analysis. Sequences of the product of the cmaS gene, which encodes the closely related SAM-dependent mycolic acid cyclopropane synthetase were also included as an outgroup, as previously described14. All arsM sequences were compared with each other using BLASTP and clustered using CLANS software81. The final visualization was obtained with an e-value threshold of 1e-40 applied in the CLANS program.
Identification of ArsP and ArsH
To identify ArsP sequences in eggNOG database, ArsP orthologs in bacteria (ENOG4105DSX) and Archaea (arCOG02712) were retrieved (no orthologs was found in eukaryotes). Only sequences with the conserved “PXCSCXXXP” motif were considered to be ArsP orthologs. To obtain putative ArsH orthologs, an initial phylogenetic tree of COGs associate with ArsH function from eukaryotes (ENOG410IITQ), bacteria (ENOG4105CAU) and archaea (arCOG04624) in the eggNOG database was built (see phylogenetic reconstruction). The sequences in the clade which includes known ArsH function was retrieved as putative ArsH orthologs (Fig. S3a). Information about oxygen requirement associated with these organisms was acquired from either NCBI or Genomes OnLine Database (GOLD).
Phylogenetic reconstruction
The selected protein sequences were aligned using several algorithms, including MUSCLE, CLUSTALX, MAFFT-L-INS-i and T-coffee. The MUSCLE aligned sequences, which yielded the highest score according to the word-oriented objective function (WOOF), were chosen for subsequent analysis82. Low quality alignment regions were removed by TrimAl to retain only those columns present in at least 30% of the sequences83. Alignments were visually inspected and manually edited when necessary. ProtTest was used to find the most suitable model for phylogeny reconstruction before phylogenetic analysis84. The maximum-likelihood (ML) phylogenetic tree was constructed using RAxML version 8.2.085 with the WAG Model (+I + G), which got the highest score in the ProtTest analysis. Support values were calculated using 100 bootstrap replicates. A Bayesian tree was constructed using MrBayes version 3.2.6 with the WAG model86. For the MrBayes consensus trees, 5,000,000 generations were completed with trees every 500 generations. To investigate the extent of selection affecting the arsM genes, SLAC (Single Likelihood Ancestor Counting) in the HyPhy software suite87 were used to perform the calculation of non-synonymous and synonymous (dN/dS) ratio, as well as statistical tests for purifying or positive selection for individual codons. Additional statistical tests in the HyPhy software suite REL (Random Effect Likelihood) and FUBAR (Fast Unbiased Bayesien Approximation) were used to confirm whether arsM codons display statistically significant signature of purifying selection.
Phylogenetic congruence Test
To determine whether the eukaryotic non-monophyly in ArsM phylogeny was statistically significant, approximately unbiased (AU) test implemented in CONSEL package88 was conducted as reported in a previous study27. Briefly, 48 alternative trees where all eukaryotic species were forced into monophyly were generated by prune and re-grafting algorithm, keeping those closest to the original maximum likelihood tree in terms of Robinson and Folds distance (as computed by treedist program in PHYLIP89 package). PHYML90 was used to optimize parameters and calculate per-site likelihoods (using option -print_site_lnl) by providing the alternative trees as user tree. Programs makemt and catpv implemented in CONSEL package were run to calculate the P value for each alternative topology using the AU test.
Molecular evolutionary analysis
To examine whether differences in the evolutionary rates among the lineages are significant, we performed a likelihood ratio test (LRT) between homogeneous rate model and heterogeneous rate model using PAML91 package. Homogeneous rate assumes a single evolutionary rate (homogeneous rate) for all compared lineages. Heterogeneous rate model assigned each compared lineage with its independent evolutionary rate (heterogeneous rate). The likelihood of two models was calculated with the codeml program in PAML and compared using the LRT. Relative rate test implemented in RRTree92 was conducted to confirm whether all compared lineages have a homogeneous rate of evolution.
Note added in proof
While this manuscript was under review, a new report was published which assessed the evolutionary origin of AS3MT93, the animal version of arsenite methyltransferase. In that study, horizontal gene transfer of AS3MT between animal and bacteria as well as between fungi and bacteria was reported. Their conclusions support our concept that recurrent horizontal transfer of arsM genes facilitated adaptation of life to arsenic.
References
Zhu, Y. G., Yoshinaga, M., Zhao, F. J. & Rosen, B. P. Earth abides arsenic biotransformations. Annu Rev Earth Pl Sc 42, 443–+, doi:10.1146/annurev-earth-060313-054942 (2014).
