Abstract
Hedgehog (Hh) signaling plays a pivotal role in animal development and its deregulation in humans causes birth defects and several types of cancer. Protein Kinase A (PKA) modulates Hh signaling activity through phosphorylating the transcription factor Cubitus interruptus (Ci) and G protein coupled receptor (GPCR) family protein Smoothened (Smo) in Drosophila, but how PKA activity is regulated remains elusive. Here, we identify a novel regulator of the Hh pathway, the capping-enzyme mRNA-cap, which positively regulates Hh signaling activity through modulating PKA activity. We provide genetic and biochemical evidence that mRNA-cap inhibits PKA kinase activity to promote Hh signaling. Interestingly, regulation of Hh signaling by mRNA-cap depends on its cytoplasmic capping-enzyme activity. In addition, we show that the mammalian homolog of mRNA-cap, RNGTT, can replace mRNA-cap to play the same function in the Drosophila Hh pathway and that knockdown of Rngtt in cultured mammalian cells compromised Shh pathway activity, suggesting that RNGTT is functionally conserved. Our study makes an unexpected link between the mRNA capping machinery and the Hh signaling pathway, unveils a new facet of Hh signaling regulation, and reveals a potential drug target for modulating Hh signaling activity.
Similar content being viewed by others
Introduction
The Hedgehog (Hh) signaling pathway is essential for embryonic development and adult tissue homeostasis in species ranging from insects to mammals1,2,3. Dysregulation of Hh signaling has been implicated in a large number of human disorders, including birth defects and a wide range of cancers4,5,6,7.
Hh signal transduction is best studied in Drosophila. In the absence of Hh, the 12-transmembrane protein Patched (Ptc) inhibits the function of the 7-transmembrane GPCR family protein Smoothened (Smo), which transduces the Hh signal across the plasma membrane to intracellular signaling components8,9,10,11,12,13,14. When Hh binds Ptc, the inhibition on Smo is relieved. Activated Smo recruits the kinesin-like protein Costal2 (Cos2) and the Ser/Thr kinase Fused (Fu) to trigger the activation of latent transcription factor Cubitus interruptus (Ci)15,16,17,18,19,20,21,22, which enters the nucleus to turn on the expression of Hh target genes including decapentaplegic (dpp), patched (ptc) and engrailed (en) 1, 23, 24. The expression of different responsive genes is dependent on the Hh dosage. Low levels of Hh are able to induce the expression of dpp, intermediate to high levels of Hh are required to activate ptc, and peak levels of Hh are required to activate en 25.
In Drosophila wing discs, when Hh levels are very low or absent in anterior (A) compartment cells far away from the anteroposterior (A/P) compartment boundary, the full-length Ci (CiF) undergoes sequential phosphorylation by PKA, GSK3 and casein kinase 1 (CK1) at multiple Ser/Thr residues, which targets CiF for SCFSlimb-mediated proteolytic processing to generate a truncated repressor form (CiR)26,27,28,29,30,31. CiR enters the nucleus to block the expression of Hh target genes, including dpp 23, 24, 28, 32,33,34,35. Among all the Ci kinases, PKA plays a primary role because its phosphorylation of Ci provides priming sites for GSK3- and CK1-mediated phosphorylation23, 27, 33, 36.
In A-cells near the A/P boundary where Hh is present, Hh induces Smo activation by promoting phosphorylation of its C-terminal tail by PKA and CK1, which increases Smo cell-surface expression and converts Smo from a closed, inactive conformation to an open and active one14, 37,38,39. Activated Smo inhibits Ci processing and converts CiF into an active form (CiA) that activates the Hh target genes including ptc and en 40, 41. Hence, PKA plays a dual role in the Hh pathway by activating Smo but inhibiting Ci depending on the availability of the Hh ligand. Despite its dual role in Hh signaling, loss of PKA results in Hh pathway activation instead of inhibition due to the depression of Ci: loss of CiR and accumulation of CiF, some of which is converted into CiA 24, 42.
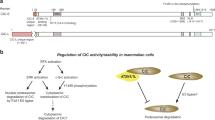
Drosophila mRNA-cap is a capping enzyme that harbors a RNA 5′-triphosphatase and mRNA 3′-guanylyltransferase domains and catalyzes the attachment of the 5′ cap to messenger RNA molecules in the nucleus during the first stage of gene expression43,44,45,46,47,48. Cap removal is considered to be a prerequisite step in the mRNA degradation pathway49, but recent findings indicate that certain portion of translational inactive mRNAs might be stored in an uncapped state and subsequently returned to an active state upon cytoplasmic re-capping50. In this case, capping enzyme cooperates with a novel kinase to generate capped ends from cleaved RNAs in cytoplasm, which is named cytoplasmic capping. Except for its capping function, other functions of mRNA-cap are elusive. In this study, we find unexpectedly that mRNA-cap regulates Hh signaling activity depending on its capping-enzyme activity in the cytoplasm. Furthermore, we find that it regulates Hh signaling through antagonizing PKA.
Results
Downregulation of mRNA-cap disrupts wing development
Given that Hh pathway plays an important role in wing development and its deregulation causes wing development defect, we sought to identify novel regulators of Hh signaling via analyzing wing phenotypes caused by either loss or gain of function of such regulators. To do that, we performed in vivo RNAi screen by silencing ~7000 genes that are conserved between Drosophila and mammals using a wing specific Gal4 driver MS1096-Gal4 and determined which genes modulated wing development. We picked up the candidate genes whose knockdown caused abnormal wing phenotypes, and further investigated whether they modulated Hh signaling activity by examining the pathway target gene ptc-lacZ via immunostaining. In our screen, we found that knockdown of mRNA-cap through its RNAi line (VDRC 108809) disrupted wing development, causing deformed wing phenotype (Fig. 1A,B). To verify the efficiency of this RNAi line and rule out any off-target effect, we made UAS-HA-mRNA-cap transgenic flies. Through co-expressing the transgene and the RNAi line, we found that mRNA-cap transgenic expression could rescue the RNAi-induced abnormal wing phenotype (Fig. 1C), suggesting that the aberrant wing phenotype is indeed caused by loss of mRNA-cap. We also employed another mRNA-cap RNAi line (Bloomington 32847), which is a relatively weaker line compared with 108809, and observed a similar wing defect (Supplementary Fig. 1A), and this defect was also rescued by overexpressing the mRNA-cap transgene (Supplementary Fig. 1B). Taken together, these results indicate that mRNA-cap is essential for normal wing formation in Drosophila.
Knockdown of mRNA-cap represses Hh signaling activity
Since mRNA-cap knockdown disrupts the wing morphology and many signaling pathways including Hh, Wingless, Notch and Hippo pathways are involved in the regulation of the wing development, we tested whether mRNA-cap knockdown affects these pathways. We found that Wingless, Notch and Hippo signaling pathway activities exhibited little if any changes when mRNA-cap was knocked down, as monitored by their cognate read-outs Dll-lacZ (Fig. 2A,B), Cut (Fig. 2C,D) and Ex-lacZ (Fig. 2E,F) respectively. However, we found that mRNA-cap RNAi downregulated the expression of Hh pathway target genes including dpp-lacZ, ptc-lacZ and En (Fig. 2H,J and L compared with Fig. 2G’,I’ and K’), suggesting that mRNA-cap selectively modulates Hh signaling activity.
