Abstract
We report a new immunodeficiency disorder in mice caused by a viable hypomorphic mutation of Snrnp40, an essential gene encoding a subunit of the U5 small nuclear ribonucleoprotein (snRNP) complex of the spliceosome. Snrnp40 is ubiquitous but strongly expressed in lymphoid tissue. Homozygous mutant mice showed hypersusceptibility to infection by murine cytomegalovirus and multiple defects of lymphoid development, stability and function. Cell-intrinsic defects of hematopoietic stem cell differentiation also affected homozygous mutants. SNRNP40 deficiency in primary hematopoietic stem cells or T cells or the EL4 cell line increased the frequency of splicing errors, mostly intron retention, in several hundred messenger RNAs. Altered expression of proteins associated with immune cell function was also observed in Snrnp40-mutant cells. The immunological consequences of SNRNP40 deficiency presumably result from cumulative, moderate effects on processing of many different mRNA molecules and secondary reductions in the expression of critical immune proteins, yielding a syndromic immune disorder.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The main data that support the findings of this study are available in the article and its supplementary figures. Data are available from the corresponding author upon reasonable request.
Code availability
The code that supports the findings of this study is available from the corresponding author upon request.
References
Rieger, M. A. & Schroeder, T. Hematopoiesis. Cold Spring Harb. Perspect. Biol. 4, a008250 (2012).
Turner, I. A., Norman, C. M., Churcher, M. J. & Newman, A. J. Roles of the U5 snRNP in spliceosome dynamics and catalysis. Biochem. Soc. Trans. 32, 928–931 (2004).
Will, C. L. & Lührmann, R. Spliceosome structure and function. Cold Spring Harb. Perspect. Biol. 3, a003707 (2011).
Achsel, T., Ahrens, K., Brahms, H., Teigelkamp, S. & Lührmann, R. The human U5-220kD protein (hPrp8) forms a stable RNA-free complex with several U5-specific proteins, including an RNA unwindase, a homologue of ribosomal elongation factor EF-2, and a novel WD-40 protein. Mol. Cell. Biol. 18, 6756–6766 (1998).
Reyes, J. L., Kois, P., Konforti, B. B. & Konarska, M. M. The canonical GU dinucleotide at the 5′ splice site is recognized by p220 of the U5 snRNP within the spliceosome. RNA 2, 213–225 (1996).
Teigelkamp, S., Whittaker, E. & Beggs, J. D. Interaction of the yeast splicing factor PRP8 with substrate RNA during both steps of splicing. Nucleic Acids Res. 23, 320–326 (1995).
Wyatt, J. R., Sontheimer, E. J. & Steitz, J. A. Site-specific cross-linking of mammalian U5 snRNP to the 5′ splice site before the first step of pre-mRNA splicing. Genes Dev. 6, 2542–2553 (1992).
Chiara, M. D., Palandjian, L., Feld Kramer, R. & Reed, R. Evidence that U5 snRNP recognizes the 3′ splice site for catalytic step II in mammals. EMBO J. 16, 4746–4759 (1997).
Teigelkamp, S., Newman, A. J. & Beggs, J. D. Extensive interactions of PRP8 protein with the 5′ and 3′ splice sites during splicing suggest a role in stabilization of exon alignment by U5 snRNA. EMBO J. 14, 2602–2612 (1995).
Umen, J. G. & Guthrie, C. A novel role for a U5 snRNP protein in 3′ splice site selection. Genes Dev. 9, 855–868 (1995).
Bartels, C., Klatt, C., Lührmann, R. & Fabrizio, P. The ribosomal translocase homologue Snu114p is involved in unwinding U4/U6 RNA during activation of the spliceosome. EMBO Rep. 3, 875–880 (2002).
Brenner, T. J. & Guthrie, C. Genetic analysis reveals a role for the C terminus of the Saccharomyces cerevisiae GTPase Snu114 during spliceosome activation. Genetics 170, 1063–1080 (2005).
