Abstract
Many herbal products have a long history of use, but there are increasing concerns over product efficacy, safety and quality in the wake of recent cases exposing discrepancies between labeling and constituents. When it comes to St. John’s wort (Hypericum perforatum L.) herbal products, there is limited oversight, frequent off-label use and insufficient monitoring of adverse drug reactions. In this study, we use amplicon metabarcoding (AMB) to authenticate 78 H. perforatum herbal products and evaluate its ability to detect substitution compared to standard methods using thin-layer chromatography (TLC) and high performance liquid chromatography coupled with mass spectrometry (HPLC-MS). Hypericum perforatum was detected in 68% of the products using AMB. Furthermore, AMB detected incongruence between constituent species and those listed on the label in all products. Neither TLC nor HPLC-MS could be used to unambiguously identify H. perforatum. They are accurate methods for authenticating presence of the target compounds, but have limited efficiency in detecting infrageneric substitution and do not yield any information on other plant ingredients in the products. Random post-marketing AMB of herbal products by regulatory agencies could raise awareness among consumers of substitution and would provide an incentive to manufacturers to increase quality control from raw ingredients to commercialized products.
Similar content being viewed by others
Introduction
St. John’s wort (Hypericum perforatum L.) herbal products are popular in complementary and alternative medicine, and are widely used to treat mild to moderate depression but have a much broader traditional use1. These products play an important role in primary healthcare, and their popularity is determined by consumer health concerns, cultural habits and by the belief that they are natural and thus safe2. Hypericum perforatum is among the top-selling herbs and is sold as over-the-counter (OTC) products in pharmacies, supermarkets, health shops and through e-commerce3. Products are typically labeled as natural foods or dietary supplements, and claims regarding their possible health benefits appear on labels and in associated advertising. In 2011, the global market for herbal products was estimated to be US$83 billion4 with Europe being the largest market.
Lack of standardized methods for quality assessment and the highly competitive market of herbal products has increased the incentive to use substitutes and unlabeled fillers5,6,7. However, adulteration is not necessarily intentional, and herbal products may be altered due to accidental adulteration, misidentification8 and confusion resulting from vernacular names9, 10. In any case, the use of unreported ingredients is a serious safety concern as adverse drug reactions cannot be associated to the product label and ingredients11, 12. Signal detection in herbal adverse drug reactions is greatly impeded if ingredients in the products are not reported on the labels10.
The European Medicines Agency (EMA) is the European Union agency that is responsible for the evaluation of medicinal products. However, differences in herbal medicine classification exist between EU/EEA member states and this complicates quality monitoring of these products11. EMA does not test the composition of herbal products or verify whether ingredients included on the label are included in the product but delegates this responsibility to the manufacturers of these products. EMA requires quality assurance of herbal substances, preparations and products and specifies the use of macroscopic and microscopic characterization, phytochemical analysis of therapeutic target compounds and markers and assays for toxic constituents such as heavy metals and toxins13. EMA suggests that identification tests specific for substitute and adulterant detection either use a combination of separate chromatographic approaches (e.g., HPLC with TLC-densitometry) or combine different approaches into a single procedure (e.g., HPLC-UV, HPLC-MS or GC-MC)13. Herbal products are usually highly processed and have numerous ingredients, and applying these methods might not enable the accurate identification of all plant ingredients, especially if target species are admixed with other species within the same genera. To complement traditional identification methods, the EMA, as well as the United States Food and Drug Administration (FDA), support the use of innovative analytical technologies such as DNA barcoding.
DNA barcoding is a validated molecular identification method that can provide species-level resolution that is commonly used in authentication of taxonomic provenance of herbal products5,6,7, 14. Sanger sequencing based DNA barcoding studies have revealed widespread levels of substitution: 6% in saw palmetto herbal dietary supplements15, 16% in ginkgo products16, 25% in black cohosh17, 33% in herbal teas18, and 50% in ginseng19. A blind test of 44 herbal products sold in North America using DNA barcoding7 found that 59% contained species not listed on the labels, and only two out of twelve screened companies had products free of substitution, contamination or unreported fillers7. High-throughput sequencing based amplicon metabarcoding (AMB)20 studies can provide insights into species composition of complex mixtures of DNA such as processed herbal products. For example, in a study by Coghlan et al.5, the species composition of 15 highly processed traditional Chinese medicines (TCM) were evaluated using high-throughput sequencing and found that these contained species and genera included on CITES appendices I and II. Other AMB studies have shown similar concerns of varying quality and product label-content fidelity. For instance, Ivanova et al.21 found that 15 tested herbal supplements contained non-listed, non-filler plant DNA and Cheng et al.22 showed that the quality of 27 tested herbal preparations was highly affected by the presence of contaminants.
In this study, we investigated complex herbal products containing St. John’s wort (Hypericum perforatum), marketed in the EU/EEA both as herbal food supplement and herbal drug. St. John’s wort has traditional indications in nervous system, psychiatric, gastrointestinal, hepatobiliary, renal and urinary, respiratory, thoracic, endocrine, musculoskeletal, metabolism and nutritional disorders, as well as in infections and infestations23. The mode of action of the major responsible bioactive compounds of H. perforatum is still not completely known, but it seems that they act in a synergetic manner to achieve the clinical effectiveness24. However, several studies showed that the antidepressant activity is associated mainly to the phloroglucinol derivative hyperforin25 and the naphthodianthrones hypericin and pseudohypericin26. The quantity and quality of active constituents in Hypericum herbal medicines are highly affected by the manufacturing process1. Treatments involving St. John’s wort are generally safe, but several studies show that use in combination with other drugs can cause potentially life-threatening adverse drug reactions due to pharmacokinetic interactions3. These adverse interactions are still not fully understood despite extensive studies in its mechanisms. Hypericum extracts can cause serious side effects when administrated simultaneously with some antidepressants27. Other studies reported adverse reactions due to high doses of hypericin in Hypericum products leading to a phototoxic effect that may induce photodermatitis and can also decrease or nullify the effect of other drugs when administrated simultaneously28. Hall et al.29 reported exacerbated vaginal breakthrough bleeding, and Murphy et al.30 observed evidence of follicle growth and probable ovulation when simultaneously administrating H. perforatum with low-dose oral contraceptives.
The aim of this study was to evaluate the ability of using AMB to detect substitution in single and multi-ingredient Hypericum herbal products compared with standard identification approaches suggested by the European Pharmacopoeia and the European Medicine Agency, such as HPLC-MS and TLC.
