Abstract
Plant leaves are a crucial organ associated closely with chloroplast development, photosynthesis rate and crop productivity. In this study, a white fine stripe leaf 1 (wfsl1) mutant was isolated and characterized from the japonica rice Zhonghua11 (ZH11) after ethyl methanesulfonate mutagenesis. The wfsl1 displayed white fine stripe leaves since tillering stage and abnormal chloroplast structure. Map-based cloning and Bioinformatic analysis indicated that WFSL1 on chromosome 1 contains an “A” to “T” substitution in protein coding region, and encodes a putative metal-dependent phosphohydrolase with HD domain at the N-terminus. WFSL1 was targeted to the chloroplasts and had higher expression in mature leaves and sheaths. RNA-seq analysis revealed that chloroplast development and photosynthesis genes were significantly affected in wfsl1 plants. Levels of WFSL1 and chloroplast encoded proteins were decreased in wfsl1 mutants via western blot analysis. Compared with WT, wfsl1 exhibits lower Chl content and defective in biogenesis of chloroplast ribosomes, which resulted in reduced grain yield. Taken together, our results show that WFSL1 is critical for chloroplast development, ribosome biogenesis, and light energy utilization, finally affects grain yield.
Similar content being viewed by others
Introduction
Chloroplast, the center for leaf cell metabolism which plays an important role in light reception and carbon sequestration in higher plants1. Chloroplast development is regulated by genes encoded of nuclear and plastids, but for the limited coding capacity of plastids, chloroplast development is mainly under nuclear control. The coordination of gene expression through nuclear and plastids is essential for chloroplasts biogenesis in plant2. The transcription of nuclear and chloroplast genes mainly depends on two RNA polymerase. One is the bacterial-type nuclear-encoded RNA polymerase (NEP). NEP transcribes plastid genes involved RpoA, B, C that are necessary for the development of plastidic genetic systems in the early stage of chloroplast development. The other bacterial-type plastid-encoded RNA polymerase (PEP) is response for the photosynthesis genes (such as RbcL, RbcS, PsbA, RCA) development at mature stage3, 4. Owing to defects in PEP activity, the formation of thylakoid membrane and photosynthesis was repressed, suggesting it is an important role of PEP in chloroplast development5. Proteins encoded by the plastid genome are synthesized by plastidic prokaryotic type 70 S ribosome that are composed of 30 S and 50 S subunits6,7,8,9. Deficiency in content of 70 S ribosome results in stunted chloroplast development. Arabidopsis konck-down mutant RH22 accumulated precursors 23 S rRNA that displayed virescent phenotype. RH22 affected ribosome assembly in rRNA metabolism10. ObgC participated in 70 S ribosome assembly. The knock down and RNAi of ObgC result in chlorotic phenotype in rice8. Thus, 70 S ribosome biogenesis is essential for chloroplast development in higher plants.
Plant variegations are characterized by the presence of white sectors and green sectors. The white sectors contain defective chloroplasts and the green sectors contain normal chloroplasts11. Immutans (im) and var2 are two types variegation mutants in Arabidopsis. The im mutant is induced by a nuclear recessive gene and the extent of variegation can be modulated by light and temperature. IM protein is a plastid homologue of the mitochondrial alternative oxidase and functions as a redox component of the phytoene desaturation pathway11. Mutations in the VAR2 locus cause variegation due to loss of a chloroplast thylakoid membrane protein that is similar to the FtsH family of AAA protein. FtsH functions in a number of diverse membrane-associated events and VAR2 protein functions in thylakoid membrane biogenesis12. Both im and var2 mutants provide an excellent system to understand the molecular mechanism of nuclear-plastid interactions in Arabidopsis. In rice, many genes were also associated with nuclear-plastid interaction. V1 encodes a chloroplast-localized protein NUS1 regulating chloroplast RNA metabolism13,14,15,16. V2 encodes a plastids/mitochondria Guanylate kinase (pt/mtGK) which catalyzes GMP to GDP in guanylate biosynthesis and affects chloroplast development. It suggests that pt/mtGK is an important part for chloroplast development17. RNRL1 and RNRS1 encode the large and small subunits of rice ribonucleotide reductase, respectively. They are necessary for the DNA replication in chloroplast18. OsDVR plays certain roles in nuclear-cytoplasmic signal transduction by which nucleus directly regulates cytoplasm19, 20. Leaf-color mutations are a crucial for dissection of regulation mechanism of with chloroplast development, photosynthesis and key agronomic traits.
Here, we isolated a w hite f ine s tripe l eaf (wfsl1) mutant in rice. The phenotype of mutant not very likely with im and var2 mutants, the green and white sectors longitudinal distributed on the blade of mutants. It’s interesting that mutant phenotype mainly first occurs on the newly grown leaves and sheath at tillering stage, continued to the mature stage. Our study discovers that map-based cloning identifies WFSL1 on chromosome 1 encoding an HD domain containing gene. The WFSL1 protein is localized to chloroplast and was decreased in wfsl1 mutant leaves. The wfsl1 mutant was defective in biogenesis of chloroplast ribosomes, and affected photosynthesis rate and grain yield. Our results suggest that WFSL1 is critical for chloroplast development and regulating chloroplast ribosome biogenesis.
