Abstract
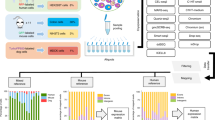
Human tissues comprise trillions of cells that populate a complex space of molecular phenotypes and functions and that vary in abundance by 4–9 orders of magnitude. Relying solely on unbiased sampling to characterize cellular niches becomes infeasible, as the marginal utility of collecting more cells diminishes quickly. Furthermore, in many clinical samples, the relevant cell types are scarce and efficient processing is critical. We developed an integrated pipeline for index sorting and massively parallel single-cell RNA sequencing (MARS-seq2.0) that builds on our previously published MARS-seq approach. MARS-seq2.0 is based on >1 million cells sequenced with this pipeline and allows identification of unique cell types across different tissues and diseases, as well as unique model systems and organisms. Here, we present a detailed step-by-step procedure for applying the method. In the improved procedure, we combine sub-microliter reaction volumes, optimization of enzymatic mixtures and an enhanced analytical pipeline to substantially lower the cost, improve reproducibility and reduce well-to-well contamination. Data analysis combines multiple layers of quality assessment and error detection and correction, graphically presenting key statistics for library complexity, noise distribution and sequencing saturation. Importantly, our combined FACS and single-cell RNA sequencing (scRNA-seq) workflow enables intuitive approaches for depletion or enrichment of cell populations in a data-driven manner that is essential to efficient sampling of complex tissues. The experimental protocol, from cell sorting to a ready-to-sequence library, takes 2–3 d. Sequencing and processing the data through the analytical pipeline take another 1–2 d.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The sequenced data analyzed in this study have been deposited in GEO: GSE123392. Published data analyzed in this paper are available at https://doi.org/10.1016/j.cell.2015.11.013. Access to all published sequenced data generated using the method described here can be found within the relevant publications.
Code availability
Users can access the code freely on our website: http://compgenomics.weizmann.ac.il/tanay/?page_id=672.
References
Tang, F. et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat. Methods 6, 377–382 (2009).
Tang, F. et al. RNA-Seq analysis to capture the transcriptome landscape of a single cell. Nat. Protoc. 5, 516–535 (2010).
Shalek, A. K. et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498, 236–240 (2013).
Moignard, V. et al. Characterization of transcriptional networks in blood stem and progenitor cells using high-throughput single-cell gene expression analysis. Nat. Cell Biol. 15, 363–372 (2013).
Deng, Q., Ramskold, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014).
Treutlein, B. et al. Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375 (2014).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Paul, F. et al. Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell 163, 1663–1677 (2015).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Pollen, A. A. et al. Low-coverage single-cell mRNA sequencing reveals cellular heterogeneity and activated signaling pathways in developing cerebral cortex. Nat. Biotechnol. 32, 1053–1058 (2014).
Grun, D. et al. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature 525, 251–255 (2015).
Petropoulos, S. et al. Single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 167, 285 (2016).
Grun, D. & van Oudenaarden, A. Design and analysis of single-cell sequencing experiments. Cell 163, 799–810 (2015).
Stegle, O., Teichmann, S. A. & Marioni, J. C. Computational and analytical challenges in single-cell transcriptomics. Nat. Rev. Genet. 16, 133–145 (2015).
Shapiro, E., Biezuner, T. & Linnarsson, S. Single-cell sequencing-based technologies will revolutionize whole-organism science. Nat. Rev. Genet. 14, 618–630 (2013).
Kolodziejczyk, A. A., Kim, J. K., Svensson, V., Marioni, J. C. & Teichmann, S. A. The technology and biology of single-cell RNA sequencing. Mol. Cell 58, 610–620 (2015).
Islam, S. et al. Characterization of the single-cell transcriptional landscape by highly multiplex RNA-seq. Genome Res. 21, 1160–1167 (2011).
Hashimshony, T., Wagner, F., Sher, N. & Yanai, I. CEL-Seq: single-cell RNA-Seq by multiplexed linear amplification. Cell Rep. 2, 666–673 (2012).
Islam, S. et al. Highly multiplexed and strand-specific single-cell RNA 5′ end sequencing. Nat. Protoc. 7, 813–828 (2012).
Ramskold, D. et al. Full-length mRNA-Seq from single-cell levels of RNA and individual circulating tumor cells. Nat. Biotechnol. 30, 777–782 (2012).
Sasagawa, Y. et al. Quartz-Seq: a highly reproducible and sensitive single-cell RNA sequencing method, reveals non-genetic gene-expression heterogeneity. Genome Biol. 14, R31 (2013).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Islam, S. et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat. Methods 11, 163–166 (2014).
Wu, A. R. et al. Quantitative assessment of single-cell RNA-sequencing methods. Nat. Methods 11, 41–46 (2014).
