Abstract
Among its other biological roles, acetylcholinesterase (AChE, EC 3.1.1.7), encoded by two ace in most insects, catalyses the breakdown of acetylcholine, thereby terminating synaptic transmission. ace1 encodes the synaptic enzyme and ace2 has other essential actions in many insect species, such as Chilo suppressalis and Plutella xylostella. The silkworm, Bombyx mori, has been domesticated for more than two thousand years and its aces have no history of pesticide exposure. Here, we investigated the functional differences between two ace genes, BmAce1 and BmAce2, in the silkworm. qPCR analysis indicated that BmAce1 is highly expressed in muscle and BmAce2 is more ubiquitously expressed among tissues and enriched in the head. Both genes were separately suppressed using chemically synthesized siRNAs. The mRNA abundance of the two ace genes was significantly reduced to about 13% – 75% of the control levels after siRNA injection. The AChE activities were decreased to 32% to 85% of control levels. Silencing BmAce2 resulted in about 26% mortality, faster and higher than the 20% in the siBmAce1-treated group. Silencing BmAce1 impacted motor control and development to a greater extent than silencing BmAce2, although both treatment groups suffered motor disability, slowed development and reduced cocoons. Both genes have essential, differing biological significance.
Similar content being viewed by others
Introduction
Acetylcholinesterase (AChE, EC 3.1.1.7) hydrolyzes the neurotransmitter acetylcholine into acetate and choline with a very high catalytic activity of about 25,000 molecules per second. Mammalian AChE is encoded by a single gene, ace, whereas some invertebrates have multiple aces. The situation is otherwise in insects because two aces were discovered in Aphis gossypii 1, Schizaphis graminum 2, and Anopheles gambiae 3. Since then, the presence of two aces have been confirmed in many insect species, including the mosquitoes Culex pipiens 4, Aedes aegypti 5, Culex tritaeniorhynchus 6, the aphids Myzus persicae 7, Rhopalosiphum padi and Sitobion avenae 8, the cockroach Blattella germanica 9, the lepidopterans Helicoverpa assulta 10, Plutella xylostella 11, Chilo suppressalis 12 and Cnaphalocrocis medinalis 13, the whitefly Bemisia tabaci 14 and the red flour beetle, Tribolium castaneum 15, to name some of them. It was suggested that two ace genes may have originated from gene duplication before insect speciation15, 16. However, outside of mosquitoes, ace2, but not ace1, have been reported in the model insect Drosophila melanogaster and other Drosophila species. Further analyses indicated that the orthologs of ace1 were lost in the Cyclorrapha, considered an unranked taxon by some authors and a dipteran suborder by others4.
The biological functions of the two ace genes have been assessed17. Generally, AChE1 is the major enzyme in insects, which is more abundant than AChE2. The expression of ace1 is higher than ace2 in some insect species9, 10, 18, 19, while the opposite is true in others. The AChE2 in B. mori and Apis mellifera is the major enzyme in synaptic transmission20, 21. Although AChE1 is regarded as the main target of organophosphorus (OP) and carbamate pesticides, resistance-associated mutations have been reported for both genes3, 10, 11, 18, 19, 21,22,23,24. Separate gene silencing experiments with B. germanica indicate that Bgace1 encodes the predominant AChE. We infer that while both genes are necessary, ace1 predominates in some, but certainly not all insect species.
AChE has many functions beyond acetylcholine hydrolysis25, 26. In mammals and zebrafish, AChE regulates cell-matrix interactions in bone27. In nervous tissues, it influences neuroblastoma cell adhesion and neurite outgrowth28. It acts in neocortical development by the alternative splicing of its own gene29. AChE mediates axonal growth30, synaptogenesis, memory formation, stress responses, cell proliferation and apoptosis31. Insect AChEs also function in development. Silencing larval H. armigera aces with dietary siRNAs led to high mortality, growth inhibition, malformation and drastically reduced fecundity, indicating to us the aces are multi-functional genes32. Silencing the two ace genes led to different sensitivity to insecticides33. Silencing the larval aces in Chilo suppressalis and P. xylostella showed that Csace1, Pxace1 and Pxace2 have non-typical functions in regulating larval growth and motor control12, 34. In our view, AChEs are pleiotropic genes with a wide range of biological significance.