Results
Phenotypic characterization of wfsl1 mutant
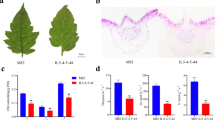
The wfsl1 mutant was isolated from an M2 population of the japonica rice Zhonghua11 (ZH11) after Ethyl methanesulfonate mutagenesis, and was designated according to the chromosome location and white fine stripe leaf phenotype. At seedlings wfsl1 showed normal phenotype (Fig. 1a) but it emerged white fine stripe leaf at tillering stage and continued to the mature stage (Fig. 1b–e). Cross-section observation showed content of chlorophyll decreased in wfsl1 mutants (Fig. 1f,i). The chlorophyll and carotenoid levels were nearly the same at seedling stage (Fig. 1j), but they remarkably decreased in wfsl1 mutants at tillering stage (Fig. 1k). Transmission electron microscopy (TEM) indicated that wfsl1 had green, green-white and white types of cells (Fig. 2a–d). The green cells have more chloroplast and well-developed thylakoid membrane systems, whereas the chloroplast were small with no thylakoid membranes in wfsl1 white cells (Fig. 2e–j). So the cells are heteroplastidic in mutants.
Phenotypes of the wild type and wfsl1 mutant. (a) Wild type and wfsl1 seedlings. (b) Wild type plants at the tillering stage. (c) White fine stripe leaf of wfsl1 at tillering stage. (d,e) The enlarged leaves of (b,c). (f,i) Cross-section of wild type and wfsl1 leaf at tillering stage. (g) The internode length of wild type (left) and wfsl1 (right). (h) The panicle pattern of wild type (left) and wfsl1 (right). (j) The chlorophyll content of wild type and wfsl1 seedlings. (k) The chlorophyll content of wild type and wfsl1 at tillering stage. Bars: (a) 10 cm; (b,c) 5 cm; (f,i) 200 μm; (g,h) 2 cm. Data are mean ± SD (n = 3). Error bars represent the SD from three independent experiments (Student’s t-test, *P < 0.05; **P < 0.01).
Leaf morphology and transmission electron microscopic images of chloroplasts of wild type and wfsl1. (a) Wild type. (b) Green leaf sectors of wfsl1. (c) Green and white leaf sectors of wfsl1. (d) White leaf sectors of wfsl1. (e,f) Wild type leaf sectors chloroplast morphology at tillering stage. (g,h) Green and white leaf sectors of chloroplast morphology at tillering stage of wfsl1. (i,j) White leaf sectors chloroplast morphology at tillering stage of wfsl1.(Ch) Chloroplast. (G) Grana. (M) Mitochondria. (N) Nucleus. (OG) Osmiophilic plastoglobuli. (S) Starch granule.
The wfsl1 plants reduced grain yield
Agronomic trait analysis indicated that the plant height, panicle length, setting rate, and thousand seed weight were remarkably reduced in wfsl1, compared with WT (Fig. 3a). The length of topmost and second topmost interondes was significantly decreased in wfsl1 (Fig. 3a). The setting rate was 70.1% of wfsl1 while 89% of wild type (Fig. 3a). The thousand seed weight reduced to 22.48 g of wfsl1 where wild type was 25.98 g (Fig. 3a). The photosynthesis rate reduced in wfsl1 (Fig. 3b). However, no difference of tiller number between wild type and wfsl1 was found (Fig. 3a). TEM suggested chloroplasts were small and thylakoid membrane impaired in wfsl1 which may reduce photosynthesis. In summary, the wfsl1 mutation resulted in the reduction of grain yield.
Agronomic traits and photosynthesis analysis of wild type and wfsl1 plants. (a) Agronomic traits analysis of wild type and wfsl1. (b) Photosynthesis rate analysis of wild type and wfsl1 at tillering stage. PH Plant height. TN Tiller number. PL Panicle length. PBN Primary branch number. SBN Second branch number. TIL Topmost internode length. SIL Secondmost internode length. SR Setting rate. TGW thousand grain weight. Error bars represent the SD from three independent experiment (Student’s t-test, *P < 0.05; **P < 0.01; n = 6).
Map-based cloning and functional confirmation of WFSL1
The WFSL1 was cloned in an F2 population derived from the cross between the wfsl1 and an indica cultivar NJ06. The segregation ratio of wild type to white fine stripe leaf phenotype among the F2 population was 3:1 (365:107, χ 2 = 1.3672 < χ2 0.05, 1; P = 0.2423 > 0.05), suggesting that the phenotype of wfsl1 mutant is controlled by a single recessive gene (Supplementary Table S2). The WFSL1 gene was primarily mapped to the region between the markers RM3252 and RM5336 on chromosome 1 (Fig. 4a). Then eight insertion-deletion polymorphism (InDel) markers were developed for further fine-mapping (Supplementary Table S1). The location of WFSL1 was narrowed down to a 50.5 kb region which includes 12 ORFs (open reading frame). The predicted region was sequenced to detect the mutation and discovered that the target ORF (LOC_Os01g01920) had an “A” to “T” substitution in protein coding region (Fig. 4a). The gene includes eighteen exons and seventeen introns. The mutation site on the seventeenth exon, with the amino acid Asn changed into Tyr (Fig. 4b). Gene prediction (Rice Genome Annotation Project) indicates that it encodes a putative metal-dependent phosphohydrolase. Sequencing alignment indicates that WFSL1 has a conserved HD domain motif (Supplementary Fig S1). To confirm whether the phenotype of wfsl1 mutant is caused by the loss function of WFSL1, complementation test was conducted. The genomic DNA fragment containing WFSL1 sequence, the upstream and downstream sequences were inserted into a binary vector to construct pCAMBIA1300-WFSL1 vector. This vector was introduced into the wfsl1 mutants by Agrobacterium-mediated transformation. More than eight transgenic lines exhibited normal green leaves (Fig. 4c), and the content of chlorophyll had no difference contrast to wild type (Fig. 4d). Besides, sequencing results confirmed the complementation plants (COM) (Fig. 4e). And a 1,743-bp fragment of full-length cDNA from the cDNA library of wild type was inserted into a binary overexpression vector to construct pCAMBIA1300S-WFSL1. The vector was transformed into wfsl1 mutants to generate overexpression plants (OE). The overexpression plants exhibited normal green leaves and relative expression increased significantly compared with wild type and mutants (Fig. 4f). Thus, the white fine stripe leaf phenotype was rescued. These results suggest that the single base substitution in LOC_Os01g01920 (WFSL1) is responsible for the phenotype of wfsl1 mutant.