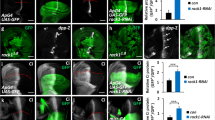
Knockdown of mRNA-cap decreases Ci protein level and downregulates Hh target genes expression. All wing imaginal discs shown in this study were oriented with anterior compartments on the left and ventral regions on the top. (A–B’) Knockdown of mRNA-cap with En-Gal4 (B,B’), which drives UAS target gene expression in the P-compartment, did not decrease the Dll-lacZ level compared with control (A,A’). (C–D’) The expression of Cut was not decreased compared with control (C,C’) upon knockdown of mRNA-cap in the P-compartment with Hh-Gal4 (D,D’). (E–F’) Knockdown of mRNA-cap with Hh-Gal4 (F,F’) did not change the expression of Ex-lacZ compared with control (E,E’). (G–M) Knockdown of mRNA-cap with MS1096 attenuated the expression of dpp-lacZ (compare G’ with H), ptc-lacZ (compare I’ with J), En (compare K’ with L) and Ci (compare M with G). Arrows indicate the decrease of dpp-lacZ, ptc-lacZ, En and Ci. (N–O”) mRNA-cap A clones with low (N–N”) and high (O–O”) magnifications of wing discs. The wing discs were immunostained to show the expression of GFP (green) and Ci (white). mRNA-cap A clones were recognized by the lack of GFP (O, arrows). Images in (O–O”) are enlarged views of the region marked by dashed lines in N’. Of note, mRNA-cap mutant cells exhibited decreased Ci level (O’, arrows). (P,P’) Wing discs expressing mRNA-cap RNAi and HA-mRNA-cap with MS1096 were immunostained to show the expression of Ci and ptc-lacZ. The reduced expression of Ci and ptc-lacZ were rescued when co-expression of HA-mRNA-cap (compare P,P’ with M and J). All images are representative of three independent experiments.
In order to future test the function of mRNA-cap in the Hh pathway, we detected the Ci and found that the level of CiF was downregulated by mRNA-cap RNAi (Fig. 2M). We obtained similar results using an independent RNAi line (32847) targeting mRNA-cap (Supplementary Fig. 1C–C”’). To confirm the function of mRNA-cap in the Hh pathway, we employed the FLP-FRT mitotic recombination technique to generate homozygous clones for an mRNA-cap mutant, mRNA-cap A. Consistent with the observation that mRNA-cap RNAi downregulated CiF level, mRNA-cap A clones, which are marked by lack of GFP expression (Fig. 2N,O), showed decreased CiF level (Fig. 2N’,O’). Furthermore, co-expression of HA-mRNA-cap with mRNA-cap RNAi rescued the expression of CiF and ptc-lacZ (Fig. 2P,P’). These observations indicate that mRNA-cap exerts a positive influence on Hh pathway activity.
Knockdown of mRNA-cap downregulates Ci but upregulates Smo
From the above results, we found that knockdown of mRNA-cap led to a decrease of CiF level, which is consistent with the observed downregulation of Hh pathway target genes expression. However, when we stained the wing discs with a Smo antibody, we found unexpectedly that knockdown of mRNA-cap resulted in Smo accumulation in A cells distant from the A/P boundary (Fig. 3B compared with Fig. 3A) even though Ci was downregulated (Fig. 3A’,B’), suggesting that mRNA-cap may regulate both Smo and Ci level but in the opposite direction.
Knockdown of mRNA-cap upregulates Smo while downregulates Ci levels. (A–A”) Control wing discs carrying MS1096 Gal4 driver were stained to show endogenous Smo (green) and full-length Ci (red). (B–B”) Wing discs expressing mRNA-cap RNAi with MS1096 were immunostained with Smo (green) and Ci (red) antibodies. Smo was accumulated in A and P compartments when mRNA-cap was knocked down. (C–C”) Wing discs expressing Myc-Smo with MS1096 were immunostained to show the expression of Myc, Ci and ptc-lacZ. (D–D”) Wing discs expressing Myc-Smo and mRNA-cap RNAi with MS1096 were stained to show the expression of Myc, Ci and ptc-lacZ. Ci and ptc-lacZ were also decreased when overexpressing Smo and mRNA-cap RNAi. (E–E”) Wing discs expressing Myc-SmoSD with MS1096 were immunostained to show the expression of Myc, Ci and ptc-lacZ. (F–F”) Wing discs expressing Myc-SmoSD with mRNA-cap RNAi by MS1096 were stained for Myc, Ci and ptc-lacZ. Ci and ptc-lacZ were also decreased when overexpressing activated Smo and mRNA-cap RNAi. (G,G’,H,H’) Wing discs expressing mRNA-cap RNAi and ci-lacZ with MS1096 were immunostained to show the expression of ci-lacZ. The expression of ci-lacZ alone was used as a control. (I) The relative mRNA levels of ci and smo of wing discs measured by RT-qPCR in mRNA-cap knockdown background. MS1096 wing discs were used as a control. All images are representative of three independent experiments. Data presented are the average of three independent experiments and error bars represent SD. ***P < 0.001, t-test.
To further confirm this notion, we co-overexpressed mRNA-cap RNAi with UAS-SmoWT or UAS-SmoSD, which is a constitutively active form of Smo with PKA sites mutated to amino acids residues Asp, and found that knockdown of mRNA-cap downregulated Ci (Fig. 3 compare C’,E’ to D’,F’) and ptc-lacZ (Fig. 3 compare C’’,E’’ to D’’,F’’) even in the presence of Smo overexpression, indicating that mRNA-cap RNAi can block the Hh signal transduction at a step downstream of Smo.
To distinguish whether Ci downregulation by mRNA-cap RNAi occurs at mRNA or protein level, we examined the transcript level of ci using the ci-lacZ reporter that expresses lacZ under the control of ci promotor. We didn’t find any difference in ci-lacZ expression between the control and mRNA-cap knockdown wing discs (Fig. 3G–H’), indicating that ci transcription is not regulated by mRNA-cap. To exclude that mRNA-cap affects Ci mRNA stability, we measured ci mRNA level by RT-qPCR in wing discs and found that mRNA-cap RNAi didn’t affect ci mRNA level (Fig. 3I). Meanwhile we found that mRNA-cap did not affect the smo mRNA level (Fig. 3I). Taken together, the RT-qPCR results imply that mRNA-cap affects Ci and Smo at protein level instead of mRNA level.
mRNA-cap regulates Hh pathway activity through PKA
Knockdown of mRNA-cap causes accumulation of Smo but downregulation of Ci and Hh target genes expression, which is reminiscent of the phenotype caused by excessive PKA activity on Hh signaling51, 52. As shown in Fig. 4A–B’, overexpression of mC*, a constitutively active form of mammalian PKA catalytic subunit, resulted in upregulation of Smo but downregulation of Ci and ptc-lacZ expression. We obtained similar results when overexpressing the Drosophila PKA catalytic subunit 1 (PKA-C1) (Fig. 4C–D’).
mRNA-cap regulates Hh signaling activity through PKA. (A–B’) A wing disc expressing mC* with MS1096 was immunostained to show PKAα, Ci, ptc-lacZ and Smo expression. Ci and ptc-lacZ levels were downregulated by mC*. However, Smo level was increased. PKAα antibody was used to show mC* expression. (C–D’) A wing disc expressing Fg-PKA-C1 with MS1096 was immunostained to show Fg, Ci, ptc-lacZ and Smo. Ci and ptc-lacZ levels were downregulated by expression of Fg-PKA-C1. But Smo level was up-regulated. (E–F”) Wing discs expressing PKA-C1 RNAi alone (E–E”) or together with mRNA-cap RNAi (F–F”) with MS1096 were stained for Smo and Ci. The accumulation of Smo and the attenuation of Ci expression caused by mRNA-cap knockdown were blocked by PKA-C1 RNAi. (G–H”) Wing discs expressing Fg-PKA-C1 K75A alone or together with mRNA-cap RNAi using MS1096 were immunostained to show Ci, ptc-lacZ and Smo expression. (I–J”) Wing discs expressing HA-Ci alone or together with mRNA-cap RNAi with MS1096 were immunostained to show Ci, ptc-lacZ and Smo expression. Ci level was unstable upon mRNA-cap knockdown. (K–L’) Wing discs expressing HA-Ci −3P alone or together with mRNA-cap RNAi using MS1096 were immunostained to show Ci and ptc-lacZ expression. The Ci−3P level was stable even mRNA-cap was knocked down. (M,N) Wing discs expressing slimb RNAi alone or together with mRNA-cap RNAi using MS1096 were immunostained to show Ci expression. All images are representative of three independent experiments.