Results
Thin Layer Chromatography
The European Pharmacopoeia sets a minimum concentration of 0.08% total hypericins in the dried drug31, and includes a TLC based identification assay intended to distinguish H. perforatum from other species, including other species within the same genus. The European Pharmacopoeia 8.031 Hyperici herba monograph TLC test yields four zones, corresponding to rutin, hyperoside, pseudohypericin and hypericin, calibrated by two reference solution compounds, rutin and hyperoside (Fig. 1). This can be used to distinguish between presence of rutin in H. perforatum and small quantities of rutin in H. maculatum Crantz. Our test control samples included H. elegans Stephan ex Willd., H. maculatum, H. olympicum L., H. patulum Thunb., H. perforatum, and H. polyphyllum Boiss. & Balansa. This testing revealed some remarkable challenges associated with this authentication method (Fig. 1). The chromatograms of H. olympicum, H. patulum, and H. polyphyllum were indistinguishable from those of H. perforatum.
Thin layer chromatogram (TLC) of Hypericum perforatum and other Hypericum species. The yellow-orange fluorescent bands from the lower third of the chromatogram correspond to rutin and hyperoside, and are used for the identification of H. perforatum. This distinguishes between presence of rutin in H. perforatum and absence or only small quantities of rutin in H. maculatum. The bands corresponding to rutin and hyperoside are found also in H. olympicum, H. patulum, and H. polyphyllum.
The product samples were all tested using the recommended test. Sample extraction for four samples (a tincture, two oils, and one juice) out of 78 failed. The test results could be grouped into three categories: (1) rutin, hyperoside, pseudohypericin and hypericin present. This could indicate presence of H. perforatum or other Hypericum species with indistinguishable chromatograms, e.g. H. olympicum, H. patulum, and H. polyphyllum; (2) hyperoside, hypericin, and pseudohypericin present, but rutin in low or undetectable concentrations. This made presence of H. perforatum or other Hypericum species with indistinguishable chromatograms unlikely, but could indicate presence of H. maculatum or other Hypericum species with indistinguishable chromatograms, for example H. elegans; (3) rutin detected or not, and hyperoside, pseudohypericin and hypericin not detected. This ruled out presence of Hypericum species. Fifty-five out of 74 (74%) samples contained H. perforatum or an indistinguishable adulterant, or a mixture of Hypericum species including H. perforatum or an indistinguishable adulterant. Nine out of 74 (12%) samples contained Hypericum species other than H. perforatum or indistinguishable adulterants, or a mixture of those other Hypericum species. Ten out of 74 (14%) samples did not contain Hypericum species in detectable amounts (Fig. 2A; Supplementary Fig. S1; Supplementary Table S2).
Presence of Hypericum perforatum within the products. (A) Detection using thin layer chromatogram (TLC). (B) Detection using high-performance liquid chromatography-mass spectrometry (HPLC-MS). (C) Detection using amplicon metabarcoding (AMB). Detection between methods is not fully comparable as the resolution of the approaches differs.
High-Performance Liquid Chromatography-Mass Spectrometry
Detection of H. perforatum using HPLC-MS is based on the presence of the two main bioactive compounds in Hyperici herba, hyperforin and hypericin. The Hyperici herba monograph in the European Pharmacopoeia 8.031 states that the dried drug should have a minimum content of total hypericins, expressed as hypericin, corresponding to 0.08% to verify the presence of H. perforatum. A literature review of 88 taxa of Hypericum with qualitative (74 taxa32) and quantitative (27 taxa33) measures of hyperforin and hypericin, together with the six test control species measured in this study (Supplementary Table S3), revealed that the hyperforin and hypericin content was not an accurate predictor of H. perforatum presence. Hypericum perforatum and its main adulterant H. maculatum had hypericin contents within similar ranges, but H. maculatum had very low content of hyperforin, <0.018%. However, several of the tested or reviewed species had both hyperforin and hypericin contents similar to those of H. perforatum, most notably H. olympicum and H. polyphyllum.
All 78 product samples were analyzed using HPLC-MS, except for four samples that could not be extracted (a tincture, two oils, and one juice). The tested products showed presence of rutin, hyperoside, pseudohypericin and hypericin in 55 out of 74 (74%) samples, which could indicate H. perforatum or other Hypericum species with indistinguishable chromatograms. In nine out of 74 (12%) samples hyperoside, hypericin, and pseudohypericin were present, but rutin was present in low or undetectable concentrations, which made presence of H. perforatum or other Hypericum species with indistinguishable chromatograms unlikely, but could indicate presence of H. maculatum or other Hypericum species with indistinguishable chromatograms, such as H. elegans, or a mixture of Hypericum species. In ten out 74 (14%) samples rutin was detected or not, and hyperoside, pseudohypericin and hypericin were not detected, which ruled out presence of Hypericum species. The test results could further be grouped into three categories: (1) hyperforin content higher than hypericin content. This could indicate presence of H. perforatum or other Hypericum species with high levels of hyperforin, e.g. H. elegans, H. olympicum or H. polyphyllum; (2) hyperforin content equal to or lower than hypericin content. This could indicate presence of H. maculatum or other species with low levels of hyperforin, such as H. barbatum Jacq., H. hirsutum L., H. humifusum L., H. linarioides Bosse, H. richeri Vill., H. rumeliacum Boiss. or H. tetrapterum Fr.; (3) no hyperforin or hypericin detected. This indicates that material from Hypericum species is absent or that these compounds are present in undetectable amounts. It should be noted that low levels of hyperforin could also indicate low content of H. perforatum in the tested product. Sixty-one out of 74 samples (82%) had hyperforin content higher than hypericin content; eight out of 74 (11%) had hyperforin content equal to or lower than hypericin content; and five out of 74 (7%) had no detectable hyperforin or hypericin (Fig. 2B; Supplementary Table S4).
DNA metabarcoding
The DNA extracted from 78 samples was highly variable in quantity and quality. Fragment Analyzer measurements gave results for 47 (60%) samples, with DNA concentration ranging from 0.01 to 140 ng/μl. Thirty-one samples (40%) did not contain measurable DNA concentrations: 11 capsules, eight tablets, eight herbal teas and four extracts. PCR amplification reactions were performed for all 78 samples and amplicons were obtained from 76% of the samples for nrITS1 and 73% for nrITS2. The highest amplification rate was obtained for herbal teas (97% for nrITS1 and 88% for nrITS2), followed by capsules (62% for nrITS1 and 62% for nrITS2), tablets (43% for nrITS1 and 45% for nrITS2) and extracts (27% for nrITS1 and 40% for nrITS2). There was no significant correlation between sample total DNA concentration and nrITS amplicon concentration or between amplicon concentration and sequenced reads or bases (Supplementary Table S5; Supplementary Fig. S6).
The raw data consisted of 9,416,033 sequences, with an average of 60,359 sequences per sample for each marker. Sequencing success rates were 49%, respectively 44% (34/78 samples) for nrITS1 and 47% (37/78 samples) for nrITS2. A dataset consisting of 1,511,356 reads, fulfilling our trimming and filtering quality criteria, was obtained, including 737,010 nrITS1 and 774,346 nrITS2 reads (on average 19,395 nrITS1 and 20,377 nrITS2 reads per sample). Forty samples out of 78 samples (51%) yielded no molecular operational taxonomic units (MOTUs) for either nrITS1 or nrITS2 and are excluded from the results and discussion (2, 7, 9, 11–14, 16, 18–20, 22, 23, 25, 26, 28, 31, 33, 34, 36, 40, 41, 45, 47, 49, 52–54, 56–58, 60, 62, 63, 64, 66–70). These included 21 herbal teas, eight capsules, nine tablets and two extracts (Supplementary Table S7). The MOTU yielding samples included 23 herbal teas, seven capsules, five tablets and three extracts.