As seen in other species, of the two B. mori genes, BmAce2 is more highly expressed than BmAce1 at the whole animal level, at different developmental stages20. The differing transcripts levels of these genes suggest to us they probably have divergent biological functions in the life cycle. In this paper, we posed the hypothesis that, in addition to synaptic transmission, BmAce1 and BmAce2 perform physiological functions in survival, larval development. Here we present and discuss the outcomes of experiments designed to test our hypothesis.
Results
Knockdown of BmAce1 and BmAce2
We designed gene-specific small interfering RNAs (siRNAs) with fluorescence-labeling to record the spread of the constructs in the silkworms under fluorescence microscopy. The injected siRNAs spread mainly to the head (Supplementary Fig. 1), where aces are highly expressed. Compared to the transcript abundances in midguts, BmAce1 transcripts accumulated to about 18-fold higher in muscle and 3.4-fold higher in heads. Similarly, the relative abundances of BmAce2 transcripts was 2.8-fold higher in muscle and 2.1-fold higher in heads, again, compared to midguts (Fig. 1). We reported that the BmAce2 had higher expression level than BmAce1 during development at the whole-animal level20. Here, at the tissue level, BmAce1 mRNA was more abundant than BmAce2 in muscle (Fig. 1A).
The relative abundance of mRNAs encoding two aces in head, midgut and muscle. Panel A: the ratios of BmAce1/BmAce2 mRNA abundances. The histogram bars represent the ratios graphically and the numbers atop the bars show the actual ratios. The error bars indicate 1 SEM. Panel B: the relative abundances of BmAce1 in the indicated tissues. Panel C: the relative abundances of BmAce2 in the indicated tissues. The histogram bars in Panels B and C depict the relative accumulation of mRNA encoding the indicated genes and the error bars indicate 1 SEM. The number shown above each bar represents the mean transcript levels of BmAce1 and BmAce2. Ribosomal Protein 49 gene (RP49) was used as the reference gene for qPCR.
To investigate the functional differences between the two aces, we injected siRNAs into the day 1, third instar larvae. Negative control siRNA was a random shuffled sequence lacking sequence identities shared with siBmAce1. The transcript abundance of BmAce1 decreased to about 52%, 64% and 75% of the control levels at 24 h, 48 h and 72 h post injection (PI; t-test, p < 0.05). siBmAce2 treatments led to decreased mRNA abundances encoding BmAce2, by approximately 13%, 16% and 21% of the control levels at 24 h, 48 h and 72 h PI (t-test, p < 0.05). Neither siRNA construct influenced accumulation of mRNA encoding the counterpart gene (Fig. 2), showing that each ace was knocked down without influencing the other.
siRNA treatments led to reduced abundances of mRNA transcripts of BmAce1 and BmAce2 in silkworm. Panel A: siAce1 treatment led to decreased mRNAs encoding BmAce1, but not BmAce2. Panel B: siAce2 treatments led to reduced accumulation of mRNAs encoding BmAce2, but not BmAce1. For both panels, the histogram bars represent mRNA abundances and the error bars indicate 1 SEM. * Significant difference at P ≤ 0.05. All experiments were repeated in triplicate, n = 30.
RNAi treatments led to decreased AChE activities
We measured AChE activities at 48 h and 96 h PI. Compared to controls, AChE activities were reduced to about 51% at 48 h (t-test, p < 0.05) and 85% at 96 h PI in the si-BmAce1 treatment group. Similarly, enzyme activities were reduced to about 32% at 48 h (t-test, p < 0.05) and 75% at 96 h PI (t-test, p < 0.05) (Fig. 3A). The RNAi efficiencies were different for two ace genes but the AChE activities were similar. We reasoned that this relates to the point that the enzyme activities are determined by the protein contents, not the mRNA levels. Among several possibilities, enzyme degradation rates may differ between the proteins and/or post-translational modifications may lead to different activity levels. The data support our view that the siRNA treatments led to reduced AChE activities, particularly at 48 h PI, despite the differences in RNAi efficiencies.
