Abstract
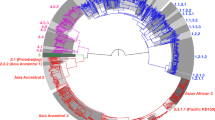
To examine the transmission dynamics of Mycobacterium tuberculosis (Mtb) isolated from tuberculosis patients in Ho Chi Minh City, Vietnam, we sequenced the whole genomes of 1,635 isolates and compared these with 3,144 isolates from elsewhere. The data identify an underlying burden of disease caused by the endemic Mtb lineage 1 associated with the activation of long-term latent infection, and a threefold higher burden associated with the more recently introduced Beijing lineage and lineage 4 Mtb strains. We find that Beijing lineage Mtb is frequently transferred between Vietnam and other countries, and detect higher levels of transmission of Beijing lineage strains within this host population than the endemic lineage 1 Mtb. Screening for parallel evolution of Beijing lineage-associated SNPs in other Mtb lineages as a signal of positive selection, we identify an alteration in the ESX-5 type VII-secreted protein EsxW, which could potentially contribute to the enhanced transmission of Beijing lineage Mtb in Vietnamese and other host populations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Zumla, A. et al. Eliminating tuberculosis and tuberculosis–HIV co-disease in the 21st century: key perspectives, controversies, unresolved issues, and needs. J. Infect. Dis. 205, S141–S146 (2012).
Global Tuberculosis Report 2017 (World Health Organization, 2017).
Casali, N. et al. Evolution and transmission of drug-resistant tuberculosis in a Russian population. Nat. Genet. 46, 279–286 (2014).
Guerra-Assuncao, J. A. et al. Large-scale whole genome sequencing of M. tuberculosis provides insights into transmission in a high prevalence area. eL ife https://doi.org/10.7554/eLife.05166 (2015).
Guerra-Assuncao, J. A. et al. Recurrence due to relapse or reinfection with Mycobacterium tuberculosis: a whole-genome sequencing approach in a large, population-based cohort with a high HIV infection prevalence and active follow-up. J. Infect. Dis. 211, 1154–1163 (2015).
Coll, F. et al. A robust SNP barcode for typing Mycobacterium tuberculosis complex strains. Nat. Commun. 5, 4812 (2014).
Anh, D. D. et al. Mycobacterium tuberculosis Beijing genotype emerging in Vietnam. Emerg. Infect. Dis. 6, 302–305 (2000).
Buu, T. N. et al. The Beijing genotype is associated with young age and multidrug-resistant tuberculosis in rural Vietnam. Int. J. Tuberc. Lung Dis. 13, 900–906 (2009).
Maeda, S. et al. Mycobacterium tuberculosis strains spreading in Hanoi, Vietnam: Beijing sublineages, genotypes, drug susceptibility patterns, and host factors. Tuberculosis 94, 649–656 (2014).
Nguyen, V. A. et al. High prevalence of Beijing and EAI4-VNM genotypes among M. tuberculosis isolates in northern Vietnam: sampling effect, rural and urban disparities. PLoS One 7, e45553 (2012).
Nguyen, V. A. et al. Mycobacterium tuberculosis lineages and anti-tuberculosis drug resistance in reference hospitals across Viet Nam. BMC Microbiol. 16, 167 (2016).
Horton, K. C., MacPherson, P., Houben, R. M., White, R. G. & Corbett, E. L. Sex differences in tuberculosis burden and notifications in low- and middle-income countries: a systematic review and meta-analysis. PLoS Med. 13, e1002119 (2016).
Malla, B. et al. First insights into the phylogenetic diversity of Mycobacterium tuberculosis in Nepal. PLoS One 7, e52297 (2012).
Lee, C. H. et al. Treatment delay and fatal outcomes of pulmonary tuberculosis in advanced age: a retrospective nationwide cohort study. BMC Infect. Dis. 17, 449 (2017).
Wlodarska, M., Johnston, J. C., Gardy, J. L. & Tang, P. A microbiological revolution meets an ancient disease: improving the management of tuberculosis with genomics. Clin. Microbiol. Rev. 28, 523–539 (2015).
Eldholm, V. et al. Four decades of transmission of a multidrug-resistant Mycobacterium tuberculosis outbreak strain. Nat. Commun. 6, 7119 (2015).
Zhang, H. et al. Genome sequencing of 161 Mycobacterium tuberculosis isolates from China identifies genes and intergenic regions associated with drug resistance. Nat. Genet. 45, 1255–1260 (2013).
Merker, M. et al. Evolutionary history and global spread of the Mycobacterium tuberculosis Beijing lineage. Nat. Genet. 47, 242–249 (2015).