To test the genetic interaction between mRNA-cap and PKA, we expressed mRNA-cap RNAi and PKA-C1 RNAi (VDRC 101524) transgenes individually or in combination. As shown in Fig. 4, we observed that knockdown of PKA-C1 either alone or in conjunction with mRNA-cap resulted in the accumulation of full-length Ci in anterior compartment and prevented the up-regulation of Smo caused by mRNA-cap RNAi (Fig. 4E–F”). Furthermore, we employed a dominant negative form of PKA-C1 (PKA-C1 K75A) with K75 mutated to alanine (A) and obtained results similar to PKA-C1 RNAi, i.e., Ci and ptc-lacZ level were high in anterior compartment but Smo level had no obvious change when Fg-PKA-C1 K75A was expressed alone (Fig. 4G–G”), and simultaneous overexpression of Fg-PKA-C1 K75A and mRNA-cap RNAi prevented the up-regulation of Smo caused by mRNA-cap RNAi but resulted in upregulation of Ci similar to expressing Fg-PKA-C1 K75A alone (Fig. 4H–H”). This epistasis analysis indicates that mRNA-cap plays the role on upstream of PKA.
To further test this idea, we overexpressed CiWT or Ci−3p (a constitutively active form of Ci with three PKA sites mutated to Ala) with mRNA-cap RNAi and found that mutating the PKA sites in Ci renders it resistant to mRNA-cap RNAi-mediated downregulation (Fig. 4I–L’). Phosphorylation of Ci by PKA targets it for Slimb-mediated degradation24, 53. We found that knockdown of slimb either alone or together with mRNA-cap upregulated Ci (Fig. 4M,N). Taken together, these results suggest that mRNA-cap regulates Smo and Ci through PKA.
mRNA-cap inhibits the PKA activity in wing discs
The above genetic evidence suggests that mRNA-cap regulates Hh signaling activity through PKA. Next, we tested whether PKA activity is altered upon knockdown of mRNA-cap. We found that the PKA activity was increased by mRNA-cap RNAi in wing discs (Fig. 5A). As PKA activity is regulated by cAMP, we further tested whether mRNA-cap regulated PKA activity through cAMP and found that cAMP level was not changed when mRNA-cap was knocked down (Fig. 5B).
mRNA-cap regulates PKA activity. (A,B) The PKA activity (A) and cAMP level (B) were measured from control wing discs or wing discs expressing mRNA-cap RNAi with MS1096. Approximately 400 discs were dissected and lysed for each assay. (C–C”’) Overexpression of HA-PKA-C2 by MS1096 was immunostained for Ci, ptc-lacZ and HA. (D–D”’) Overexpression of HA-PKA-C3 by MS1096 was immunostained for Ci, ptc-lacZ and HA. The levels of Ci were decreased. (E–F”’) Overexpression of HA-PKA-R1 (E–E”’) or Myc-PKA-R2 (F–F”’) with MS1096 was immunostained for Ci, ptc-lacZ and HA or Myc. The levels of Ci and ptc-lacZ were up-regulated. (G) The relative mRNA levels of PKA-C1, PKA-C3, PKA-R1 and PKA-R2 of wing discs were measured by RT-qPCR upon knockdown of mRNA-cap. MS1096 acted as a control. (H) Western blot analysis of lysates from control wing discs or wing discs expressing mRNA-cap RNAi with MS1096. Approximately 100 discs were dissected, lysed, and blotted with anti-PKA-C1 antibody, respectively. (I) Western blot analysis of PKA-R1 levels from control wing discs or wing discs expressing mRNA-cap RNAi. Approximately 100 discs were used. (J,K) Knockdown of mRNA-cap did not affect the dimer formation of PKA-R1 (J) or PKA-C1 (K). (L,M) mRNA-cap didn’t change the interaction between PKA-C1 and PKA-R1. All images are representative of three independent experiments. (A,B,G,H and I) Data presented are the average of three independent experiments and error bars represent SD. ***P < 0.001, t-test.
In Drosophila, there are two PKA regulatory subunits: PKA-R1, R2 and three PKA catalytic subunits: PKA-C1, C2 and C3. We generated transgenic flies expressing these PKA subunits under the control of UAS promoter. Except for PKA-C2, over-expression of all other transgenes had the expected effects on Hh signaling; however, the effect of PKA-C3 overexpression on Hh signaling was relatively weak compared with PKA-C1 (Fig. 5C–F”’). We further examined the endogenous mRNA levels of these PKA subunits upon knockdown of mRNA-cap, and found no obvious changes despite of efficient knockdown of mRNA-cap (Fig. 5G).
As PKA-C1 and PKA-R1 are predominantly expressed in wing discs compared with other PKA subunits (Fig. 5G), their protein levels were further investigated. Comparing the MS1096 > mRNA-cap RNAi with control wing discs, we found that neither PKA-C1 nor PKA-R1 exhibited significant changes in their protein levels (Fig. 5H,I). Taken together, these results suggest that mRNA-cap regulates PKA activity independent of cAMP and the protein levels of PKA regulatory and catalytic subunits.
We next examined whether mRNA-cap affects the interaction between the regulatory and catalytic subunits and found that mRNA-cap didn’t affect R1/R1, C1/C1, C1/R1 dimer formation (Fig. 5J–M; Supplementary Fig. 2A–C). All these results indicate that mRNA-cap may regulate PKA activity through a non-canonical mechanism.
mRNA-cap regulates Hh signaling through its enzyme activity
To test which domain of mRNA-cap is essential for its regulation of Hh signaling, we generated two transgenes: HA-mRNA-cap-N (aa1-272) and HA-mRNA-cap-C (aa248-616), which harbor the N-terminal 5′-triphosphatase domain and the C-terminal guanylyltransferase domain, respectively. We co-expressed mRNA-cap RNAi with HA-mRNA-cap, HA-mRNA-cap-N or HA-mRNA-cap-C and found that only HA-mRNA-cap rescued the adult wing phenotype (Fig. 6B”’) as well as the expression of Ci, ptc-lacZ and Smo to wild type levels (Fig. 6B–B”), whereas neither HA-mRNA-cap-N nor HA-mRNA-cap-C could rescue the RNAi phenotypes (Fig. 6C–D”’), indicating that both N- and C-terminus of mRNA-cap are essential for its function in the Hh signaling pathway.