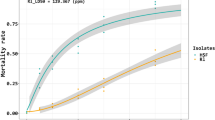
AChE activities and mortality in experimental larvae. Panel A: siRNA treatments led to reduced AChE activities at the indicated times PI. The histogram bars represent mean percent enzyme activity normalized to zero h PI. * Significant difference at P ≤ 0.05. Panel B: Mortality as a function of time PI. At all times after 24 h PI, siBmAce1- and siBmAce2-treated silkworms had higher mortalities compared to controls. Each data point indicates mean mortality and the error bars show 1 SEM. The asterisks above the bars indicate significant differences. All experiments were repeated in triplicate, n = 30.
Mortality in siRNA-treated silkworms
Silencing the two genes led to tremors, paralysis and death in some of the tested silkworms. At 24 h to 96 h PI, mortalities increased from 0 to 20% in the siBmAce1 treatment group, and from 7% to 26% in the siBmAce2 group (Fig. 3B). Because BmAce2 was more highly expressed in the head than BmAce1, knockdown of BmAce2 resulted in higher mortalities. It remains possible that the higher mortalities induced by siBmAce2 is related to its better RNAi efficiency.
Ace gene knockdown impaired motor ability
We conducted a movement test to investigate the motor ability of survivors in the experimental groups. At 120 h PI, ten silkworms from each treatment group were starved for 24 h and then offered fresh mulberry leaves placed 1.5 cm away from them (Supplementary Fig. 2). After 20 mins, 40% of the controls larvae, 85% of the siBmAce1-treated larvae and 65% of the siBmAce2-treated individuals were unable to move to the mulberry leaves (Fig. 4A). For the larvae that reached the leaves, the mean time required to consume them was 129 s in the control group, 303 s in the si-BmAce1 treated group and 226 s in the si-BmAce2 group (Fig. 4B).
siRNA treatments led to reduced motor activity at 120 h PI. Panel A: Percent of larvae not able to move to the mulberry leaves after 20 mins. The histogram bars indicate percent of treated larvae that were unable to move to the leaves. The error bars indicate 1 SEM. * Significant difference at P ≤ 0.05. Panel B: The mean time (s) required to move to the mulberry leaves. The histogram bars indicate mean times and the error bars indicate 1 SEM. The asterisks indicate significant differences. All experiments were repeated in triplicate, n = 10.
Ace knockdown led to arrested larval and pupal development
We recorded body lengths and weights of larvae at 4, 7, and 14 d PI. These parameters did not change at 4 and 7 d PI, and were decreased relative to controls at 14 d PI. Mean body lengths were about 35 mm (siBmAce1-treated group) and 38 mm (siBmAce2-treated group), significantly shorter compared to the control group (49 mm; Fig. 5A). Similarly, mean larvae weights were reduced in both experimental groups (about 571 mg in the siBmAce1 group and 505 mg in the siBmAce2 treated group at 14 days PI, lower than the control group (689 mg; Fig. 5B). The siRNA treatments led to smaller cocoons in the siBmAce1-treated group and to silk-lacking cocoons in the siBmAce2-treated group (Fig. 6).
The siRNA treatments influenced larval development. Panel A: Mean larval body lengths of siRNA-treated silkworms at the indicated d PI. The histogram bars indicate mean body lengths and the error bars indicate 1SEM. * Significant difference at P ≤ 0.05. (B) Mean larval weight of siRNA treated silkworms at 4, 7 and 14 d PI. The histogram bars indicate mean body wet weights and the error bars indicate 1SEM. * Significant difference at P ≤ 0.05. All experiments were repeated in triplicate, n = 30.