Comas, I. et al. Population genomics of Mycobacterium tuberculosis in Ethiopia contradicts the virgin soil hypothesis for human tuberculosis in sub-Saharan Africa. Curr. Biol. 25, 3260–3266 (2015).
Phelan, J. et al. Mycobacterium tuberculosis whole genome sequencing and protein structure modelling provides insights into anti-tuberculosis drug resistance. BMC Med. 14, 31 (2016).
Stucki, D. et al. Mycobacterium tuberculosis lineage 4 comprises globally distributed and geographically restricted sublineages. Nat. Genet. 48, 1535–1543 (2016).
Hanekom, M. et al. Mycobacterium tuberculosis Beijing genotype: a template for success. Tuberculosis 91, 510–523 (2011).
Parwati, I., van Crevel, R. & van Soolingen, D. Possible underlying mechanisms for successful emergence of the Mycobacterium tuberculosis Beijing genotype strains. Lancet Infect. Dis. 10, 103–111 (2010).
Coscolla, M. & Gagneux, S. Consequences of genomic diversity in Mycobacterium tuberculosis. Semin. Immunol. 26, 431–44 (2014).
van Laarhoven, A. et al. Low induction of proinflammatory cytokines parallels evolutionary success of modern strains within the Mycobacterium tuberculosis Beijing genotype. Infect. Immun. 81, 3750–3756 (2013).
Farhat, M. R. et al. Genomic analysis identifies targets of convergent positive selection in drug-resistant Mycobacterium tuberculosis. Nat. Genet. 45, 1183–1189 (2013).
Hazbon, M. H. et al. Convergent evolutionary analysis identifies significant mutations in drug resistance targets of Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 52, 3369–3376 (2008).
Knudsen, N. P. et al. Tuberculosis vaccine with high predicted population coverage and compatibility with modern diagnostics. Proc. Natl Acad. Sci. USA 111, 1096–1101 (2014).
Baldwin, S. L. et al. Intradermal immunization improves protective efficacy of a novel TB vaccine candidate. Vaccine 27, 3063–3071 (2009).
Baldwin, S. L. et al. Protection and long-lived immunity induced by the ID93/GLA-SE vaccine candidate against a clinical Mycobacterium tuberculosis isolate. Clin. Vaccine Immunol. 23, 137–147 (2015).
Bertholet, S. et al. A defined tuberculosis vaccine candidate boosts BCG and protects against multidrug-resistant Mycobacterium tuberculosis. Sci. Transl. Med. 2, 53ra74 (2010).
Bertholet, S. et al. Identification of human T cell antigens for the development of vaccines against Mycobacterium tuberculosis. J. Immunol. 181, 7948–7957 (2008).
Uplekar, S., Heym, B., Friocourt, V., Rougemont, J. & Cole, S. T. Comparative genomics of Esx genes from clinical isolates of Mycobacterium tuberculosis provides evidence for gene conversion and epitope variation. Infect. Immun. 79, 4042–4049 (2011).
Lewinsohn, D. M. et al. Human Mycobacterium tuberculosis CD8 T cell antigens/epitopes identified by a proteomic peptide library. PLoS One 8, e67016 (2013).
Kim, Y. et al. Immune epitope database analysis resource. Nucleic Acids Res. 40, W525–W530 (2012).
Mortimer, T. D., Weber, A. M. & Pepperell, C. S. Evolutionary thrift: Mycobacteria repurpose plasmid diversity during adaptation of type VII secretion systems. Genome Biol. Evol. 9, 398–413 (2017).
Gey Van Pittius, N. C. et al. The ESAT-6 gene cluster of Mycobacterium tuberculosis and other high G + C Gram-positive bacteria. Genome Biol. 2, RESEARCH0044 (2001).
Abdallah, A. M. et al. PPE and PE_PGRS proteins of Mycobacterium marinum are transported via the type VII secretion system ESX-5. Mol. Microbiol. 73, 329–340 (2009).
Fishbein, S., van Wyk, N., Warren, R. M. & Sampson, S. L. Phylogeny to function: PE/PPE protein evolution and impact on Mycobacterium tuberculosis pathogenicity. Mol. Microbiol. 96, 901–916 (2015).
Brennan, M. J. The enigmatic PE/PPE multigene family of Mycobacteria and tuberculosis vaccination. Infect. Immun. 85, e00969-16 (2017).
Groschel, M. I., Sayes, F., Simeone, R., Majlessi, L. & Brosch, R. ESX secretion systems: mycobacterial evolution to counter host immunity. Nat. Rev. Microbiol. 14, 677–691 (2016).
