Abstract
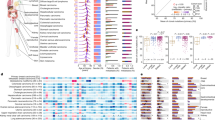
Genomic profiling is critical for the identification of treatment options for patients with metastatic cancer, but it remains unclear how frequently this procedure should be repeated during the course of the disease. To address this, we analyzed whole-genome sequencing (WGS) data of 250 biopsy pairs, longitudinally collected over the treatment course of 231 adult patients with a representative variety of metastatic solid malignancies. Within the biopsy interval (median, 6.4 months), patients received one or multiple lines of (mostly) standard-of-care (SOC) treatments, with all major treatment modalities being broadly represented. SOC biomarkers and biomarkers for clinical trial enrollment could be identified in 23% and 72% of biopsies, respectively. For SOC genomic biomarkers, we observed full concordance between the first and the second biopsy in 99% of pairs. Of the 219 biomarkers for clinical trial enrollment that were identified in the first biopsies, we recovered 94% in the follow-up biopsies. Furthermore, a second WGS analysis did not identify additional biomarkers for clinical trial enrollment in 91% of patients. More-frequent genomic evolution was observed when considering specific genes targeted by small-molecule inhibitors or hormonal therapies (21% and 22% of cases, respectively). Together, our data demonstrate that there is limited evolution of the actionable genome of treated metastases. A single WGS analysis of a metastatic biopsy is generally sufficient to identify SOC genomic biomarkers and to identify investigational treatment opportunities.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw and processed genomics data (for sample identifiers, see Supplementary Table 1) and clinical data are freely available from the Hartwig Medical Foundation through standardized procedures and request forms (www.hartwigmedicalfoundation.nl). Additional clinical data collected for this study, including detailed information about treatments, treatment outcomes and biopsy sites, are available as Supplementary Tables 1–5 of this manuscript.
Code availability
Bioinformatics analyses of WGS data were performed with an optimized pipeline based on open-source tools, which are freely available on GitHub (https://github.com/hartwigmedical/pipeline5).
References
Hyman, D. M., Taylor, B. S. & Baselga, J. Implementing genome-driven oncology. Cell 168, 584–599 (2017).
Stockley, T. L. et al. Molecular profiling of advanced solid tumors and patient outcomes with genotype-matched clinical trials: the Princess Margaret IMPACT/COMPACT trial. Genome Med. 8, 109 (2016).
Larkin, J. et al. Combined vemurafenib and cobimetinib in BRAF-mutated melanoma. N. Engl. J. Med. 371, 1867–1876 (2014).
Marabelle, A. et al. Efficacy of pembrolizumab in patients with noncolorectal high microsatellite instability/mismatch repair-deficient cancer: results from the phase II KEYNOTE-158 study. J. Clin. Oncol. 38, 1–10 (2020).
Riaz, N. et al. Pan-cancer analysis of bi-allelic alterations in homologous recombination DNA repair genes. Nat. Commun. 8, 857 (2017).
Okamura, R. et al. Analysis of NTRK alterations in pan-cancer adult and pediatric malignancies: implications for NTRK-targeted therapeutics. JCO Precis. Oncol. 2, 1–20 (2018).
Bonneville, R. et al. Landscape of microsatellite instability across 39 cancer types. JCO Precis. Oncol. 1, 1–15 (2017).
van der Velden, D. L. et al. The Drug Rediscovery protocol facilitates the expanded use of existing anticancer drugs. Nature 574, 127–131 (2019).
Hyman, D. M. et al. HER kinase inhibition in patients with HER2- and HER3-mutant cancers. Nature 554, 189–194 (2018).
Hyman, D. M. et al. Vemurafenib in multiple nonmelanoma cancers with BRAF V600 mutations. N. Engl. J. Med. 373, 726–736 (2015).
Subbiah, V. et al. Pan-cancer efficacy of vemurafenib in BRAFV600-mutant non-melanoma cancers. Cancer Discov. 10, 657–663 (2020).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Drilon, A. et al. Efficacy of larotrectinib in TRK fusion-positive cancers in adults and children. N. Engl. J. Med. 378, 731–739 (2018).
Aitken, M., Villa, P., Bennett, K., Tewary, V. & Lech, C. Optimizing Oncology Care Through Biomarker Adoption: Barriers and Solutions (IQVIA Institute for Human Data Science, 2020).
Roepman, P. et al. Clinical validation of whole genome sequencing for cancer diagnostics. J. Mol. Diagnostics 23, 816–833 (2021).