Discussion
AChE has been widely used as a pharmaceutical and insecticide target. It is the target of therapeutic agents against helminth parasites35. Tacrine, a cholinesterase inhibitor, used for the treatment of Alzheimer’s disease symptoms36. Many insecticides and acaricides target AChE to minimize crop loss. A large number of these compounds have been withdrawn because of their high environmental damage, including loss of non-target organisms and high pest insect resistance. Investigating insect ace genes and the proteins they encode is necessary to develop pesticides that are more target-selective and compatible with resistance management programs.
There are four genes encoding AChE in the nematodes Caenorhabditis elegans and C. briggsae 37. Three of them encode distinct enzymes whereas the fourth is likely to be non-catalytic35. AChE-3 acts in resistance to phoxim, an OP used in veterinary medicine and the other AChEs presumably operate in other areas of C. elegans physiology38. The nematode aces have different expression patterns in muscle, sensory neurons and motor neurons, from which we infer they have non-overlapping functions. The two insect aces also have different expression patterns and physiological functions9, 12, 17, 21, 34. Knockdown of either gene led to high death rates in experimental insects, indicating both genes have essential, and separate, actions. In rice stem borer, C. suppressalis, CsAce1 encodes an AChE that acts in sensory, motor and larval growth physiology. Silencing CsAce1 resulted in high mortality, motor disability and retarded development. CsAce2 may make specific contributions to sensory physiology because suppressing it led to mortality without influencing larval growth or motor ability12. However, in the diamondback moth, P. xylostella, PxAce2 acts in motor ability and larval growth although mRNA encoding it is less abundant than mRNA encoding PxAce1. Knockdown of either gene led to death, motor disability and slowed larval development34, indicating the biological significance of these enzymes.
B. mori has been domesticated for more than two thousand years. Here, we infer that domestication influenced the biological assignments of the aces. Based on expression patterns and RNAi results, the BmAce1 transcript was abundant specifically in muscle, where it acts in motor control and larval growth. BmAce2 was expressed in all tissues we analyzed and most likely acts in sensory physiology. This is consistent with our reports that ace2 encodes the major AChE in silkworms, whereas ace1 encodes the major AChE in other non-domesticated lepidopterans, such as C. suppressalis and P. xylostella. We hold the view that these differences between B. mori and other lepidopterans emerged after the silkworms were domesticated.
We found that the average body lengths of larvae treated with siBmAce1 and siBmAce2 were similar at 7 days PI, but, in terms of weight accumulation, siBmAce1-treated larvae were lighter at 7 d PI. SiBmAce2-treated larvae were longer, but lighter than siBmAce1-treated group. These results indicate to us that the body length, based on the photographic results, did not positively correlate with weights. These results further support our conclusion that knockdown of ace genes disrupts metabolic processes broadly. Overall, the ace1 orthologs have similar functions over a broad range of insect taxa and ace2 have generally evolved different biological functions. We found ace knockdown led to thinner and smaller cocoons, we take this finding to indicate the AChEs also act in silk production.
The RNAi efficiencies of two ace genes at the transcriptional level differed and knockdown of BmAce2 seemed to be more effective. However, we found the AChE activities of siBmAce1- and siBmAce2-treated insects were similar. We speculate the differences between siRNA efficiencies could be balanced by differences in protein stability over time and/or differences in AChE protein catabolism. Looking more broadly, we expect that understanding the molecular mechanisms of how these genes influence cocoon production will lead to new hypotheses and research results of value in the silk production industry.
Methods
Insects
The silkworms used in the experiments were breed 7091, kindly provided by Sericultural Research Institute (Zhenjiang, Jiangsu), Chinese Academy of Agricultural Sciences. First and 2nd instar larvae were maintained at 28 °C, remaining stages at 24–26 °C under 16 L:8D photoperiod and 60–80% relative humidity.
siRNA design and injection
siBmAce1 and siBmAce2 were designed for gene-specific silencing experiments. To avoid off-target effects, the siRNAs were aligned with other aces and were used to interrogate the silkworm genome by BLAST (http://www.ncbi.nlm.nih.gov/BLAST). The siRNAs that contained more than 11 contiguous base pairs of homology to the other ace or 16–17 contiguous base pairs of homology to other AChE protein coding sequences were discarded. siRNAs with shuffled siBmAce1 and siBmAce2 sequences were used as the negative control (Table 1).