Shah, S., Cannon, J. R., Fenselau, C. & Briken, V. A duplicated ESAT-6 region of ESX-5 is involved in protein export and virulence of Mycobacteria. Infect. Immun. 83, 4349–4361 (2015).
Kumar, A., Chandolia, A., Chaudhry, U., Brahmachari, V. & Bose, M. Comparison of mammalian cell entry operons of Mycobacteria: in silico analysis and expression profiling. FEMS Immunol. Med. Microbiol. 43, 185–195 (2005).
Bukka, A., Price, C. T., Kernodle, D. S. & Graham, J. E. Mycobacterium tuberculosis RNA expression patterns in sputum bacteria indicate secreted Esx factors contributing to growth are highly expressed in active disease. Front. Microbiol. 2, 266 (2012).
Coppola, M. et al. New genome-wide algorithm identifies novel in-vivo expressed mycobacterium tuberculosis antigens inducing human T-cell responses with classical and unconventional cytokine profiles. Sci. Rep. 6, 37793 (2016).
Abdallah, A. M. et al. Mycobacterial secretion systems ESX-1 and ESX-5 play distinct roles in host cell death and inflammasome activation. J. Immunol. 187, 4744–4753 (2011).
Ford, C. B. et al. Mycobacterium tuberculosis mutation rate estimates from different lineages predict substantial differences in the emergence of drug-resistant tuberculosis. Nat. Genet. 45, 784–790 (2013).
Fox, G. J., Barry, S. E., Britton, W. J. & Marks, G. B. Contact investigation for tuberculosis: a systematic review and meta-analysis. Eur. Respir. J. 41, 140–156 (2013).
Lonnroth, K. et al. Systematic screening for active tuberculosis: rationale, definitions and key considerations. Int. J. Tuberc. Lung Dis. 17, 289–298 (2013).
Thai, P. V. K. et al. Bacterial risk factors for treatment failure and relapse among patients with isoniazid resistant tuberculosis. BMC Infect. Dis. 18, 112 (2018).
Caws, M. et al. The influence of host and bacterial genotype on the development of disseminated disease with Mycobacterium tuberculosis. PLoS Pathog. 4, e1000034 (2008).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–9 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Comas, I. et al. Out-of-Africa migration and Neolithic coexpansion of Mycobacterium tuberculosis with modern humans. Nat. Genet. 45, 1176–1182 (2013).
Pepperell, C. S. et al. The role of selection in shaping diversity of natural M. tuberculosis populations. PLoS Pathog. 9, e1003543 (2013).
Bradley, P. et al. Rapid antibiotic-resistance predictions from genome sequence data for Staphylococcus aureus and Mycobacterium tuberculosis. Nat. Commun. 6, 10063 (2015).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2 – approximately maximum-likelihood trees for large alignments. PLoS One 5, e9490 (2010).
Ashkenazy, H. et al. FastML: a web server for probabilistic reconstruction of ancestral sequences. Nucleic Acids Res. 40, W580–W584 (2012).
Bollback, J. P. SIMMAP: stochastic character mapping of discrete traits on phylogenies. BMC Bioinformatics 7, 88 (2006).
Revell, L. phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2012).
Vijay, S. V. et al. Influence of stress and antibiotic resistance on cell-length distribution in Mycobacterium tuberculosis clinical isolates. Front. Microbiol. 8, 1–12 (2017).
Rohde, K. H., Abramovitch, R. B. & Russell, D. G. Mycobacterium tuberculosis invasion of macrophages: linking bacterial gene expression to environmental cues. Cell Host Microbe 2, 352–364 (2007).
Carver, T. et al. BamView: visualizing and interpretation of next-generation sequencing read alignments. Brief Bioinform. 14, 203–212 (2013).
Jafri, M. et al. Germline mutations in the CDKN2B tumor suppressor gene predispose to renal cell carcinoma. Cancer Discov. 5, 723–729 (2015).
Usher, J. L. et al. Analysis of HGD gene mutations in patients with alkaptonuria from the United Kingdom: identification of novel mutations. JIMD Rep. 24, 3–11 (2015).
Jubb, H. C. et al. Mutations at protein–protein interfaces: Small changes over big surfaces have large impacts on human health. Prog. Biophys. Mol. Biol. 128, 3–13 (2017).
Kano, F. S. et al. The presence, persistence and functional properties of Plasmodium vivax Duffy binding protein II antibodies are influenced by HLA class II allelic variants. PLoS Negl. Trop. Dis. 10, e0005177 (2016).