A total of 219 different species were identified using BLAST from the retained MOTUs (Supplementary Table S7; Fig. 3 for MOTUs with >1% normalized reads). For nrITS1 we detected a total of 143 different species and for nrITS2 137 species (Supplementary Fig. S8, Supplementary Fig. S9). Analysis of the BLAST identified MOTUs showed that a total of 34 genera (24%) and 76 species (36%) were detected exclusively with nrITS1. Analysis of nrITS2 showed that 82 (39%) species, but no unique genera, were detected exclusively with this marker. A total of 106 genera (76%) and 55 (26%) species were detected with both markers. Identified MOTUs and their read numbers were merged for both markers per sample for further analysis.
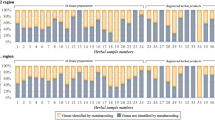
Species identified within the products using amplicon metabarcoding (AMB). Only MOTUs with >1% normalized read numbers per sample are shown. Species are colored according to relative abundance of normalized read numbers. Products are grouped by product form: herbal teas, capsules, tablets/pills/pastilles and extracts/tinctures/oils.
The number of species detected per sample ranged from one to 57, with an average of 18.7 species per sample. Out of the 17 successfully analyzed single ingredient samples, those containing only H. perforatum according to the label, none contained only H. perforatum, 13 contained more than one species, and four did not contain H. perforatum. Out of the 21 successfully analyzed multiple ingredient samples, those containing H. perforatum and other species according to the label, none contained only the ingredients listed on the label, two contained all species listed on the label plus additional species not listed on the label, 19 contained fewer species than listed on the label plus additional species, and eight did not contain H. perforatum (Fig. 2). The fidelity for H. perforatum in single ingredient products was 76% (13 out of 17), and for multi ingredient products, 62% (13 out of 21). The overall ingredient fidelity (detected species from product label/total number of species on label) for multi ingredient products was 41% and for all products 57%. The following four species were found in more than 50% of the samples: Hypericum perforatum (68%), Convolvulus arvensis L. (63%), Achillea millefolium L. (53%) and Urtica dioica L. (50%). Plant taxa present in more than 20% of the samples are listed in Table 1. Eleven out of the 78 products were commercialized and registered as herbal medicines, and 4 of these yielded AMB results. Hypericum perforatum was found in all four products, and in addition an average of 20.3 other species.
In addition to the target species, H. perforatum, that was detected in 68% (26) of the MOTU yielding samples, several other Hypericum species were also detected: H. humifusum in 21% (8), H. tetrapterum in 13% (5) and H. hirsutum in 3% (1). Other Hypericum species were never detected without H. perforatum suggesting that adulteration by admixture is more widespread than complete substitution. The five most common species detected with AMB, but which were not present on the label of the products were: Agrostis gigantea Roth (Poaceae) detected in 24%, Centaurea cyanus L. (Compositae) in 21%, Vaccinium myrtillus L. (Ericaceae) in 21%, Lolium perenne L. (Poaceae) in 18%, and Apera spica-venti (L.) P. Beauv. (Poaceae) in 16% of the samples. A total of 34 anemophilous (wind-pollinated) species were detected, including 31 grasses and the woody species Fraxinus excelsior L., Humulus lupulus L., and Juglans regia L. (Supplementary Table S7).
Comparative results
In Fig. 2 the detection rates of the three authentication methods are summarized. The detection rate of AMB was much lower than that of the other approaches, but neither TLC nor HPLC-MS could be used to unambiguously identify H. perforatum, and it was thus difficult to make an overall comparison of the three methods. Considering only the 26 samples in which H. perforatum was detected using AMB, TLC could be used to detect rutin, hyperoside, hypericin and pseudohypericin in 21 samples (81%), and HPLC-MS could be used to detect higher hyperforin than hypericin content in 23 samples (88%). All three methods were in agreement for 19 samples (73%). For the 12 samples that yielded MOTUs using AMB but in which H. perforatum was not detected, TLC could be used to detect rutin, hyperoside, hypericin and pseudohypericin in six samples (50%), and HPLC-MS could be used to detect higher hyperforin than hypericin content in six samples (50%). For five samples (42%), the results from TLC and HPLC-MS were in agreement.
Discussion
Misidentification of H. perforatum and/or adulteration of products containing H. perforatum with other Hypericum species, and the common and hyperforin-less H. maculatum in particular, requires the use of accurate analytical methods for the quality control of herbal products of H. perforatum. Identification of Hypericum species is feasible using taxonomic identification keys, but recognition of species in the field is challenging and many Hypericum species have superficially similar morphology. Positive identification requires the researchers to study flower, leaf and stem morphology including diagnostic characters that might be absent early or late in the flowering season. Identification is further complicated as H. perforatum may occasionally hybridize with other species in the genus, resulting in hybrids that have intermediate morphology and secondary metabolite spectra34, 35. Eastern Europe is a significant source of Hypericum material for European herbal products36, and here material is mostly wild-harvested and several different species co-occur, which could easily lead to intentional and/or accidental picking of the different species.
Using the TLC based identification assay included in the European Pharmacopoeia on a limited number of reference species included to check the accuracy of the test, show that several species have indistinguishable chromatograms from that of H. perforatum (Fig. 1). In addition to the ambiguous results with regard to different Hypericum species, the test would show the same chromatogram for an admixture of H. perforatum and H. maculatum as for an unadulterated H. perforatum sample. In addition, the TLC assay does not provide useful information about the concentrations of the main bioactive compounds, hyperforin and hypericin.
An alternative to testing for indicator compounds using TLC is to use HPLC-MS to detect and measure the content of the main bioactive secondary metabolites. This method does not claim to distinguish H. perforatum from other Hypericum species. Reviews state that the hyperforin content in H. perforatum is 2.0–4.5%, and the hypericin content 0.1–0.15% in fresh material37, 38. This is supported by several recent studies that have found hyperforin and hypericin contents of 1.4% and 0.5%, respectively, in the flowers and of 0.4–1.4% and 0.015–0.17%, respectively, in the vegetative parts39, 40. Both the hyperforin and hypericin contents vary greatly between studies, probably based on the nature and quality of the analyzed plant material41. To check the accuracy of HPLC-MS for distinguishing H. perforatum, we analyzed seven samples representing six Hypericum species, and compiled literature on hyperforin and hypericin values from 27 species33, 39, 40, 42. The results show that the hyperforin and hypericin contents vary considerably between and within species (Supplementary Table S3). Hypericum perforatum and its main adulterant H. maculatum have hypericin contents within similar ranges, but H. maculatum has very low content of hyperforin, 0.004–0.018%. This compilation highlights the low predictive value of hyperforin and hypericin content in determining substitution in H. perforatum preparations. Several species have similar hyperforin and hypericin contents to H. perforatum, most notably H. olympicum and H. polyphyllum.
