Abstract
The interplay between corticotropin-releasing hormone (CRH) and the dopaminergic system has predominantly been studied in addiction and reward, while CRH–dopamine interactions in anxiety are scarcely understood. We describe a new population of CRH-expressing, GABAergic, long-range-projecting neurons in the extended amygdala that innervate the ventral tegmental area and alter anxiety following chronic CRH depletion. These neurons are part of a distinct CRH circuit that acts anxiolytically by positively modulating dopamine release.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Refojo, D. et al. Science 333, 1903–1907 (2011).
Swanson, L. W., Sawchenko, P. E., Rivier, J. & Vale, W. W. Neuroendocrinology 36, 165–186 (1983).
Henckens, M. J. A. G., Deussing, J. M. & Chen, A. Nat. Rev. Neurosci. 17, 636–651 (2016).
Lemos, J. C. et al. Nature 490, 402–406 (2012).
Grieder, T. E. et al. Nat. Neurosci. 17, 1751–1758 (2014).
Silberman, Y. & Winder, D. G. Front. Psychiatry 4, 42 (2013).
Wanat, M. J., Bonci, A. & Phillips, P. E. M. Nat. Neurosci. 16, 383–385 (2013).
George, O., Le Moal, M. & Koob, G. F. Physiol. Behav. 106, 58–64 (2012).
Sanford, C. A. et al. Neuron 93, 164–178 (2017).
Haubensak, W. et al. Nature 468, 270–276 (2010).
Jennings, J. H. et al. Nature 496, 224–228 (2013).
Fadok, J. P. et al. Nature 542, 96–100 (2017).
Rodaros, D., Caruana, D. A., Amir, S. & Stewart, J. Neuroscience 150, 8–13 (2007).
Tagliaferro, P. & Morales, M. J. Comp. Neurol. 506, 616–626 (2008).
Kubota, Y. Curr. Opin. Neurobiol. 26, 7–14 (2014).
Yan, X., Toth, Z., Schultz, L., Ribak, C.E. & Baram, T.Z. 243, 231–243 (1998).
Robison, A. J. Trends Neurosci. 37, 653–662 (2014).
Robison, A. J. et al. J. Neurosci. 33, 4295–4307 (2013).
Wanat, M. J., Hopf, F. W., Stuber, G. D., Phillips, P. E. M. & Bonci, A. J. Physiol. (Lond.) 586, 2157–2170 (2008).
Garcia, I. et al. Dev. Cell 30, 645–659 (2014).
Taniguchi, H. et al. Neuron 71, 995–1013 (2011).
Madisen, L. et al. Nat. Neurosci. 15, 793–802 (2012).
Goebbels, S. et al. Genesis 44, 611–621 (2006).
Erdmann, G., Schütz, G. & Berger, S. BMC Neurosci. 8, 63 (2007).
Brust, R. D., Corcoran, A. E., Richerson, G. B., Nattie, E. & Dymecki, S. M. Cell Reports 9, 2152–2165 (2014).
Bang, S. J., Jensen, P., Dymecki, S. M. & Commons, K. G. Eur. J. Neurosci. 35, 85–96 (2012).
Miyoshi, G. et al. J. Neurosci. 30, 1582–1594 (2010).
Kühne, C. et al. J. Comp. Neurol. 520, 3150–3180 (2012).
Rodríguez, C. I. et al. Nat. Genet. 25, 139–140 (2000).
Farley, F. W., Soriano, P., Steffen, L. S. & Dymecki, S. M. Genesis 28, 106–110 (2000).
Grinevich, V., Brecht, M. & Osten, P. J. Neurosci. 25, 8250–8258 (2005).
Pilpel, N., Landeck, N., Klugmann, M., Seeburg, P. H. & Schwarz, M. K. J. Neurosci. Methods 182, 55–63 (2009).
Nielsen, S. M., Nielsen, L. Z., Hjorth, S. A., Perrin, M. H. & Vale, W. W. Proc. Natl Acad. Sci. USA 97, 10277–10281 (2000).
Anderzhanova, E. A. et al. Eur. J. Pharmacol. 708, 95–104 (2013).
Chen, Y., Molet, J., Gunn, B. G., Ressler, K. & Baram, T. Z. Endocrinology 156, 4769–4780 (2015).
Muglia, L., Jacobson, L., Dikkes, P. & Majzoub, J. A. Nature 373, 427–432 (1995).
Dedic, N. et al. Mol. Psychiatry 23, 533–543 (2018).
Hartmann, J. et al. Mol. Psychiatry 22, 466–475 (2017).
Kamprath, K. & Wotjak, C. T. Learn. Mem. 11, 770–786 (2004).
Siegmund, A., Langnaese, K. & Wotjak, C. T. Behav. Brain Res. 157, 291–298 (2005).
Thoeringer, C. K. et al. Genes Brain Behav. 9, 947–957 (2010).
Kamprath, K. et al. J. Neurosci. 26, 6677–6686 (2006).
Kamprath, K. et al. Genes Brain Behav. 8, 203–211 (2009).
Siegmund, A. & Wotjak, C. T. J. Psychiatr. Res. 41, 848–860 (2007).
Siegmund, A. et al. J. Psychiatr. Res. 43, 1156–1165 (2009).
Dine, J. et al. Front. Cell. Neurosci. 10, 108 (2016).
Vogl, A. M. et al. Nat. Neurosci. 18, 239–251 (2015).
Wang, X. D. et al. Nat. Neurosci. 16, 706–713 (2013).
Walsh, J. J. et al. Nat. Neurosci. 17, 27–29 (2014).
Pomrenze, M. B. et al. Front. Neurosci. 9, 487 (2015).