Map-based cloning of the WFSL1 gene. (a) Fine mapping of WFSL1. Markers used for mapping are indicated. AP002863 and AP002845 are accession of BACs. (b) Structure of WFSL1. ATG and TGA represent the start and stop codons, respectively. Black boxes indicate the exons and the lines indicate introns. A single nucleotide replacement ‘A’ to ‘T’ led to an amino acid change. (c) Leaf phenotype at tillering stage wild type and wfsl1 (left), complementation plants (COM) (middle), overexpression plants (OE) (right). (d) The chlorophyll content of wild type and complementation plants at tillering stage. (e) Sequencing results indicate wild type genome DNA fragment transfer into wfsl1 plants. (f) The expression level of wild type, wfsl1, complementation and overexpression at tillering stage. The relative expression level of wfsl1 represents “1” Data are mean ± SD (n = 3). Bars: (c) 5 cm. Error bars represent the SD from three independent experiments.(Student’s t-test, *P < 0.05; **P < 0.01)
Location of WFSL1 to chloroplast
To investigate the subcellular localization of WFSL1, the full-length WFSL1 cDNA was amplified from the cDNA library of wild type. The WFSL1 cDNA was ligated to GFP sequence, and injected into rice protoplasts. Its expression was under the control of 35 S promoter. The GFP fluorescence in transformed protoplasts was examined using confocal fluorescence microscope, which showed that WFSL1 was located to chloroplast (Fig. 5a,b).
Subcellular localization and expression pattern analysis of WFSL1. (a,b) Localization of GFP (control) and WFSL1 protein in rice protoplasts. Green fluorescence shows GFP, red fluorescence shows chlorophyll, orange indicates the two types of florescence merged. (c) qRT-PCR analysis WFSL1 expression in young root, stem, young leaf, mature leaf, sheath, panicle and mature root of wild type. The relative expression level of stem represents “1”. (d,e) Relative expression levels of some genes of chloroplast synthesis, degradation and development. Error bars represent the SD from three independent experiments. (Student’s t-test, *P < 0.05; **P < 0.01).
Analysis of WFSL1 and plastid-encoded genes expression pattern
qRT-PCR was used to examine the tissue-specific expression pattern of WFSL1. RNA was extracted from young leaves and young roots (three leaf stage), mature leaves and mature roots (heading stage), stems, sheaths and panicles, respectively. WFSL1 was highly expressed in sheaths and mature leaves, but low expression in young leaves, roots, stems and panicles (Fig. 5c). Next, we investigated the transcription levels of the genes associated with chlorophyll biosynthesis, chlorophyll degradation or chloroplast development in wfsl1 mutant. Ten genes were selected, including chlorophyll biosynthesis-related genes OsPORA 21, 22 and OsPORB 22, OsCAO1 and OsCAO2 23, chlorophyll degradation-related genes NOL 24 and SGR 25. Chloroplast developmental genes V1 13,14,15,16, V2 17, OsDVR 19, 20 and OsChlH 26. UBQ5 (Actin) was used as control27 (Supplementary Table S3). qRT-PCR analysis showed that the transcriptions of OsPORB, NOL, V1, OsDVR, OsChlH were significantly suppressed in wfsl1 mutant at seedling stage. The expression level of OsPORA, OsPORB, SGR were increased and OsCAO1, OsCAO2, V1, V2, OsDVR, OsChlH were decreased in mutant at tillering stage (Fig. 5d,e). The relative expression levels of plastid-encoded genes PsaA, PsbA, AtpB and Rubisco large subunit (RbcL), Rubisco activase (RCA) were all decreased in wfsl1. while the expression levels of nuclear-encoded genes RpoA, RpoB, RpoC1, RpoC2 increased in wfsl1. These results suggest wfsl1 is defective in biogenesis of plastid-encoded genes and affects chloroplast development.
Expression of photosynthesis related genes is repressed in wfsl1
RNA-seq was performed to analyze the effect of the wfsl1 mutation on gene expression. More than 40 million reliable clean reads were obtained from wild type and wfsl1. About 355 up regulated genes while 536 repressed genes in wfsl1 (Fig. 6a–d, Supplementary Table S4). We randomly selected 10 down-regulated and 10 up-regulated genes, and using qRT-PCR methods to verify the results of RNA-seq. The qRT-PCR results were consistent with the RNA-seq (Fig. 6e). Go and KEGG enrichment analysis indicated that genes encoding photosynthesis, light reaction, chloroplast envelope, PSI and PSII, chlorophyll binding, carbon fixation were remarkably reduced in wfsl1 (Supplementary Figs S2 and S3).