Williams, R. J. P. & Da Silva, J. J. R. F. Evolution was chemically constrained. J Theor Biol 220, 323–343, doi:10.1006/jtbi.2003.3152 (2003).
Anbar, A. D. Oceans elements and evolution. Science 322, 1481–1483, doi:10.1126/science.1163100 (2008).
Fru, E. C. et al. Arsenic stress after the Proterozoic glaciations. Sci Rep 5, doi:ARTN 1778910.1038/srep17789 (2015).
Mukhopadhyay, R., Rosen, B. P., Pung, L. T. & Silver, S. Microbial arsenic: from geocycles to genes and enzymes. FEMS Microbiol Rev 26, 311–325, doi:PII S0168-6445(02)00112-2DOI 10.1111/j.1574-6976.2002.tb00617.x (2002).
Harel, A., Bromberg, Y., Falkowski, P. G. & Bhattacharya, D. Evolutionary history of redox metal-binding domains across the tree of life. Proc Natl Acad Sci USA 111, 7042–7047, doi:10.1073/pnas.1403676111 (2014).
David, L. A. & Alm, E. J. Rapid evolutionary innovation during an Archaean genetic expansion. Nature 469, 93–96, doi:10.1038/nature09649 (2011).
Dupont, C. L., Yang, S., Palenik, B. & Bourne, P. E. Modern proteomes contain putative imprints of ancient shifts in trace metal geochemistry. Proc Natl Acad Sci USA 103, 17822–17827, doi:10.1073/pnas.0605798103 (2006).
Dupont, C. L., Butcher, A., Valas, R. E., Bourne, P. E. & Caetano-Anolles, G. History of biological metal utilization inferred through phylogenomic analysis of protein structures. Proc Natl Acad Sci USA 107, 10567–10572, doi:10.1073/pnas.0912491107 (2010).
Rosen, B. P. Families of arsenic transporters. Trends Microbiol 7, 207–212, doi:10.1016/S0966-842x(99)01494-8 (1999).
Yang, H. C., Fu, H. L., Lin, Y. F. & Rosen, B. P. Pathways of arsenic uptake and efflux. Curr Top Membr 69, 325–358, doi:10.1016/B978-0-12-394390-3.00012-4 (2012).
Dheeman, D. S., Packianathan, C., Pillai, J. K. & Rosen, B. P. Pathway of human AS3MT arsenic methylation. Chem Res Toxicol 27, 1979–1989, doi:10.1021/tx500313k (2014).
Marapakala, K. et al. A disulfide-bond cascade mechanism for arsenic(III) S-adenosylmethionine methyltransferase. Acta Crystallogr D 71, 505–515, doi:10.1107/S1399004714027552 (2015).
Saunders, J. K. & Rocap, G. Genomic potential for arsenic efflux and methylation varies among global Prochlorococcus populations. ISME J 10, 197–209, doi:10.1038/ismej.2015.85 (2016).
Thomas, D. J. & Rosen, B. P. In Encyclopedia of Metalloproteins (eds Robert H. Kretsinger, Vladimir N. Uversky, & Eugene A. Permyakov) 138-143 (Springer New York, 2013).
Treangen, T. J. & Rocha, E. P. Horizontal transfer, not duplication, drives the expansion of protein families in prokaryotes. PLoS Genet 7, e1001284, doi:10.1371/journal.pgen.1001284 (2011).
Yue, J. P., Hu, X. Y., Sun, H., Yang, Y. P. & Huang, J. L. Widespread impact of horizontal gene transfer on plant colonization of land. Nat Commun 3, doi:ARTN 115210.1038/ncomms2148 (2012).
Moran, Y., Fredman, D., Szczesny, P., Grynberg, M. & Technau, U. Recurrent horizontal transfer of bacterial toxin genes to eukaryotes. Mol Biol Evol 29, 2223–2230, doi:10.1093/molbev/mss089 (2012).
Keeling, P. J. Functional and ecological impacts of horizontal gene transfer in eukaryotes. Curr Opin Genet Dev 19, 613–619, doi:10.1016/j.gde.2009.10.001 (2009).
Schonknecht, G. et al. Gene transfer from bacteria and archaea facilitated evolution of an extremophilic eukaryote. Science 339, 1207–1210, doi:10.1126/science.1231707 (2013).