Maeder, C., Kutach, A. K. & Guthrie, C. ATP-dependent unwinding of U4/U6 snRNAs by the Brr2 helicase requires the C terminus of Prp8. Nat. Struct. Mol. Biol. 16, 42–48 (2009).
Small, E. C., Leggett, S. R., Winans, A. A. & Staley, J. P. The EF-G-like GTPase Snu114p regulates spliceosome dynamics mediated by Brr2p, a DExD/H box ATPase. Mol. Cell 23, 389–399 (2006).
Wang, T. et al. Real-time resolution of point mutations that cause phenovariance in mice. Proc. Natl Acad. Sci. USA 112, E440–E449 (2015).
Biron, C. A., Nguyen, K. B., Pien, G. C., Cousens, L. P. & Salazar-Mather, T. P. Natural killer cells in antiviral defense: function and regulation by innate cytokines. Annu. Rev. Immunol. 17, 189–220 (1999).
Laggerbauer, B. et al. The human U5 snRNP 52K protein (CD2BP2) interacts with U5-102K (hPrp6), a U4/U6.U5 tri-snRNP bridging protein, but dissociates upon tri-snRNP formation. RNA 11, 598–608 (2005).
Wang, T. et al. Probability of phenotypically detectable protein damage by ENU-induced mutations in the Mutagenetix database. Nat. Commun. 9, 441 (2018).
Dickinson, M. E. et al. High-throughput discovery of novel developmental phenotypes. Nature 537, 508–514 (2016).
Ayadi, A. et al. Mouse large-scale phenotyping initiatives: overview of the European Mouse Disease Clinic (EUMODIC) and of the Wellcome Trust Sanger Institute Mouse Genetics Project. Mamm. Genome 23, 600–610 (2012).
White, J. K. et al. Genome-wide generation and systematic phenotyping of knockout mice reveals new roles for many genes. Cell 154, 452–464 (2013).
Růžičková, Š. & Staněk, D. Mutations in spliceosomal proteins and retina degeneration. RNA Biol. 14, 544–552 (2017).
Verma, B., Akinyi, M. V., Norppa, A. J. & Frilander, M. J. Minor spliceosome and disease. Semin. Cell Dev. Biol. 79, 103–112 (2018).
Kurtovic-Kozaric, A. et al. PRPF8 defects cause missplicing in myeloid malignancies. Leukemia 29, 126–136 (2015).
Park, J. W., Parisky, K., Celotto, A. M., Reenan, R. A. & Graveley, B. R. Identification of alternative splicing regulators by RNA interference in Drosophila. Proc. Natl Acad. Sci. USA 101, 15974–15979 (2004).
Pleiss, J. A., Whitworth, G. B., Bergkessel, M. & Guthrie, C. Transcript specificity in yeast pre-mRNA splicing revealed by mutations in core spliceosomal components. PLoS Biol. 5, e90 (2007).
Papasaikas, P., Tejedor, J. R., Vigevani, L. & Valcárcel, J. Functional splicing network reveals extensive regulatory potential of the core spliceosomal machinery. Mol. Cell 57, 7–22 (2015).
Inoue, D., Bradley, R. K. & Abdel-Wahab, O. Spliceosomal gene mutations in myelodysplasia: molecular links to clonal abnormalities of hematopoiesis. Genes Dev. 30, 989–1001 (2016).
Wahl, M. C., Will, C. L. & Lührmann, R. The spliceosome: design principles of a dynamic RNP machine. Cell 136, 701–718 (2009).
Kakugawa, K. et al. A novel gene essential for the development of single positive thymocytes. Mol. Cell. Biol. 29, 5128–5135 (2009).
Fu, G. et al. Themis controls thymocyte selection through regulation of T cell antigen receptor-mediated signaling. Nat. Immunol. 10, 848–856 (2009).
Johnson, A. L. et al. Themis is a member of a new metazoan gene family and is required for the completion of thymocyte positive selection. Nat. Immunol. 10, 831–839 (2009).
Lesourne, R. et al. Themis, a T cell-specific protein important for late thymocyte development. Nat. Immunol. 10, 840–847 (2009).