Priestley, P. et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature 575, 210–216 (2019).
Reiter, J. G. et al. Minimal functional driver gene heterogeneity among untreated metastases. Science 361, 1033–1037 (2018).
Pich, O. et al. The mutational footprints of cancer therapies. Nat. Genet. 51, 1732–1740 (2019).
Kucab, J. E. et al. A compendium of mutational signatures of environmental agents. Cell 177, 821–836 (2019).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
McGranahan, N. & Swanton, C. Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168, 613–628 (2017).
Johnson, B. E. et al. Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 343, 189–193 (2014).
Visakorpi, T. et al. In vivo amplification of the androgen receptor gene and progression of human prostate cancer. Nat. Genet. 9, 401–406 (1995).
Waltering, K. K., Urbanucci, A. & Visakorpi, T. Androgen receptor (AR) aberrations in castration-resistant prostate cancer. Mol. Cell. Endocrinol. 360, 38–43 (2012).
Robinson, D. R. et al. Activating ESR1 mutations in hormone-resistant metastatic breast cancer. Nat. Genet. 45, 1446–1451 (2013).
Toy, W. et al. ESR1 ligand-binding domain mutations in hormone-resistant breast cancer. Nat. Genet. 45, 1439–1445 (2013).
Angus, L. et al. The genomic landscape of metastatic breast cancer highlights changes in mutation and signature frequencies. Nat. Genet. 51, 1450–1458 (2019).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature 578, 122–128 (2020).
Levine, A. J. p53: 800 million years of evolution and 40 years of discovery. Nat. Rev. Cancer 20, 471–480 (2020).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. 1, 1–16 (2017).
Tamborero, D. et al. Cancer Genome Interpreter annotates the biological and clinical relevance of tumor alterations. Genome Med. 10, 25 (2018).
Griffith, M. et al. CIViC is a community knowledgebase for expert crowdsourcing the clinical interpretation of variants in cancer. Nat. Genet. 49, 170–174 (2017).
Mateo, J. et al. A framework to rank genomic alterations as targets for cancer precision medicine: the ESMO scale for clinical actionability of molecular targets (ESCAT). Ann. Oncol. 29, 1895–1902 (2018).
Fribbens, C. et al. Plasma ESR1 mutations and the treatment of estrogen receptor-positive advanced breast cancer. J. Clin. Oncol. 34, 2961–2968 (2016).
Samsom, K. G. et al. Study protocol: whole genome sequencing implementation in standard diagnostics for every cancer patient (WIDE). BMC Med. Genomics 13, 169 (2020).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. Preprint at bioRxiv https://doi.org/10.1101/201178 (2017).
Kim, S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods 15, 591–594 (2018).
Cameron, D. L. et al. GRIDSS, PURPLE, LINX: unscrambling the tumor genome via integrated analysis of structural variation and copy number. Preprint at bioRxiv https://doi.org/10.1101/781013 (2019).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041 (2017).
Forbes, S. A. et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res. 45, D777–D783 (2017).
Ni Huang, M. et al. MSIseq: software for assessing microsatellite instability from catalogs of somatic mutations. Sci. Rep. 5, 13321 (2015).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082 (2018).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Acknowledgements
We acknowledge the Hartwig Medical Foundation and the Center of Personalised Cancer Treatment (CPCT) for providing the data used in this the study. This work was supported by funding from the Josephine Nefkens Foundation and the Oncode Institute.
Author information
Authors and Affiliations
Contributions
Original concept was developed by J.v.d.H., L.R.H. and E.E.V. Coordination of clinical data collection was carried out by L.R.H. Representatives of main patient accrual sites: M.P.L., H.M.W.V., H.G., A.J.d.L., E.F.S. and E.E.V. Primary data analyses were performed by J.v.d.H., L.R.H. and P.R. Coordination of genomics data collection was done by E.C. The initial draft was written by J.v.d.H., L.R.H., L.F.A.W. and E.E.V. Figures were created by J.v.d.H. All authors critically reviewed all analyses and results of this work and edited and approved the final manuscript. This work was jointly supervised by L.F.A.W. and E.E.V.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Medicine thanks Mark Cowley, Funda Meric-Bernstam and the other, anonymous, reviewer(s) for their contribution to the peer review this work. Javier Carmona was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Tumor type distribution of the cohort.
Percentage of biopsies per anatomical site of primary tumor, for the paired biopsy cohort (red; n = 481) and full HMF cohort as published (gray; n = 2921).