Acknowledgements
We thank S. Bauer, C. Eggert, D. Harbich, M. Holzapfel, R. Hüttl, A. Mederer, A. Parl, A. Ressle, R. Stoffel, L. Tietze and S. Unkmeir for excellent technical support, M. Eder for Core Unit infrastructure and J. Keverne for manuscript proofreading. This work was partially supported by the Max Planck Society, Volkswagen Foundation, German Federal Ministry of Education and Research (FKZ:01ZX1314H to J.M.D., W.W.), Helmholtz Alliance ICEMED (to W.W.), Chica and Heinz Schaller Foundation (to V.G.), DFG (SFB 1134 to V.G.) and FAPESP (2013/03445-3 to K.S.G.).
Author information
Authors and Affiliations
Contributions
N.D. and C.K. designed and performed experiments and analyzed data. N.D. and J.M.D. wrote the manuscript. M.J., J.H., K.S.G., M.L.P., S.C., A.K., A.M.V., M.W.M., B.S. and R.C.A. assisted with behavioral experiments and tracing and imaging experiments. E.A., J.D. and A.J.G. conducted microdialysis, electrophysiology and optogenetic experiments, respectively. K.J.R., C.T.W., V.G., A.C., M.V.S., W.W. and D.R. contributed to methodology and resources. J.M.D. designed experiments, analyzed data and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 CRH neurons in the aBNST and CeA directly project to the VTA and SNc.
Extension of Fig. 1a,b. AAVs expressing a Cre-dependent Synaptophysin (Syp)-GFP fusion protein (AAV-Ef1a::DIO-Syp-GFP) were unilaterally injected into the aBNST or CeA of Crh-ires-Cre mice. Presynaptic Syp-GFP puncta are shown in green, immunostaining for tyrosine hydroxylase (TH), a dopaminergic marker, in red. The anterogradely labeled presynaptic terminals (Syp-GFP puncta) represent direct projection sites of aBNST and CeA CRH neurons. Prominent Syp-GFP labeling is observed throughout the VTA and SNc following AAV-Ef1a::DIO-Syp-GFP injections into the aBNST or CeA of Crh-ires-Cre mice. The density of Syp-GFP puncta in the VTA was higher in aBNST-injected compared to CeA-injected mice. The opposite was observed for the SNc, which received stronger input from CeA CRH neurons (a,b, bottom panels). Additional projection sites of aBNST and CeA CRH neurons, as well as of other CRH-expressing brain regions, are listed in Supplementary Table 1. Monosynaptic innervation of the VTA by CRH-expressing neurons of the CeA and aBNST has also been demonstrated by others13,14,50. (c) Accuracy of the tracing approach was verified by co-injecting AAV-Ef1a::DIO-Syp-GFP and AAV-Ef1a::DIO-RFP into the PVN. AAV-Ef1a::DIO-RFP helped to identify correctly targeted neurons at the injection site, since most Syp-GFP is transported to presynaptic terminals and thus often not visible in the cell soma. The highest Syp-GFP puncta density originating from the PVN was observed in the zona externa of the median eminence, the most prominent projection site of parvocellular CRH-positive PVN neurons. From there CRH is released into the hypophysial portal vasculature to initiate activation of the hypothalamic-pituitary-adrenal axis. (d) Projections from CeA and BNST to the VTA were further verified by retrograde tracing using Fluorogold (FG). FG injected to the VTA of Crh-ires-Cre;Ai9 mice revealed a significant number of CRH/tdTomato positive neurons in the BNST and CeA co-labeled with FG. (e) Similarly, injection of retrobeads into the VTA of Crh-ires-Cre;Ai9 mice resulted in CRH/tdTomato neurons co-labeled with retrobeads. All tracing experiments were independently replicated three times. Abbreviations: Anterior BNST (aBNST), anterior dorsal BNST (adBNST), anterior ventral BNST (avBNST), basolateral amygdala (BLA), central amygdala (CeA), median eminence (ME), paraventricular nucleus of the hypothalamus (PVN), substantia nigra pars compacta (SNc), substantia nigra pars reticularis (SNr), synaptophysin (Syp), third ventricle (3V), Ventral tegmental area (VTA). Scale bars represent 100 µm.
Supplementary Figure 2 Crh-ires-Cre-driven reporter expression faithfully reproduces native Crh expression patterns.
Crh-ires-Cre mice were generated by inserting an ires-Cre expression cassette immediately after the translational STOP codon in the 3’ UTR of the endogenous Crh gene21. As a result, Cre expression is regulated by endogenous Crh promoter/enhancer elements, without compromising Crh expression. (a) Left row depicts the endogenous Crh mRNA expression pattern in adult wild-type mice (C57BL/6J) determined by ISH. Right row shows representative coronal sections of Crh-ires-Cre mice crossed with Ai9 reporter mice (R26::loxP-STOP-loxP-tdTomato). tdTomato fluorescence (red) mirrors the native Crh mRNA expression pattern. Regions of interest are highlighted with arrowheads or dashed lines. Representative images of three independent experiments. (b) High specificity of Cre recombinase expression in CRH neurons was additionally confirmed by double ISHs against endogenous Crh (white) and Crh-ires-Cre-driven tdTomato (red). (c) Quantification of double positive cells in (b); n = 3 mice, 2-3 sections/mouse. Most tdTomato-expressing neurons co-expressed Crh, and likewise the majority of Crh-positive neurons also expressed tdTomato (arrowheads). Upon Cre/loxP recombination-mediated removal of the transcriptional STOP cassette from the R26CAG::loxP-STOP-loxP-tdTomato allele, tdTomato-reporter expression is driven by a CAG promoter and continuously detected in all neurons that have expressed CRH at any given time point within their lineage. Thus, tdTomato-positive neurons that lack Crh expression (arrows) could represent cells that expressed CRH during the course of development, but ceased to do so in adulthood. The presence of Crh-positive, but tdTomato-negative cells (dashed arrows), could be explained by lack of activity of the CAG promoter (that drives tdTomato expression) in those particular cells. Abbreviations: anterior dorsal BNST (adBNST), anterior ventral BNST (avBNST), central nucleus of the amygdala (CeA), cortex (Ctx), hippocampus (Hip), piriform cortex (Pir), paraventricular nucleus of the hypothalamus (PVN). Error bars represent s.e.m.