RNA-seq analysis of wild type and wfsl1. mRNA was purified from total RNA isolated from tillering stage plants of wild type and wfsl1 using poly-T oligo-attached magnetic beads. cDNA was synthesized using random hexamer primers. The library was constructed and sequenced using an Illumina Hisequation 2000. (a) Numbers of genes sorted according to their expression level. (b) Read numbers of wild type and wfsl1 sequences. (c) Volcano plot showing the overall alterations in gene expression in wild type and wfsl1. (d) Cluster analysis of differently expressed genes in wild type and wfsl1 Red represents high expression genes. Blue represents low expression genes. (e) qRT-PCR analysis differently expression genes of RNA-seq. 20 up-regulated or down-regulated genes were tested. Error bars represent the SD from three independent experiments (Student’s t-test, *P < 0.05; **P < 0.01)
Defects of WFSL1 and plastid proteins in wfsl1
We tested the accumulation of WFSL1 protein in wild type and wfsl1 mutants using western-blot analysis. However, the accumulation of WFSL1 protein was decreased in wfsl1 mutant at tillering stage (Fig. 7a). SDS-PAGE gel and western-blot suggest the protein levels of the large subunit of Rubisco (RbcL) and Rubisco activase (RCA) were decreased in wfsl1 (Fig. 7b,c).Other plastidic proteins including ATP synthase subunit beta (AtpB), A1 of PSI, D1 of PSII, “alpha and beta” subunits of RNA polymerase were also tested. The results showed that the levels of plastid-encoded proteins were significantly decreased in wfsl1 (Fig. 7c). qRT-PCR results suggest the expression levels of class I genes RbcL, PsbA, AtpB were remarkably decreased, while class III genes including RpoA, RpoC2 increased (Fig. 7d). RNA-seq results indicated that expression levels of many plastidic genes changed between wild type and wfsl1. Class I, II and III genes are three types of plastidic genes in plant. Class I genes are transcribed by PEP, class II genes are transcribed by NEP and PEP, and class III genes are mainly transcribed by NEP. The expression levels of class III genes were increased while class I genes decreased (Fig. 8d). These results indicate that wfsl1 was defective in PEP activity and chloroplast protein biosynthesis.
Protein levels analysis of WFSL1 and some representative proteins. (a) Western blot analysis of wild type and wfsl1 at seedling and tillering stage. (b) SDS-PAGE gel shows Rubisco using Coomassie Brilliant Blue stained in wild type and wfsl1 at tillering stage. (c) Western blot analysis of chloroplast proteins and RCA. (d) qRT-PCR analysis the relative expression level of plastic encoding genes in wild type and wfsl1 at tilering stage. Hsp90 was used as an internal control. Error bars represent the SD from three independent experiments (Student’s t-test, *P < 0.05; **P < 0.01).
Analysis of rRNA and chloroplast ribosomal gene expression in wild type and wfsl1. (a) Total RNA from wild type and wfsl1 at tillering stage. (b,c) rRNA analysis from wild type and wfsl1 using Agilent 2100, respectively. (d) qRT-PCR analysis of the expression level of 16 S, 23 S, 18 S and 25 S in wild type and wfsl1 plants. (e) RNA-seq analysis of chloroplast ribosomal genes. RNA was isolated from wild type and wfsl1 at tillering stage. Error bars represent the SD from three independent experiments (Student’s t-test, *P < 0.05; **P < 0.01).
Reduction of chloroplast ribosome activity in wfsl1
Chloroplast ribosome is composed of 30 S small subunit and 50 S large subunit. The 30 S and 50 S subunits are mainly comprised of 16 S and 23 S rRNAs and ribosomal genes9. We found that the 16 S and 23 S rRNAs were decreased in wfsl1 (Fig. 8a). We used an Agilent 2100 to analyze the composition and content of rRNAs from wild type and wfsl1 at tillering stage. We found that the 16 S and 23 S rRNAs were reduced to one half of wild type levels (Fig. 8b–d). RNA-seq analysis indicates that the expression levels of some ribosomes 30 S genes RPS1, RPS5, RPS17 and 50 S genes RPL5, RPL11, RPL13, RPL18 were all decreased in wfsl1 (Fig. 8e). These results indicated that wfsl1 mutant was defective in biogenesis of chloroplast ribosomes.
Discussion
Rice leaf-color mutants are key for investigation of chloroplast development, light energy utilization rate and reproduction. In this study, we isolated the wfsl1 mutant displayed white fine stripe leaves at tillering stage (Fig. 1b,c). Compared with the wild type, wfsl1 mutant exhibited pleiotropic phenotypes including reduction of plant height, panicle length, seed setting rate, and thousand seed weight. Also the chlorophyll content decreased in wfsl1 plants since tillering stage compared with WT (Fig. 1f,i,j,k). wfsl1 has three types of cells (green, green and white, white) and its chloroplasts were defective in tillering stage (Fig. 2e–j). TEM results identified the pattern of the white fine stripe phenotype, suggesting that wfsl1 cells are heteroplastidic containing white cell sector with abnormal chloroplasts (Fig. 2a–d). The wfsl1 mutant only has single-base mutation and leads to a series of phenotypic changes. These phenotypic differences between the wfsl1 and previously reported mutants may be due to the different genetic backgrounds and the different mutation sites in the target genes28,29,30. So determination of the pattern of white fine stripe leaf using the wfsl1 mutant is an important to understand the mechanism of white fine stripe and the function of WFSL1 protein. The reduction of the content of chlorophyll in wfsl1 is useful to study the relationship among WFSL1 and chloroplast development, light energy utilization and grain yield.
We cloned WFSL1 using map-based cloning method and confirmed its function in transgenic plants (Fig. 4). WFSL1 has a HD domain motif31 (Supplementary Fig. S1) with a H…HD…H…H…D sequence motif. The histidines or aspartates in HD domain are highly conserved and coordinate with the metal ion to regulate the activity of protein. HD domain is globular in dGTPases32 and has the (Guanosine 5′-triphosphate 3′-diphosphate) ppGpp hydrolase activity of SpoT protein in E.coli, which supports that the protein containing HD-domain is a phosphohydrolase33. Cyclic nucleotide Phosphodiesterase (PDEs) has a HD domain, and is regulated by Zn2+ coordination. Other metals, such as Mn2+, Co2+ and Mg2+, also have catalytic activity34. RelA and SpoT regulate ppGpp levels in E.coli 35. In SpoT protein, HD domain has ppGppase activity33, and the His-Asp doublet is involved in the hydrolysis for ppGpp36. Arabidopsis has four RelA/SpoT homologues AtRSH1, AtRSH2, AtRSH3 and AtCRSH. They are all targeted to plastids and expressed in green tissues and flowers which have important functions in chloroplast development and reproduction via ppGpp synthetase activity37, 38. The transcripts encoding the RC and LHC subunits of PSI and PSII, the small and large subunits of Rubisco was decreased in OX:RSH2 and OX:RSH3 plants. Plastid-encoded genes were affected in OX:RSH2 and OX:RSH3 plants39.
