Abstract
UV-DDB, a key protein in human global nucleotide excision repair (NER), binds avidly to abasic sites and 8-oxo-guanine (8-oxoG), suggesting a noncanonical role in base excision repair (BER). We investigated whether UV-DDB can stimulate BER for these two common forms of DNA damage, 8-oxoG and abasic sites, which are repaired by 8-oxoguanine glycosylase (OGG1) and apurinic/apyrimidinic endonuclease (APE1), respectively. UV-DDB increased both OGG1 and APE1 strand cleavage and stimulated subsequent DNA polymerase β-gap filling activity by 30-fold. Single-molecule real-time imaging revealed that UV-DDB forms transient complexes with OGG1 or APE1, facilitating their dissociation from DNA. Furthermore, UV-DDB moves to sites of 8-oxoG repair in cells, and UV-DDB depletion sensitizes cells to oxidative DNA damage. We propose that UV-DDB is a general sensor of DNA damage in both NER and BER pathways, facilitating damage recognition in the context of chromatin.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Friedberg, E. C. et al. DNA repair: from molecular mechanism to human disease. DNA Repair (Amst.) 5, 986–996 (2006).
Hill, J. W., Hazra, T. K., Izumi, T. & Mitra, S. Stimulation of human 8-oxoguanine-DNA glycosylase by AP-endonuclease: potential coordination of the initial steps in base excision repair. Nucleic Acids Res. 29, 430–438 (2001).
Rodriguez, Y. & Smerdon, M. J. The structural location of DNA lesions in nucleosome core particles determines accessibility by base excision repair enzymes. J. Biol. Chem. 288, 13863–13875 (2013).
Odell, I. D., Wallace, S. S. & Pederson, D. S. Rules of engagement for base excision repair in chromatin. J. Cell Physiol. 228, 258–266 (2013).
Odell, I. D., Newick, K., Heintz, N. H., Wallace, S. S. & Pederson, D. S. Non-specific DNA binding interferes with the efficient excision of oxidative lesions from chromatin by the human DNA glycosylase, NEIL1. DNA Repair (Amst.) 9, 134–143 (2010).
Prasad, A., Wallace, S. S. & Pederson, D. S. Initiation of base excision repair of oxidative lesions in nucleosomes by the human, bifunctional DNA glycosylase NTH1. Mol. Cell Biol. 27, 8442–8453 (2007).
Hinz, J. M. Impact of abasic site orientation within nucleosomes on human APE1 endonuclease activity. Mutat. Res. 766-767, 19–24 (2014).
Bilotti, K., Tarantino, M. E. & Delaney, S. Human oxoguanine glycosylase 1 removes solution accessible 8-oxo-7,8-dihydroguanine lesions from globally substituted nucleosomes except in the dyad region. Biochemistry 57, 1436–1439 (2018).
Bilotti, K., Kennedy, E. E., Li, C. & Delaney, S. Human OGG1 activity in nucleosomes is facilitated by transient unwrapping of DNA and is influenced by the local histone environment. DNA Repair (Amst.) 59, 1–8 (2017).
Maher, R. L. et al. Human cells contain a factor that facilitates the DNA glycosylase-mediated excision of oxidized bases from occluded sites in nucleosomes. DNA Repair (Amst.) 57, 91–97 (2017).
Czaja, W., Mao, P. & Smerdon, M. J. The emerging roles of ATP-dependent chromatin remodeling enzymes in nucleotide excision repair. Int. J. Mol. Sci. 13, 11954–11973 (2012).
Rodriguez, Y., Hinz, J. M. & Smerdon, M. J. Accessing DNA damage in chromatin: preparing the chromatin landscape for base excision repair. DNA Repair (Amst.) 32, 113–119 (2015).
Balliano, A. J. & Hayes, J. J. Base excision repair in chromatin: Insights from reconstituted systems. DNA Repair (Amst.) 36, 77–85 (2015).