Supplementary Figure 3 Neurochemical identity of Crh-expressing neurons in the mouse brain.
(a) The first left column depicts dark-field photomicrographs of the Crh mRNA expression pattern in brain sections of adult wild-type (C57BL/6) mice determined by ISH. Regions of interest are highlighted with arrowheads. Additional columns represent bright field photomicrographs of coronal wild-type brain sections showing double ISH of Crh mRNA (silver grains) together with different neuronal markers (red staining) including glutamic acid decarboxylase 65 and 67 (Gad65/67), vesicular glutamate transporters 1 and 2 (Vglut1, Vglut2) and calcium/calmodulin-dependent protein kinase 2 alpha (Camk2a). Black arrowheads indicate cells only expressing Crh (silver grains). Grey arrowheads indicate cells co-expressing Crh and the respective marker. All images are representatives of three independent experiments. (b) Quantifications of Crh co-expression with different neuronal markers (Gad65/67 n = 4 mice; Vglut1, Vglut2, Camk2a n = 3 mice; adBNST, avBNST, CeA, Pir and Hip = 3 sections/mouse, Ctx = 5 sections/mouse). Crh neurons in the cortex and limbic regions such as the CeA, BNST and hippocampus are predominately GABAergic which coincides with previous reports15,16,50. Most CRH neurons in the piriform cortex are glutamatergic, and a few Vglut1- and Vglut2-expressing Crh neurons were also detected throughout the remaining cortical layers. Considering that Camk2a is primarily expressed in glutamatergic neurons suggests that CRH/CAMK2A-positive neurons in the piriform cortex and cortex most likely represent excitatory, Vglut1 or Vglut2-positive cells. Contrastingly, Crh/Camk2a co-expressing neurons in the aBNST and CeA are GABAergic (see Main Text). Abbreviations: anterior dorsal BNST (adBNST), anterior ventral BNST (avBNST), central nucleus of the amygdala (CeA), cortical layers (CtxII/III, CtxV/VI), hippocampus (Hip), paraventricular nucleus of the hypothalamus (PVN), piriform cortex (Pir). Error bars represent s.e.m.
Supplementary Figure 4 Characterization of CRH neurons in the CeA and morphological assessment of cortical and hippocampal CRH neurons.
(a) Representative bright field photomicrographs of coronal wild-type brain sections covering the rostral to caudal extension of the CeA. Depicted are double ISHs of Crh mRNA (silver grains) together with CeA-specific markers (red staining) protein kinase C delta (Pkcδ) and somatostatin (Som). Black arrowheads indicate cells only expressing Crh. Red arrowheads indicate cells only expressing Pkcδ or Som, respectively. Double positive cells are indicated by black arrowheads enframed by a red line. In accordance with others9,10, Crh neurons in the CeA appear to constitute a distinct GABAergic population as they show limited overlap with CeA markers Pkcδ or Som. (b) Quantification of Crh co-expression with Pkcδ at different CeA levels from bregma −0.82 to −1.82 (n = 3 mice from independent experiments). (c) Quantification of Crh co-expression with Som at different CeA levels from bregma −0.82 to −1.82 (n = 3 mice from independent experiments, 1-2 sections/mouse). (d-f) Overview of confocal images of the cortex (Ctx) and CA1 region of the hippocampus (Hip) in Crh-ires-Cre;Ai32 (d,e top) and Crh-ires-Cre;Ai9 (f, top) mice showing native ChR2-EYFP and tdTomato fluorescence (both in green) in CRH neurons. DAPI staining shown in blue. Higher power images illustrate that cortical (d, e middle) and hippocampal (f, middle) CRH neurons display bipolar and multipolar morphologies with aspiny dendrites (d-f, bottom) characteristic of GABAergic interneurons. Images in d-f are representatives from four independent experiments. Pyramidal cell layer (pyr), stratum radiatum (sr).
Supplementary Figure 5 Combining dual fate mapping with immunohistochemistry identifies triple-positive GABAergic/CAMK2A/CRH neurons in the anterior bed nucleus of the stria terminalis (aBNST) and central amygdala (CeA).
(a) Schematic illustration of the recombinase-based intersectional strategy used to genetically label double-positive GABAergic/CAMK2A neurons. RC::FrePE double-reporter mice25,26 were initially crossed with Dlx5/6-Flp mice, leading to the excision of the frt-flanked STOP cassette and consequent expression of mCherry in Dlx5/6-positve (GABAergic) neurons. Subsequent crossing of RC::FrePE;Dlx5/6 mice to Camk2a-CreERT2 mice leads to the excision of the loxP-flanked mCherry-STOP sequence, which results in the activation of EGFP expression within cells that have expressed both Dlx5/6-Flp and Camk2a-CreERT2 at any time and in any order within their lineage. Thus, EGFP fluorescence reports the presence of double-positive GABAergic/CAMK2A neurons, while mCherry labeling is observed in cells that only express Dlx5/6-Flp (exclusively GABAergic, non-CAMK2A-expressing neurons). Cre activity alone (solely CAMK2A-expressing neurons) is not reported. Immunohistochemistry against CRH was performed on coronal vibratome sections of RC::FrePE;Dlx5/6;Camk2a-CreERT2 mice. (b) Representative lower magnification confocal images showing native EGFP (green) and mCherry (gray) expression, as well as CRH immunostaining (red) in the aBNST and CeA of RC::FrePE;Dlx5/6-Cre;Camk2a-CreERT2 mice. The merged images demonstrate the presence of triple-positive GABAergic/CAMK2A/CRH neurons in the aBNST and CeA. The accuracy and specificity of this dual-fate mapping approach is demonstrated by the fact that only a few single GABAergic (mCherry, gray) and GABAergic/CAMK2A-positive (EGFP, green) neurons were detected in the basolateral amygdala (BLA), a structure that primarily contains excitatory glutamatergic neurons. (c, d) Representative higher magnification images of triple-positive GABAergic/CAMK2A/CRH neurons in the aBNST and CeA. Expectedly (and further proof of the accuracy and specificity of this dual-fate mapping approach) no co-localization is detected between the mutually exclusive mCherry- and EGFP-labeled neurons. All representative images were derived from three independent experiments. Scale bars in b-c = 50 µm; d = 20 µm.