Ajees, A. A., Marapakala, K., Packianathan, C., Sankaran, B. & Rosen, B. P. Structure of an As(III) S-Adenosylmethionine methyltransferase: insights into the mechanism of arsenic biotransformation. Biochemistry 51, 5476–5485, doi:10.1021/bi3004632 (2012).
Bentley, R. & Chasteen, T. G. Microbial methylation of metalloids: Arsenic, antimony, and bismuth. Microbiol Mol Biol R 66, 250-+, doi:10.1128/Mmbr.66.2.250-271.2002 (2002).
Baldauf, S. L. An overview of the phylogeny and diversity of eukaryotes. J Syst Evol 46, 263–273 (2008).
Verma, S. et al. A novel arsenic methyltransferase gene of Westerdykella aurantiaca isolated from arsenic contaminated soil: phylogenetic, physiological, and biochemical studies and its role in arsenic bioremediation. Metallomics 8, 344–353, doi:10.1039/c5mt00277j (2016).
Lin, S. et al. A novel S-adenosyl-L-methionine: arsenic(III) methyltransferase from rat liver cytosol. J Biol Chem 277, 10795–10803, doi:10.1074/jbc.M110246200 (2002).
Sanderson, M. J. & Shaffer, H. B. Troubleshooting molecular phylogenetic analyses. Annu Rev Ecol Syst 33, 49–72, doi:10.1146/annurev.ecolsys.33.010802.150509 (2002).
Ku, C. et al. Endosymbiotic origin and differential loss of eukaryotic genes. Nature 524, 427–432, doi:10.1038/nature14963 http://www.nature.com/nature/journal/v524/n7566/abs/nature14963.html#supplementary-information (2015).
Latysheva, N., Junker, V. L., Palmer, W. J., Codd, G. A. & Barker, D. The evolution of nitrogen fixation in cyanobacteria. Bioinformatics 28, 603–606, doi:10.1093/bioinformatics/bts008 (2012).
Schirrmeister, B. E., Gugger, M. & Donoghue, P. C. J. Cyanobacteria and the Great Oxidation Event: evidence from genes and fossils. Palaeontology 58, 769–785, doi:10.1111/pala.12178 (2015).
Schirrmeister, B. E., de Vos, J. M., Antonelli, A. & Bagheri, H. C. Evolution of multicellularity coincided with increased diversification of cyanobacteria and the Great Oxidation Event. Proc Natl Acad Sci USA 110, 1791–1796, doi:10.1073/pnas.1209927110 (2013).
Rye, R. & Holland, H. D. Paleosols and the evolution of atmospheric oxygen: A critical review. Am J Sci 298, 621–672 (1998).
Canfield, D. E., Rosing, M. T. & Bjerrum, C. Early anaerobic metabolisms. Philos T R Soc B 361, 1819–1834, doi:10.1098/rstb.2006.1906 (2006).
Li, J. J., Pawitwar, S. S. & Rosen, B. P. The organoarsenical biocycle and the primordial antibiotic methylarsenite. Metallomics 8, 1047–1055, doi:10.1039/c6mt00168h (2016).
Chen, J., Madegowda, M., Bhattacharjee, H. & Rosen, B. P. ArsP: a methylarsenite efflux permease. Mol Microbiol 98, 625–635, doi:10.1111/mmi.13145 (2015).
Chen, J., Bhattacharjee, H. & Rosen, B. P. ArsH is an organoarsenical oxidase that confers resistance to trivalent forms of the herbicide monosodium methylarsenate and the poultry growth promoter roxarsone. Mol Microbiol 96, 1042–1052, doi:10.1111/mmi.12988 (2015).
Ayala, F. J., Rzhetsky, A. & Ayala, F. J. Origin of the metazoan phyla: Molecular clocks confirm paleontological estimates. Proc Natl Acad Sci USA 95, 606–611, doi:10.1073/pnas.95.2.606 (1998).
Battistuzzi, F. U., Feijao, A. & Hedges, S. B. A genomic timescale of prokaryote evolution: insights into the origin of methanogenesis, phototrophy, and the colonization of land. BMC Evol Biol 4, doi:Artn 4410.1186/1471-2148-4-44 (2004).
Ochman, H., Elwyn, S. & Moran, N. A. Calibrating bacterial evolution. Proc Natl Acad Sci USA 96, 12638–12643, doi:10.1073/pnas.96.22.12638 (1999).