The categorization of hyperforin and hypericin content per product into three groups made it possible to conclude that five products out of 74 (7%) have no detectable hyperforin or hypericin, eight out of 74 (11%) have low content or absence of H. perforatum, and 61 out of 74 samples (82%) have spectra typical of H. perforatum but also several other species. The 74 analyzed samples had hypericin contents ranging from 0–0.03%, and none passed the minimum threshold set by the European Pharmacopoeia (Supplementary Table S4). Wurglics et al.43 showed that commercial products bought in Germany have total hypericin contents ranging from 0.16–0.30%, and thus exceeding the European Pharmacopoeia minimum. The reason for the discrepancy between these and our results is not entirely clear, especially as both have been measured using similar methodology, but could be a result of lower product quality in our study material. In summary, HPLC-MS is not an accurate method for detection of substitution, adulteration or admixture, but suitable for control of bioactive compound content in products, and thus important in quality control for consumer safety. A superior, but more cost-intensive approach, than TLC and HPLC-MS is NMR metabolomics that enables chemical fingerprinting encompassing a huge range of target molecules44.
Several studies have shown the resolution and efficacy of DNA metabarcoding20 for identifying plant species diversity in a range of products5, 6, 21, 22, 45,46,47. Comparative identification of processed food and pharmaceutical products is challenging as compared to substrates that can be used for morphological identification, such as pollen clumps46, 48 and pollen in honey45. The advantage of pharmaceutical products and traditional and complementary alternative medicines is that these have their putative contents printed on the package5, 6, 21, 22. Galimberti et al.48, Richardson et al.46, and Hawkins et al.45 used rbcL and trnH-psbA, and nrITS2 and rbcL, respectively, to analyze DNA from pollen in pollen grains and honey to investigate honey bee foraging preferences. Cheng et al.22 used amplicon metabarcoding to analyze nine traditional Chinese medicines (TCMs) and detected on average 4.8 species using nrITS2 and 2.8 using trnL. Coghlan et al.5, 6 analyzed TCMs for presence of both animal and plant ingredients and found over 68 plant families and eight vertebrate genera in these products. Ivanova et al.21 used universal nrITS primers and found a host of plant species in eight herbal supplements, as well as many fungi due to specificity of these primers in amplifying fungal nrITS. In this study, plant specific nrITS primers were used to amplify nrITS1 and nrITS2, and an average of 18.7 species were detected per sample. In addition, the presence of H. perforatum was detected in 26 out of 38 sequenced samples (68%). These findings corroborate previous results that amplicon metabarcoding is an effective way to investigate species composition in products that contain a mixture of DNA from different species. Looking at the subset of registered herbal medicines that yielded AMB data, all four were found to contain H. perforatum, and this was supported by TLC results for all four, and by HPLC results in only two cases.
The relatively low success rate (49%) after applying strict read quality and filtering criteria makes this method challenging to use for routine screening at this time. Ivanova et al.21 reported a slightly higher success rate with eight out of 15 samples (53%) for nrITS2 AMB of herbal products from Echinacea, Gingko, Hypericum, Trigonella, and Valeriana. Cheng et al.22 reported a 100% success rate for 30 individual samples of TCM, all of which were unprocessed crude drugs. The varying degrees of success probably reflect the quality and type raw material, but also the many details in the analysis that can be varied to optimize the results, roughly in order of significance: extraction procedures and purification, primers, markers, identification approach, clustering, MOTU thresholds, sequencing platform, filtering, quality thresholds and chimera removal, library preparation, and amplification protocols.
Quantifying contamination is important when focusing on the tolerated levels of foreign matter in herbal pharmaceuticals. Quantifying relative species abundances based on sequence read numbers from samples with unknown ingredients is hampered by several factors. Firstly, AMB relies on the availability of DNA, but plant DNA can be removed or highly degraded during the harvesting, drying, storage, transportation, and processing (e.g., mode of extraction, irradiation, ultraviolet light exposure, heat or pressure, filtration, extractive distillation or supercritical fluid extraction)49. Secondly, AMB is a PCR-based method, and variation in nrITS copy number, primer annealing, and amplification bias all influence the number of taxon-specific reads50. Thirdly, incomplete reference databases and sequences with incorrect species names, can render taxonomic identifications prone to uncertainties.
AMB detected the target species, H. perforatum, in 68% of the samples, but in addition other Hypericum species were detected, H. humifusum in 21% (8), H. tetrapterum in 13% (5) and H. hirsutum in 3% (1) of the samples. Hypericum tetrapterum belongs to the same taxonomic section as H. perforatum but is more closely related to H. maculatum, whereas H. humifusum and H. hirsutum belong to sections Oligostema and Hirtella s.l., respectively51, 52. These species can co-occur in natural habitats from which H. perforatum is wild crafted. These other Hypericum species were never detected without H. perforatum suggesting that adulteration by admixture is more widespread than complete substitution.
The overlooked species diversity through poor primer fit and amplification bias is difficult to quantify but some diversity is likely missed53. The detection of additional plant species, other that the ones from the label or those that can be expected as substitutes, contaminants or fillers, may be explained by (1) amplified PCR chimeras; (2) false-positive BLAST identifications due to incomplete or error-prone reference databases; or (3) presence of pollen from anemophilous (wind-pollinated) species. Thirty-four of the MOTUs belong to anemophilous species, including 31 grasses of which Agrostis gigantea Roth (Poaceae) was detected in 24% of samples, Lolium perenne L. (Poaceae) in 18%, and Apera spica-venti (L.) P.Beauv. (Poaceae) in 16%. DNA from pollen from these species can end up in the products through co-occurrence with the collected material in natural habitats as well as during other steps in the process chain.
Conclusions
The metabarcoding results confirm that AMB can be used to test for the presence of H. perforatum and simultaneously to detect substitution, adulteration and/or admixture of other species. These results corroborate with previous results that show the usefulness of metabarcoding for use in complementing traditional methods of quality control for consumer safety5, 6, 21, 22. It should be emphasized however that the relatively low success rate of generating sequence reads per product makes this method challenging to use for routine screening, as only 49% of the tested products yielded sequences as compared to 95% of products that could be used for TLC and HPLC-MS analyses. Moreover, the high sensitivity of AMB in detecting everything from grass pollen on field-collected plants to plant dust left in production equipment requires a careful consideration of the concept of contamination. AMB results for herbal pharmaceutical authentication should be interpreted with a focus on presence and or absence of target species, i.e. the labeled ingredients, but alarms need not be raised over trace contaminations from species plausibly present in the cultivation, transport or production chain. However, the TLC and HPLC-MS results show that these methods are of limited applicability with regard to detecting species substitution, but may be used efficiently to detect target compounds. The clear advantage of HPLC-MS over TLC is the ability to quantify constituents and to screen for a vastly larger number of compounds. If product safety relies on threshold levels of specific bioactive compounds, absence of toxins, allergens and admixed pharmaceuticals, then chemical analysis methods are more relevant than DNA based composition analysis, but if product fidelity, species substitution or adulteration is suspected then the latter method outperforms in terms of resolution. Development of novel molecular markers and approaches for genomic barcoding are likely to increase the resolution of DNA barcoding in species-level identification. Several others have advocated the use of DNA barcoding and metabarcoding in herbal product authentication and herbal pharmacovigilance5, 7, 9, 21, 22, 54,55,56 and adoption of standards for quality control by regulatory agencies could raise product quality and increase consumer confidence.