Supplementary Figure 6 CAMK2A-expressing CRH neurons in the aBNST and CeA project to the VTA.
Representative confocal images of Crh-ires-Cre mice injected with AAV-Camk2a::DIO-EYFP into the aBNST (a) or CeA (b). EYFP expression is driven by the Camk2a promoter specifically in CRH neurons. EYFP-labeled projections that originate from CAMK2A-expressing neurons of the aBNST or CeA (top panels) are detected in the VTA and SNc (bottom panels). Bottom right depict higher magnification images of the VTA/SN midbrain region. Representative images were derived from three independent experiments each. Abbreviations: anterior commissure (ac), anterior dorsal BNST (adBNST), basolateral amygdala (BLA), central amygdala (CeA), periaqueductal gray (PAG), substantia nigra compacta (SNc), substantia nigra reticularis (SNr), ventral tegmental area (VTA). Scale bars represent 100 µm.
Supplementary Figure 7 Generation of conditional Crh knockout mice.
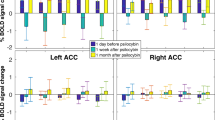
Strategy for targeted manipulation of the Crh locus. (a) Partial restriction maps of the wild-type locus and targeting vector. For better visualization, loxP, frt and attP sites are not to scale. (b) Recombined reporter allele (Crhtz) following homologous recombination in embryonic stem (ES) cells. (c) Recombined floxed allele (Crhflox) following removal of the frt-flanked reporter-selection cassette, and (d) conditional knockout allele (Crh-) following Cre-mediated excision of the loxP-flanked exon 2. (e) Southern blot analysis of wild-type (+/+) and targeted ES cell clones (+/tZ). The external Crh 5’-probe was hybridized to EcoRI-digested genomic ES cell DNA. The targeted allele is indicated by the presence of an additional mutant (mt) 8.9 kb fragment. The Crh 3’-probe was hybridized to BamHI-digested DNA from the same ES cell clone confirming homologous recombination by detection of an additional mutant fragment at 8.3 kb. Homologous recombination was confirmed with two external probes in two independently prepared genomic DNA samples. (f) Functionality of the conditional Crh allele was confirmed by breeding Crhflox mice to Cre-Deleter mice resulting in ubiquitous ablation of Crh mRNA expression throughout the brain determined by ISH (representative images from four independent experiments). (g) Quantification of Crh mRNA expression in (f), shown as percent change compared to littermate controls. Unpaired two-tailed t-test; OB (t(6) = 2.9, *p = 0.032); Pir (t(6) = 24.3, ***p < 0.0001); adBNST (t(6) = 5.6, **p = 0.0014); avBNST (t(6) = 10.7, ***p = 0.0013); CeA (t(6) = 5.7, **p = 0.0013); PVN (t(6) = 7.3, ***p = 0.0003); n = 4 mice/group, 1-2 sections/mouse). (h) Expectedly, ablation of CRH in Crh-/- mice also resulted in strongly diminished hypothalamic-pituitary-adrenal axis activity evident by significantly reduced corticosterone levels measured in the morning (am), evening (pm), and following 10 min (stress) and 90 min (recovery) of restraint stress. Unpaired two-tailed t-test - am (t(20) = 2.6, p = *0.017, n = 11 mice/group); pm (t(20) = 4.1, ***p = 0.0005, n = 13 Crhflox/flox, 9 Crh-/-); stress (t(22) = 8.9, ***p < 0.0001, n = 13 Crhflox/flox, 11 Crh-/-); recovery (t(22) = 3.7, **p = 0.0013, n = 13 Crhflox/flox, 11 Crh-/-). Error bars represent s.e.m.
Supplementary Figure 8 Additional molecular and behavioral assessment of CrhCKO-Camk2a mice.