Fu, G. et al. Themis sets the signal threshold for positive and negative selection in T-cell development. Nature 504, 441–445 (2013).
Bar, E., Whitney, P. G., Moor, K., Reis e Sousa, C. & LeibundGut-Landmann, S. IL-17 regulates systemic fungal immunity by controlling the functional competence of NK cells. Immunity 40, 117–127 (2014).
Hosoya, T. et al. GATA-3 is required for early T lineage progenitor development. J. Exp. Med. 206, 2987–3000 (2009).
Pai, S. Y. et al. Critical roles for transcription factor GATA-3 in thymocyte development. Immunity 19, 863–875 (2003).
Ting, C. N., Olson, M. C., Barton, K. P. & Leiden, J. M. Transcription factor GATA-3 is required for development of the T-cell lineage. Nature 384, 474–478 (1996).
Wang, L. et al. Distinct functions for the transcription factors GATA-3 and ThPOK during intrathymic differentiation of CD4+ T cells. Nat. Immunol. 9, 1122–1130 (2008).
Barton, K. et al. The Ets-1 transcription factor is required for the development of natural killer cells in mice. Immunity 9, 555–563 (1998).
Clements, J. L., John, S. A. & Garrett-Sinha, L. A. Impaired generation of CD8+ thymocytes in Ets-1-deficient mice. J. Immunol. 177, 905–912 (2006).
Higuchi, T. et al. Thymomegaly, microsplenia, and defective homeostatic proliferation of peripheral lymphocytes in p51-Ets1 isoform-specific null mice. Mol. Cell. Biol. 27, 3353–3366 (2007).
Nuez, B., Michalovich, D., Bygrave, A., Ploemacher, R. & Grosveld, F. Defective haematopoiesis in fetal liver resulting from inactivation of the EKLF gene. Nature 375, 316–318 (1995).
White, R. A. et al. Hematologic characterization and chromosomal localization of the novel dominantly inherited mouse hemolytic anemia, neonatal anemia (Nan). Blood Cells Mol. Dis. 43, 141–148 (2009).
Foudi, A. et al. Distinct, strict requirements for Gfi-1b in adult bone marrow red cell and platelet generation. J. Exp. Med. 211, 909–927 (2014).
Vassen, L. et al. Growth factor independence 1b (gfi1b) is important for the maturation of erythroid cells and the regulation of embryonic globin expression. PLoS ONE 9, e96636 (2014).
Saleque, S., Cameron, S. & Orkin, S. H. The zinc-finger proto-oncogene Gfi-1b is essential for development of the erythroid and megakaryocytic lineages. Genes Dev. 16, 301–306 (2002).
Mayle, A., Luo, M., Jeong, M. & Goodell, M. A. Flow cytometry analysis of murine hematopoietic stem cells. Cytometry A 83, 27–37 (2013).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Carbon, S. et al. Expansion of the Gene Ontology knowledgebase and resources. Nucleic Acids Res. 45, D331–D338 (2017).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Acknowledgements
This work was supported by National Institutes of Health grants (nos. AI100627, AI125581 and GM067759 to B.B.). Funding for this project was provided by the Rheumatology Research Foundation Tobé and Stephen E. Malawista, MD Endowment in Academic Rheumatology (to E.N.-G.).
Author information
Authors and Affiliations
Contributions
D.Z. and B.B. designed the research. D.Z., T.Y., J.H.C., E.N.-G., X. Zhong, K.-w.W., X. Zhan, X.L., M.C., M.T. and J.Q. performed the research. S.H. performed the computational analysis. D.Z. and B.B. analyzed the data. D.Z., E.M.Y.M. and B.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Laurie Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Immune phenotypes detected in skywarp mice from 17 unrelated pedigrees derived from the C57BL/6J stock.