Methods
Sample collection
Seventy-eight herbal products that included Hypericum perforatum according to the label were randomly purchased in European countries (and Turkey), including Romania (51), Germany (5), Poland (4), Turkey (4), Slovakia (3), Spain (2), UK (2), Austria (2), Czech Republic (1), France (1), Italy (1), Sweden (1) and the Netherlands (1). The samples were bought from pharmacies (44), herbal shops (25), super markets (2) or via e-commerce (7), and were sold as herbal teas (44), capsules (15), tablets (14) and extracts (5). According to the label information, the products included 38 single ingredient products, 33 products contained between two and ten ingredients and seven products contained more than ten ingredients. These medicinal products for scientific analyses were imported into Norway under Norwegian Medicines Agency license no. 16/04551–2. An overview of the samples including label information, but not the producer/importer name, lot number, expiration date or any other information that could lead to the identification of that specific product can be found in Supplementary Table S10.
For the phytochemical analysis, aerial parts of H. elegans (voucher ARNHM01He), H. maculatum (ARNHM01Hm), H. maculatum (92151 UMF CLUJ), H. olympicum (ARNHM01Ho), H. patulum (ARNHM01Hpa), H. perforatum (92141 UMF CLUJ) and H. polyphyllum (ARNHM01Hpo) were used as references for the identification and quantification of the main compounds. The reference species were selected to include the main adulterant, H. maculatum (two geographically isolated samples from Romania and Norway), species with known high levels of hyperforin and hypericin, H. polyphyllum and H. olympicum, as well as random species collected during fieldwork in Romania, H. patulum and H. elegans. All voucher specimens are deposited in the Herbarium of the Alexander Borza Botanical Garden (CL) of Babes-Bolyai University, Cluj-Napoca, Romania.
Phytochemical analysis
Thin layer chromatography (TLC)
Samples were processed according to the Hyperici herba monograph in the European Pharmacopoeia 8.031. Control solutions were prepared by mixing 500 mg of ground identified and vouchered material of selected Hypericum species with 10 ml of methanol (Analytical grade, Chimreactiv SRL, Romania) for 10 min at 60 °C. After cooling, the obtained solution was filtered. The reference solution was obtained by dissolving 5 mg of hyperoside (Analytical grade, Sigma-Aldrich) and 5 mg of rutin (Analytical grade, Sigma-Aldrich) in 5 ml of methanol. The test solutions of the Hypericum products were prepared in the same way as the control solution. Herbal products were processed depending on the pharmaceutical formulation and following the principles of the same extraction procedure, which were adapted to each pharmaceutical formulation. The analysis of the samples was performed in triplicate on SilicaGel plates (60 G F254, 20 × 20 cm, Merck), in twin bands of 10 mm, consisting of a 10 µl test sample and a 5 µl reference solution. As a mobile phase, a mixture of formic acid (Analytical grade, Nordic):distilled water:ethyl acetate (Analytical grade, Lachner) (6:9:90 V/V/V) was used. After migration of the principal components, the plates were dried at 100–105 °C for 10 min. Detection was performed in UV light, at 365 nm, after spraying with a mixture of 1% methanolic diphenylboryloxyetylamine (Sigma-Aldrich) and 5% methanolic polyethylene glycol 400 (Sigma-Aldrich) and 30 min incubation.
High-Performance Liquid Chromatography-Mass Spectrometry (HPLC-MS)
Hypericum extracts were prepared by adding 750 mg of powdered plant material of Hypericum species, H. perforatum and H. maculatum, to 15 ml methanol in a glass tube31, 57. The tubes were capped and agitated in the dark at 25 °C for 3 hours on a digital ceramic hotplate stirrer (Arec. X. Velp Scientifica). The extracts were filtered and diluted (1:100) in the mobile phase consisting of a mixture of 1 mM acetonitrile (Merck)/ammonium acetate (Merck) 45/50 (V/V) in double distilled, deionised water (Infusion Solution Laboratory of the University of Medicine and Pharmacy Cluj-Napoca - Romania), and 1 µl of the mixture was injected into the HPLC chromatographic system. Quantities of the herbal product test samples were individually adapted in order to identify and quantify the reference compounds and the extraction procedure was subsequently followed as described above for the references. The HPLC system used was an 1100 series Agilent Technologies model (Darmstadt, Germany) consisting of a G1312A binary pump, an in-line G1379A degasser, a G1329A autosampler, a G1316A column thermostat and an Agilent Ion Trap Detector 1100 SL. Chromatographic separation was performed on a Zorbax SB-C18 (50 mm × 2.1 mm i.d., 3.5 µm) column (Agilent Technologies) equipped with a Zorbax SB-C18 precolumn with the mobile phase above, at 45 °C with a flow rate of 0.6 ml/min. The detection of analytes was performed in triplicate in non-reactive MS2 mode for the quantification of hypericin (Hwi Analytik Gmbh) or in reactive MS2 mode for hyperforin (Sigma), negative ion ionisation, using an ion trap mass spectrometer equipped with an electrospray ionisation ion source (ESI): capillary +2500 V, nebulizer 40 psi (nitrogen), dry gas nitrogen at 8 l/min, dry gas temperature 350 °C.
Standard calibration curves were obtained by plotting the peak areas of standard concentrations of hypericin (10, 20, 50, 100, 200 and 500 ng/ml) and hyperforin (2, 4, 10, 20, 40 and 100 ng/ml) against their nominal concentrations. Two linear regression equations (R2 > 0.998) were obtained. Positive identification of the target compounds was performed by mass-spectrometry, and quantification of hypericin and hyperforin was based on peak area (RT, retention time of 1.1 and 2.3 min, respectively) in comparison with the standard curves.
Genetic analysis
DNA extraction and quantification
Total DNA was extracted using a modified CTAB extraction method from small amounts of each herbal product (about 300 mg)58. The substrate was homogenized using 2–3 zirconium grinding beads in a Mini-Beadbeater-1 (Biospec Products Inc., USA). The final elution volume was 100 μl, and extracted DNA was quantified using a Fragment Analyzer™ (Advanced Analytical Technologies, Inc., USA) and a DNF-488-33 HS Genomic DNA Reagent Kit (50 bp–40,000 bp).