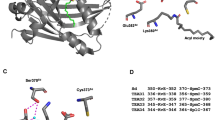
Regulation of Hh signaling by mRNA-cap depends on its enzymatic activity. (A) Schematic drawings show the domains and motifs in wild type mRNA-cap and its mutant forms used in transgenic flies and immunofluorescence assays. White bars denoted the RNA 5′-triphosphatase and 3′-guanylyltransferase domains. (B–B”’) Wing discs expressing HA-mRNA-cap and mRNA-cap RNAi were stained for Ci, ptc-lacZ and Smo. The attenuation of Ci and ptc-lacZ caused by mRNA-cap knockdown was restored by the expression of HA-mRNA-cap. Meanwhile, the accumulation of Smo was suppressed. Adult wing phenotype was also rescued by the expression of HA-mRNA-cap. (C–D”’) Wing discs expressing HA-mRNA-cap-N or HA-mRNA-cap-C with mRNA-cap RNAi were stained for Ci, ptc-lacZ and Smo. Neither the defects in Ci, ptc-lacZ and Smo expression nor the adult wing phenotypes were rescued. (E–F”’) Wing discs expressing HA-mRNA-cap MP or HA-mRNA-cap MG with mRNA-cap RNAi were stained for Ci, ptc-lacZ and Smo. The Ci, ptc-lacZ, Smo and adult wing phenotypes could not be restored. (G–G”’) Wing discs expressing HA-mRNA-cap R569K with mRNA-cap RNAi were stained for Ci, ptc-lacZ and Smo. Ci, ptc-lacZ, Smo and adult wing phenotype also could not be rescued. (H–M) The expression levels of HA-mRNA-cap (H), HA-mRNA-cap-N (I), HA-mRNA-cap-C (J), HA-mRNA-cap MP (K), HA-mRNA-cap MG (L) and HA-mRNA-cap R569K (M) were comparable in the wing discs expressing these transgenes with MS1096. All images are representative of three independent experiments.
We next generated transgenic flies expressing two mutant forms of mRNA-cap, mRNA-cap MP and mRNA-cap MG, which mutated the essential residues in the triphosphatase and guanylyltransferase domains, respectively (Fig. 6A)48. We found that neither mRNA-cap MP nor mRNA-cap MG could rescue the abnormal wing phenotypes as well as the abnormal expression of Ci, ptc-lacZ and Smo caused by mRNA-cap RNAi (Fig. 6E–F”’). These results suggest that modulation of Hh signaling by mRNA-cap may depend on its capping enzyme activity. This notion was further supported by the fact that a single amino acid substitution in mRNA-cap, mRNA-cap R569K, which affects its interaction with poly (U) and disrupts the capping enzyme activity in mammalian48, affected its ability to rescue Hh signaling defects caused by mRNA-cap RNAi (Fig. 6G–M). Taken together, these results suggest that mRNA-cap regulates Hh signaling activity depending on its capping enzyme activity.
To define where mRNA-cap exerts its influence on Hh signaling, we transfected HA-mRNA-cap in S2 cells and found that it is located in both cytoplasm and nucleus (Fig. 7A–A”). Treating the cells with the nuclear export inhibitor LMB resulted in an exclusive nuclear localization of HA-mRNA-cap (Fig. 7B–B”), suggesting that mRNA-cap shuttles between the nucleus and cytoplasm and may modulate PKA activity in the cytoplasm, nucleus or both.
Cytoplasmic mRNA-cap regulates the Hh pathway. (A–A”) S2 cells expressing HA-mRNA-cap were stained to show DAPI and HA. (B–B”) LMB treatment resulted in nuclear localization of HA-mRNA-cap in S2 cells. (C–D”) S2 cells and wing discs expressing HA-NES-RNGTT△NLS were stained for DAPI and HA. (E–E””) Wing discs expressing HA-NES-RNGTT △NLS with mRNA-cap RNAi were stained for HA, Ci, ptc-lacZ and Smo. The detects in Ci, ptc-lacZ and Smo levels as well as adult wing phenotype were rescued. (F–G”) S2 cells and wing discs expressing HA-NES-RNGTT△NLSK294A were stained for DAPI and HA. (H–H””) Wing discs expressing HA-NES-RNGTT△NLS K294A under knockdown of mRNA-cap background were stained for HA, Ci, ptc-lacZ and Smo. The defects in Ci, ptc-lacZ, Smo levels and adult wing phenotype could not be rescued. All images are representative of three independent experiments.
To determine where mRNA-cap acts, we overexpressed an mRNA-cap variant, HA-NES-RNGTT△NLS, which is exclusively localized in the cytoplasm both in the S2 cells and in the wing discs (Fig. 7C–D”), and found that it could rescue the Ci, ptc-lacZ, Smo levels (Fig. 7E–E”’) as well as adult wing phenotype in mRNA-cap RNAi background (Fig. 7E””). On the other hand, a capping enzyme-dead cytoplasmic form of mRNA-cap, HA-NES-RNGTT△NLSK294A (Fig. 7F–G”) failed to rescue the Hh pathway defects caused by mRNA-cap RNAi (Fig. 7H–H””). These results suggest that mRNA-cap can regulate Hh signaling in the cytoplasm through its capping enzyme activity.
Mammalian mRNA-cap is functionally conserved
Our above results show mRNA-cap regulates Hh signaling activity. To test whether mammalian mRNA-cap is functionally conserved throughout evolution, we sub-cloned RNGTT, the homolog of mRNA-cap in humans, into the UAS vector and generated transgenic flies carrying UAS-RNGTT. Co-expression of UAS-RNGTT with the mRNA-cap RNAi transgene rescued the wing phenotype and restored the normal levels of Ci, ptc-lacZ and Smo (Fig. 8A–A”’), suggesting that RNGTT could substitute Drosophila mRNA-cap to regulate Hh signaling, indicating that mRNA-cap is functionally conserved between Drosophila and mammals. Furthermore, mutating the essential residues in the 5′-triphosphatase (MP) or the 3′-guanylyltransferase (MG) domain of human RNGTT or a single point mutation (K294A) in RNGTT, which eliminated its guanylylation function54, all prevented it from rescuing the Hh signaling defects and adult wing phenotypes caused by mRNA-cap RNAi (Fig. 8B–D”’). Taken together, the results suggest that RNGTT is functionally conserved during evolution, and it can substitute mRNA-cap to modulate Hh singling depending on the capping enzyme activity.
A conserved function of human mRNA-cap in Hh pathway regulation. (A–A”) A wing disc expressing HA-h-RNGTT and mRNA-cap RNAi with MS1096 was immunostained for Ci, ptc-lacZ and Smo. Defects in Ci, ptc-lacZ and Smo level were rescued by HA-h-RNGTT overexpression. (A”’,B”’,C”’,D”’) Adult wings expressing HA-h-RNGTT (A”’), HA-h-RNGTT MP (B”’), HA-h-RNGTT MG (C”’) or HA-h-RNGTT K294A (D”’) in mRNA-cap knockdown background. Only expression of HA-h-RNGTT could rescue the adult wing defect induced by mRNA-cap RNAi. (B–D”) Wing discs expressing mRNA-cap RNAi in conjunction with HA-h-RNGTT MP (B–B”) or HA-h-RNGTT MG (C–C”) or HA-h-RNGTT K294A (D–D”) were immunostained for Ci, ptc-lacZ and Smo. The defects in Ci, ptc-lacZ and Smo levels were not rescued by the expression of HA-h-RNGTT MP, HA-h-RNGTT MG or HA-h-RNGTT K294A. (E–H) Gli-luciferase (Gli-luc) reporter assays in NIH/3T3 cells transfected with the indicated constructs in the absence (E,F) or presence (G,H) of Shh treatment. Gli luciferase activities were normalized to Renilla luciferase activities. RNGTT increased whereas RNGTTK294A or knockdown of RNGTT decreased Shh pathway activity. Co-transfection with PRKAR1A restored the pathway activity. The knockdown efficiency of RNGTT was assessed through PCR. Actin acts as a loading control. All images are representative of three independent experiments. Data presented are the average of three independent experiments and error bars represent SD. *P < 0.05, **P < 0.01, ***P < 0.001, t-test.