Also WFSL1 was located to chloroplast (Fig. 5a,b), and the same that the HD domain containing proteins OsCRSH1, OsCRSH2 and OsCRSH3 were located to chloroplast38. Go term analysis found that genes of photosynthesis, photosystem II assembly, and plastid were highly expressed in wfsl1 (Supplementary Fig. S1). KEGG analysis found that photosynthesis, and carbon fixation metabolism were significantly difference in WT and wfsl1 (Supplementary Fig. S2). These suggest WFSL1 was essential in chloroplast development. The expression of chlorophyll synthesis genes OsCAO1 and OsCAO2, chloroplast development genes V1, V2, OsDVR and OsChlH and plastid encoded genes PsaA, PsbA, AtpB, RCA were all decreased in wfsl1 at tillering stage. The expression level of nuclear-encoded genes RpoA, RpoB, RpoC1, RpoC2 was increased in wfsl1 (Fig. 7d). These suggest plastid encoded genes were defective in wfsl1. WFSL1 is highly expressed in mature leaves and sheaths, indicating that it functions at tillering and mature stages (Fig. 5d).
The expression class I genes decreased and class III genes increased in wfsl1. It is similar to these PEP-related mutants obgc 8, rh3 40, and PPR proteins mutants41, 42. So we further conducted western blot analysis, and found WFSL1 protein levels were decreased in wfsl1 at tillering stage (Fig. 7a). These results in decreased expression levels of WFSL1 at tillering stage (Fig. 5h,i). The rubisco large subunit, plastid-encoded proteins and Rubisco activase (RCA) were decreased in wfsl1. Nuclear-encoded protein RpoB increased in wfsl1. qRT-PCR results indicate that the expression levels of plastid-encoded genes decreased and nuclear-encoded genes increased in wfsl1. These are very likely previously reported mutant wp1 9. The wp1 mutant decreased plastid-encoded proteins and defective in chloroplast developmental. Since levels of plastid-encoded proteins decreased in wfsl1. We analyzed the content and composition of rRNA using an Agilent 2100. The results showed little difference between 18 S and 25 S rRNA levels in wild type and wfsl1. However, we found that wfsl1 16 S and 23 S rRNAs contents were dropped one half that of wild type (Fig. 8a–d). RNA-seq analysis suggest expression levels of ribosomal genes, including 50 S ribosomal genes RPL5, RPL10, RPL18, RPL21 and 30 S ribosomal genes RPS1, RPS5, RPS9, RPS17 were all decreased in wfsl1 (Fig. 8e). These results indicated that wfsl1 was defective in chloroplast ribosome biogenesis
Chloroplast plays an important role in light reception in higher plants1. In this study, the wfsl1 displayed white fine stripe leaves and reduced Chl content at tillering stage (Fig. 1f,i,k). The thylakoid was abnormal in wfsl1 (Fig. 2e–j), and leads to the reduced of photosynthesis rate. Some plastid-encoded proteins A1 of PSI (PsaA), D1 of PSII (PsbA), Rubisco large subunit and Rubisco activase (RCA) were decreased in wfsl1 (Fig. 7a–c). PSI and PSII are two pathways that responsible for electron transfer during photosynthesis. Ribulose-1,5-bisphosphate (RuBP) carboxylase/oxygenase (Rubisco), which constitute of RbcL and RbcS subunit that catalyzes the first step in net photosynthetic CO2 assimilation and photorespiratory carbon oxidation43, 44. PSI and PSII may inhibit protein levels of RbcL in wfsl1, suggesting that the photosynthesis was affected in wfsl1 plants. The decreased setting rate and thousand seed weight (Fig. 3a) were associated with decreased photosynthesis in wfsl1 plants. These results showed that defective developmental of chloroplast affected the photosynthesis rate and grain yield in wfsl1.
In conclusion, this study suggested that WFSL1, which is essential in expression of plastid genes and plastid ribosome biogenesis, is important for chloroplast development.
Materials and Methods
Plant materials and growth conditions
The wfsl1 mutant was isolated from a M2 population of the japonica rice ZH11 (wild type) after Ethyl methanesulfonate mutagenesis. The japonica rice ZH11 and wfsl1 mutant were grown in the paddy fields of Zhejiang (30°03′N, summer season, temperate climate) and Hainan (18°48′N, winter season, subtropical climate) provinces in China under local growing conditions.
Chlorophyll analysis
The leaves from different stages of wild type and wfsl1 were collected and weighted. Then, the leaves were soaked in an acetone and ethanol mixture solution at 26 °C in dark for 24 h. The content of chl a, chl b and carotenoids were calculated as described previously45.
Transmission electron microscopy (TEM) analysis
The leaves of wild type and wfsl1 were selected at tillering stages and cut into small pieces. They were fixed by 2.5% glutaraldehyde (PH 7.2) and vacuumed until fully sinking to the bottom. Subsequently, samples were successively washed three times with 0.2 mol/L sodium cacodylate buffer for 30 min, fixed in 10% osmic acid for 1 h, distilled three times with deionized water for 45 min, dehydrated with ethanol, treated with acetone and embedded in epoxy resins and polymerized at 70 °C. The samples were then cut into about 500–800 Ǻ thick with a slicer and stained by the mixture of uranyl acetate dihydrate and lead citrate. The sections were washed with deionized water and visualized using HITACHI Transmission Electron Microscope (HT7700).