siRNAs were chemically synthesized by Shanghai GenePharma Co., Ltd (Shanghai, China). To monitor the spread of siRNA in larval bodies under fluorescence microscopy, both constructs were fluorescently labelled with fluorescein amidite at the 5′end of the sense strand. A final siRNA concentration (1 μg/μl) was obtained by dissolving them in RNase-free water (Milli-Q grade). We used an Eppendorf InjectMan NI 2 microinjection system (Eppendorf, Hamburg, Germany) to inject 1 μl siRNA between the third and fourth segments of 1 d, 3rd instar larvae. The injection pressure was 200 Pa and injection time was 0.2 s. The needles were made by a micropipette puller (Model P-87, Sutter Instruments Co., Novato, CA, USA) with parameters: Heat = 596, Pull = 0, Vel = 30–40, Time = 250 s. To avoid construct leakage, needles were kept still at the injection point for 30 s. All experiments were repeated in triplicate, n = 30 larvae/treatment.
Total RNA isolation and cDNA synthesis
The tested larvae were frozen in liquid nitrogen and stored at −80 °C for further use. Total RNA was isolated with TRIzol reagent (Takara, Kyoto, Japan) following the recommended procedures. Genomic DNA was removed by treating total RNAs with DNAse I following the manufacturer’s protocol (Ambion, Austin, TX, USA). The RNA quality was checked on a 1.5% agarose gel. The first strand of the cDNA template was synthesized with Maloney murine leukemia virus reverse transcriptase (Promega, Madison, WI, USA), using Oligo (dT18) as the anchor primer and 1 mg of total RNA as the template.
qPCR
The qPCR reactions were carried out using an ABI Prism 7000 with SYBR Premix Ex TaqTM (Takara). The qPCR primers (Table 1) were designed using PrimerQuest (http://www.idtdna.com/Scitools/Applications/Primerquest/). The stably expressed Ribosomal Protein 49 (RP49) was used as the reference gene39. The qPCR program included one cycle of 95 °C for 10 s, 40 cycles of 95 °C for 5 s and then annealed at 60 °C for 31 s, followed by one cycle of 95 °C for 15 s, 60 °C for 60 s, 95 °C for 15 s and 60 °C for 15 s. The qPCR specificity was monitored with melting curve analysis and gel electrophoresis. Amplification efficiencies were determined by a series of template dilutions. The relative abundances of mRNAs was calculated using the 2−△△CT method40.
AChE activity
Enzyme activities were measured using the kinetic method with minor modifications20. Two silkworms were homogenized on ice in 1 ml phosphate buffer (0.02 M, pH 7.5, 0.1% Triton X-100) in a glass tissue grinder. The homogenate was centrifuged at 10,000 g for 20 m at 4 °C and the supernatants were used as the enzyme source. 50 μl enzyme preparation was added to well of a microplate, along with 50 μl phosphate buffer (pH 7.5, 0.02 M) and 50 μl Ellman’s reagent (5,5′-dithiobis-(2-nitrobenzoic acid; 45 μM). The reactions were started by adding 50 μl acetylthiocholine iodide (1.56 mM) at 25 °C. The reactions were monitored using a Bio-Rad microplate reader at 405 nm every 30 s for 20 min. The reaction linearity was > 0.9. Protein concentration was measured with the Bradford method using bovine serum albumin as the quantitative standard41.
Motor experiment
At five d PI silkworms were fasted for 24 h, then fresh mulberry leaves were placed 1.5 cm away from 10 silkworms for each experiment. The time required to move to the mulberry leaves and numbers of silkworms that did not reach the leaves within 20 mins were recorded. All experiments were repeated in triplicate.
Larval growth and death rates
Silkworm body lengths and larval weights were recorded at 4, 7, and 14 d PI. At 21 d PI individuals that did not move when touched with a Chinese brush were recorded as dead. The mortalities of siRNA-treated silkworms were recorded at 24 h, 48 h, 72 h, and 96 h PI.