(a) Overview images and quantification (b) of Crh mRNA expression assessed by ISH in control and CrhCKO-Camk2a mice. Regions of interest are marked with arrowheads. Deletion of Crh in CrhCKO-Camk2a mice was primarily detected in the piriform cortex (Pir), and in a subset of neurons within the BNST and CeA. Thus, the deletion pattern in CrhCKO-Camk2a mice supports the expression maps obtained with double ISH (Supplementary Fig. 3). Unpaired two-tailed t-test; Pir (t(8) = 13.8, ****p < 0.0001); adBNST (t(8) = 2.7, *p = 0.028); avBNST (t(8) = 3.0, *p = 0.018); CeA (t(8) = 3.3, *p = 0.01); for Pir, adBNST and avBNST n = 5 mice/group, 1-2 sections/mouse; for CeA n = 4 mice/group, 1-2 sections/mouse. (c) Hypothalamic-pituitary-adrenal (HPA) axis activity was similar between control and CrhCKO-Camk2a mice, evident by comparable corticosterone levels measured in the morning (am), evening (pm), and following 10 min (stress) and 90 min (recovery) of restraint stress. Unpaired two-tailed t-test; am (t(22) = 1.1, p = 0.3, n = 13 CrhCtrl, 11 CrhCKO-Camk2a); pm (t(23) = 0.45, p = 0.7, n = 13 CrhCtrl, 12 CrhCKO-Camk2a); stress (t(22) = 0.95, p = 0.35, n = 11 CrhCtrl, 13 CrhCKO-Camk2a), recovery (t(22) = 0.8, p = 0.43, n = 13 CrhCtrl, 11 CrhCKO-Camk2a). (d) CrhCKO-Camk2a mice showed increased anxiety in the open field test (unpaired two-tailed t-test / inner zone time: t(23) = 2.8, *p = 0.01 / number of inner zone entries: t(23) = 2.1, *p = 0.044), without exhibiting changes in locomotion (unpaired two-tailed t test / distance travelled: t(23) = 1.6, p = 0.13; n = 12 CrhCtrl, 13 CrhCKO-Camk2a). (e) Freezing time was similar between CrhCKO-Camk2a and control mice following contextual fear conditioning (RM ANOVA - time genotype x interaction: F(2,52) = 0.63, p = 0.54; n = 13 CrhCtrl, 15 CrhCKO-Camk2a). (f) Foot shock perception, tested in a new cohort of mice, was not different between CrhCKO-Camk2a mice and littermate controls. Both, control and CrhCKO-Camk2a mice showed first signs of discomfort (backwards moving; PT1) and pain (jumping and vocalization; PT2) at similar foot shock current intensities (RM ANOVA - genotype: F(1,21) = 1.91, p = 0.18; genotype x current intensity: F1,21 = 0.46, p = 0.505; n = 11 CrhCtrl, 12 CrhCKO-Camk2a). This was also the case if only PT2 was considered (CrhCtrl: 0.26 ± 0.01 mA; CrhCKO-Camk2a: 0.23 ± 0.01 mA; unpaired two-tailed t-test: t22 = 1.5, p = 0.15). (g) CrhCKO-Camk2a mice display higher levels of sensitized fear. After termination of PT2 in (f), all mice received an unsignalled foot shock (1.5 mA for 2s). The next day all animals were placed into a neutral context for a 3 min baseline period followed by a 3 min tone presentation. This procedure allows for the measurement of non-associative memory components of the conditioning procedure39,42,43,44,45. CrhCKO-Camk2a mice showed similarly low levels of freezing before and after tone presentation. However, CrhCKO-Camk2a mice displayed a sustained freezing response in particular towards the end of the (unconditioned) tone presentation (RM ANOVA - time: F(6,132) = 44.31, **p < 0.0001; Genotype: F(1,22) = 4.93, *p = 0.037; Genotype x Time: F(6,132) = 2.33, *p = 0.036; Bonferroni post-hoc test, *p < 0.05; n = 12 CrhCtrl, 12 CrhCKO-Camk2a) suggesting impairments in fear adaptation. (h) CrhCKO-Camk2a mice show no changes in generalization of contextual fear. Compared to the freezing levels in the shock context before shock presentation on day 0, and to the novel context before tone presentation on day 1, both groups showed a significantly higher freezing response upon re-exposure to the shock context on day 2. This validates the results in (e) and demonstrates again similar levels of contextual fear between control and CrhCKO-Camk2a mice (RM ANOVA - context: F(2,22) = 41.06, ***p < 0.0001; Genotype: F(1,22) = 0.085, p = 0.773; context x genotype interaction: F(2,22) = 1.18, p = 0.32; n = 12 CrhCtrl, 12 CrhCKO-Camk2a). (i) Representative coronal brain section (right) and corresponding atlas images (left) depicting the correct placement of microdialysis probes within the prefrontal cortex. (j) In vivo microdialysis showing a similar magnitude in footshock-induced dopamine release between control and CrhCKO-Camk2a mice (dopamine levels normalized to baseline levels (fractions 1-6) and depicted in percent) / RM ANOVA - genotype x time interaction F(11,187) = 0.89, p = 0.55; genotype F(1,17) = 1.18, p = 0.29; time F(11,187) = 7.06, ***p < 0.0001; n = 10 CrhCtrl, 9 CrhCKO-Camk2a. (k) Representative coronal brain section (right) and corresponding atlas images (left) depicting the micropunched area of the nucleus accumbens. (l) Basal dopamine tissue content in the nucleus accumbens was similar between control and CrhCKO-Camk2a mice (unpaired two-tailed t-test; t(10) = 0.28, p = 0.79; n = 4 CrhCtrl, 8 CrhCKO-Camk2a). (m-o) mRNA expression of Ucn1, Crhr1 (determined by real-time qPCR; unpaired two tailed t-test; VTA t(10) = 2.28, *p = 0.046; BLA t(10) = 2.0, p = 0.075; PFC, adBNST, avBNST, BLA, CeA, VTA and SNc n = 6 mice/group; pBNST n = 6 CrhCtrl, 5 CrhCKO-Camk2a; Hip n = 5 CrhCtrl, 4 CrhCKO-Camk2a) and Crhr2 (determined by ISH; unpaired two tailed t test; MeA t(6) = 2.1, p = 0.076;n = 4 CrhCtrl, 4 CrhCKO-Camk2a; LS, BNST, DRn and VMH n = 5 CrhCtrl, 5 CrhCKO-Camk2a) across different brain regions. . Error bars represent s.e.m.
Supplementary Figure 9 Behavioral effects in CrhCKO-Camk2a mice are independent of Crh deletion in glutamatergic neurons.