Genetic analysis and map-based cloning
For genetic analysis, we constructed the crosses between wfsl1 and the indica cultivars TN1, 93–11, SH527 (Shuhui527) to analyze whether a dominant/recessive single/multiple gene control(s) the wfsl1 phenotype. The segregation population of F2 was examined by χ 2 test (Additional file 1: Table S2).
To map the genomic location of WFSL1, 1,900 mutants were selected from the F2 population which was derived from the cross between wfsl1 and the indica cultivar NJ06 (Nanjing06). A total of 117 pairs of rice chromosome markers were used for primary mapping, and the sequence-tagged-site (STS) markers for fine mapping were developed based on the gap difference between the contig sequences of the japonica cultivar Nipponbare and the indica cultivar 93–11 (http://ensemblgenomes.org). WFSL1 was ultimately mapped to a 50.5 kb region on chromosome 1. The predicted region was PCR-amplified and sequenced to detect the mutation.
Sequence alignment of WFSL1 protein
Using blastp program to search the protein sequence database at the NCBI with an E-value cut-off of 0.001. Using the Clustal Omega (http://www.ebi.ac.uk/Tools/msa/clustalo/) to identify motif that are conserved in the alignment sequence.
Complementation test and overexpression
A 11,296 bp genomic DNA fragment containing WFSL1 sequence, the upstream and downstream sequences was amplified by two pairs of PCR primers: 5′-ggatcccttgggtgtgccgtcgatgtgagc-3′, 5′-tatccacaacagtgaaaggatatgtggttaac-3′ and 5′-gcattgtgtcattcaggctgcgggatctaaaac-3′, 5′-ccaagcttcggtggtaggaggactcccgttgg-3′. The 11,296 bp PCR product was inserted into the binary vector pCAMBIA1300 (containing CaMV 35 S promoter) to construct the pCAMBIA1300-WFSL1 vector. This vector was then introduced into the wfsl1 mutants by Agrobacterium-mediated transformation using the Agrobacterium tumefacien EHA105. For the WFSL1 overexpression construct, a 1,743-bp fragment of full-length cDNA was amplified by two pairs of PCR primers: 5′-GGGGTACCATGAAACATCCCTCCCGCATTAAATTGGC-3′ and 5′-GCTCTAGATCAGTTGTAGGTTCTCGAAGGCTTCTG-3′ from the cDNA library of wild type and inserted into the binary overexpression vector pCAMBIA1300S (pCAMBIA1300 containing 35 S promoter) into pCAM-BIA1300S-WFSL1. The vector was transformed into wfsl1 mutants to generate overexpression plants.
Identifying the subcellular location of WFSL1
To investigate the subcellular location of WFSL1, a 1,743-bp fragment of full-length cDNA was amplified by PCR and ligated into pCAMBIA1300-GFP (containing 35 s promoter and GFP reporter protein) vector to generate the pCAMBIA1300-WFSL1-GFP construct. The PCR primers for WFSL1 were: 5′-gctctagaatgaaacatccctcccgcattaaattggc-3′ and 5′-cgggatcccggttgtaggttctcgaaggcttctg-3′. The pCAMBIA1300-WFSL1-GFP vector, as well as the control were transformed into rice protoplasts according to the protocol described previously46. The transformed rice protoplasts cells were incubated for 16 h at 28 °C under dark conditions. The GFP fluorescence in transformed protoplasts was examined by confocal fluorescence microscope (Carl Zeiss, LSM 780).
RNA extraction and qRT-PCR
Total RNA was extracted from the seedling and tillering stage of wild type and wfsl1 mutants using a Total RNA Extraction Kit (Axygene, cat No, AP-MN-MS-RNA-250) according to the manufacturer’s instructions. The complementary DNA was synthesized using a ReverTra Ace qPCR-RT Kit (TOYOBA, Japan). RT-PCR was run in Applied Biosystems 7900HT Real-time System using 2 × SYBR Green PCR Master Mix (Applied Biosystems). The RT-PCR program was as follows: initial denaturation at 95 °C for 10 min, followed by 40 cycles at 95 °C for 10 s and 60 °C for 1 min. The analysis of each sample was based on three technical replicates and biological replicates. The relative expression level of each transcript was compared with that of UBQ5 and quantified with the 2−ΔΔC T method47. The primers for the genes are listed (Supplementary Table S3).
RNA-seq analysis
Total RNA was extracted from wild type and wfsl1 at tillering stage. mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. cDNA was synthesized using random hexamer primers. The library was constructed and sequenced using an Illumina Hisequation 2000 (Novogene). A total of 45 million reads genes from wild type and 40 million from wfsl1 were obtained. The significance of differentially expressed genes (DEGs) were using log2 (fold change) > 1 and q values < 0.05. Gene ontology analysis was performed on GOseq48. Pathway enrichment analysis was using the Kyoto Encyclopedia of Genes and Genomes database49.