Statistics
All experiments were repeated in three biologically independent replicates. The data were analyzed with Student’s t-test using SPSS v22.0 software (SPSS, Chicago, IL, USA). Differences were considered statistically significant at p < 0.05.
References
Li, F. & Han, Z. J. Two different genes encoding acetylcholinesterase existing in cotton aphid (Aphis gossypii). Genome 45, 1134–1141 (2002).
Gao, J. R., Kambhampati, S. & Zhu, K. Y. Molecular cloning and characterization of a greenbug (Schizaphis graminum) cDNA encoding acetylcholinesterase possibly evolved from a duplicate gene lineage. Insect Biochem Mol Biol 32, 765–775 (2002).
Weill, M. et al. A novel acetylcholinesterase gene in mosquitoes codes for the insecticide target and is non-homologous to the ace gene in Drosophila. Proc Biol Sci 269, 2007–2016, doi:10.1098/rspb.2002.2122 (2002).
Huchard, E. et al. Acetylcholinesterase genes within the Diptera: takeover and loss in true flies. Proc Biol Sci 273, 2595–2604, doi:10.1098/rspb.2006.3621 (2006).
Mori, A., Lobo, N. F., deBruyn, B. & Severson, D. W. Molecular cloning and characterization of the complete acetylcholinesterase gene (Ace1) from the mosquito Aedes aegypti with implications for comparative genome analysis. Insect Biochem Mol Biol 37, 667–674, doi:10.1016/j.ibmb.2007.03.014 (2007).
Nabeshima, T. et al. An amino acid substitution attributable to insecticide-insensitivity of acetylcholinesterase in a Japanese encephalitis vector mosquito, Culex tritaeniorhynchus. Biochem Biophys Res Commun 313, 794–801 (2004).
Nabeshima, T., Kozaki, T., Tomita, T. & Kono, Y. An amino acid substitution on the second acetylcholinesterase in the pirimicarb-resistant strains of the peach potato aphid. Myzus persicae. Biochem Biophys Res Commun 307, 15–22 (2003).
Chen, M. & Han, Z. Cloning and sequence analysis of 2 different acetylcholinesterase genes in Rhopalosiphum padi and Sitobion avenae. Genome 49, 239–243, doi:10.1139/g05-104 (2006).
Kim, J. I., Jung, C. S., Koh, Y. H. & Lee, S. H. Molecular, biochemical and histochemical characterization of two acetylcholinesterase cDNAs from the German cockroach Blattella germanica. Insect Mol Biol 15, 513–522, doi:10.1111/j.1365-2583.2006.00666.x (2006).
Lee, D. W., Kim, S. S., Shin, S. W., Kim, W. T. & Boo, K. S. Molecular characterization of two acetylcholinesterase genes from the oriental tobacco budworm, Helicoverpa assulta (Guenee). Biochim Biophys Acta 1760, 125–133, doi:10.1016/j.bbagen.2005.10.009 (2006).
Lee, D. W. et al. Mutations of acetylcholinesterase1 contribute to prothiofos-resistance in Plutella xylostella (L.). Biochem Biophys Res Commun 353, 591–597, doi:10.1016/j.bbrc.2006.12.088 (2007).
Hui, X. M. et al. RNA interference of ace1 and ace2 in Chilo suppressalis reveals their different contributions to motor ability and larval growth. Insect Mol Biol 20, 507–518, doi:10.1111/j.1365-2583.2011.01081.x (2011).
Wang, D. M. et al. MOLECULAR CHARACTERIZATION OF TWO ACETYLCHOLINESTERASE GENES FROM THE RICE LEAFFOLDER, Cnaphalocrocis medinalis (LEPIDOPTERA: PYRALIDAE). Arch Insect Biochem Physiol 93, 129–142, doi:10.1002/arch.21347 (2016).