Doolittle, W. E. You are what you eat: a gene transfer ratchet could account for bacterial genes in eukaryotic nuclear genomes. Trends Genet 14, 307–311 (1998).
Richter, D. J. & King, N. The genomic and cellular foundations of animal origins. Annu Rev Genet 47, 509–537, doi:10.1146/annurev-genet-111212-133456 (2013).
Bowler, C. et al. The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 456, 239–244, doi:10.1038/nature07410 (2008).
Morelli, E., Mascherpa, M. & Scarano, G. Biosynthesis of phytochelatins and arsenic accumulation in the marine microalga Phaeodactylum tricornutum in response to arsenate exposure. Biometals 18, 587–593, doi:10.1007/s10534-005-2998-1 (2005).
Duncan, E. G., Maher, W. A., Foster, S. D. & Krikowa, F. Influence of culture regime on arsenic cycling by the marine phytoplankton Dunaliella tertiolecta and Thalassiosira pseudonana. Environ Chem 10, 91–101, doi:10.1071/En12191 (2013).
Taleshi, M. S., Seidler-Egdal, R. K., Jensen, K. B., Schwerdtle, T. & Francesconi, K. A. Synthesis and characterization of arsenolipids: naturally occurring arsenic compounds in fish and algae. Organometallics 33, 1397–1403, doi:10.1021/om4011092 (2014).
Dembitsky, V. M. & Levitsky, D. O. Arsenolipids. Prog Lipid Res 43, 403–448, doi:10.1016/j.plipres.2004.07.001 (2004).
Fitzpatrick, D. A. Horizontal gene transfer in fungi. FEMS Microbiol Lett 329, 1–8, doi:10.1111/j.1574-6968.2011.02465.x (2012).
Marcet-Houben, M. & Gabaldon, T. Acquisition of prokaryotic genes by fungal genomes. Trends Genet 26, 5–8, doi:10.1016/j.tig.2009.11.007 (2010).
Kennedy, S. P., Ng, W. V., Salzberg, S. L., Hood, L. & DasSarma, S. Understanding the adaptation of Halobacterium species NRC-1 to its extreme environment through computational analysis of its genome sequence. Genome Res 11, 1641–1650, doi:10.1101/gr.190201 (2001).
Oremland, R. S. & Stolz, J. F. The ecology of arsenic. Science 300, 939–944, doi:10.1126/science.1081903 (2003).
Oremland, R. S. & Stolz, J. F. Arsenic, microbes and contaminated aquifers. Trends Microbiol 13, 45–49, doi:10.1016/j.tim.2004.12.002 (2005).
Nelson-Sathi, S. et al. Acquisition of 1,000 eubacterial genes physiologically transformed a methanogen at the origin of Haloarchaea. Proc Natl Acad Sci USA 109, 20537–20542, doi:10.1073/pnas.1209119109 (2012).
Nelson-Sathi, S. et al. Origins of major archaeal clades correspond to gene acquisitions from bacteria. Nature 517, 77–U185, doi:10.1038/nature13805 (2015).
Leander, B. S. Did trypanosomatid parasites have photosynthetic ancestors? Trends Microbiol 12, 251–258, doi:10.1016/j.tim.2004.04.001 (2004).
Halter, D. et al. Arsenic hypertolerance in the protist Euglena mutabilis is mediated by specific transporters and functional integrity maintenance mechanisms. Environ Microbiol 17, 1941–1949, doi:10.1111/1462-2920.12474 (2015).
da Silva, M. T. A., Caldas, V. E. A., Costa, F. C., Silvestre, D. A. M. M. & Thiemann, O. H. Selenocysteine biosynthesis and insertion machinery in Naegleria gruberi. Mol Biochem Parasit 188, 87–90, doi:10.1016/j.molbiopara.2013.04.002 (2013).
Mariotti, M. et al. Evolution of selenophosphate synthetases: emergence and relocation of function through independent duplications and recurrent subfunctionalization. Genome Res 25, 1256–1267, doi:10.1101/gr.190538.115 (2015).
Tomitani, A., Knoll, A. H., Cavanaugh, C. M. & Ohno, T. The evolutionary diversification of cyanobacteria: Molecular-phylogenetic and paleontological perspectives. Proc Natl Acad Sci USA 103, 5442–5447, doi:10.1073/pnas.0600999103 (2006).