Peterson, C. L. & Almouzni, G. Nucleosome dynamics as modular systems that integrate DNA damage and repair. Cold Spring Harbor Perspect. Biol. https://doi.org/10.1101/cshperspect.a012658 (2013).
Polo, S. E. & Almouzni, G. Chromatin dynamics after DNA damage: The legacy of the access-repair-restore model. DNA Repair (Amst.) 36, 114–121 (2015).
Hinz, J. M. & Czaja, W. Facilitation of base excision repair by chromatin remodeling. DNA Repair (Amst.) 36, 91–97 (2015).
Gong, F., Kwon, Y. & Smerdon, M. J. Nucleotide excision repair in chromatin and the right of entry. DNA Repair (Amst.) 4, 884–896 (2005).
Waters, R., van Eijk, P. & Reed, S. Histone modification and chromatin remodeling during NER. DNA Repair (Amst.) 36, 105–113 (2015).
Sugasawa, K. UV-DDB: a molecular machine linking DNA repair with ubiquitination. DNA Repair (Amst.) 8, 969–972 (2009).
Rapic-Otrin, V., McLenigan, M. P., Bisi, D. C., Gonzalez, M. & Levine, A. S. Sequential binding of UV DNA damage binding factor and degradation of the p48 subunit as early events after UV irradiation. Nucleic Acids Res. 30, 2588–2598 (2002).
Fischer, E. S. et al. The molecular basis of CRL4DDB2/CSA ubiquitin ligase architecture, targeting, and activation. Cell 147, 1024–1039 (2011).
Kapetanaki, M. G. et al. The DDB1-CUL4ADDB2 ubiquitin ligase is deficient in xeroderma pigmentosum group E and targets histone H2A at UV-damaged DNA sites. Proc. Natl Acad. Sci. USA 103, 2588–2593 (2006).
Kondo, S. et al. Assignment of three patients with xeroderma pigmentosum to complementation group E and their characteristics. J. Invest Dermatol 90, 152–157 (1988).
Rapic Otrin, V. et al. Relationship of the xeroderma pigmentosum group E DNA repair defect to the chromatin and DNA binding proteins UV-DDB and replication protein A. Mol. Cell Biol. 18, 3182–3190 (1998).
Dualan, R. et al. Chromosomal localization and cDNA cloning of the genes (DDB1 and DDB2) for the p127 and p48 subunits of a human damage-specific DNA binding protein. Genomics 29, 62–69 (1995).
Rapic-Otrin, V. et al. True XP group E patients have a defective UV-damaged DNA binding protein complex and mutations in DDB2 which reveal the functional domains of its p48 product. Hum. Mol. Genet. 12, 1507–1522 (2003).
Oh, K. S. et al. Nucleotide excision repair proteins rapidly accumulate but fail to persist in human XP-E (DDB2 mutant) cells. Photochem Photobio. 87, 729–733 (2011).
Wittschieben, B. O., Iwai, S. & Wood, R. D. DDB1-DDB2 (xeroderma pigmentosum group E) protein complex recognizes a cyclobutane pyrimidine dimer, mismatches, apurinic/apyrimidinic sites, and compound lesions in DNA. J. Biol. Chem. 280, 39982–39989 (2005).
Wakasugi, M. et al. DDB accumulates at DNA damage sites immediately after UV irradiation and directly stimulates nucleotide excision repair. J. Biol. Chem. 277, 1637–1640 (2002).
Reardon, J. T. et al. Comparative analysis of binding of human damaged DNA-binding protein (XPE) and Escherichia coli damage recognition protein (UvrA) to the major ultraviolet photoproducts: T[c,s]T, T[t,s]T, T[6-4]T, and T[Dewar]T. J. Biol. Chem. 268, 21301–21308 (1993).
Yeh, J. I. et al. Damaged DNA induced UV-damaged DNA-binding protein (UV-DDB) dimerization and its roles in chromatinized DNA repair. Proc. Natl Acad. Sci. USA 109, E2737–E2746 (2012).
Scrima, A. et al. Structural basis of UV DNA-damage recognition by the DDB1-DDB2 complex. Cell 135, 1213–1223 (2008).