Alon, M., Alon, F., Nauen, R. & Morin, S. Organophosphates’ resistance in the B-biotype of Bemisia tabaci (Hemiptera: Aleyrodidae) is associated with a point mutation in an ace1-type acetylcholinesterase and overexpression of carboxylesterase. Insect Biochem Mol Biol 38, 940–949, doi:10.1016/j.ibmb.2008.07.007 (2008).
Lu, Y. et al. Genome organization, phylogenies, expression patterns, and three-dimensional protein models of two acetylcholinesterase genes from the red flour beetle. PLoS One 7, e32288, doi:10.1371/journal.pone.0032288 (2012).
Cha, D. J. & Lee, S. H. Evolutionary origin and status of two insect acetylcholinesterases and their structural conservation and differentiation. Evol Dev 17, 109–119, doi:10.1111/ede.12111 (2015).
Kim, Y. H. & Lee, S. H. Which acetylcholinesterase functions as the main catalytic enzyme in the Class Insecta? Insect Biochem Mol Biol 43, 47–53 (2013).
Jiang, X. et al. Mutation in acetylcholinesterase1 associated with triazophos resistance in rice stem borer, Chilo suppressalis (Lepidoptera: Pyralidae). Biochem Biophys Res Commun 378, 269–272, doi:10.1016/j.bbrc.2008.11.046 (2009).
Lee, S. H., Kim, Y. H., Kwon, D. H., Cha, D. J. & Kim, J. H. Mutation and duplication of arthropod acetylcholinesterase: Implications for pesticide resistance and tolerance. Pestic Biochem Physiol 120, 118–124, doi:10.1016/j.pestbp.2014.11.004 (2015).
Chen, H. J. et al. Ace2, rather than ace1, is the major acetylcholinesterase in the silkworm. Bombyx mori. Insect Science 16, 297–303 (2009).
Kim, Y. H., Cha, D. J., Jung, J. W., Kwon, H. W. & Lee, S. H. Molecular and kinetic properties of two acetylcholinesterases from the western honey bee. Apis mellifera. PLoS One 7, e48838, doi:10.1371/journal.pone.0048838 (2012).
Li, F. & Han, Z. Mutations in acetylcholinesterase associated with insecticide resistance in the cotton aphid, Aphis gossypii Glover. Insect Biochem Mol Biol 34, 397–405, doi:10.1016/j.ibmb.2004.02.001 (2004).
Chen, Z., Newcomb, R., Forbes, E., McKenzie, J. & Batterham, P. The acetylcholinesterase gene and organophosphorus resistance in the Australian sheep blowfly. Lucilia cuprina. Insect Biochem Mol Biol 31, 805–816 (2001).
Temeyer, K. B. & Chen, A. C. Identification and characterization of a cDNA encoding the acetylcholinesterase of Haematobia irritans (L.) (Diptera: Muscidae). DNA Seq 18, 85–91, doi:10.1080/10425170601060558 (2007).
Jiang, H. & Zhang, X. J. Acetylcholinesterase and apoptosis. A novel perspective for an old enzyme. FEBS J 275, 612–617, doi:10.1111/j.1742-4658.2007.06236.x (2008).
Seibt, K. J. et al. Typical and atypical antipsychotics alter acetylcholinesterase activity and ACHE expression in zebrafish (Danio rerio) brain. Comp Biochem Physiol C Toxicol Pharmacol 150, 10–15 (2009).
Inkson, C. A., Brabbs, A. C., Grewal, T. S., Skerry, T. M. & Genever, P. G. Characterization of acetylcholinesterase expression and secretion during osteoblast differentiation. Bone 35, 819–827, doi:10.1016/j.bone.2004.05.026 (2004).
Johnson, G. & Moore, S. W. Cholinesterases modulate cell adhesion in human neuroblastoma cells in vitro. Int J Dev Neurosci 18, 781–790 (2000).
Dori, A., Cohen, J., Silverman, W. F., Pollack, Y. & Soreq, H. Functional manipulations of acetylcholinesterase splice variants highlight alternative splicing contributions to murine neocortical development. Cereb Cortex 15, 419–430, doi:10.1093/cercor/bhh145 (2005).