Amplicon library preparation
All amplicon libraries were prepared using fusion PCR based on two nuclear ribosomal target sequences, internal transcribed spacers nrITS1 and nrITS2. PGM fusion primers were based on 17SE and 5.8 I1, and 5.8 I2 and 26 SE, respectively59. The forward primers were labeled with unique 10 bp multiplex identifier (MID) tags and the reverse primers with uniform truncated P1 (trP1) tags. Thermal cycling was carried out in 25 μl reaction volumes, and each reaction contained 5 μl 5X Q5 reaction buffer (New England Biolabs Inc, UK), 1.5 μl 10 μM of each primer (Biolegio, the Netherlands), 0.5 μl 10 mM dNTPs, 0.25 μl 20 U/μl Q5 High-Fidelity DNA Polymerase (New England Biolabs Inc, UK), 5 μl 5X Q5 High GC enhancer, 10.75 μl of Milli-Q ultrapure water and 0.5 μl of template DNA. The following thermocycling protocol was used: 30 s of initial denaturation at 98 °C, followed by 35 cycles of denaturation at 98 °C for 10 s, annealing at 30 s, and elongation at 72 °C for 30 s, followed by a final elongation step at 72 °C for 2 min. The annealing temperature was 56 °C for nrITS1, and 71 °C for nrITS2.
Equimolar pool preparation
The size, purity and the molar concentration (nmol/l) of each amplicon library was measured using a Fragment Analyzer™ (Advanced Analytical Technologies, Inc., USA) and a DNF-910 dsDNA Reagent Kit (35 bp–1,500 bp). An equimolar pool (2 ng/μl/library) was prepared from the amplicon libraries using the Biomek 4000 Laboratory Automation Workstation (Beckman Coulter, USA). Agencourt AMPure XP (Beckman Coulter, USA) was used for removal of unincorporated primers and nucleotides using the manufacturer’s instructions (Agencourt AMPure XP v. B37419AA). The total concentration of the purified pooled amplicon library stock and three serial dilutions (undiluted, 1/5, 1/10) were analyzed using the Fragment Analyzer™ (Advanced Analytical Technologies, Inc., USA) and DNF-488 High Sensitivity Genomic DNA Analysis Kit in order to identify the optimum concentration range for the template preparation.
High throughput sequencing
An Ion Chef (Life Technologies (LT), Thermo-Fisher Scientific, USA) was used to prepare pooled Ion AmpliSeq libraries (LT) for emulsion PCR and to load the sequencing chips. The input DNA template concentration was adjusted to the number of Ion Sphere Particles (ISPs) and added to the emulsion PCR master mix. The emulsion PCR was done using the Ion Chef, and template-positive ISPs were enriched and loaded on an Ion 318 v2 Chip (LT) and sequenced on an Ion Torrent Personal Genome Machine (LT) using an Ion PGM Sequencing 400 kit (LT). Sequencing read data was analyzed and demultiplexed into FASTQ files per sample using Torrent Suite version 5.0.4 (LT).
Bioinformatics analysis
FASTQ read files were processed using the HTS-barcode-checker pipeline60 available as a Galaxy pipeline at the Naturalis Biodiversity Center (http://145.136.240.164:8080/). Using the HTS pipeline, nrITS1 and nrITS2 primer sequences were used to demultiplex the sequencing reads per sample and to filter out reads that did not match any of the primers. PRINSEQ61 was used to determine filtering and trimming values based on read lengths and Phred read quality62. All reads with a mean Phred quality score of less than 26 were filtered out, as well as reads with a length of less than 300 bp. Remaining reads were trimmed to a maximum length of 440 bp for nrITS1 and 350 bp for nrITS2. CD-HIT-EST63 was used to cluster reads into molecular operational taxonomic units (MOTUs) defined by a sequence similarity of >99% and a minimum number of ten reads. The consensus sequences of non-singleton MOTUs were queried using BLAST64 against a reference nucleotide sequence database, with a maximum e-value of 0.05, a minimum hit length of 100 bp and sequence identity of >97%. The number of reads per MOTU, as well as the BLAST results per MOTU, were compiled using custom scripts from the HTS Barcode Checker pipeline60. The reference sequence database consisted of a local copy of the NCBI/GenBank nucleotide database.
Data availability
Ion-Torrent amplicon read data is deposited in DRYAD: doi:10.5061/dryad.32j7r.
References
Linde, K. St. John’s Wort – an overview. Forsch. Komplementärmedizin Res. Complement. Med. 16, 146–155 (2009).
Lynch, N. & Berry, D. Differences in perceived risks and benefits of herbal, over-the-counter conventional, and prescribed conventional, medicines, and the implications of this for the safe and effective use of herbal products. Complement. Ther. Med. 15, 84–91, doi:10.1016/j.ctim.2006.06.007 (2007).
Borrelli, F. & Izzo, A. A. Herb–drug Interactions with St John’s wort (Hypericum perforatum): an update on clinical observations. AAPS J. 11, 710–727, doi:10.1208/s12248-009-9146-8 (2009).
WHO. The World medicines situation 2011—Traditional medicines: global situation, issues and challenges. WHO Press, Geneva, Switzerland (2011).
Coghlan, M. L. et al. Deep sequencing of plant and animal DNA contained within traditional Chinese medicines reveals legality issues and health safety concerns. PLOS Genet. 8, e1002657, doi:10.1371/journal.pgen.1002657 (2012).
Coghlan, M. L. et al. Combined DNA, toxicological and heavy metal analyses provides an auditing toolkit to improve pharmacovigilance of traditional Chinese medicine (TCM). Sci. Rep. 5, 17475, doi:10.1038/srep17475 (2015).
Newmaster, S. G., Grguric, M., Shanmughanandhan, D., Ramalingam, S. & Ragupathy, S. DNA barcoding detects contamination and substitution in North American herbal products. BMC Med. 11, 222, doi:10.1186/1741-7015-11-222 (2013).
Saslis-Lagoudakis, C. H. et al. Identification of common horsetail (Equisetum arvense L.; Equisetaceae) using thin layer chromatography versus DNA barcoding. Sci. Rep 5, 11942, doi:10.1038/srep11942 (2015).
De Boer, H. J., Ichim, M. C. & Newmaster, S. G. DNA barcoding and pharmacovigilance of herbal medicines. Drug Saf. 38, 611–620, doi:10.1007/s40264-015-0306-8 (2015).
Farah, M. H. et al. Botanical nomenclature in pharmacovigilance and a recommendation for standardisation. Drug Saf. 29, 1023–1029, doi:10.2165/00002018-200629110-00002 (2006).
Gilbert, N. Regulations: Herbal medicine rule book. Nature 480, S98–S99, doi:10.1038/480S98a (2011).