Western blot analysis
Total proteins extraction was performed as previously described50 isolated from wild type and wfsl1 at seedling and tillering stage. The tissues were ground in liquid nitrogen and thawed in extraction buffer [50 mM Tris–HCl pH 7.5, 150 mM NaCl, 10% glycerol (v/v), 0.1% Nonidet P-40, 1 mM DTT, 1 mM PMSF, and 1x complete protease inhibitor cocktail (Roche)] for 15 min on ice. The supernatant was collected by centrifugation at 12,000 g for 10 min at 4 °C. Total proteins were separated by SDS-PAGE gels (8%), transferred to the polyvinylidene difluoride (PVDF) membranes (GE Healthcare), blotted with different primary antibodies, detected with ECL prime (GE Healthcare). Anti-WFSL1 antibody was obtained from Shanghai Youke Biotechnology (http://www.youke-ab.cn/) and other antibodies such as anti-PsaA (LOC_Osp1g00340.1, Cat:AbP80033-A-SE), anti-AtpB (LOC_Os10g21266.1,Cat:AbP80331-A-SE), anti-RCA (LOC_Os11g47970.1, Cat:AbP80246-A-SE) and anti-RpoA (LOC_Osp1g00660.1, Cat:AbP80103-A-SE) from Beijing Protein Innovation (http://www.proteomics.org.cn/) and anti-PsbA (LOC_Osp1g00110.1, Cat:AS05084) from Agrisera.
References
Zhen, X. H. et al. Photosynthetic characteristics of flag leaves in rice white stripe mutant 6001 during senescence process. Rice Sci 21, 335–342 (2014).
Koussevitzky, S. et al. Signals from chloroplasts converge to regulate nuclear gene expression. Science 316, 715–719 (2007).
Mullet, J. E. Dynamic regulation of chloroplast transcription. Plant Physiol 103, 309–313 (1993).
Hedtke, B., Borner, T. & Weihe, A. Mitochondrial and chloroplast phage type RNA polymerases in Arabidopsis. Science 277, 809–811 (1997).
Liere, K., Weihe, A. & Börner, T. The transcription machineries of plant mitochondria and chloroplasts: composition, function, and regulation. J Plant Physiol 168, 1345–1360 (2011).
Scott, J. M. & Haldenwang, W. G. Obg, an essential GTP binding protein of Bacillus subtilis, is necessary for stress activation of transcriptionfactor sigma(B). J. Bacteriol 181, 4653–4660 (1999).
Manuell, A. L., Quispe, J. & Mayfield, S. P. Structure of the chloroplast ribosome: novel domains for translation regulation. PLoS Biol 5, e209 (2007).
Bang, W. Y. et al. Functional characterization of ObgC in ribosome biogenesis during chloroplast development. Plant J 71, 122–134 (2012).
Wang, Y. et al. WHITE PANICLE1, a Val-tRNA synthetase regulating chloroplast ribosome biogenesis in rice, is essential for early chloroplast development. Plant Physiol 170, 2110–2123 (2016).
Chi, W. et al. The function of RH22, a DEAD RNA helicase, in the biogenesis of the 50 S ribosomal subunits of Arabidopsis chloroplasts. Plant Physiol 158, 693–707 (2012).
Aluru, M. R., Yu, F., Fu, A. & Rodermel, S. Arabidopsis variegation mutants: new insights into chloroplast biogenesis. J Exp Bot 57, 1871–1881 (2006).
Chen, M., Choi, Y., Voytas, D. F. & Rodermel, S. Mutations in the Arabidopsis VAR2 locus cause leaf variegation due to the loss of a chloroplast FtsH protease. Plant J 22, 303–313 (2000).
Kusumi, K., Mizutani, A., Nishimura, M. & Iba, K. A virescent gene V1 determines the expression timing of plastid genes for transcription/translation apparatus during early leaf development in rice. Plant J 12, 1241–1250 (1997).
Kusumi, K., Yara, A., Mitsui, N., Tozawa, Y. & Iba, K. Characterization of a rice nuclear-encoded plastid RNA polymerase gene OsRpoTp. Plant Cell Physiol 45, 1194–1201 (2004).
Kusumi, K. et al. Contribution of chloroplast biogenesis to carbon-nitrogen balance during early leaf development in rice. J Plant Res 123, 617–622 (2010).
Kusumi, K. et al. A plastid protein NUS1 is essential for build-up of the genetic system for early chloroplast development under cold stress conditions. Plant J 68, 1039–1050 (2011).
Sugimoto, H. et al. The rice nuclear gene, VIRESCENT 2, is essential for chloroplast development and encodes a novel type of guanylate kinase targeted to plastids and mitochondria. Plant J 52, 512–527 (2007).
Su, N. et al. Disruption of a rice pentatricopeptide repeat protein causes a seedling-specific albino phenotype and its utilization to enhance seed purity in hybrid rice production. Plant Physiol 159, 227–238 (2012).
Wang, P. R. et al. Divinyl chlorophyll(ide) a can be converted to monovinyl chlorophyll(ide) a by a divinyl reductase in rice. Plant Physiol 153, 994–1003 (2010).
Wang, P. R. et al. One divinyl reductase reduces the 8-vinyl groups in various intermediates of chlorophyll biosynthesis in a given higher plant species, but the isozyme differs between species. Plant Physiol 161, 521–534 (2013).
Yang, Q. et al. NOA1 functions in a temperature-dependent manner to regulate chlorophyll biosynthesis and Rubisco formation in rice. Plos One 6, e20015 (2011).
Sakuraba, Y. et al. The rice faded green leaf locus encodes protochlorophyllide oxidoreductase B and is essential for chlorophyll synthesis under high light conditions. Plant J 74, 122–133 (2013).
Lee, S. et al. Differential regulation of chlorophyll a oxygenase genes in rice. Plant Mol Biol 57, 805–818 (2005).
Kusaba, M. et al. Rice NON-YELLOW COLORING1 is involved in light-harvesting complex II and grana degradation during leaf senescence. Plant Cell 19, 1362–1375 (2007).
Morita, R., Sato, Y., Masuda, Y., Nishimura, M. & Kusaba, M. Defect in non-yellow coloring 3, an alpha/beta hydrolase-fold family protein, causes a stay-green phenotype during leaf senescence in rice. Plant J 59, 940–952 (2009).