Zhaxybayeva, O., Gogarten, J. P., Charlebois, R. L., Doolittle, W. F. & Papke, R. T. Phylogenetic analyses of cyanobacterial genomes: Quantification of horizontal gene transfer events. Genome Res 16, 1099–1108, doi:10.1101/gr.5322306 (2006).
Andersson, A. F., Riemann, L. & Bertilsson, S. Pyrosequencing reveals contrasting seasonal dynamics of taxa within Baltic Sea bacterioplankton communities. ISME J 4, 171–181, doi:10.1038/ismej.2009.108 (2010).
Townsend, J. P., Bohn, T. & Nielsen, K. M. Assessing the probability of detection of horizontal gene transfer events in bacterial populations. Front Microbiol 3, doi:ARTN 2710.3389/fmicb.2012.00027 (2012).
Santosa, S. J., Wada, S. & Tanaka, S. Distribution and cycle of arsenic compounds in the ocean. Appl Organomet Chem 8, 273–283, doi:10.1002/aoc.590080319 (1994).
Wurl, O., Zimmer, L. & Cutter, G. A. Arsenic and phosphorus biogeochemistry in the ocean: Arsenic species as proxies for P-limitation. Limnol Oceanogr 58, 729–740, doi:10.4319/lo.2013.58.2.0729 (2013).
Yin, X. X. et al. Biotransformation and volatilization of arsenic by three photosynthetic cyanobacteria. Plant Physiol 156, 1631–1638, doi:10.1104/pp.111.178947 (2011).
Schlebusch, C. M. et al. Human adaptation to arsenic-rich environments. Mol Biol Evol 32, 1544–1555, doi:10.1093/molbev/msv046 (2015).
Caro-Quintero, A., Ritalahti, K. M., Cusick, K. D., Loffler, F. E. & Konstantinidis, K. T. The chimeric genome of sphaerochaeta: nonspiral Spirochetes that break with the prevalent dogma in Spirochete biology. mBio 3, doi:ARTN e00025-1210.1128/mBio.00025-12 (2012).
Caro-Quintero, A. & Konstantinidis, K. T. Inter-phylum HGT has shaped the metabolism of many mesophilic and anaerobic bacteria. ISME J 9, 958–967, doi:10.1038/ismej.2014.193 (2015).
Bekker, A. et al. Dating the rise of atmospheric oxygen. Nature 427, 117–120, doi:10.1038/nature02260 (2004).
Arnold, G. L., Anbar, A. D., Barling, J. & Lyons, T. W. Molybdenum isotope evidence for widespread anoxia in mid-proterozoic oceans. Science 304, 87–90, doi:10.1126/science.1091785 (2004).
Anbar, A. D. & Knoll, A. H. Proterozoic ocean chemistry and evolution: A bioinorganic bridge? Science 297, 1137–1142, doi:10.1126/science.1069651 (2002).
Lyons, T. W., Reinhard, C. T. & Planavsky, N. J. The rise of oxygen in Earth’s early ocean and atmosphere. Nature 506, 307–315, doi:10.1038/nature13068 (2014).
Reinhard, C. T. et al. Proterozoic ocean redox and biogeochemical stasis. Proc Natl Acad Sci USA 110, 5357–5362, doi:10.1073/pnas.1208622110 (2013).
Moriguchi, K., Yamamoto, S., Ohmine, Y. & Suzuki, K. A fast and practical yeast transformation method mediated by Escherichia coli based on a trans-kingdom conjugal transfer System: just mix two cultures and wait one hour. PloS One 11, doi:ARTN e014898910.1371/journal.pone.0148989 (2016).
Conaco, C. et al. Detection of prokaryotic genes in the amphimedon queenslandica genome. PloS One 11, doi:ARTN e015109210.1371/journal.pone.0151092 (2016).
Wisecaver, J. H. & Rokas, A. Fungal metabolic gene clusters-caravans traveling across genomes and environments. Front Microbiol 6, doi:ARTN 16110.3389/fmicb.2015.00161 (2015).
Huang, J. L. Horizontal gene transfer in eukaryotes: The weak-link model. Bioessays 35, 868–875, doi:10.1002/bies.201300007 (2013).
Wallden, K., Rivera-Calzada, A. & Waksman, G. Type IV secretion systems: versatility and diversity in function. Cell Microbiol 12, 1203–1212, doi:10.1111/j.1462-5822.2010.01499.x (2010).