Levine, J. H. et al. Data-driven phenotypic dissection of AML reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Gierahn, T. M. et al. Seq-Well: portable, low-cost RNA sequencing of single cells at high throughput. Nat. Methods 14, 395–398 (2017).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Regev, A. et al. The Human Cell Atlas. Elife 6, e27041 (2017).
Trapnell, C. Defining cell types and states with single-cell genomics. Genome Res. 25, 1491–1498 (2015).
Ballas, S. K. Erythrocyte concentration and volume are inversely related. Clin. Chim. Acta 164, 243–244 (1987).
Giladi, A. et al. Single-cell characterization of haematopoietic progenitors and their trajectories in homeostasis and perturbed haematopoiesis. Nat. Cell Biol. 20, 836–846 (2018).
Medaglia, C. et al. Spatial reconstruction of immune niches by combining photoactivatable reporters and scRNA-seq. Science 358, 1622–1626 (2017).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Jaitin, D. A. et al. Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-seq. Cell 167, 1883–1896.e15 (2016).
Paul, F. & Amit, I. Plasticity in the transcriptional and epigenetic circuits regulating dendritic cell lineage specification and function. Curr. Opin. Immunol. 30, 1–8 (2014).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s Disease. Cell 169, 1276–1290.e17 (2017).
Bornstein, C. et al. Single-cell mapping of the thymic stroma identifies IL-25-producing tuft epithelial cells. Nature 559, 622–626 (2018).
Cohen, M. et al. Lung single-cell signaling interaction map reveals basophil role in macrophage imprinting. Cell 175, 1031–1044.e18 (2018).
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 (2016).
Steuerman, Y. et al. Dissection of influenza infection in vivo by single-cell RNA sequencing. Cell Syst. 6, 679–691.e4 (2018).
Sebe-Pedros, A. et al. Cnidarian cell type diversity and regulation revealed by whole-organism single-cell RNA-seq. Cell 173, 1520–1534.e20 (2018).
Sebe-Pedros, A. et al. Early metazoan cell type diversity and the evolution of multicellular gene regulation. Nat. Ecol. Evol. 2, 1176–1188 (2018).
Ledergor, G. et al. Single cell dissection of plasma cell heterogeneity in symptomatic and asymptomatic myeloma. Nat. Med. 24, 1867–1876 (2018).
Lavin, Y. et al. Innate immune landscape in early lung adenocarcinoma by paired single-cell analyses. Cell 169, 750–765.e17 (2017).
Li, H. et al. Dysfunctional CD8 T cells form a proliferative, dynamically regulated compartment within human melanoma. Cell 176, 775–789.e18 (2019).
Halpern, K. B. et al. Paired-cell sequencing enables spatial gene expression mapping of liver endothelial cells. Nat. Biotechnol. 36, 962–970 (2018).
Halpern, K. B. et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature 542, 352–356 (2017).
Kang, H. M. et al. Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat. Biotechnol. 36, 89–94 (2018).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Han, K. Y. et al. SIDR: simultaneous isolation and parallel sequencing of genomic DNA and total RNA from single cells. Genome Res. 28, 75–87 (2018).
Biase, F. H. et al. Rainbow-Seq: combining cell lineage tracing with single-cell RNA sequencing in preimplantation embryos. iScience 7, 16–29 (2018).
Giladi, A. & Amit, I. Single-cell genomics: a stepping stone for future immunology discoveries. Cell 172, 14–21 (2018).
Wu, Y. E., Pan, L., Zuo, Y., Li, X. & Hong, W. Detecting activated cell populations using single-cell RNA-seq. Neuron 96, 313–329.e6 (2017).
Grun, D., Kester, L. & van Oudenaarden, A. Validation of noise models for single-cell transcriptomics. Nat. Methods 11, 637–640 (2014).
Kivioja, T. et al. Counting absolute numbers of molecules using unique molecular identifiers. Nat. Methods 9, 72–74 (2011).
Rais, Y. et al. Deterministic direct reprogramming of somatic cells to pluripotency. Nature 502, 65–70 (2013).
Acknowledgements
We thank the J. H. Hanna lab for the donation of v6.5 mouse embryonic stem (ES) cells, DR4 mouse embryonic fibroblast (MEF) cells and WIBR3 human ES cells. We thank G. Brodsky for help with the artwork. This work was funded by the Chan Zuckerberg Initiative (CZI) (A.T. and I.A.), European Research Council consolidator grants (A.T. and I.A.), ERC-COG (724824-scAssembly; A.T.), ERC-COG (724471-HemTree2.0; I.A.) and the I-Core program (A.T. and I.A.).