Heubl, G. New aspects of DNA-based authentication of Chinese medicinal plants by molecular biological techniques. Planta Med. 76, 1963–1974, doi:10.1055/s-0030-1250519 (2010).
European Medicines Agency. EMEA/CVMP/815/00 - Guideline on specifications: test procedures and acceptance criteria for herbal substances, herbal preparations and herbal medicinal products, http://www.ema.europa.eu/docs/en_GB/document_library/Scientific_guideline/2009/09/WC500003393.pdf (2006).
De Boer, H. J., Ouarghidi, A., Martin, G., Abbad, A. & Kool, A. DNA barcoding reveals limited accuracy of dentifications based on folk taxonomy. PLOS ONE 9, e84291, doi:10.1371/journal.pone.0084291 (2014).
Little, D. P. & Jeanson, M. L. DNA barcode authentication of Saw Palmetto herbal dietary supplements. Sci. Rep. 3, 3518, doi:10.1038/srep03518 (2013).
Little, D. P. Authentication of Ginkgo biloba herbal dietary supplements using DNA barcoding. Genome 57, 513–516, doi:10.1139/gen-2014-0130 (2014).
Baker, D. A., Stevenson, D. W. & Little, D. P. DNA barcode identification of black cohosh herbal dietary supplements. J. AOAC Int. 95, 1023–1034, doi:10.5740/jaoacint.11-261 (2012).
Stoeckle, M. Y. et al. Commercial teas highlight plant DNA barcode identification successes and obstacles. Sci. Rep. 1, 42, doi:10.1038/srep00042 (2011).
Wallace, L. J. et al. DNA barcodes for everyday life: Routine authentication of natural health products. Food Res. Int. 49, 446–452, doi:10.1016/j.foodres.2012.07.048 (2012).
Taberlet, P., Coissac, E., Pompanon, F., Brochmann, C. & Willerslev, E. Towards next-generation biodiversity assessment using DNA metabarcoding. Mol. Ecol. 21, 2045–2050, doi:10.1111/j.1365-294X.2012.05470.x (2012).
Ivanova, N. V., Kuzmina, M. L., Braukmann, T. W. A., Borisenko, A. V. & Zakharov, E. V. Authentication of herbal supplements using next-generation sequencing. PLOS ONE 11, e0156426, doi:10.1371/journal.pone.0156426 (2016).
Cheng, X. et al. Biological ingredient analysis of traditional Chinese medicine preparation based on high-throughput sequencing: the story for Liuwei Dihuang Wan. Sci. Rep. 4, 5147, doi:10.1038/srep05147 (2014).
European Medicines Agency. EMA/HMPC/101303/2008 - Assessment report on Hypericum perforatum L., herba http://www.ema.europa.eu/docs/en_GB/document_library/Herbal_-_HMPC_assessment_report/2010/01/WC500059144.pdf (2009).
Butterweck, V. & Schmidt, M. St. John’s wort: Role of active compounds for its mechanism of action and efficacy. Wien. Med. Wochenschr. 157, 356–361, doi:10.1007/s10354-007-0440-8 (2007).
Wonnemann, M., Singer, A., Siebert, B. & Müller, W. E. Evaluation of synaptosomal uptake inhibition of most relevant constituents of St. John’s wort. Pharmacopsychiatry 34, 148–151, doi:10.1055/s-2001-15465 (2001).
Butterweck, V., Korte, B. & Winterhoff, H. Pharmacological and endocrine effects of Hypericum perforatum and hypericin after repeated treatment. Pharmacopsychiatry 34, 2–7, doi:10.1055/s-2001-15508 (2001).
Bonetto, N., Santelli, L., Battistin, L. & Cagnin, A. Serotonin syndrome and rhabdomyolysis induced by concomitant use of triptans, fluoxetine and hypericum. Cephalalgia Int. J. Headache 27, 1421–1423, doi:10.1111/j.1468-2982.2007.01430.x (2007).
Parker, V., Wong, A. H., Boon, H. S. & Seeman, M. V. Adverse reactions to St John’s wort. Can. J. Psychiatry Rev. Can. Psychiatr. 46, 77–79, doi:10.1177/070674370104600112 (2001).
Hall, S. D. et al. The interaction between St John’s wort and an oral contraceptive. Clin. Pharmacol. Ther. 74, 525–535, doi:10.1016/j.clpt.2003.08.009 (2003).
Murphy, P. A., Kern, S. E., Stanczyk, F. Z. & Westhoff, C. L. Interaction of St. John’s Wort with oral contraceptives: effects on the pharmacokinetics of norethindrone and ethinyl estradiol, ovarian activity and breakthrough bleeding. Contraception 71, 402–408, doi:10.1016/j.contraception.2004.11.004 (2005).
Council of Europe. European Pharmacopoeia. 8th Edition. Strasbourg: Council Of Europe (2014).
Crockett, S. L., Schaneberg, B. & Khan, I. A. Phytochemical profiling of new and old world Hypericum (St. John’s Wort) species. Phytochem. Anal. 16, 479–485, doi:10.1002/(ISSN)1099-1565 (2005).
Smelcerovic, A., Zuehlke, S., Spiteller, M., Raabe, N. & Özen, T. Phenolic constituents of 17 Hypericum species from Turkey. Biochem. Syst. Ecol. 36, 316–319, doi:10.1016/j.bse.2007.09.002 (2008).
Mártonfi, P., Repčák, M. & Mihoková, L. Hypericum maculatum Crantz subsp. maculatum x H. perforatum L. (Hypericaceae): Corroboration of natural hybridization by secondary metabolite analysis. Folia Geobot. 31, 245–250, doi:10.1007/BF02812067 (1996).
Noack, K. L. Geschlechtsverlust und Bastardierung beim Johanniskraut. Forsch Fortschr 17, 13–15 (1941).
Kathe, W., Honnef, S. & Heym, A. Medicinal and aromatic plants in Albania, Bosnia-Herzegovina, Bulgaria, Croatia and Romania. Bundesamt für Naturschutz-Skripten 91. Bonn: Federal Agency for Nature Conservation (2003).
Barnes, J., Anderson, L. A. & Phillipson, J. D. St John’s wort (Hypericum perforatum L.): a review of its chemistry, pharmacology and clinical properties. J. Pharm. Pharmacol. 53, 583–600, doi:10.1211/0022357011775910 (2001).
Greeson, J. M., Sanford, B. & Monti, D. A. St. John’s wort (Hypericum perforatum): a review of the current pharmacological, toxicological, and clinical literature. Psychopharmacology (Berl.) 153, 402–414, doi:10.1007/s002130000625 (2001).
Smelcerovic, A. & Spiteller, M. Phytochemical analysis of nine Hypericum L. species from Serbia and the F.Y.R. Macedonia. Int. J. Pharm. Sci. 61, 251–252 (2006).
Umek, A., Kreft, S., Kartnig, T. & Heydel, B. Quantitative phytochemical analyses of six Hypericum species growing in Slovenia. Planta Med. 65, 388–390, doi:10.1055/s-2006-960798 (1999).