a-d, Lower panels, frequency of (a) CD3+ T cells, (b) CD4+ T cells, (c) CD8+ T cells, and (d) B220+ B cells among total white blood cells isolated from the blood of mice from 17 unrelated pedigrees carrying ENU-induced mutations. Upper panels, Manhattan (Mht) plots showing P values of association between the phenotypes in the lower panels and mutations (ENU-induced and spontaneous) identified in the 17 pedigrees, calculated using a recessive model of inheritance. e, Lower panel, B cell to T cell ratio in the blood of mice from the same 17 pedigrees. Upper panel, Mht plot showing the P values of association between the elevated B:T ratio and mutations identified in the 17 pedigrees, calculated using a recessive model of inheritance. f, Lower panel, MCMV titer in the spleen of mice from 5 of the 17 pedigrees five days after infection with MCMV. Upper panel, Mht plot showing the P values of association between elevated MCMV titer and mutations identified in the 5 pedigrees, calculated using a recessive model of inheritance. For scatter plots: C57BL/6J, REF (Snrnp40+/+), HET (Snrnp40swp/+), and VAR (Snrnp40swp/swp). Each data point represents one mouse. Mean (μ) and SD (σ) are indicated. For Mht plots: The −Log10 P values (Y axis) were plotted versus the chromosomal positions of mutations (X axis). Horizontal red and purple lines represent thresholds of P = 0.05 with or without Bonferroni correction, respectively. REF n = 168, HET n = 212, VAR n = 29 in a-e; REF n = 56, HET n = 55, VAR n = 8 in f. Ten to twenty C57BL/6J wild-type mice (total n in scatter plots not counted) were included as controls in each screening experiment but were not used for P value calculation (a-f). All P values in upper panels (a-f) were calculated using generalized linear model. P = 3.274 × 10−24 in a (raw data, with G2, recessive model), P = 8.547 × 10−19 in b (raw data, with G2, recessive model), P = 1.044 × 10−26 in c (raw data, with G2, recessive model), P = 0.045939 in d (raw data, with G2, recessive model), P = 3.22 × 10−29 in e (normalized data, with G2, recessive model), P = 1.614 × 10−19 in f (log data, with G2, recessive model).
Supplementary Figure 2 Detection of a spontaneous mutation in Snrnp40 in the C57BL/6J stock.
a, DNA sequencing chromatograms. Direct capillary sequencing of 250 non-mutagenized C57BL/6J stock mice from JAX detected heterozygosity for the swp mutation in two animals. The region containing the swp mutation in the two C57BL/6J stock mice is shown. b, Multiple sequence alignment of Snrnp40 amino acid sequence in 11 species. Alignment was performed using Clustal Omega program, and reformatted by MView tool. cov, percent coverage; pid, percent identity with respect to the first sequence (human). Identities normalized by aligned length. Colors indicate identical sequence. *Red box denotes the missing sequence caused by the swp mutation. Equivelence
Supplementary Figure 3 Snrnp40 expression profile and the Snrnp40rplc allele.
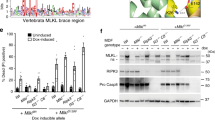
a,b, RT−qPCR analysis of Snrnp40 mRNA levels in various mouse tissues normalized by Gapdh mRNA levels using 2–ΔΔCt method (a), or by total RNA levels used for reverse transcription using 2–ΔCt method (b). n = 3 wild-type C57BL/6J littermate mice in a,b. c, Immunoblot analysis of SNRNP40 in various mouse tissues. d, Immunoblot analysis of Snrnp40 in splenic B cells, T cells, and NK cells. e, DNA sequencing chromatograms. The region containing the swp mutation (C→G) and the marker mutation (C→T) in Snrnp40+/+, Snrnp40rplc/+, and Snrnp40swp/rplc mice is shown. f, RT−PCR amplification across exon 5 of Snrnp40 mRNA from splenocytes of Snrnp40+/+, Snrnp40rplc/+, and Snrnp40rplc/swp mice. g, Immunoblot analysis of SNRNP40 in splenocytes of the indicated genotypes. Data are representative of five independent experiments with similar results. h, Body weight of female and male mice at 9 weeks of age (n = 4 for each genotype). P values were determined by one-way ANOVA with Sidak’s multiple comparisons test. *P = 0.031, **P = 0.0015, ***P = 0.0009, ****P < 0.0001. i, Representative images of Snrnp40swp/rplc mice and Snrnp40swp/+ littermates. Snrnp40swp/rplc mice display reduced body size, some white back hairs, and a patch of white hair on the belly. Data are representative of two (a-c), three (d,f,h),or five (g) independent experiments (mean ± s.d. in a,b,h).