Nemethova, M. et al. Twelve novel HGD gene variants identified in 99 alkaptonuria patients: focus on ‘black bone disease’ in Italy. Eur. J. Hum. Genet. 24, 66–72 (2016).
Silvino, A. C. et al. Variation in human cytochrome P-450 drug-metabolism genes: a gateway to the understanding of Plasmodium vivax relapses. PLoS One 11, e0160172 (2016).
White, R. R. et al. Ubiquitin-dependent modification of skeletal muscle by the parasitic nematode, Trichinella spiralis. PLoS Pathog. 12, e1005977 (2016).
Albanaz, A. T. S., Rodrigues, C. H. M., Pires, D. E. V. & Ascher, D. B. Combating mutations in genetic disease and drug resistance: understanding molecular mechanisms to guide drug design. Expert Opin. Drug Discov. 12, 553–563 (2017).
Casey, R. T. et al. SDHA related tumorigenesis: a new case series and literature review for variant interpretation and pathogenicity. Mol. Genet. Genomic Med. 5, 237–250 (2017).
Pandurangan, A. P., Ascher, D. B., Thomas, S. E. & Blundell, T. L. Genomes, structural biology and drug discovery: combating the impacts of mutations in genetic disease and antibiotic resistance. Biochem. Soc. Trans. 45, 303–311 (2017).
Soardi, F. C. et al. Familial STAG2 germline mutation defines a new human cohesinopathy. NPJ Genom. Med. https://doi.org/10.1038/s41525-017-0009-4 (2017).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Pires, D. E., Chen, J., Blundell, T. L. & Ascher, D. B. In silico functional dissection of saturation mutagenesis: interpreting the relationship between phenotypes and changes in protein stability, interactions and activity. Sci. Rep. 6, 19848 (2016).
Pandurangan, A. P., Ochoa-Montano, B., Ascher, D. B. & Blundell, T. L. SDM: a server for predicting effects of mutations on protein stability. Nucleic Acids Res. 45(W1), W229–W235 (2017).
Pires, D. E., Ascher, D. B. & Blundell, T. L. mCSM: predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics 30, 335–42 (2014).
Pires, D. E., Ascher, D. B. & Blundell, T. L. DUET: a server for predicting effects of mutations on protein stability using an integrated computational approach. Nucleic Acids Res. 42, W314–W319 (2014).
Mahmood, A. et al. Molecular characterization of secretory proteins Rv3619c and Rv3620c from Mycobacterium tuberculosis H37Rv. FEBS J. 278, 341–53 (2011).
Ascher, D. B. et al. Potent hepatitis C inhibitors bind directly to NS5A and reduce its affinity for RNA. Sci. Rep. 4, 4765 (2014).
Chan, L. J. et al. Conjugation of 10 kDa linear PEG onto trastuzumab Fab’ is sufficient to significantly enhance lymphatic exposure while preserving in vitro biological activity. Mol. Pharm. 13, 1229–1241 (2016).
Chan, L. J. et al. PEGylation does not significantly change the initial intravenous or subcutaneous pharmacokinetics or lymphatic exposure of trastuzumab in rats but increases plasma clearance after subcutaneous administration. Mol. Pharm. 12, 794–809 (2015).
Acknowledgements
We would like to thank the clinical staff who recruited patients into our study from the following District TB Units (DTUs) in HCMC, Vietnam: Districts 1, 4, 5, 6 and 8, Tan Binh, Binh Thanh and Phu Nhuan; and also our colleagues from Pham Ngoc Thach Hospital for Tuberculosis and Lung Disease, HCMC Vietnam. This work was supported by the National Health and Medical Research Council, Australia (project grant no. 1056689 to S.J.D., Fellowship no. 1061409 to K.E.H., Fellowship no. 1061435 to M.I., Fellowship no. 1072476 to D.B.A.), A*STAR Biomedical Research Council, Singapore (12/1/21/24/6689 to Y.Y.T.) and the Wellcome Trust UK (research training fellowship no. 081814/Z/06/Z to M.C.) and as part of their Major Overseas Program in Vietnam (089276/Z/09/Z to J.F. and 106680/B/14/Z to G.T.).