To further confirm the conservation of mRNA-cap function in the Hh signaling pathway, we performed the Gli-luciferase reporter assay in NIH/3T3 cells, which was regulated by Sonic hedgehog (Shh) pathway components. Results from the reporter assay showed that overexpression of RNGTT promoted Gli-luciferase activity whereas RNGTTK294A decreased the Hh pathway activity (Fig. 8E). Consistently, knockdown of endogenous RNGTT suppressed Gli-luciferase activity (Fig. 8F), indicating that RNGTT is a functionally conserved positive regulator in the Shh pathway. Meanwhile, we found that expression of the mammalian homolog of PKA-R1, PRKAR1A, increased the expression of Gli-luciferase reporter gene in both control and RNGTT knockdown cells (Fig. 8F). We obtained the similar results when cells were treated with ShhN (Fig 8G,H). Taken together, these results support that RNGTT regulates Shh pathway activity through PKA.
Discussion
Hh signaling plays essential roles in embryonic development and adult tissue homeostasis, and its misregulation causes numerous human diseases including birth defects and cancers. PKA modulates Hh signaling activity through phosphorylation of the transcription factor Ci/Gli and the GPCR family protein and Hh signal transducer Smo. Besides the regulation of PKA-substrate interaction55 and possible involvement of Gαi56, how PKA activity is regulated remains poorly understood. In this study, we identify a novel regulator of the Hh pathway, the capping-enzyme mRNA-cap, which positively regulates Hh signaling activity through modulating PKA activity (Fig. 9). Interestingly, we demonstrate that the regulation of Hh signaling by mRNA-cap can be achieved by its cytoplasmic capping-enzyme activity. Finally, we show that the mammalian homolog of mRNA-cap, RNGTT, plays a conserved role in the regulation of Hh signaling. Interestingly, it has been reported recently that mRNA capping machinery can regulate specific gene expression, and is involved in the maintenance of pluripotency and differentiation of embryonic stem cell (ESC), the regulation of G1 Phase transcripts, and the regulation of the Wnt/β-catenin signaling activity57,58,59. Here, our study links the mRNA capping machinery to another important developmental signaling pathway, Hh signaling pathway, unveils a new facet of Hh signaling regulation and uncovers a potential new target for modulating Hh signaling activity in the treatment of cancer.
As shown in Flybase, Drosophila has only one capping enzyme encoded by mRNA-cap, which harbors an RNA 5′-triphosphatase domain and an mRNA 3′-guanylyltransferase domain and catalyzes the attachment of the 5′ cap to messenger RNA molecules in the nucleus during the first stage of gene expression. Ubiquitous knockdown of mRNA-cap causes fly lethality. In addition, in situ hybridization assay reveals that its expression is ubiquitous in Drosophila wing discs (Supplementary Fig. 1D,D’). These observations suggest that mRNA-cap is an essential gene for fly development. Surprisingly, knockdown of mRNA-cap can selectively affect Hh signaling without affecting several other signaling pathways we have tested. Our results indicate that mRNA-cap shuttles between the cytoplasm and the nucleus, implying that mRNA-cap may also play a role in the cytoplasm in addition to its well-characterized function in nucleus. As mentioned before, mRNA-cap can recap a selective fraction of translationally inactive mRNAs to return them to an active state through cytoplasmic re-capping mechanism50. In this case, there may be only a small number of genes that are controlled by mRNA-cap through cytoplasmic capping function, one of which could be involved in regulating PKA and Hh signaling. As shown in Fig. 7 and Supplementary Fig. 3A,B, the results in both Drosophila and the NIH/3T3 cells support this idea.
How mRNA-cap affects PKA activity is not resolved and remains an important issue to address by future study. The PKA holoenzyme consists of two catalytic (C) subunits and two regulatory (R) subunits, cAMP binding to R subunit and the subsequent dissociation between R and C subunits are needed for PKA activity. Our results indicate that knockdown of mRNA-cap upregulated PKA activity, but did not affect cAMP level, PKA-R1, R2, C1, C3 mRNA levels and R1, C1 protein levels. Neither did it affect PKA-R1/R1, C1/C1 and R1/C1 interaction. In addition, the regulation of Hh signaling by mRNA-cap is dependent on its cytoplasmic capping-enzyme activity, and knockdown of the mRNA cap binding protein, Cbp80, showed similar Hh signaling defects (Supplementary Fig. 4A–C), indicating that the mRNA capping machinery is involved in the regulation of Hh signaling activity. Since PKA can directly bind Smo and Cos236, 55, we also carried out experiments to exclude the possibility that mRNA-cap regulates the interactions of Cos2-PKA and Smo-PKA (Supplementary Fig. 5A,B). Taken together, our results suggest that mRNA-cap may regulate an unidentified factor that is involved in the inhibition of PKA activity through a non-canonical mechanism.
Importantly, our results indicate that mRNA-cap function is conserved during evolution. Given that PKA plays essential roles in developmental and adult homeostasis as well as a variety of physiological conditions, our study suggests an alternative way to modulate PKA activity and indicates that mRNA-cap and its relevant target genes may be the potential drug targets for the treatment of Hh- or PKA-related diseases.
Materials and Methods
Constructs, Mutants and Transgenes
The constructs for S2 cell transfection experiments are as follows: Myc/Fg/HA-mRNA-cap, Fg/HA-PKA-C1, Myc/HA-PKA-R1, Myc-Cos2 and Myc-Smo, the corresponding cDNA fragments of which were amplified, and cloned into the pUAST vectors. Similarly, the constructs of Myc-RNGTT, Myc-RNGTTK294A, Myc-PRKAR1A, HA-NES-RNGTT△NLS and HA-NES-RNGTT△NLSK294A used in mammal cells were cloned into the pcDNA3.1 vectors. The mutants of mRNA-cap/RNGTT, including HA-mRNA-capMP, HA-mRNA-capMG, HA-mRNA-cap-N (aa1-272), HA-mRNA-cap-C (aa248-616), HA-mRNA-capR569K, HA-RNGTTMP, HA-RNGTTMG and HA-RNGTTK294A were constructed in the backbones of mRNA-cap or RNAGTT through the deletion of the corresponding regions or PCR-based site-directed mutagenesis. We also added the nuclear export signal (NES) LQLPPLERLTLD and deleted the nuclear localization signal (NLS) KRKH to generate the HA-NES-RNGTT△NLS and HA-NES-RNGTT△NLSK294A plasmids in the background of RNGTT and RNGTTK294A. The RNAi lines that targeted mRNA-cap (v108809, BL32847), PKA-C1 (v101524), Cbp80 (7035R-3) and slimb (3412R-1) were obtained from the Vienna Drosophila RNAi Center (VDRC), the Bloomington Stock Center and National Institute of Genetics (NIG), respectively. Flies of MS1096, C765-Smo -PKA12, Hh-Gal4, En-Gal4, HA-Ci, HA-Ci −3P, Myc-Smo, Myc-SmoSD, ptc-lacZ, dpp-lacZ, Dll-lacZ, Ex-lacZ, ci-lacZ, mC*, Fg-PKA-C1 (BL35554) and Fg-PKA-C K75A (35559) have been described (Flybase)60. mRNA-cap A is an mRNA-cap mutant allele, whose tyrosine (Y) at 346 is replaced by alanine (A) (Flybase). The transgenic flies of HA-mRNA-cap, HA-mRNA-capMP, HA-mRNA-capMG, HA-mRNA-cap-N, HA-mRNA-cap-C, HA-mRNA-capR569K, HA-RNGTT, HA-RNGTTMP, HA-RNGTTMG, HA-RNGTTK294A, HA-PKA-C2, HA-PKA-C3, HA-PKA-R1, Myc-PKA-R2, HA-NES-RNGTT△NLS and HA-NES-RNGTT△NLSK294A were generated by injection of corresponding constructs into Drosophila embryos according to the methods described previously61. The parental strain for all germline transformations was w 1118. All stocks used in this study were maintained and raised under standard conditions.