Hölzl, J. & Ostrowski, H. Johanniskraut (Hypericum perforatum L.) HPLC-Analyse der wichtigen Inhaltsstoffe und deren Variabilität in einer Population. Dtsch. Apoth.-Ztg. 127, 1227–1230 (1987).
Smelcerovic, A. et al. Phytochemical analysis and genetic characterization of six Hypericum species from Serbia. Phytochemistry 67, 171–177, doi:10.1016/j.phytochem.2005.10.021 (2006).
Wurglics, M. et al. Comparison of German St. John’s wort products according to hyperforin and total hypericin content. J. Am. Pharm. Assoc. Washington DC 1996 41, 560–566, doi:10.1016/S1086-5802(16)31280-3 (2001).
Kim, H. K., Choi, Y. H. & Verpoorte, R. NMR-based metabolomic analysis of plants. Nat. Protoc. 5, 536–549, doi:10.1038/nprot.2009.237 (2010).
Hawkins, J. et al. Using DNA metabarcoding to identify the floral composition of honey: a new tool for investigating honey bee foraging preferences. PLOS ONE 10, e0134735, doi:10.1371/journal.pone.0134735 (2015).
Richardson, R. T. et al. Application of ITS2 metabarcoding to determine the provenance of pollen collected by honey bees in an agroecosystem. Appl. Plant Sci. 3, 1400066, doi:10.3732/apps.1400066 (2015).
Galimberti, A. et al. DNA barcoding as a new tool for food traceability. Food Res. Int. 50, 55–63, doi:10.1016/j.foodres.2012.09.036 (2013).
Sickel, W. et al. Increased efficiency in identifying mixed pollen samples by meta-barcoding with a dual-indexing approach. BMC Ecol. 15, 20, doi:10.1186/s12898-015-0051-y (2015).
Novak, J., Grausgruber-Gröger, S. & Lukas, B. DNA-based authentication of plant extracts. Food Res. Int. 40, 388–392, doi:10.1016/j.foodres.2006.10.015 (2007).
Pawluczyk, M. et al. Quantitative evaluation of bias in PCR amplification and next-generation sequencing derived from metabarcoding samples. Anal. Bioanal. Chem. 407, 1841–1848, doi:10.1007/s00216-014-8435-y (2015).
Meseguer, A. S., Aldasoro, J. J. & Sanmartín, I. Bayesian inference of phylogeny, morphology and range evolution reveals a complex evolutionary history in St. John’s wort (Hypericum). Mol. Phylogenet. Evol. 67, 379–403, doi:10.1016/j.ympev.2013.02.007 (2013).
Nürk, N. M., Madriñán, S., Carine, M. A., Chase, M. W. & Blattner, F. R. Molecular phylogenetics and morphological evolution of St. John’s wort (Hypericum; Hypericaceae). Mol. Phylogenet. Evol. 66, 1–16, doi:10.1016/j.ympev.2012.08.022 (2013).
Aylagas, E., Borja, Á., Irigoien, X. & Rodríguez-Ezpeleta, N. Benchmarking DNA Metabarcoding for biodiversity-based monitoring and assessment. Front. Mar. Sci. 3, 679–688, doi:10.3389/fmars.2016.00096 (2016).
Palhares, R. M. et al. Medicinal plants recommended by the World Health Organization: DNA barcode Identification associated with chemical analyses guarantees their quality. PLOS ONE 10, e0127866, doi:10.1371/journal.pone.0127866 (2015).
Techen, N., Crockett, S. L., Khan, I. A. & Scheffler, B. E. Authentication of medicinal plants using molecular biology techniques to compliment conventional methods. Curr. Med. Chem. 11, 1391–1401, doi:10.2174/0929867043365206 (2004).
Techen, N., Parveen, I., Pan, Z. & Khan, I. A. DNA barcoding of medicinal plant material for identification. Curr. Opin. Biotechnol. 25, 103–110, doi:10.1016/j.copbio.2013.09.010 (2014).
Cossuta, D. et al. Extraction of hyperforin and hypericin from St. John’s wort (Hypericum perforatum L.) with different solvents. J. Food Process Eng. 35, 222–235, doi:10.1111/jfpe.2012.35.issue-2 (2012).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19, 11–15 (1987).
Sun, Y., Skinner, D. Z., Liang, G. H. & Hulbert, S. H. Phylogenetic analysis of Sorghum and related taxa using internal transcribed spacers of nuclear ribosomal DNA. Theor. Appl. Genet. 89, 26–32, doi:10.1007/BF00226978 (1994).
Lammers, Y., Peelen, T., Vos, R. A. & Gravendeel, B. The HTS barcode checkerpipeline, a tool for automated detection of illegally traded species from high-throughput sequencing data. BMC Bioinformatics 15, 44, doi:10.1186/1471-2105-15-44 (2014).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864, doi:10.1093/bioinformatics/btr026 (2011).
Ewing, B. & Green, P. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8, 186–194, doi:10.1101/gr.8.3.186 (1998).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659, doi:10.1093/bioinformatics/btl158 (2006).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410, doi:10.1016/S0022-2836(05)80360-2 (1990).
Acknowledgements
The authors acknowledge funding from the Romanian - EEA Research Programme operated by the MECS-ANCSI PO under the EEA Financial Mechanism 2009–2014 and project contract number 2SEE/2014.
Author information
Authors and Affiliations
Contributions
A.C.R., A.K.B., M.C.I., H.d.B. conceived the experiment. A.C.R. carried out the molecular lab work. R.P., L.V., G.C. carried out the chemical lab work and analysis. A.C.R., A.L., V.M. carried out the analysis of the molecular lab work. A.C.R. and H.d.B. wrote the manuscript, and all authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Raclariu, A.C., Paltinean, R., Vlase, L. et al. Comparative authentication of Hypericum perforatum herbal products using DNA metabarcoding, TLC and HPLC-MS. Sci Rep 7, 1291 (2017). https://doi.org/10.1038/s41598-017-01389-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-01389-w
This article is cited by
-
Molecular approach for identification and validation of Eurycoma longifolia, Labisia pumila, and Orthosiphon stamineus in herbal products
Beni-Suef University Journal of Basic and Applied Sciences (2024)
-
Authentication of milk thistle commercial products using UHPLC-QTOF-ESI + MS metabolomics and DNA metabarcoding
BMC Complementary Medicine and Therapies (2023)
-
Comprehensive analysis using DNA metabarcoding, SCAR marker based PCR assay, and HPLC unveils the adulteration in Brahmi herbal products
Molecular Biology Reports (2023)
-
Application of DNA Barcoding for Quality Control of Herbal Drugs and Their Phytopharmaceuticals
Revista Brasileira de Farmacognosia (2021)
-
Quality control of the traditional herbs and herbal products: a review
Future Journal of Pharmaceutical Sciences (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.