Supplementary Figure 4 Analysis of immune cells in the blood of Snrnp40swp/rplc mice.
a-l, Hematological analysis of blood from Snrnp40swp/rplc and Snrnp40swp/+ littermates (n = 6 mice per genotype). Neutrophil (a), eosinophil (b), basophil (c), red blood cells (RBC) (d), hemoglobin (e), hematocrit (HCT) (f), red cell distribution width (RDW) (g), mean cell volume (MCV) (h), mean cell hemoglobin (MCH) (i), mean corpuscular hemoglobin concentration (MCHC) (j), platelet (k), and mean platelet volume (MPV) (l) were determined. The indicated P values were determined by unpaired, two-tailed Student’s t-test. m,n, Flow cytometry of cells in samples containing equal volumes of blood isolated from Snrnp40swp/rplc and Snrnp40swp/+ mice, assessing the absolute number ratio of B cells (B220+), T cells (CD3ε+), and NK cells (NK1.1+CD3ε–) (m), or macrophages (CD11b+F4/80+) and neutrophils (CD11b+F4/80–) (n) in Snrnp40swp/rplc and Snrnp40swp/+ littermates. Numbers indicate percent cells in outlined areas. o, Frequency of macrophages and neutrophils in the mixed blood samples. n.s., not significant (n = 5 mice per genotype; P values indicated; unpaired, two-tailed Student’s t-test). Data are representative of two (n,o), three (m), or four (a-l) independent experiments (mean ± s.d. in a-l,o).
Supplementary Figure 5 Defective cell cycle progression of Snrnp40swp/swp T cells and normal adaptive immune responses in Snrnp40swp/swp mice.
a, Mitotic cell cycle analysis. Flow cytometry of splenic T cells activated by TCR stimulation for three days and then labeled with EdU for 6 h, assessed for EdU incorporation and 7-AAD staining. Dotted line denotes the average size of T cells at G0/G1 stage. Numbers indicate percent cells in outlined areas. b, Quantification of the percentage of T cells in the indicated stage of the cell cycle, or the percentage of apoptotic cells determined as in (a). n.s., not significant (P = 0.80), *P = 0.021, **P = 0.0039, *P = 0.016; unpaired, two-tailed Student’s t-test. c, Wild-type recipient mice (C57BL/6J, CD45.1) adoptively transferred with CellTrace Far Red-labeled Snrnp40swp/swp or wild-type OT-I CD8+ T cells (CD45.2). Flow cytometry of CellTrace Far Red dilution in Snrnp40swp/swp or wild-type OT-I CD8+ T cells isolated from the spleens of recipient mice 60 or 72 h after immunization with OVA or PBS. d, Wild-type recipient mice (C57BL/6J, CD45.1) adoptively transferred with CellTrace Far Red-labeled Snrnp40swp/swp or wild-type OT-II CD4+ T cells (CD45.2). Flow cytometry of CellTrace Far Red dilution in Snrnp40swp/swp or wild-type OT-II CD4+ T cells isolated from the spleens of recipient mice 72 h after immunization with OVA or PBS. e, Quantification of the percentage of OT-I (CD8+) and OT-II (CD4+) cells that underwent proliferation in response to OVA immunization measured 72 h after immunization (n = 4 mice per genotype; n.s., not significant (P = 0.94, 0.95); unpaired, two-tailed Student’s t-test). f,g, Antibody responses to immunization. T-dependent antibody response to rSFV-β-gal (f) and T-independent antibody response to NP-Ficoll (g) in Snrnp40+/+ (n = 6), Snrnp40swp/+ (n = 10), and Snrnp40swp/swp (n = 6) mice were measured 13 days and 6 days after immunization, respectively. n.s., not significant (P = 0.87 in f, P = 0.05 in g); one-way ANOVA with Sidak’s multiple comparisons. h, In vivo CTL cytotoxicity in Snrnp40+/+ (n = 6), Snrnp40swp/+ (n = 10), and Snrnp40swp/swp (n = 6) mice. n.s., not significant (P = 0.19); one-way ANOVA with Sidak’s multiple comparisons. Data are representative of two (f-h) and three (a-e) independent experiments (mean ± s.d. in b,e,f-h).