Immunostaining and in situ hybridization
Immunostaining and in situ hybridization of imaginal discs were performed according to the standard protocols53, 62. Antibodies were used in this study as follows: rat anti-Ci (2 A) (DSHB, 1:50), mouse anti-Flag (M2) (Sigma,1:200), mouse anti-HA (F7) (Santa Cruz, 1:200), mouse anti-Myc (9E10) (Santa Cruz, 1:5000), mouse anti-β-galactosidase (Sigma, 1:500), mouse anti-En (DSHB, 1:50), mouse anti-Smo (DSHB, 1:50), mouse anti-Cut (DSHB, 1:50), 4,6-iamidino-2-phenylindole dihydrochloide (DAPI) (Santa Cruz, 1:1000). Secondary antibodies used in this study were bought from Jackson ImmunoResearch, and then were diluted at 1:500. For LMB (Sigma) treatment, S2 cells were treated with LMB at a final concentration of 5 nM for 2 h before cells were harvested for mRNA-cap localization assay. For in situ hybridization assay, the primers for mRNA-cap sub-cloning as follows: mRNA-cap upstream 5′-GTTATATGATGCTCATCGAT-3′; mRNA-cap downstream 5′-TTAATTTCCTAATGTCGGGT-3′; the corresponding of cDNA of mRNA-cap was cloned into pBluescript vector. mRNA-cap probe was prepared according to the instruction of the DIG RNA labeling kits (Roche).
Cell culture, Transfection, RNAi interference, Immunoprecipitation, and Western blot assay
S2 cells were maintained at 25 °C in Schneider′s Drosophila Medium (S9895, Sigma) supplemented with 10% FBS (F0718, Gibco) and 1% penicillin/streptomycin (P0781, Sigma). Transfection was performed using calcium phosphate according to the manufacturer’s instructions (Invitrogen). Usually S2 cells are transfected in 10-cm plates with no more than 20 μg of total DNA for an ubiquitin-Gal4 construct and other co-transfected pUAST expression vectors. 48 hrs after transfection, cells are harvested for immunoprecipitation and western blot analysis with standard protocols as previously described40. NIH/3T3 cells were cultured at 37 °C in an atmosphere of 5% CO2, in Dulbecco’s modified Eagle’s medium (DMEM; Life Technologies) supplemented with 10% FBS (F0718, Gibco) and 1% penicillin/streptomycin (P0781, Sigma). Indicated plasmids were transfected into cells with Lipofectamine 2000 (Invitrogen) according to manufacturer’s instructions. The following antibodies were used for immunoprecipitation and immunoblotting: mouse anti-Myc (9E10) (Santa Cruz, 1:5000), mouse anti-HA (Santa Cruz, 1:5000), mouse anti-Flag (M2) (Sigma, 1:5000), mouse anti-β-actin (GenScript, 1:10000), rabbit anti-PKAα (Santa Cruz, 1:500), rabbit anti-PKAIβ (Santa Cruz, 1:500), goat anti-rabbit HRP (1:10000; Jackson ImmunoResearch) and goat anti-mouse HRP (1:10000; Jackson ImmunoResearch). For RNA interference experiment, dsRNA was generated through in vitro transcription by using the MEGAscript T7 kit (Ambion). S2 cells were cultured in serum-free medium containing dsRNA for 12 hrs at 25 °C before transfection with DNA constructs. Then the culture medium was changed to serum medium and additional culturing for 48 hrs63. The primer sequences of mRNA-cap dsRNA are as follows: upstream 5′-GTTATATGATGCTCATCGAT-3′, downstream 5′-TTAATTTCCTAATGTCGGGT-3′. Western blot was performed as the standard protocols (described in Molecular Cloning). To silence RNGTT expression in 3T3 cells, the RNGTT siRNA was used (RiboBio). The sequence was 5′-GGAACCAUUUAGCGUCAGAdTdT-3′. All siRNA duplexes were transfected at a final concentration of 100 nM and maintained in serum-free medium for 4–6 hrs at 37 °C before transfection with indicated DNA constructs.
Real-time quantitative PCR (RT-qPCR)
For RT-qPCR, RNA of wing discs was isolated by using Trizol reagent (Invitrogen) and then reverse-transcribed by Prime-Script RT reagent kit (TaKaRa). Finally, the real-time PCR was done using the SYBR Premix Ex Taq (TaKaRa) according to the instrument of StepOnePlus (Applied Biosystem). Standard RT-qPCR primers for Drosophila mRNA-cap (upstream: 5′-AAAAACCATTTAAAGCGC-3′ and downstream 5′-GCTTTGCGCGATCACTAGC-3′), ci (upstream: 5′-CCTCTTGCGTATTCTGAATT-3′ and downstream: 5′-GAATCTGATGTTCCACCCGT-3′), smo (upstream: 5′-CAGCTATACAGCCCTTTTTG-3′ and downstream: 5′-CACTGGCCAGTTCCGTTGAA-3′), PKA-C1 (upstream: 5′-CACGAAAGACTATTATGCC-3′ and downstream: 5′-CGCGTAGAAGCGCGAGTGCGGCTCCG-3′), PKA-C3 (upstream: 5′-GCCGCTCGAACGGCCGAAG-3′ and downstream: 5′-GGTGGTGGTGGTGGTGGCGG-3′), PKA-R1 (upstream: 5′-GCGTTCTGAGCAGGGCGAGG-3′ and downstream: 5′-CCCAACACGCGTTCAAATC-3′), PKA-R2 (upstream: 5′-ATATGCCCAGAGCGGCCACCGTGC-3′ and downstream: 5′-GGCGGC GTCACCCTGTTTGATG-3′) and actin (upstream: 5′-GTACCCCATTGAGCACGGTA-3′ and downstream: 5′-CGAACATGATCTGGGCATC-3′) were synthesized by GenScript. actin was used as a control.
Luciferase reporter assay
In mammalian cells, Hh reporter assay was performed as previously described64. 48 hrs after transfection, cells were harvested and washed once with PBS, lysed in passive lysis buffer and luciferase activity was measured using a Dual Luciferase Assay Kit (Promega) as per manufacturer′s instructions. All luciferase activity data are presented as means ± SD of values from at least three experiments.