Ghodke, H. et al. Single-molecule analysis reveals human UV-damaged DNA-binding protein (UV-DDB) dimerizes on DNA via multiple kinetic intermediates. Proc. Natl Acad. Sci. USA 111, E1862–E1871 (2014).
D’Errico, M. et al. New functions of XPC in the protection of human skin cells from oxidative damage. EMBO J. 25, 4305–4315 (2006).
Parlanti, E. et al. The cross talk between pathways in the repair of 8-oxo-7,8-dihydroguanine in mouse and human cells. Free Radic. Biol. Med 53, 2171–2177 (2012).
He, J. et al. A genetically targetable near-infrared photosensitizer. Nat. Methods 13, 263–268 (2016).
Fouquerel, E. et al. Targeted and persistent 8-oxoguanine base damage at telomeres promotes telomere loss and crisis. Mol. Cell. https://doi.org/10.1016/j.molcel.2019.04.024 (2019).
Fujiwara, Y. et al. Characterization of DNA recognition by the human UV-damaged DNA-binding protein. J. Biol. Chem. 274, 20027–20033 (1999).
Prasad, R., Shock, D. D., Beard, W. A. & Wilson, S. H. Substrate channeling in mammalian base excision repair pathways: passing the baton. J. Biol. Chem. 285, 40479–40488 (2010).
Sidorenko, V. S., Nevinsky, G. A. & Zharkov, D. O. Mechanism of interaction between human 8-oxoguanine-DNA glycosylase and AP endonuclease. DNA Repair (Amst.) 6, 317–328 (2007).
Mokkapati, S. K., Wiederhold, L., Hazra, T. K. & Mitra, S. Stimulation of DNA glycosylase activity of OGG1 by NEIL1: functional collaboration between two human DNA glycosylases. Biochemistry 43, 11596–11604 (2004).
Prasad, R., Dyrkheeva, N., Williams, J. & Wilson, S. H. Mammalian base excision repair: functional partnership between PARP-1 and APE1 in AP-site repair. PloS ONE 10, e0124269 (2015).
Gembka, A. et al. The checkpoint clamp, Rad9-Rad1-Hus1 complex, preferentially stimulates the activity of apurinic/apyrimidinic endonuclease 1 and DNA polymerase beta in long patch base excision repair. Nucleic Acids Res. 35, 2596–2608 (2007).
de Souza-Pinto, N. C. et al. The recombination protein RAD52 cooperates with the excision repair protein OGG1 for the repair of oxidative lesions in mammalian cells. Mol. Cell Biol. 29, 4441–4454 (2009).
Giuntoli, R. D. et al. DNA-segment-facilitated dissociation of Fis and NHP6A from DNA detected via single-molecule mechanical response. J. Mol. Biol. 427, 3123–3136 (2015).
Hadizadeh, N., Johnson, R. C. & Marko, J. F. Facilitated dissociation of a nucleoid protein from the bacterial chromosome. J. Bacteriol. 198, 1735–1742 (2016).
Freudenthal, B. D., Beard, W. A., Cuneo, M. J., Dyrkheeva, N. S. & Wilson, S. H. Capturing snapshots of APE1 processing DNA damage. Nat. Struct. Mol. Biol. 22, 924–931 (2015).
Liu, L. et al. PARP1 changes from three-dimensional DNA damage searching to one-dimensional diffusion after auto-PARylation or in the presence of APE1. Nucleic Acids Res. 45, 12834–12847 (2017).
Reardon, J. T., Bessho, T., Kung, H. C., Bolton, P. H. & Sancar, A. In vitro repair of oxidative DNA damage by human nucleotide excision repair system: possible explanation for neurodegeneration in xeroderma pigmentosum patients. Proc. Natl Acad. Sci. USA 94, 9463–9468 (1997).
Muniandy, P. A., Thapa, D., Thazhathveetil, A. K., Liu, S. T. & Seidman, M. M. Repair of laser-localized DNA interstrand cross-links in G1 phase mammalian cells. J. Biol. Chem. 284, 27908–27917 (2009).