Jung, K. H. Characterization of a rice chlorophyll-deficient mutant using the T-DNA gene-trap system. Plant Cell Physiol 44, 463–472 (2003).
Jain, M., Nijhawan, A., Tyagi, A. K. & Khurana, J. P. Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem Biophy Res Co 345, 646–651 (2006).
Ren, D. et al. Regulatory role of OsMADS34 in the determination of glumes fate, grain yield and quality in rice. Front Plant Sci 7, 1853 (2016).
Chen, X. et al. SPL5, a cell death and defense-related gene, encodes a putative splicing factor 3b subunit 3 (SF3b3) in rice. Mol Breed 30, 939–949 (2012).
Ge, C. et al. Map-based cloning of a spotted-leaf mutant gene OsSL5 in Japonica rice. Plant Growth Regul 75, 595–603 (2015).
Aravind, L. & Koonin, E. V. The HD domain defines a new superfamily of metal-dependent phosphohydrolases. Trends Biochem Sci 23, 469–472 (1998).
Wootton, J. C. Non-globular domains in protein sequences: automated segmentation using complexity measures. Comput Chem 18, 269–285 (1994).
Gentry, D. R. & Cashel, M. Mutational analysis of the Escherichia coli spoT gene identifies distinct but overlapping regions involved in ppGpp synthesis and degradation. Mol Microbiol 19, 1373–1384 (1996).
Francis, S. H., Colbran, J. L., McAllister-Lucas, L. M. & Corbin, J. D. Zinc interactions and conserved motifs of the cGMP-binding cGMP-specific phosphodiesterase suggest that it is a zinc hydrolase. J Biol Chem 269, 22477–22480 (1994).
Potrykus, K. & Cashel, M. (p)ppGpp: still magical? Annu Rev Microbiol 62, 35–51 (2008).
Braeken, K., Moris, M., Daniels, R., Vanderleyden, J. & Michiels, J. New horizons for (p)ppGpp in bacterial and plant physiology. Trends Microbiol 14, 45–54 (2006).
Mizusawa, K., Masuda, S. & Ohta, H. Expression profiling of four RelA/SpoT-like proteins, homologues of bacterial stringent factors. In Arabidopsis thaliana. Planta 228, 553–562 (2008).
Tozawa, Y. et al. Calcium-activated (p)ppGpp synthetase in chloroplasts of land plants. J Biol Chem 282, 35536–35545 (2007).
Sugliani, M. et al. An ancient bacterial signaling pathway regulates chloroplast function to influence growth and development in Arabidopsis. Plant Cell 28, 661–679 (2016).
Asakura, Y., Galarneau, E., Watkins, K. P., Barkan, A. & van Wijk, K. J. Chloroplast RH3 DEAD box RNA helicases in maize and Arabidopsis function in splicing of specific group II introns and affect chloroplast ribosome biogenesis. Plant Physiol 159, 961–974 (2012).
Chateigner-Boutin, A. L. et al. CLB19, a pentatricopeptide repeat protein required for editing of rpoA and clpP chloroplast transcripts. Plant J 56, 590–602 (2008).
Chateigner-Boutin, A. L. et al. OTP70 is a pentatricopeptide repeat protein of the E subgroup involved in splicing of the plastid transcript rpoC1. Plant J 65, 532–542 (2011).
Spreitzer, R. J. & Salvucci, M. E. Rubisco: structure, regulatory interactions, and possibilities for a better enzyme. Annu Rev Plant Biol 53, 449–475 (2002).
Chen, F. et al. A nucleus-encoded chloroplast protein YL1 is lnvolved in chloroplast development and efficient biogenesis of chloroplast ATP synthase in rice. Sci Rep 6, 32295 (2016).
Arnon, D. I. Copper enzymes in isolated chloroplasts. polyphenoloxidase in beta vulgaris. Plant Physiol 24, 1–15 (1949).
Chen, S. et al. A highly efficient transient protoplast system for analyzing defence gene expression and protein-protein interactions in rice. Mol Plant Pathol 7, 417–427 (2006).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408 (2001).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11, R14 (2010).
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucleic acids res 36, D480–D484 (2008).
Rubio, V. et al. An alternative tandem affinity purification strategy applied to Arabidopsis protein complex isolation. Plant J 41, 767–778 (2005).
Acknowledgements
This work was supported by National Natural Science Foundation of China (31461143014, 31661143006, 91535205); Hangzhou Scientific and Technological Program (20170432B03). China Postdoctoral Science Foundation (2016T90155). The authors are grateful to the editors and the anonymous reviewers for their valuable comments and help.
Author information
Authors and Affiliations
Contributions
C.G. carried out the mapping, cloning and wrote the manuscript; L.W. performed the rice transformation; W.Y. and L.W. revised the manuscript; Y.C., P.C., J.P. and D.Z. investigated agronomic traits; J.H., D.Z., G.D. and Q.Q. helped perform the analysis with constructive discussions; L.G. and D.X. conceived the project and corrected the manuscript. All authors read and approve the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ge, C., Wang, L., Ye, W. et al. Single-point Mutation of an Histidine-aspartic Domain-containing Gene involving in Chloroplast Ribosome Biogenesis Leads to White Fine Stripe Leaf in Rice. Sci Rep 7, 3298 (2017). https://doi.org/10.1038/s41598-017-03327-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-03327-2
This article is cited by
-
Characterization and fine mapping of a white stripe leaf mutant in rice
Genetic Resources and Crop Evolution (2024)
-
A natural variation in the promoter of GRA117 affects carbon assimilation in rice
Planta (2023)
-
Disruption of a Rice Chloroplast-Targeted Gene OsHMBPP Causes a Seedling-Lethal Albino Phenotype
Rice (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.