Bigbee, J. W. & Sharma, K. V. The adhesive role of acetylcholinesterase (AChE): detection of AChE binding proteins in developing rat spinal cord. Neurochem Res 29, 2043–2050 (2004).
Jin, Q. H., He, H. Y., Shi, Y. F., Lu, H. & Zhang, X. J. Overexpression of acetylcholinesterase inhibited cell proliferation and promoted apoptosis in NRK cells. Acta Pharmacol Sin 25, 1013–1021 (2004).
Kumar, M., Gupta, G. P. & Rajam, M. V. Silencing of acetylcholinesterase gene of Helicoverpa armigera by siRNA affects larval growth and its life cycle. J Insect Physiol 55, 273–278, doi:10.1016/j.jinsphys.2008.12.005 (2009).
Revuelta, L. et al. RNAi of ace1 and ace2 in Blattella germanica reveals their differential contribution to acetylcholinesterase activity and sensitivity to insecticides. Insect Biochem Mol Biol 39, 913–919, doi:10.1016/j.ibmb.2009.11.001 (2009).
He, G., Sun, Y. & Li, F. RNA interference of two acetylcholinesterase genes in Plutella xylostella reveals their different functions. Arch Insect Biochem Physiol 79, 75–86, doi:10.1002/arch.21007 (2012).
Selkirk, M. E., Lazari, O. & Matthews, J. B. Functional genomics of nematode acetylcholinesterases. Parasitology 131(Suppl), S3–18, doi:10.1017/S0031182005008206 (2005).
Camps, P. et al. Tacrine-based dual binding site acetylcholinesterase inhibitors as potential disease-modifying anti-Alzheimer drug candidates. Chem Biol Interact 187, 411–415, doi:10.1016/j.cbi.2010.02.013 (2010).
Combes, D., Fedon, Y., Grauso, M., Toutant, J. P. & Arpagaus, M. Four genes encode acetylcholinesterases in the nematodes Caenorhabditis elegans and Caenorhabditis briggsae. cDNA sequences, genomic structures, mutations and in vivo expression. J Mol Biol 300, 727–742, doi:10.1006/jmbi.2000.3917 (2000).
Han, Y., Song, S., Guo, Y., Zhang, J. & Ma, E. ace-3 plays an important role in phoxim resistance in Caenorhabditis elegans. Ecotoxicology 25, 835–844, doi:10.1007/s10646-016-1640-z (2016).
He, L., Shen, G., Li, F. & Huang, S. Automatic extraction of reference gene from literature in plants based on texting mining. Int J Data Min Bioinform 12, 400–416 (2015).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3, 1101–1108 (2008).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72, 248–254 (1976).
Acknowledgements
We thank Professor Muwang Li, Sericultural Research Institute (Zhenjiang, Jiangsu), Chinese Academy of Agricultural Sciences for kindly providing the silkworms. This work was partially supported by the National Key Research and Development Program [2016YFC1200600], National Basic Research Program of China [2013CB127600] and The Science and Technology Research Project of the Ministry of Education [V201308]. Mention of trade names or commercial products in this article is solely for providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. All programs and services of the U.S. Department of Agriculture are offered on a nondiscriminatory basis without regard to race, color, national origin, religion, sex, age, marital status, or handicap.
Author information
Authors and Affiliations
Contributions
X.Y. analyzed the data, made the figures and table, drafted the manuscript. L.Y. carried out experiments and analyzed the data. D.S. wrote the manuscript. F.L. designed this project, analysed the data and wrote the manuscript, Q.F. participated in discussion and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ye, X., Yang, L., Stanley, D. et al. Two Bombyx mori acetylcholinesterase genes influence motor control and development in different ways. Sci Rep 7, 4985 (2017). https://doi.org/10.1038/s41598-017-05360-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-05360-7
This article is cited by
-
Identification and validation of potential reference gene for effective dsRNA knockdown analysis in Chilo partellus
Scientific Reports (2019)
-
RNAi in Tuta absoluta management: effects of injection and root delivery of dsRNAs
Journal of Pest Science (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.