Crh deletion specifically in glutamatergic neurons (CrhCKO-Glu) was achieved by crossing Crhflox mice with Nex-Cre mice. Overview images (a) and quantification (b) of Crh mRNA expression assessed by ISH in control and CrhCKO-Glu mice. Unpaired two-tailed t-test; Pir (t(4) = 6.9, **p = 0.002); adBNST (t(4) = 0.5, p = 0.65); avBNST (t(4) = 1.1, p = 0.34); CeA (t(4) = 0.21, p = 0.84); n = 3 mice/group, 1-2 sections/mouse. Deletion of Crh in CrhCKO-Glu mice was primarily detected in the piriform cortex (Pir, arrow, bottom panel) and a few single neurons in cortical layers II/III and V/VI (not visible in the overview images). Thus, the deletion pattern in CrhCKO-Glu mice supports the expression maps obtained with double ISH (Supplementary Fig. 3) and further confirms the absence of glutamatergic CRH neurons in the aBNST and CeA. (c-e) Locomotion (unpaired two-tailed t-test; t(28) = 0.29, p = 0.77), and anxiety-related behavior in the open field (unpaired two-tailed t-test; inner zone time (%): t(28) = 1.3, p = 0.22 / no. inner zone entries: t(28) = 0.48, p = 0.64), as well as anxiety-related behavior in the dark-light box test (unpaired two-tailed t-test; lit zone time (%): t(28) = 0.54, p = 0.59 / no. lit zone entries: t(28) = 0.35, p = 0.72) and EPM (unpaired two-tailed t-test; t(28) = 0.23, p = 0.82) were not altered in CrhCKO-Glu mice compared to littermate controls; n = 15 mice/group. (f) HPA axis activity was similar between control and CrhCKO-Glu mice, evident by comparable corticosterone levels measured in the morning (am), evening (pm), and following 10 min (stress) and 90 min (recovery) of restraint stress. Unpaired two-tailed t-test / am (t(19) = 0.58, p = 0.57; n = 9 CrhCtrl, 12 CrhCKO-Glu) / pm (t(19) = 0.57, p = 0.58; n = 10 CrhCtrl, 11 CrhCKO-Glu) / stress (t(19) = 1.1, p = 0.3; n = 10 CrhCtrl, 11 CrhCKO-Glu) / recovery (t(18) = 0.76, p = 0.46; n = 9 CrhCtrl, 11 CrhCKO-Glu). Error bars represent s.e.m.
Supplementary Figure 10 Generation of Crhr1-ires-Cre mice.
Strategy for targeted manipulation of the Crhr1 locus. (a) Wild-type Crhr1 locus depicting the position of integration of the docking site for recombinase-mediated cassette exchange (RMCE) in Intron 2. For better visualization different scales were used for exons and introns to illustrate the Crhr1 gene structure. (b) Crhr1tZ-ΔloxP allele containing the RMCE docking site in intron 2. A magnification of the docking site is provided above the Crhr1tZ-ΔloxP allele. The targeting vector for RMCE equipped with respective attB sites is depicted below. Of note, loxP, frt, attB, attP, and T2A sites are not to scale. (c) ΦC31-mediated cassette exchange in embryonic stem (ES) cells resulted in integration of the targeting vector into the right attP site. ES cells were used to generate mice, which were subsequently bred to FLPeR mice to simultaneously remove the reporter and selection cassettes. (c) The correct integration and removal of selection and reporter cassettes was verified by PCR. (d) Detection of Flp recombinase-mediated removal of Tau-lacZ reporter cassette with PCR. (e) Confirmation of simultaneous Flp recombinase-mediated removal of the hygromycin selection cassette. (f) Detection of Cre recombinase expression cassette in Crhr1 locus. PCRs in d-f were independently replicated in each individual animal (within the respective mouse line) that was genotyped (n > 20).
Supplementary Figure 11 Validation of Cre expression and recombination specificity in Crhr1-ires-Cre mice.
(a) Cre mRNA expression fully recapitulates the endogenous Crhr1 mRNA expression pattern. The first row depicts endogenous Crhr1 mRNA expression in Crhr1+/+ mice (rostral to caudal), determined by ISH using a radiolabeled riboprobe detecting the Crhr1 3’UTR which is not expressed from the Crhr1-ires-Cre allele. The second row shows a similar hybridization signal in Crhr1Cre/Cre littermates, using a Cre-specific riboprobe. Riboprobe specificity and targeting accuracy is further illustrated by the absence of a Crhr1 3’UTR signal in Crhr1Cre/Cre mice (see targeting approach in Supplementary Figure 10), and absence of a Cre signal in their wildtype littermates. The third row shows representative images of Crhr1Cre/Cre mice crossed with Ai9 reporter mice (R26::loxP-STOP-loxP-tdTomato). tdTomato fluorescence (red staining) mirrors the native Crhr1 expression pattern (silver grains). Images are representatives from three independent experiments. (b) High specificity of Cre recombinase expression in CRHR1 neurons was additionally confirmed by double ISHs against endogenous Crhr1 (silver grains) and Crhr1-ires-Cre-driven tdTomato (red). Images are representatives from three independent experiments. (c) Quantification of double positive cells in (b); n = 3 mice from independent experiments, 1-2 sections/mouse. Most tdTomato-expressing neurons co-expressed Crhr1, and likewise the majority of Crhr1-positive neurons also expressed tdTomato (red-framed, black arrowheads). Upon Cre recombinase-mediated removal of the transcriptional STOP cassette from the R26CAG::loxP-STOP-loxP-tdTomato allele, tdTomato-reporter expression is driven by a CAG promoter and continuously detected in all neurons that have expressed CRHR1 at any given time point within their lineage. Thus, tdTomato-positive neurons that lack Crhr1 expression (red arrowheads) could represent cells that expressed CRHR1 during the course of development, but ceased to do so in adulthood. The presence of Crhr1-positive, but tdTomato-negative cells (black arrowheads), could be explained by lack of activity of the CAG promoter (that drives tdTomato expression) in those particular cells. (d) HPA axis activity was similar between Crhr1+/+ and Crhr1Cre/Cre mice, evident by comparable corticosterone levels measured in the morning (am), evening (pm), and following 10 min of restraint stress (stress). Thus, in addition to its expression, functionality of CRHR1 activity is not compromised in Crhr1Cre/Cre mice (unpaired two-tailed t-test: am t(12) = 0.86, p = 0.41; n = 6 Crhr1+/+, 8 Crhr1Cre/Cre / pm t(11) = 0.41, p = 0.69; n = 5 Crhr1+/+, 8 Crhr1Cre/Cre / stress t(12) = 0.42, p = 0.68; n = 6 Crhr1+/+, 8 Crhr1Cre/Cre). Abbreviations: cortex (Ctx), globus pallidus (GP), hippocampus (Hip), lateral septum (LS), pontine nuclei (PN), prefrontal cortex (PFC), reticular thalamic nucleus, red nucleus (RMC), substantia nigra, pars compacta (SNc), olfactory tubercle (Tu), ventral tegmental area (VTA). Error bars represent s.e.m.