Protein kinase and cAMP Assay
For measuring PKA activity, about 400 wing discs were used as a sample, which were dissected in the 1.5 ml tube with PBS on ice and kept at 4 °C. After centrifugation at 2,000 × g for 3–5 min at 4 °C, the PBS was removed, instead 500ul cold sample preparation buffer was added. Then the discs were sonicated for 30–60 seconds or 15 seconds x 3 times. The samples were kept cool and cell suspensions were avoided foaming during the sonication. Next, cytosol fraction was separated by centrifugation at 100,000 × g for 1 hour at 4 °C. Finally, the supernatant was used to measure the PKA activities according to the protocol of MESACUP Protein Kinase Assay kit (MBL, No.5230). The cAMP amount was measured by cAMP Direct Immunoassay Kit (Biovision, No.K371-100) with about 400 wing discs, which were collected as described above and frozen quickly with liquid nitrogen. Then 200ul 0.1 M HCl was added, the sample was blended with homogenizer on ice. After centrifugation at 100,000 × g for 5 min, the supernatant was used to test the cAMP amount. The PKA activity and cAMP level were measured from MS1096 control wing discs or wing discs expressing mRNA-cap RNAi with MS1096 driver.
Statistical analysis
Imaging data were analyzed using Image J. Statistical tests were performed in GraphPad Prism 5. Data presented are the average of three independent experiments and error bars represent SD. A value of P < 0.05 is considered statistically significant by using Student′s t test.
References
Ingham, P. W. & McMahon, A. P. Hedgehog signaling in animal development: paradigms and principles. Genes & Development 15, 3059–3087 (2001).
Jia, J. & Jiang, J. Decoding the Hedgehog signal in animal development. Cellular and molecular life sciences: CMLS 63, 1249–1265 (2006).
Lum, L. & Beachy, P. A. The Hedgehog Response Network: Sensors, Switches, and Routers. Science 304, 1755–1759 (2004).
Ingham, P. W., Nakano, Y. & Seger, C. Mechanisms and functions of Hedgehog signalling across the metazoa. Nature reviews. Genetics 12, 393–406 (2011).
Jiang, J. & Hui, C. C. Hedgehog signaling in development and cancer. Developmental cell 15, 801–812 (2008).
Pasca di Magliano, M. & Hebrok, M. Hedgehog signalling in cancer formation and maintenance. Nature reviews. Cancer 3, 903–911 (2003).
Taipale, J. & Beachy, P. A. The Hedgehog and Wnt signalling pathways in cancer. Nature 411, 349–354 (2001).
Casali, A. & Struhl, G. Reading the Hedgehog morphogen gradient by measuring the ratio of bound to unbound Patched protein. Nature 431, 76–80 (2004).
Corbit, K. C. et al. Vertebrate Smoothened functions at the primary cilium. Nature 437, 1018–1021 (2005).
Stone, D. et al. The tumour-suppressor gene patched encodes a candidate receptor for Sonic hedgehog. Nature 384, 129–134 (1996).
Hooper, J. E. & Scott, M. P. The Drosophila patched gene encodes a putative membrane protein required for segmental patterning. Cell 59, 751–765 (1989).
Rohatgi, R., Milenkovic, L. & Scott, M. P. Patched1 Regulates Hedgehog Signaling at the Primary Cilium. Science 317, 372–377 (2007).
Marigo, V., Davey, R., Zuo, Y., Cunningham, J. & Tabin, C. Biochemical evidence that Patched is the Hedgehog receptor. Nature 384, 176–179 (1996).
Zhao, Y., Tong, C. & Jiang, J. Hedgehog regulates smoothened activity by inducing a conformational switch. Nature 450, 252–258 (2007).
Alcedo, J., Ayzenzon, M., Ohlen, T. V., Noll, M. & Hooper, J. The Drosophila smoothened Gene Encodes a Seven-Pass Membrane Protein, a Putative Receptor for the Hedgehog Signal. Cell 86, 221–232 (1996).
Denef, N., Neubuser, D., Perez, L. & Cohen, S. M. Hedgehog Induces Opposite Changes in Turnover and Subcellular Localization of Patched and Smoothened. Cell 102, 521–531 (2000).
Heuvel, Mvd & Ingham, P. Smoothened encodes a receptor-like serpentine protein required for hedgehog signalling. Nature 382, 547–551 (1996).
Jia, J., Tong, C. & Jiang, J. Smoothened transduces Hedgehog signal by physically interacting with Costal2/Fused complex through its C-terminal tail. Genes & Development 17, 2709–2720 (2003).
Lum, L. et al. Hedgehog Signal Transduction via Smoothened Association with a Cytoplasmic Complex Scaffolded by the Atypical Kinesin, Costal-2. Molecular Cell 12, 1261–1274 (2003).
Sisson, J. C., Ho, K. S., Suyama, K. & Scott, M. P. Costal2, a Novel Kinesin-Related Protein in the Hedgehog Signaling Pathway. Cell 90, 235–245 (1997).
Zhang, Y. et al. Transduction of the Hedgehog signal through the dimerization of Fused and the nuclear translocation of Cubitus interruptus. Cell Research 21, 1436–1451 (2011).
Zhou, Q. & Kalderon, D. Hedgehog activates fused through phosphorylation to elicit a full spectrum of pathway responses. Developmental cell 20, 802–814 (2011).
Jia, J. et al. Phosphorylation by double-time/CKIepsilon and CKIalpha targets cubitus interruptus for Slimb/beta-TRCP-mediated proteolytic processing. Developmental cell 9, 819–830 (2005).
Smelkinson, M. G., Zhou, Q. & Kalderon, D. Regulation of Ci-SCF Slimb binding, Ci proteolysis, and hedgehog pathway activity by Ci phosphorylation. Developmental cell 13, 481–495 (2007).
Strigini, M. & Cohen, S. M. A Hedgehog activity gradient contributes to AP axial patterning of the Drosophila wing. Development 124, 4697–4705 (1997).
Chen, C. et al. Nuclear Trafficking of Cubitus interruptus in the Transcriptional Regulation of Hedgehog Target Gene Expression. Cell 98, 305–316 (1999).
Price, M. & Kalderon, D. Proteolysis of the Hedgehog Signaling Effector Cubitus interruptus Requires Phosphorylation by Glycogen Synthase Kinase 3 and Casein Kinase 1. Cell 108, 823–835 (2002).
Aza-Blanc, P., Ramírez-Weber, F., Laget, M., Schwartz, C. & Kornberg, T. Proteolysis That Is Inhibited by Hedgehog Targets Cubitus interruptus Protein to the Nucleus and Converts It to a Repressor. Cell 89, 1043–1053 (1997).
Wang, Q. & Holmgren, R. The subcellular localization and activity of Drosophila Cubitus interruptus are regulated at multiple levels. Development 126, 5097–5106 (1999).
Tian, L., Holmgren, R. A. & Matouschek, A. A conserved processing mechanism regulates the activity of transcription factors Cubitus interruptus and NF-kappaB. Nature structural & molecular biology 12, 1045–1053 (2005).
Wang, B. & Li, Y. Evidence for the direct involvement of {beta}TrCP in Gli3 protein processing. Proc Natl Acad Sci USA 103, 33–38 (2006).
Jiang, J. & Struhl, G. Regulation of the Hedgehog and Wingless signalling pathways by the F-box/WD40-repeat proteinSlimb. Nature 391, 493–496 (1998).