Author information
Authors and Affiliations
Contributions
H.K.-S. developed the experimental protocols; designed, performed, and analyzed experiments; and wrote the manuscript. E.K. designed experiments, developed the computational methods, and performed bioinformatics analysis. D.A.J. developed experimental protocols. E.D. performed bioinformatic analysis. F.P. contributed to the experimental design. A.T. directed the project and developed computational methods. I.A. directed the project and designed the experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Jaitin, D. A. et al. Cell 167, 1883–1896.e15 (2016): https://doi.org/10.1016/j.cell.2016.11.039
Keren-Shaul, H. et al. Cell 169, 1276–1290.e17 (2017): https://doi.org/10.1016/j.cell.2017.05.018
Medaglia, C. et al. Science 358, 1622–1626 (2017): https://doi.org/10.1126/science.aao4277
Integrated supplementary information
Supplementary Figure 1 Statistics for 1 million cells analyzed by MARS-seq2.0.
UMI and gene counts distributions on 1,033,944 mouse cells using genome assembly mm9 and 16,488 empty wells analyzed by MARS-seq2.0. Cells analyzed were either immune, epithelial, fibroblast, stromal, hepatocytes or cancer mouse cells as well as cell lines (mouse ES and MEF). (A-B) A histogram showing the fraction of cells (black) and empty wells (red) binned by logarithmic intervals of UMI-counts (A) or by gene-counts (B). (C) Empty wells are enriched with lowly detected genes. Histogram showing the fraction of cells/wells (similar to A-B) binned by the average number of UMIs observed per gene (log2 scale).
Supplementary Figure 2 Optimization of MARS-seq.
(A) Background noise estimation for high and low volume protocol versions. Shown is the average number of UMIs in wells containing single mouse ES (y-axis) vs. an average number of UMIs in negative control wells (x-axis) for two protocol versions with high (green) and low (black) liquid volumes in the RT step. (B) Background noise estimation for different second strand synthesis enzymes mixtures. Shown is the average number of UMIs in common myeloid progenitor cells (CMP; y-axis) vs. an average number of UMIs in negative controls wells (x-axis). Second strand synthesis was performed with ‘A’ composition (green symbols), with addition of RNaseH or without DNA polymerase, or with ‘B’ composition (black symbols) in dilutions of 1:4 or 1:8. Dashed lines demarcate background noise levels of 5% (red) and 10% (magenta).
Supplementary Figure 3 MARS-seq2.0: method overview.
(A) Molecular yield. Shown is a cumulative cell percentage (y-axis) vs. the total number of detected molecules per cell (x-axis). (B) Sequencing depth. Shown is a cumulative cell percentage (y-axis) vs. the total number of reads per cell (x-axis) for amplification batches of FACS sorted mouse ES (black curves) and MEF (red curves). (C) Total number of molecules detected vs. flow cytometry forward scatter values. Data was analyzed from 5500 bone marrow myeloid progenitors index sorted single cells (Lineage- c-Kit+ Sca1+) generated as previously reported9. (D) Average gene expression (number of detected molecules per gene divided by the total number of molecules) for two randomly selected groups of 100 mouse ES single-cells (x and y axis) is shown for 7567 genes with average expression higher than 10−6 (E) Similarly to D, showing average gene expression over groups of MEF (y-axis) vs. mouse ES (x-axis) single cells for 2350 differentially expressed genes (fold change >4 and FDR<10-5; red dots) and other genes (black dots). (F-I) Absolute molecules count for TPM1 (y-axis) vs. KLF5 (x-axis) (F), Id3 vs. Oct4 (G), Lgals1 vs. Dppa4 (H), and Actn1 vs. Dphs1 (I) in 1128 mouse ES cells (black dots) and 752 MEF cells (red dots).
Supplementary Figure 4 Evaluation of doublet rate in MARS-seq2.0.
Total number of molecules identified for each species in a 1:1 mouse/human mixing experiment (mouse ES/ human ES). Two out of 1041 single cells analyzed were identified as doublets.
Supplementary Figure 5 Modeling of single-cell RNA-seq data.
A schematic diagram of MARS-seq2.0 Library preparation is shown as a sequential sampling process. We model our RNA-seq data as a sequential process of multiple samplings and amplifications steps. During the first RT conversion and second strand synthesis, the mRNA molecules (labeled mRNA A, B, C, D, E and F) of the cell (transcriptome) are sampled and tagged with a barcode and a unique molecular identifier (UMI, ‘tagged mRNA pool’). Tagged molecules which are schematically represented by UMI A, C, and D are stochastically linearly amplified by IVT to produce multiple copies of each molecule (‘IVT products’) that are sheared at random positions (see Pos 1–6). IVT products will be further exponentially amplified using PCR which involves repeated cycles of sampling and amplification that will be followed by an additional random sampling of ~10K PCR products (reads) per cell that will be sequenced (x5, x10, x1, and x2 represent the number of PCR products of each IVT product). The pipeline quantifies the number of tagged mRNA molecules per gene that went through this complex sampling process while eliminating several types of experimental artifacts.