Author information
Authors and Affiliations
Contributions
S.J.D., K.E.H., M.C., M.I., Y.Y.T. and C.C.K. are the study principal investigators who conceived and obtained funding for the project. S.J.D. provided overall project coordination; M.I. organized and supervised the DNA sequencing and K.E.H. devised the overall analysis plan and wrote the first draft of the manuscript along with P.M. M.C. and S.J.D. established the TB cohort for this genetics study by working with P.V.K.T., D.T.M.H., N.N.L., N.H.L., N.T.Q.N., N.T.T.T., G.T. and J.F. to coordinate the collection of clinical samples and phenotypes. K.P. performed DNA quality checks and genome sequencing on all Vietnamese samples, while V.T.N.H. performed Sanger sequencing on selected samples. D.B.A. performed protein structure analyses, and H.T.H. and N.T.T.T. performed the macrophage growth and infection experiments of EsxW variants. K.E.H., P.M., M.I., D.J.E. and A.P.N. analyzed the data. All authors critically reviewed manuscript revisions and contributed intellectual input to the final submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Association between Mtb genetic distance and spatial proximity of patients.
For all pairs of Mtb isolates, the probability of both originating from patients living in the same district was calculated (y-axis) and stratified according to genetic distance between the Mtb genomes (expressed in number of SNPs, x-axis). P-values indicate result of testing for difference in proportions of isolate pairs originating from same district, for isolate pairs with SNP distance <10 vs >20, calculated separately for Beijing and non-Beijing isolates. (Note it was not possible to further stratify non-Beijing pairs into L1 pairs vs L4 pairs for statistical tests, due to low sample size.).
Supplementary Figure 2 Proportion of clusters not explained by transmission of antimicrobial resistance mutations.
Each cluster (defined at various thresholds of maximum patristic distance between tips) was checked to determine whether all members of the cluster shared any of the antimicrobial resistance mutations identified by Mykrobe Predictor. Clusters in which no known antimicrobial resistance mutation was conserved in all members of the cluster were considered not driven by antimicrobial resistance. For each lineage and at each maximum patristic distance threshold value for clustering, the proportion of clusters not explained by antimicrobial resistance are shown below. Colour legend for lineages is the same as throughout the paper: pink, lineage 1; dark blue, lineage 2.1; orange, lineage 2.2.1; light blue, lineage 2.2.2; red, lineage 4.
Supplementary Figure 3 Homoplasic mutations detected in the HCMC Mtb population and located in known drug resistance-associated genes.
Mutations arising on more than one branch in the HCMC Mtb tree in the following genes are included: embB (ethambutol resistance), gidB (streptomycin resistance), gyrA (fluoroquinolone resistance); inhA (isoniazid resistance), rpoB (rifampicin resistance).
Supplementary Figure 4 Homoplasic nsSNPs that are fixed within Beijing lineage and also arise independently in other branches in the HCMC phylogeny.
Tree shown is the ML phylogeny for 1,635 HCMC isolates with lineages highlighted, reproduced from Figure 1. Outer rings indicate the presence of derived alleles for the three nsSNPs that were found to be fixed within the Beijing lineage (2.2; including 2.2.1 and 2.2.2).
Supplementary Figure 5 Map of HCMC showing location of district TB units where patients were recruited.
Yellow regions show districts included in the study; red dots indicate location of the district TB units where participants were recruited; red star indicates location of Pham Ngoc Thach Hospital for Tuberculosis and Lung Disease. Blue regions indicate other urban HCMC districts, green are greater HCMC districts (not included in recruitment).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 3 and 4
Supplementary Table 1
Details of the M. tuberculosis isolates sequenced for this study
Supplementary Table 2
Details of the publicly available M. tuberculosis genomes included in this study
Supplementary Table 5
Repetitive regions of the M. tuberculosis genome that were excluded from SNP analysis
Rights and permissions
About this article
Cite this article
Holt, K.E., McAdam, P., Thai, P.V.K. et al. Frequent transmission of the Mycobacterium tuberculosis Beijing lineage and positive selection for the EsxW Beijing variant in Vietnam. Nat Genet 50, 849–856 (2018). https://doi.org/10.1038/s41588-018-0117-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0117-9
This article is cited by
-
Parallel signatures of Mycobacterium tuberculosis and human Y-chromosome phylogeography support the Two Layer model of East Asian population history
Communications Biology (2023)
-
Evolution and spread of a highly drug resistant strain of Mycobacterium tuberculosis in Papua New Guinea
BMC Infectious Diseases (2022)
-
Whole-genome single nucleotide variant phylogenetic analysis of Mycobacterium tuberculosis Lineage 1 in endemic regions of Asia and Africa
Scientific Reports (2022)
-
A genome epidemiological study of mycobacterium tuberculosis in subpopulations with high and low incidence rate in Guangxi, South China
BMC Infectious Diseases (2021)
-
Assessment of closely related Mycobacterium tuberculosis variants with different transmission success and in vitro infection dynamics
Scientific Reports (2021)