Supplementary Figure 6 Normal Snrnp40swp/swp NK cell degranulation and BMDC cytokine production.
a-d, Flow cytometry of NK cells from the spleens of Snrnp40swp/swp and wild-type mice assessing the expression of CD107a (LAMP-1) (a), granzyme B (b), TNF (c), and IFN-γ (d) in response to stimulation with PMA/ionomycin (50 ng/ml PMA and 1 μg/ml ionomycin) or no stimulation. e-h, Concentration of TNF (e), IL-6 (f), IFN-α (g), and IFN-β (h) in the supernatants of Snrnp40swp/swp and wild-type BMDCs 16 hours after infection with the indicated viruses and multiplicities of infection (MOI) (n = 4 independent BMDCs, each generated from independent mice; *P = 0.015, *P = 0.014, ***P = 0.00066, *P = 0.022; unpaired, two-tailed Student’s t-test). Data are representative of three independent experiments (a-h; mean ± s.d. in e-h).
Supplementary Figure 7 Hematopoietic cell-intrinsic defect in lymphoid development in Snrnp40swp/rplc mutant mice.
a-h, Flow cytometry of immune cells in the blood of mixed bone marrow chimeras reconstituted with a 1:1 mixture of wild-type (CD45.1) and Snrnp40swp/rplc (CD45.2) bone marrow cells, assessing the percentage of B cells (a, B220+), CD3+ T cells (b, CD3ε+), CD4+ T cells (c, CD3ε+CD4+), CD8+ T cells (d, CD3ε+CD8α+), NK cells (e, NK-1.1+CD3ε–), NKT cells (f, NK-1.1+CD3ε+), macrophages (g, CD11b+F4/80+), and neutrophils (h, CD11b+F4/80–) derived from wild-type (CD45.1) and Snrnp40swp/rplc (CD45.2) bone marrows respectively. Representative results from one of ten chimeras in two experiments are shown (a-h). i-j, Gating strategy for flow cytometry analysis of hematopoietic stem and progenitor cells in Snrnp40swp/+ (i) and Snrnp40swp/rplc (j) bone marrow. Single cells negative for Lineage expression (Lineage–) were further classified into three populations based on c-Kit and Sca-1 expression: LKS (c-Kit+Sca-1+), LK (c-Kit+Sca-1–), and KloSlo (c-KitloSca-1lo). The LKS population was classified into three populations based on Flk-2 and CD34 expression: long-term hematopoietic stem cells (LT-HSCs, CD34–Flk-2–), short-term hematopoietic stem cells (ST-HSCs, CD34+Flk-2–), and multipotent progenitor cells (MPPs, CD34+Flk-2+). The LK population was classified into three populations based on CD16/32 and CD34 expression: megakaryocyte–erythroid progenitors (MEPs, CD34–CD16/32–), common myeloid progenitors (CMPs, CD34+CD16/32–), and granulocyte-macrophage progenitors (GMPs, CD34+CD16/32+). The KloSlo population expressing IL-7Rα was defined as common lymphoid progenitors (CLPs, IL-7Rα+). Representative results from one of three independent experiments are shown (i,j).