Extended Data Fig. 2 Tumor purity differences among paired biopsies.
Barplot denoting the absolute difference (Δ) of tumor purity for each paired biopsy (n = 250), where positive values indicate an increase in the second biopsy as compared to the first. The samples are ranked from high to low Δ purity. The P-value for the two-sided paired Wilcoxon test is shown.
Extended Data Fig. 3 The influence of intertumoral heterogeneity on the discordance of genome-wide somatic profiles between paired biopsies.
a, Boxplot of the fraction of genomic alterations private to the first biopsy (lost alterations, as a fraction of the total number of alterations in the first biopsy), stratified for pairs obtained from the same lesion (n = 83 pairs) or pairs obtained from different lesions (n = 92 pairs). Different classes of genomic alterations are plotted separately: nucleotide substitutions (all, clonal, subclonal), indels (all, clonal, subclonal), structural variants (SVs). Clonal alterations have a probability of clonality above >95%, whereas subclonal alterations have a probability of subclonality >95%. Box center lines, box ranges, whiskers, and dots indicate medians, quartiles, 1.5 interquartile ranges, and outliers, respectively. Two-sided Wilcoxon rank sum test-based P-values are shown (red for significant associations). Only paired biopsies for which it was known whether or not the two biopsies were obtained from the same lesion were used for this analysis (n = 175). b, As A, but the y-axis represents the fraction of alterations private to the second biopsy (gained alterations, as a fraction of the total number of alterations in the second biopsy). c, As A, but the y-axis indicates the Pearson correlation coefficient for the copy number profiles of biopsy pairs.
Extended Data Fig. 4 The influence of time between the biopsies on the discordance of genome-wide somatic profiles between paired biopsies.
a, Dot plots of the ascsin-transformed fractions of genomic alterations private to the first biopsy (lost alterations, as a fraction of the total number of alterations in the first biopsy; top row of plots; y-axis), or the ascsin-transformed fraction of genomic alterations private to the second biopsy (gained alterations, as a fraction of the total number of alterations in the second biopsy; bottom row of plots; y-axis) for all biopsy pairs (n = 250), against the time between biopsies (x-axis). Different classes of somatic alterations are plotted separately: nucleotide substitutions (clonal, subclonal), indels (clonal, subclonal), and structural variants (SVs), as indicated at each subplot’s title. Clonal alterations have a probability of clonality above >95%, whereas subclonal alterations have a probability of subclonality >95%. Arcsin transformation was performed to achieve normality of the inherently non-normally distributed fractions. Linear fits and bootstrapping-obtained 95% confidence intervals are plotted. Pearson correlation coefficients (r) and accompanying two-sided P-values are shown (red for significant associations). b, As A, but the y-axis indicates the Pearson correlation coefficient (r) for the copy number profiles of biopsy pairs. Analysis is based on all paired biopsies (n = 250).
Extended Data Fig. 5 The number of standard of care (left) or investigational (right) biomarkers per biopsy, for patients included in the DRUP study (blue) vs all others (orange).
The y-axis denotes the percentage of biopsies relative to the total number of biopsies from patients in the DRUP study (blue; n = 140 biopsies) or all other patients (orange; n = 341 biopsies). Some patients included in the DRUP study lack actionable biomarkers, for two reasons. First, some patients were included in the DRUP study based on a high tumor mutational burden, which is not considered as an actionable biomarker in the current study. Second, some patients were included in the DRUP study based on the absence of a particular resistance marker (for example, the absence of a KRAS mutation), without the need for the presence of a particular biomarker. Wilcoxon rank sum test-based two-sided P-values are shown.
Supplementary information
Supplementary Tables
Supplementary Table 1 | Treatment information and clinical outcome of all regimens administered within the biopsy interval. Supplementary Table 2 | Somatic genomic drivers detected in the cohort. Supplementary Table 3 | Comparison of actionable genomic alterations between paired biopsies. Supplementary Table 4 | Comparison of genomic alterations in drug targets between paired biopsies, for treatments with small-molecule inhibitors, hormonal therapies and monoclonal antibodies. Supplementary Table 5 | Overview of all drugs and their targets used in this study.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
van de Haar, J., Hoes, L.R., Roepman, P. et al. Limited evolution of the actionable metastatic cancer genome under therapeutic pressure. Nat Med 27, 1553–1563 (2021). https://doi.org/10.1038/s41591-021-01448-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-021-01448-w