Itoh, T., Cado, D., Kamide, R. & Linn, S. DDB2 gene disruption leads to skin tumors and resistance to apoptosis after exposure to ultraviolet light but not a chemical carcinogen. Proc. Natl Acad. Sci. USA 101, 2052–2057 (2004).
Pines, A. et al. Differential activity of UV-DDB in mouse keratinocytes and fibroblasts: impact on DNA repair and UV-induced skin cancer. DNA Repair (Amst.) 8, 153–161 (2009).
Yoon, T. et al. Tumor-prone phenotype of the DDB2-deficient mice. Oncogene 24, 469–478 (2005).
Matsumoto, S. et al. DNA damage detection in nucleosomes involves DNA register shifting. Nature https://doi.org/10.1038/s41586-019-1259-3 (2019).
Kovtun, I. V. et al. OGG1 initiates age-dependent CAG trinucleotide expansion in somatic cells. Nature 447, 447–452 (2007).
Nunez, N. N., Majumdar, C., Lay, K. T. & David, S. S. Fe-S Clusters and MutY base excision repair glycosylases: purification, kinetics, and DNA affinity measurements. Methods Enzymol. 599, 21–68 (2018).
Strauss, P. R., Beard, W. A., Patterson, T. A. & Wilson, S. H. Substrate binding by human apurinic/apyrimidinic endonuclease indicates a Briggs-Haldane mechanism. J. Biol. Chem. 272, 1302–1307 (1997).
Beard, W. A. & Wilson, S. H. Purification and domain-mapping of mammalian DNA polymerase beta. Methods Enzymol. 262, 98–107 (1995).
Slupphaug, G. et al. Properties of a recombinant human uracil-DNA glycosylase from the UNG gene and evidence that UNG encodes the major uracil-DNA glycosylase. Biochemistry 34, 128–138 (1995).
Chen, X. et al. Human DNA ligases I, III, and IV-purification and new specific assays for these enzymes. Methods Enzymol. 409, 39–52 (2006).
Schofield, M. J., Lilley, D. M. J. & White, M. F. Dissection of the sequence specificity of the holliday junction endonuclease CCE1. Biochemistry 37, 13042 (1998).
Pollard, T. D. A guide to simple and informative binding assays. Mol. Biol. Cell 21, 4061–4067 (2010).
Prasad, R., Williams, J. G., Hou, E. W. & Wilson, S. H. Pol beta associated complex and base excision repair factors in mouse fibroblasts. Nucleic Acids Res. 40, 11571–11582 (2012).
Kong, M., Beckwitt, E. C., Springall, L., Kad, N. M. & Van Houten, B. Single-molecule methods for nucleotide excision repair: building a system to watch repair in real time. Methods Enzym. 592, 213–257 (2017).
Zhao, R. et al. DDB2 modulates TGF-beta signal transduction in human ovarian cancer cells by downregulating NEDD4L. Nucleic Acids Res. 43, 7838–7849 (2015).
Acknowledgements
We thank G. Gibson and C. Wallace for help with troubleshooting related to imaging. We thank W. Vermeulen (Erasmus MC) for the generous gift of the mCherry-DDB2 vector. We greatly appreciate N. Kad, C. Kisker and J. Kuper for helpful discussions and comments on the manuscript. This work was supported by funding from the National Institutes of Health including R01ES019566, R01ES028686 (B.V.H.) and R33ES025606 (B.V.H. and P.L.O.), P30CA047904 (Hillman Cancer Center), R01EB017268 (M.P.B.), R01CA067985 (S.S.D.) and 1ZIAES050158 and 1ZIAES050159 (S.H.W.). E.C.B. was supported on a T32 training grant (T32GM088119). C.K. was supported by an NIEHS T32 training grant (ES007059).