Supplementary Figure 12 Virally mediated expression of a constitutively active (CA) CRHR1-EGFP fusion construct in VTA neurons of Crhr1-ires-Cre mice.
(a) Double ISH against GFP (red staining) and endogenous Crhr1 (silver grains) in Crhr1-ires-Cre mice following VTA-injection of AAV-Ef1α::(CA)CRHR1-GFP (representative coronal sections are shown from rostral to caudal; corresponding atlas images are shown on the left). The vast majority of cells co-express GFP and native Crhr1 (red-framed black arrow), further demonstrating recombination-specificity of the Crhr1-Cre driver. Images are representatives from three independent experiments. (b) Crhr1-ires-Cre mice expressing (CA)CRHR1-EGFP in the VTA exhibit increased locomotion in the OF (unpaired two-tailed t-test: t(15) = 2.45, *p = 0.027), and a trend towards decreased anxiety compared to controls injected with AAV-Ef1α::DIO-mCherry (unpaired two-tailed t-test: inner zone time (%) - t(15) = 1.5, p = 0.15; no. inner zone entries - t(15) = 1.8, p = 0.09; n = 8 Ctrl, 10 (CA)CRHR1). Although locomotion in the open field was increased it is unlikely to compromise the interpretation of the other tests (Fig. 3) given that mice expressing CA(CRHR1) also displayed reduced anxiety in the marble burying task (Fig. 3l), an assay in which confounding increases in locomotion would have produced the opposite effect. (c) No changes between groups were observed during contextual fear conditioning (RM ANOVA – time x genotype interaction: F(2,34) = 0.45, p = 0.64; genotype: F(1,17) = 0.05, p = 0.82; n = 8 Ctrl, 11 (CA)CRHR1). Error bars represent s.e.m.
Supplementary Figure 13 CRH microinjections into the VTA decrease anxiety at specific doses.
Bilateral injections of CRH into the VTA of wildtype C57BL/6J mice at (a) 0.4, (b) 40 and (c) 400 ng/side. No behavioral alterations were observed at the lowest CRH dose (unpaired two-tailed t-test / OF: distance t(16) = 0.71, p = 0.49; inner zone time (%) t(16) = 0.18, p = 0.86; n = 8 Vehicle, 10 CRH / DaLi: lit zone time (%) t(15) = 0.75, p = 0.46; no. lit zone entries t(15) = 0.23, p = 0.82; n = 8 vehicle, 9 CRH / EPM: open arm time (%) t(11) = 0.58, p = 0.57; open arm entries (%) t(11) = 0.14, p = 0.89; n = 7 vehicle, 6 CRH). Microinjections of 40 ng/side induced decreased anxiety-related behavior specifically in the EPM (unpaired two-tailed t-test: t(9) = 3.95, **p = 0.003, n = 5 vehicle, 6 CRH) without alerting locomotion or cued fear conditioning (unpaired two-tailed t-test / OF: distance t(11) = 0.52, p = 0.66; inner zone time (%) t(11) = 0.60, p = 0.56; n = 6 vehicle, 7 CRH / DaLi: lit zone time (%) t(13) = 1.4, p = 0.19; no. lit zone entries t(13) = 0.93, p = 0.37; n = 7 vehicle, 8 CRH / EPM: open arm entries (%) t(11) = 0.66, p = 0.53; n = 5 vehicle, 6 CRH / fear conditioning (RM ANOVA) – tone/auditory: time x genotype interaction F6,72 = 1.83, p = 0.11; genotype F1,12 = 0.42, p = 0.69 / context: time x genotype interaction F2,24 = 0.53, p = 0.59; genotype F1,12 = 0.40, p = 0.75; n = 7 vehicle, 7 CRH). At the highest dose of 400 ng/side, CRH induced a pronounced decrease in locomotion in the OF (unpaired two-tailed t-test / OF: distance - t(15) = 3.5, **p = 0.003; inner zone time (%) t(15) = 1.83, p = 0.09; n = 8 vehicle, 9 CRH) and lead to a decrease in number of lit zone entries in the DaLi (unpaired two-tailed t-test / DaLi: lit zone time (%) t(16) = 0.43, p = 0.67; no. lit zone entries t(16) = 2.5, *p = 0.025; n = 9 vehicle, 9 CRH). (d) Representative coronal brain section (right) and corresponding atlas image (left) depicting the correct placement of the injection cannula in the VTA. Error bars represent s.e.m.
Supplementary Figure 14 Acute optogenetic activation of VTA-targeting CRH terminals does not alter anxiety.