Supplementary Figure 6 A representative diagnostics report for an amplification batch of mouse ES cells.
(A) Sequencing depth per cell. Shown is a cumulative cell percentage (y-axis) vs. the total number of reads per cell (x-axis). (B) Mapping analysis. Shown are the fractions of reads per cell mapped to exonic loci, spike-ins, or unmapped due to multiple mapping or low MAPQ score. Cells are ordered according to gene mapping fractions. (C) Oligo contamination gauge. Shown are fractions of RT primer, poly A sequences and other oligo sequences within the unmapped reads pool. (D) UMI nucleotide composition. Nucleotide composition (y-axis) for all UMI positions (x-axis). (E-G) Error distributions. Cumulative cell percentage (y-axis) vs. the fraction of molecules that were filtered (x-axis) due to sequencing errors in the UMI (E), cell barcode (F) or template switching errors (G). (H) Negative control wells. The number of unique UMIs mapped to genes (blue) and spike-ins (red) that have a cell barcode associated with four negative control wells. (I) Molecular yield vs. technical efficiency. Shown is the number of detected mouse mRNA molecules (y-axis) vs. the number of spike-ins molecules (x-axis) detected in four wells following sorting of single cells (black dots) and four negative control wells that do not contain single cells (red signs). This visualization highlights potential problems of background noise and failed sorting. (J) Proportion molecules with a single IVT product. The percentage of detected molecules with a single IVT product (single offset; y-axis) vs. the total number of detected molecules (x-axis) per cell (black dots) or in empty wells (red signs). (K-L) Plate visualization. Normalized number of extracted gene (K) or spike-in (L) molecules (blue - low, red - high) in wells (ordered according to the physical plate positions) that were pooled and amplified together (single amplification batch). This visualization allows identification of sorting or robot related problems. (M-N) Molecules per cell. Shown is a cumulative cell percentage (y-axis) vs. the total number of detected gene (M) or spike-in (N) molecules per cell (x-axis). (O) IVT products per molecule. A histogram of the number of IVT products (y-axis, logarithmic scale) per UMI. (P) Reads per molecule. A histogram of the number of reads (y-axis, logarithmic scale) per UMI. (Q) Highly expressed genes. The average number of detected molecules (log2, y-axis) for the 25 genes (x-axis) with the highest expression levels. (R) Highly variable genes. The variance of detected number of molecules divided by the average number of detected molecules (log2, y-axis) for the 25 genes (x-axis) with the highest variance/mean score.
Supplementary Figure 7 A representative diagnostics report for an amplification batch of MEF cells.
Same as supplementary figure 6.
Supplementary Figure 8 A representative diagnostics report for an amplification batch of mixed mouse ES and MEF cells.
Same as supplementary figure 6.
Supplementary Figure 9 Sequencing depth of mouse ES single cells.
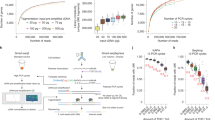
(A) Marginal sequencing efficiency. Shown is the number of molecules detected (y-axis) vs. the total number of reads following logarithmic down-sampling of the reads (black dots) as well as exponential fit (dashed red curve). Dashed gray line demarcates the saturation level of the library (8250 molecules). (B) Similarly to A, showing saturation curves for six cell groups divided according to their saturation level (color coded lines), indicating sequencing depth that was sufficient to sequence 50%, 75%, and 90% of the molecules in the library (circle, triangle and diamond, respectively). Inferred number of molecules per cell assuming saturation is indicated on the right margin for the six cell groups. A magnified view (right panel) of the marked area in the bottom is highlighting saturation curves for cells with low complexity.
Supplementary Figure 10 A representative successful MARS-seq2.0 library profile.
TapeStation profile of a MARS-seq library generated from mouse ES cells. ‘Lower’ and ‘Upper’ represent internal instrument markers. The average library size (384 bp) is indicated.
Supplementary information
Supplementary Information
Supplementary Figures 1–10, Supplementary Manuals 1–3 and Supplementary Methods
Supplementary Software
Bravo robot scripts for MARS-seq2.0
Supplementary Table 1
List of RT1 primers.
Supplementary Table 2
List of ligation adaptor primers.
Rights and permissions
About this article
Cite this article
Keren-Shaul, H., Kenigsberg, E., Jaitin, D.A. et al. MARS-seq2.0: an experimental and analytical pipeline for indexed sorting combined with single-cell RNA sequencing. Nat Protoc 14, 1841–1862 (2019). https://doi.org/10.1038/s41596-019-0164-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0164-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.