Author information
Authors and Affiliations
Contributions
Conceptualization: S.J. and B.V.H.; methodology: S.J., N.K., E.C.B., S.S.D., C.K., C.M., M.K., E.F., V.R.-O., R.P., S.C.W., S.H.W., M.P.B., P.L.O., and B.V.H.; investigation: S.J., S.S.D., C.K., C.M., N.K., E.C.B., M.K., E.F.,V.R.-O., R.P., S.C.W., S.H.W., P.L.O. and B.V.H.; writing of the original draft: S.J. and B.V.H.; writing, reviewing and editing: S.J., N.K., E.C.B, M.K., E.F., V. R-O., P.L.O., and B.V.H; funding acquisition: S.H.W, P.L.O., S.S.D. and B.V.H; resources: S.J., E.F., V.R.-O., R.P., S.C.W., S.H.W., M.P.B., P.L.O. and B.V.H.; supervision: S.H.W., S.C.W, P.L.O. and B.V.H.
Corresponding author
Ethics declarations
Competing interests
M.P.B. is a founder in Sharp Edge Labs, a company applying the FAP-fluorogen technology commercially.
Additional information
Peer review information: Beth Moorefield was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Purified proteins and UV-DDB EMSA experiments, Related to Methods and Fig. 1.
(a) Coomassie stain of SDS-PAGE showing purified proteins used in this study. (b-f) Representative native gels for EMSA experiments, quantification and binding isotherm fitting shown in Fig. 1a. UV-DDB was mixed with different 37 bp fluorescein-labeled dsDNA substrates with different lesions/modifications: THF, tetrahydrofuran; CPD, cyclobutane pyrimidine dimer; 8-oxoG:C; 8-oxoG:A; and UD37, non-damaged.
Supplementary Figure 2 OGG1 glycosylase/lyase, MUTYH glycosylase, and APE1 endonuclease activities in the absence or presence of UV-DDB, Related to Methods and Fig. 1.
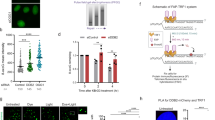
(a) Comparison of APE1 incision activity on 37 bp DNA with a THF moiety (left) and an authentic abasic site (right). AP37 substrate is the product of a reaction between dU37 and Uracil DNA glycosylase. (b) AP lyase activity of OGG1 on 8-oxoG37(G:C) with increasing amount of OGG1, as indicated. DNA products were separated by denaturing polyacrylamide electrophoresis. (c) Chemical structure of 8-oxoG:C. (d) Effect of UV-DDB concentration on stimulation of OGG1 incision. 8-oxoG37(G:C) was incubated with OGG1 and/or increasing amounts of UV-DDB at 37°C for 1.5hrs and separated by denaturing polyacrylamide electrophoresis. (e) Quantification of (d). Percent of total DNA that was incised by OGG1 plotted as a function of UV-DDB concentration. Data shown as the mean of three experiments ± s.d. (f) Effect of UV-DDB on stimulation of OGG1 incision. 8-oxoG37(G:C) was incubated with OGG1 at 37°C for 2hrs and UV-DDB (16nM) was added. Incised DNA was separated by denaturing polyacrylamide electrophoresis. (g) Quantification of (f). Percent of total DNA that was incised by OGG1 plotted as a function of time. Data shown as the mean of three experiments ± s.d. (h) Effect of APE1 and UV-DDB on stimulation of OGG1 incision. 8-oxoG37(G:C) was incubated with OGG1 only, OGG1+APE1, or OGG1+APE1+UV-DDB for 2hrs at 37°C and separated by denaturing polyacrylamide electrophoresis. (i) Quantification of (h). Percent of total DNA that was incised by OGG1 plotted as a function of time. Data shown as the mean of three experiments ± s.d. (** p< 0.01). (j) Effect of UV-DDB concentration on stimulation of MUTYH glycosylase activity. 