Gogarten, J. P. & Townsend, J. P. Horizontal gene transfer, genome innovation and evolution. Nat Rev Microbiol 3, 679–687, doi:10.1038/nrmicro1204 (2005).
Lemire, J. A., Harrison, J. J. & Turner, R. J. Antimicrobial activity of metals: mechanisms, molecular targets and applications. Nat Rev Microbiol 11, 371–384, doi:10.1038/nrmicro3028 (2013).
Powell, S. et al. eggNOG v4.0: nested orthology inference across 3686 organisms. Nucleic Acids Res 42, D231–D239, doi:10.1093/nar/gkt1253 (2014).
Ye, J., Rensing, C., Rosen, B. P. & Zhu, Y. G. Arsenic biomethylation by photosynthetic organisms. Trends Plant Sci 17, 155–162, doi:10.1016/j.tplants.2011.12.003 (2012).
Frickey, T. & Lupas, A. CLANS: a Java application for visualizing protein families based on pairwise similarity. Bioinformatics 20, 3702–3704, doi:10.1093/bioinformatics/bth444 (2004).
Beiko, R. G., Chan, C. X. & Ragan, M. A. A word-oriented approach to alignment validation. Bioinformatics 21, 2230–2239, doi:10.1093/bioinformatics/bti335 (2005).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973, doi:10.1093/bioinformatics/btp348 (2009).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165, doi:10.1093/bioinformatics/btr088 (2011).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313, doi:10.1093/bioinformatics/btu033 (2014).
Ronquist, F. et al. MrBayes 3.2: efficient bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61, 539–542, doi:10.1093/sysbio/sys029 (2012).
Pond, S. L. K., Frost, S. D. W. & Muse, S. V. HyPhy: hypothesis testing using phylogenies. Bioinformatics 21, 676–679, doi:10.1093/bioinformatics/bti079 (2005).
Shimodaira, H. & Hasegawa, M. CONSEL: for assessing the confidence of phylogenetic tree selection. Bioinformatics 17, 1246–1247, doi:10.1093/bioinformatics/17.12.1246 (2001).
Felsenstein, J. Inferring phylogenies from protein sequences by parsimony, distance, and likelihood methods. Method Enzymol 266, 418–427 (1996).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59, 307–321, doi:10.1093/sysbio/syq010 (2010).
Yang, Z. H. PAML 4: Phylogenetic analysis by maximum likelihood. Mol Biol Evol 24, 1586–1591, doi:10.1093/molbev/msm088 (2007).
Robinson-Rechavi, M. & Huchon, D. RRTree: Relative-Rate tests between groups of sequences on a phylogenetic tree. Bioinformatics 16, 296–297, doi:10.1093/bioinformatics/16.3.296 (2000).
Palmgren, M. et al. AS3MT-mediated tolerance to arsenic evolved by multiple independent horizontal gene transfers from bacteria to eukaryotes. PloS One 12, e0175422, doi:10.1371/journal.pone.0175422 (2017).
Acknowledgements
This work is supported by the Strategic Priority Research Program of Chinese Academy of Sciences (XDB15020402, XDB15020302). BPR was supported by NIH grants R01 ES023779 and R01 GM55425.
Author information
Authors and Affiliations
Contributions
Y.G.Z. designed research; S.C.C., G.X.S. and B.K.Z., performed research; S.C.C., S.Y.Z. and Y.D. contributed to data analysis; S.C.C., G.X.S., B.P.R., C.R. and Y.G.Z. wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, SC., Sun, GX., Rosen, B.P. et al. Recurrent horizontal transfer of arsenite methyltransferase genes facilitated adaptation of life to arsenic. Sci Rep 7, 7741 (2017). https://doi.org/10.1038/s41598-017-08313-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-08313-2
This article is cited by
-
Lipophilic arsenic compounds in the cultured green alga Chlamydomonas reinhardtii
Analytical and Bioanalytical Chemistry (2024)
-
Two-tiered mutualism improves survival and competitiveness of cross-feeding soil bacteria
The ISME Journal (2023)
-
Discovery of a novel bacterial class with the capacity to drive sulfur cycling and microbiome structure in a paleo-ocean analog
ISME Communications (2023)
-
Genome-wide signatures of adaptation to extreme environments in red algae
Nature Communications (2023)
-
Understanding the diversified microbial operon framework coupled to arsenic transformation and expulsion
Biologia (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.