Supplementary Figure 8 Coimmunoprecipitation of SNRNP40-interacting proteins for analysis by mass spectrometry.
a, Immunoblot analysis of SNRNP40, SNRNP200, EFTUD2 and CD2BP2 in mouse splenocytes of the indicated genotypes. Experiment was repeated three times independently with similar results. b-d, Identification of SNRNP40-interacting proteins by mass spectrometry using mouse EL4 cells. b, Silver stained gel containing proteins immunoprecipitated from EL4 cells stably expressing FLAG-tagged mouse SNRNP40 (FLAG−SNRNP40) or vector alone. Arrows indicate bands present exclusively or with greater intensity in FLAG−SNRNP40 immunoprecipitate. c, Analysis of FLAG−SNRNP40 expression in EL4 cells by immunoblot of whole cell lysate or by immunoprecipitation and immunoblot. d, Venn diagram for top 83 proteins identified by mass spectrometry that coimmunoprecipitated with FLAG−SNRNP40 (top 25 candidates present only in SNRNP40 sample, and top 58 candidates with spectral index ratio > 7 in SNRNP40 vs. vector samples). 80.7% (67 proteins) were known splicing factors or regulators. e-g, Identification of SNRNP40−interacting proteins by mass spectrometry using human NK-92 cells. e, Silver stained gel containing proteins immunoprecipitated from NK-92 cells stably expressing FLAG-tagged human SNRNP40 (FLAG−SNRNP40) or vector alone. Arrows indicate bands present exclusively or with greater intensity in FLAG−SNRNP40 immunoprecipitate. f, Analysis of FLAG−SNRNP40 expression in NK-92 cells by immunoblot of whole cell lysate or by immunoprecipitation and immunoblot. g, Venn diagram for top 111 proteins identified by mass spectrometry that coimmunoprecipitated with FLAG−SNRNP40 (top 63 candidates present only in SNRNP40 sample, and top 48 candidates with spectral index ratio > 24 in SNRNP40 vs. vector samples). 64.0% (71 proteins) were known splicing factors or regulators. Experiment was performed one time (b-g).
Supplementary information
Supplementary Information
Supplementary Figs. 1–11 and Supplementary Tables 1–5
Supplementary Dataset 1
Mass spectrometric identification of Snrnp40-binding proteins in mouse EL4 cells
Supplementary Dataset 2
Mass spectrometric identification of SNRNP40-binding proteins in human NK-92 cells
Supplementary Dataset 3
Splicing analysis of RNA-seq data from Snrnp40-KO and wild-type EL4 cells
Supplementary Dataset 4
Validated intron retention in Snrnp40-mutant cells
Supplementary Dataset 5
Splicing analysis of RNA-seq data from Snrnp40+/+, Snrnp40swp/+ and Snrnp40swp/rplc HSPCs
Supplementary Dataset 6
Splicing analysis of RNA-seq data from Snrnp40+/+, Snrnp40swp/+ and Snrnp40swp/rplc T cells
Supplementary Dataset 7
Differentially expressed genes in Snrnp40-KO versus wild-type EL4 cells
Supplementary Dataset 8
DAVID analysis of differentially expressed genes in Snrnp40-KO versus wild-type EL4 cells
Supplementary Dataset 9
Differentially expressed genes in Snrnp40+/+, Snrnp40swp/+ and Snrnp40swp/rplc HSPCs
Supplementary Dataset 10
DAVID analysis of differentially expressed genes in Snrnp40+/+, Snrnp40swp/+ and Snrnp40swp/rplc HSPCs
Supplementary Dataset 11
Differentially expressed genes in Snrnp40+/+, Snrnp40swp/+ and Snrnp40swp/rplc T cells
Supplementary Dataset 12
DAVID analysis of differentially expressed genes in Snrnp40+/+, Snrnp40swp/+ and Snrnp40swp/rplc T cells
Rights and permissions
About this article
Cite this article
Zhang, D., Yue, T., Choi, J.H. et al. Syndromic immune disorder caused by a viable hypomorphic allele of spliceosome component Snrnp40. Nat Immunol 20, 1322–1334 (2019). https://doi.org/10.1038/s41590-019-0464-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0464-4
This article is cited by
-
OVOL2 sustains postnatal thymic epithelial cell identity
Nature Communications (2023)
-
Emerging roles of spliceosome in cancer and immunity
Protein & Cell (2022)