8-oxoG37(G:A) was incubated with MUTYH and/or increasing amount of UV-DDB for 80mins at 37°C. The reaction was immediately stopped by adding 2X loading dye with 0.1M NaOH followed by heating 95°C for 5mins then quickly chilling on ice for 5mins. (k) Quantification of (j). Percent of total DNA that was incised by MUTYH plotted as a function of UV-DDB concentration. Data shown as the mean of three experiments ± s.d. (l) Effect of UV-DDB concentration on stimulation of APE1 incision. THF37 was incubated with OGG1 and/or increasing amounts of UV-DDB at RT for 1hr and separated by denaturing polyacrylamide electrophoresis. (m) Quantification of (l). Percent of total DNA that was incised by APE1 plotted as a function of UV-DDB concentration. Data shown as the mean of three experiments ± s.d. (n) Stimulation of APE1 activity by UV-DDB. THF-containing DNA (200 nM) was incubated with APE1 (1 nM) in the absence (-) (Lanes 1-3) or presence (+) (Lanes 4-6) of UV-DDB (50 nM) for the periods indicated (i.e., 2, 5 and 10 min). Aliquots (4.5 μl) were removed for analysis at the indicated times. The reaction was terminated by addition of an equal volume of DNA gel loading buffer. The reaction products were analyzed as described in methods. The migration positions of the substrate and product are indicated. The results shown are representative of three experiments. (o) Quantification of (n). Solid line represents mean value of three experiments at 10 mins, normalized to APE1 alone. Black circles represent APE1 activity in the presence of UV-DDB, normalized to reference APE1 activity (right, set to 1.0).
Supplementary Figure 3 Binding isotherms of OGG1 and APE1 and UV-DDB displacement of OGG1 from abasic DNA, Related to Fig. 2.
(a) Binding isotherms of OGG1 binding to THF37 (filled circles) and 8-oxoG37(G:C) (open circles). Binding data from three independent experiments are plotted as mean percent of DNA bound ± s.e.m. Data were globally fit to a single equilibrium dissociation constant (Kd), shown in the table as best fit value ± s.e. to the fit. (b) Binding isotherm of catalytically dead mutant APE1 (K87E/E96Q/D210N) to THF37. Binding data from three independent experiments are plotted as mean percent of DNA bound ± s.e.m. Data were globally fit to a single equilibrium dissociation constant (Kd), shown in the table as best fit value ± s.e to the fit. (c) Displacement of OGG1 on abasic sites by UV-DDB, shown by EMSA. Binding reactions of THF37 and increasing amounts of UV-DDB with or without OGG1 were separated by native PAGE. Protein-DNA complexes were identified based on band migration and labeled accordingly. Representative gel shown, N=2. (d) Quantification of (c) from lane 8 to 14. Bound Percent of total DNA by UV-DDB or OGG1 are plotted as a function of UV-DDB concentration. Data shown as the mean of two experiments ± s.d.
Supplementary Figure 4 DNA tightrope assay showing co-localization of UV-DDB and OGG1 or APE1, related to Fig 3.
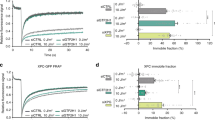
(a - f) Dual-color tightrope assays were performed with UV-DDB (red) and OGG1 or APE1 (green) on abasic (THF) substrates. Individual and co-localized particles were observed and their behavior was recorded. Bar graphs show motile (M) or non-motile (NM) percentages of the total number of proteins observed. Percentage that dissociated (D) is also shown. Data are plotted as weighted mean ± weighted s.d. (a) OGG1 in the presence of UV-DDB, but not co-localized. N=99, 49.5%. (b) Co-localized particles of OGG1 and UV-DDB. N=19, 9.5%. (c) UV-DDB in the presence of OGG1, but not co-localized. N=82, 41.0%. (d) APE1 in the presence of UV-DDB, but not co-localized. N=107, 47.8%. (e) Co-localized particles of APE1 and UV-DDB. N=21, 9.4%. (f) UV-DDB in the presence of APE1, but not co-localized. N=96, 42.8%. (g) Additional still frames and corresponding kymographs of co-localized OGG1 and UV-DDB (OGG1: green, UV-DDB: red, and merge: yellow). Top, scale bar represents 2.5 μm; arrows point to co-localized particles. Bottom, horizontal and vertical scale bars represent 50s and 2kb, respectively. (h) Additional still frames and corresponding kymographs of co-localized APE1 and UV-DDB (OGG1: green, UV-DDB: red, and merge: yellow). Top, scale bar represents 2.5 μm; arrows point to co-localized particles. Bottom, horizontal and vertical scale bars represent 50s and 2kb, respectively.