Optogenetic activation of CRH-positive terminals within the VTA was performed in Crh-ires-Cre mice bred to ChR2-expressing Ai32 mice. This enabled targeting of all VTA-projecting CRH fibers without having to perform multiple viral injections and optic fiber placements in the BNST and CeA. Lack of local CRH-expressing cell bodies in the VTA ensured the predominant activation of VTA-targeting CRH fibers. (a) Electrophysiological confirmation of optogenetic activation of a subset of ChR2-expressing CRH neurons of the CeA in acute brain slices. Current-clamp recordings show that ChR2-expressing CRH neurons fire an action potential to single blue laser pulses (488 nm, 2 msec, 5 mW/mm², 1 Hz) and can follow 10 Hz and 20 Hz laser stimulation (representative of n = 4 animals). (b) Representative coronal image showing the optic fiber tract (white dashed line) above the VTA. (c) Individual, bilateral placements of optic fiber tips in the VTA of Crh-ires-Cre mice bred to ChR2-expressing Ai32 animals (green) and Crhr-ires-Cre mice bred to Ai9 reporter mice (red). (d-f) VTA-targeting CRH fibers were stimulated by delivering blue (460 nm) light (20 Hz, 15 ms pulses) via bilateral optical fibers implanted over the VTA.. (d) Photostimulation of VTA-projecting CRH fibers in the ChR2 group did not affect anxiety in the DaLi (light ON during the entire test) or (e) cued fear conditioning (1 min light epoch shown in blue) compared to tdTom-expressing controls (unpaired two-tailed t-test for DaLi – lit zone time (%) t(18) = 0.82, p = 0.43; no. lit zone entries t(18) = 0.84, p = 0.41; n = 9 Ctrl, 11 ChR2 / RM ANOVA for FC – genotype x time interaction F(20,340) = 0.75, p = 0.78; genotype F(1,17) = 0.30, p = 0.59; n = 8 Ctrl, 11 ChR2). (f) 15 min session in the EPM with alternating OFF-ON-OFF illumination epochs (5 min each). No significant effect of light stimulation was detected in ChR2 mice for open arm entries or open arm time (RM ANOVA / open arm time (%): group x epoch interaction F(2,34) = 0.22, p = 0.81; group F(1,17) = 1.1, p = 0.31; epoch F(2,34) = 2.7, p = 0.08: open arm entries (%): group x epoch interaction F(2,34) = 0.5, p = 0.61; group F(1,17) = 0.45, p = 0.51; epoch F(2,34) = 4.7, *p = 0.014; n = 8 Ctrl, 11 ChR2). Both, Ctrl and CrhR2 mice appeared to exhibit time-dependent habituation to the EPM, as seen by a gradual decrease in distance travelled/exploration already during the first 5 min (RM ANOVA: group x time interaction F(14, 238) = 2.2, **p = 0.008; genotype F(1,17) = 0.7, p = 0.42; time F(14,238) = 8.7, ***p < 0.0001; n = 8 Ctrl, 11 ChR2). Error bars represent s.e.m.
Supplementary Figure 15 Working model of the anxiolytic CRH circuit.
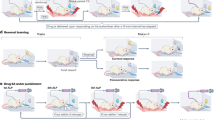
(a) In control mice under baseline conditions, GABAergic/CAMK2A/CRH neurons in the anterior bed nucleus of the stria terminalis (aBNST) and central amygdala (CeA) project monosynaptically to the ventral tegmental area (VTA). There, presynaptic CRH release activates postsynaptic CRHR1 on VTA-to-prefrontal cortex (PFC) projecting (mesocortical) dopaminergic neurons to modulate dopamine release, and thereby alter anxiety. (b) Conditional deletion of Crhr1 in dopaminergic neurons (Crhr1CKO-DA mice) results in diminished dopamine release in the PFC and increased anxiety behavior1. Similar effects are observed when Crh is deleted from VTA-targeting GABAergic/CAMK2A/CRH neurons in the aBNST and CeA (Crhr1CKO-Camk2α mice).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Table 1
Supplementary Video 1 – CLARITY-based visualization of VTA/SN-targeting axonal fibers originating from CAMK2A-positive CRH neurons in the aBNST.
CLARITY-processed brain of a Crh-ires-Cre mouse injected with AAV-Camk2a::DIO-EYFP into the aBNST. The video shows a horizontal z-stack movie (scan depth = 1,410 µm) scanning from dorsal to ventral. CAMK2A-positive CRH neurons (EYFP-expressing cell bodies) can be seen in the aBNST. CAMK2A/CRH-expressing axons originating in the aBNST and targeting the VTA/SN (EYFP-labeled fibers) are marked by arrows. Abbreviations: anterior bed nucleus of the stria terminalis (aBNST), ventral tegmental area (VTA), substantia nigra (SN).
Rights and permissions
About this article
Cite this article
Dedic, N., Kühne, C., Jakovcevski, M. et al. Chronic CRH depletion from GABAergic, long-range projection neurons in the extended amygdala reduces dopamine release and increases anxiety. Nat Neurosci 21, 803–807 (2018). https://doi.org/10.1038/s41593-018-0151-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0151-z
This article is cited by
-
CRF regulates pain sensation by enhancement of corticoaccumbal excitatory synaptic transmission
Molecular Psychiatry (2024)
-
Stress-induced plasticity of a CRH/GABA projection disrupts reward behaviors in mice
Nature Communications (2023)
-
Electroacupuncture Inhibits Pain Memory and Related Anxiety-Like Behaviors by Blockading the GABAB Receptor Function in the Midcingulate Cortex
Molecular Neurobiology (2023)
-
Single-cell reconstruction reveals input patterns and pathways into corticotropin-releasing factor neurons in the central amygdala in mice
Communications Biology (2022)
-
Single-cell morphological characterization of CRH neurons throughout the whole mouse brain
BMC Biology (2021)