Jia, J. et al. Shaggy/GSK3 antagonizes Hedgehog signalling by regulating Cubitus interruptus. Nature 416, 548–552 (2002).
Méthot, N. & Basler, K. Hedgehog Controls Limb Development by Regulating the Activities of Distinct Transcriptional Activator and Repressor Forms of Cubitus interruptus. Cell 96, 819–831 (1999).
Smelkinson, M. G. & Kalderon, D. Processing of the Drosophila hedgehog signaling effector Ci-155 to the repressor Ci-75 is mediated by direct binding to the SCF component Slimb. Current biology: CB 16, 110–116 (2006).
Zhang, W. et al. Hedgehog-regulated Costal2-kinase complexes control phosphorylation and proteolytic processing of Cubitus interruptus. Developmental cell 8, 267–278 (2005).
Apionishev, S., Katanayeva, N. M., Marks, S. A., Kalderon, D. & Tomlinson, A. Drosophila Smoothened phosphorylation sites essential for Hedgehog signal transduction. Nature Cell Biology 7, 86–92 (2005).
Chen, Y. et al. G protein-coupled receptor kinase 2 promotes high-level Hedgehog signaling by regulating the active state of Smo through kinase-dependent and kinase-independent mechanisms in Drosophila. Genes & Development 24, 2054–2067 (2010).
Zhang, C., Williams, E. H., Guo, Y., Lum, L. & Beachy, P. A. Extensive phosphorylation of Smoothened in Hedgehog pathway activation. Proc Natl Acad Sci USA 101, 17900–17907 (2004).
Jiang, J. & Struhl, G. Protein Kinase A and Hedgehog Signaling in Drosophila Limb Development. Cell 80, 563–572 (1995).
Ohlmeyer, J. & Kalderon, D. Hedgehog stimulates maturation of Cubitus interruptus into a labile transcriptional activator. Nature 396, 749–753 (1998).
Chen, Y., Gallaher, N., Goodman, R. & Smolik, S. Protein kinase A directly regulates the activity and proteolysis of cubitus interruptus. Proc Natl Acad Sci USA 95, 2349–2354 (1998).
Changela, A., Ho, C., Martins, A., Shuman, S. & Mondragón, A. Structure and mechanism of the RNA triphosphatase component of mammalian mRNA capping enzyme. The EMBO Journal 20, 2575–2586 (2001).
Liu, L. Functional characterization of the C-terminal domain of mouse capping enzyme. Cell Biochem Funct 24, 95–102 (2006).
Cong, P. & Shuman, S. Mutational Analysis of mRNA Capping Enzyme Identifies Amino Acids Involved in GTP Binding, Enzyme-Guanylate Formation, and GMP Transfer to RNA. Molecular and Cellular Biology 15, 6222–6231 (1995).
Wang, S., Deng, L., Ho, C. & Shuman, S. Phylogeny of mRNA capping enzymes. Proc Natl Acad Sci USA 94, 9573–9578 (1997).
Takagi, T., Moore, C., Diehn, F. & Buratowski, S. An RNA 5′-Triphosphatase Related to the Protein Tyrosine Phosphatases. Cell 89, 867–873 (1997).
Wen, Y., Yue, Z. & Shatkin, A. Mammalian capping enzyme binds RNA and uses protein tyrosine phosphatase mechanism. Proc Natl Acad Sci USA 95, 12226–12231 (1998).
Parker, R. & Song, H. The enzymes and control of eukaryotic mRNA turnover. Nature structural & molecular biology 11, 121–127 (2004).
Schoenberg, D. R. & Maquat, L. E. Re-capping the message. Trends in biochemical sciences 34, 435–442 (2009).
Price, M. A. & Kalderon, D. Proteolysis of Cubitus interruptus in Drosophila requires phosphorylation by Protein Kinase A. Development 126, 4331–4339 (1999).
Ranieri, N., Therond, P. P. & Ruel, L. Switch of PKA substrates from Cubitus interruptus to Smoothened in the Hedgehog signalosome complex. Nature communications 5, 5034–5049 (2014).
Wang, G., Wang, B. & Jiang, J. Protein kinase A antagonizes Hedgehog signaling by regulating both the activator and repressor forms of Cubitus interruptus. Genes & Development 13, 2828–2837 (1999).
Yue, Z. et al. Mammalian capping enzyme complements mutant Saccharomyces cerevisiae lacking mRNA guanylyltransferase and selectively binds the elongating form of RNA polymerase II. Proc Natl Acad Sci USA 94, 12898–12903 (1997).
Li, S., Ma, G., Wang, B. & Jiang, J. Hedgehog induces formation of PKA-Smoothened complexes to promote Smoothened phosphorylation and pathway activation. Science signaling 7, ra62 (2014).
Ogden, S. K. et al. G protein Galphai functions immediately downstream of Smoothened in Hedgehog signalling. Nature 456, 967–970 (2008).
Posternak, V., Ung, M. H., Cheng, C. & Cole, M. D. MYC Mediates mRNA Cap Methylation of Canonical Wnt/β-Catenin Signaling Transcripts By Recruiting CDK7 and RNA Methyltransferase. Molecular Cancer Research 15, 213–224 (2017).
Grasso, L. et al. mRNA Cap Methylation in Pluripotency and Differentiation. Cell Reports 16, 1352–1365 (2016).
Aregger, M. et al. CDK1-Cyclin B1 Activates RNMT, Coordinating mRNA Cap Methylation with G1 Phase Transcription. Molecular Cell 61, 734–746 (2016).
Zhang, Q. et al. A hedgehog-induced BTB protein modulates hedgehog signaling by degrading Ci/Gli transcription factor. Developmental cell 10, 719–729 (2006).
Rubin, G. & Spradling, A. Genetic transformation of Drosophila with transposable element vectors. Science 218, 348–353 (1982).
Zhou, Z. et al. Deubiquitination of Ci/Gli by Usp7/HAUSP Regulates Hedgehog Signaling. Developmental cell 34, 58–72 (2015).
Liu, C. et al. Hedgehog signaling downregulates suppressor of fused through the HIB/SPOP-Crn axis in Drosophila. Cell Research 24, 595–609 (2014).
Di Marcotullio, L. et al. Numb is a suppressor of Hedgehog signalling and targets Gli1 for Itch-dependent ubiquitination. Nat Cell Biol 8, 1415–1423 (2006).
Acknowledgements
We thank Fly Stocks of National Institute of Genetics of Japan (NIG-Fly), Vienna Drosophila RNAi Center (VDRC), the Bloomington Stock Center and Developmental Studies Hybridoma Bank at the University of Iowa for providing fly stocks and reagents. This work was supported by grants from the National Key Scientific Program of China (2011CB943902), and the National Natural Science Foundation of China (30971679, 31071264 and 31271531) to QZ, and Welch foundation grant (I-1603) and NIH grant (GM118063) to JJ.
Author information
Authors and Affiliations
Contributions
Q.Z. designed the experiments. P.C., Z.Z., X.Y., S.P., M.L. and W.J. performed the experiments. P.C., Q.Z. and J.J. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, P., Zhou, Z., Yao, X. et al. Capping Enzyme mRNA-cap/RNGTT Regulates Hedgehog Pathway Activity by Antagonizing Protein Kinase A. Sci Rep 7, 2891 (2017). https://doi.org/10.1038/s41598-017-03165-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-03165-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.