Supplementary Figure 5 Loss of UV-DDB increases sensitivity to oxidant induced 8-oxoG, Related to Fig. 4.
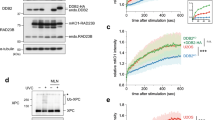
(a) Effect of UV-DDB on DNA ligase III activity in BER. A phosphorimage of the ligation step in BER is shown. The ligation reaction was performed in a 10 μl volume containing 50 mM HEPES, pH 7.5, 20 mM KCl, 5 mM MgCl2, 0.5 mM EDTA, 2 mM DTT, 2 mM ATP. The radiolabeled nicked DNA substrate (200 nM) was incubated with varying concentrations (0 to 400 nM) of DNA ligase III in the absence (-) or presence (+) of UV-DDB (50 nM). The incubation was at 37°C for 5 min. The reaction products were analyzed by denatured polyacrylamide gel electrophoresis. (b) Cell growth curves of normal and XPE lymphoblastoid cells treated with increasing concentrations of KBrO3. Data represents mean +/- SEM from three independent experiments, each performed in quadruplicates. (c) Western blot of BJ-hTERT cells transfected with scrambled or DDB2 siRNA and probed for DDB2, APE1 and OGG1, 72 hours post transfection. (d) Immunofluorescence images depicting mCherry-DDB2 recruitment to and departure from telomeres up to three hours after inducing damage. Graphs show DDB2 and OGG1 co-localization at telomeres. Scale bar: 5μm.
Supplementary information
Supplementary Information
Supplementary Figures 1–5
Supplementary Video 1
Co-localization of OGG1 and UV-DDB on an abasic DNA tightrope, Related to Fig. 3c,d. The movie shows co-localization (yellow) of 605 nm Qdot-labeled OGG1 (green) and 705 nm Qdot-labeled UV-DDB (red) bound to a DNA molecule containing THF sites. OGG1 particle dissociated during observation. The data are collected at 0.68 fps and are played back at 12 fps.
Supplementary Video 2
Co-localization of APE1 and UV-DDB on an abasic DNA tightrope, Related to Fig. 3f,g. The movie shows co-localization (yellow) of 605 nm Qdot-labeled APE1 (green) and 705 nm Qdot-labeled UV-DDB (red) bound to a DNA molecule containing THF sites. APE1 and UV-DDB particles move together during observation. The data are collected at 0.59 fps and are played back at 12 fps.
Supplementary Video 3
UV-DDB recruitment to sites of damage in living cells, Related to Fig. 4d,e. mCherry-DDB2 is recruited to the damaged telomeres minutes after singlet oxygen generation.
Supplementary Data Set 1
Uncropped images for western blots, EMSA gels sequencing gels and SDS-PAGE gels.
Rights and permissions
About this article
Cite this article
Jang, S., Kumar, N., Beckwitt, E.C. et al. Damage sensor role of UV-DDB during base excision repair. Nat Struct Mol Biol 26, 695–703 (2019). https://doi.org/10.1038/s41594-019-0261-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0261-7
This article is cited by
-
Structural basis for APE1 processing DNA damage in the nucleosome
Nature Communications (2022)
-
Global and transcription-coupled repair of 8-oxoG is initiated by nucleotide excision repair proteins
Nature Communications (2022)
-
Biology of aging: Oxidative stress and RNA oxidation
Molecular Biology Reports (2022)
-
XPG: a multitasking genome caretaker
Cellular and Molecular Life Sciences (2022)
-
Current perspectives on the clinical implications of oxidative RNA damage in aging research: challenges and opportunities
GeroScience (2021)