Abstract
The rich genetic diversity in Oryza sativa and Oryza rufipogon serves as the main sources in rice breeding. Large-scale resequencing has been undertaken to discover allelic variants in rice, but much of the information for genetic variation is often lost by direct mapping of short sequence reads onto the O. sativa japonica Nipponbare reference genome. Here we constructed a pan-genome dataset of the O. sativa–O. rufipogon species complex through deep sequencing and de novo assembly of 66 divergent accessions. Intergenomic comparisons identified 23 million sequence variants in the rice genome. This catalog of sequence variations includes many known quantitative trait nucleotides and will be helpful in pinpointing new causal variants that underlie complex traits. In particular, we systemically investigated the whole set of coding genes using this pan-genome data, which revealed extensive presence and absence of variation among rice accessions. This pan-genome resource will further promote evolutionary and functional studies in rice.
Similar content being viewed by others
Introduction
Rice is one of the world’s major crops and the primary source of carbohydrate intake. Cultivated rice (O. sativa) and its closest wild relative O. rufipogon have a broad geographical distribution with adaptations to many kinds of ecological and agronomic conditions1. The rich genetic diversity in rice has played important roles in both domestication and modern breeding, and it will be a crucial resource to respond to the growth in food demand and the future genetic improvement associated with the rapid climate changes globally.
With the application of high-throughput sequencing technologies, diverse rice accessions have been resequenced and phenotyped during recent years, with the aim of exploring genomic diversity to look for the gene loci under human selection and to uncover the molecular basis of many agronomic traits2,3,4,5,6,7,8,9. However, in these resequencing efforts, characterizations of the genetic variants all rely on high levels of sequence similarity to map the short reads (typically, ~100 bp) onto the rice reference genome10, which means that the information from highly polymorphic regions would often be inevitably lost. Moreover, previous studies have found that there are functionally important genes that are absent in the reference Nipponbare genome but present in other rice varieties11,12,13, indicating that one or a few rice genomes cannot include all of the important genomic content. Hence, to comprehensively capture the genomic diversity in rice, it is necessary to de novo construct the complete genomic sequences for dozens of diverse accessions14,15,16,17,18. Particularly, the genomic sequences of three divergent rice varieties were de novo assembled, and many of the genome-specific loci that were absent from the reference genome were identified, illustrating the utility of de novo assemblies for biological discovery in rice19.
Previously, we collected ~1,500 diverse accessions of O. sativa and O. rufipogon and generated a genome-variation map to reveal the molecular evolutionary history in rice6. From the large collection, a total of 66 accessions were selected and used for deep sequencing and whole-genome de novo assembly, independently of the Nipponbare reference. Comparative analyses and genome annotations of the assemblies enabled the identification of diverse alleles and the functional consequences of various polymorphisms at a fine-scale level. The pan-genome data provided not only the whole set of genes that was shared among rice but also new insights into intra- and inter-species differentiation. The establishment of a rice pan-genome will be helpful in utilizing the various alleles within the gene pools for genetic studies and breeding.
Results
Genome assemblies of 66 rice accessions
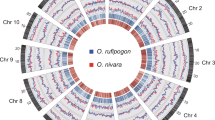
According to the phylogenetic tree of 1,529 rice accessions6, we selected 57 divergent accessions in the O. sativa–O. rufipogon species complex (Supplementary Figs. 1 and 2) for the rice pan-genome study. Moreover, nine widely used modern cultivars (for example, Koshihikari in Japan, Basmati in India, Kongyu-131 in northeast China and Guangluai-4 in southern China) were also included in this collection. The samples that we selected in this study included 22 accessions of O. sativa temperate japonica, 5 of O. sativa tropical japonica, 1 of O. sativa aromatic, 19 of O. sativa indica, 6 of O. sativa aus and 13 of O. rufipogon, all of which represented all of the major genetically distinct clusters in O. sativa and O. rufipogon. The genomic DNA of each rice accession was sequenced with an average of 115-fold depth using Illumina technology, generating a total of 3.1 Tb of raw sequence reads. The 66 rice genomes were all de novo assembled, resulting in final assemblies with contig N50 sizes (where N50 size refers to the size of the contig that, along with the larger contigs in the assembly, contains half of the rice genome sequence) that ranged from 21 to 75 kb in different accessions (Supplementary Fig. 3 and Supplementary Data 1), which was much higher than the average size of rice genes (~2.9 kb)10. The draft sequence of one rice accession, Guangluai-4 (GLA4), was validated using 22 Mb of high-quality BAC-based sequences as a gold standard, and the comparisons showed that there were very few assembly errors (Supplementary Fig. 4 and Supplementary Data 2). We also sequenced and assembled the Nipponbare genome using the same method. The genome assembly of Nipponbare was compared with the reference sequence for quality control, and the sequence identity between them was >99.96%, with error rates in genes and intergenic regions of 0.0218% and 0.0352%, respectively. We also plotted the error rates across the 12 rice chromosomes (Supplementary Fig. 5) and found that a small fraction (~3%) of genomic regions were enriched with errors (Supplementary Fig. 6). Using the Nipponbare reference genome as a standard, our Nipponbare assembly showed a genome coverage of 84.86%, with many gaps in repetitive regions. Moreover, the collections of full-length cDNAs for two rice accessions, W1943 and GLA4, were mapped onto the corresponding genome assemblies20,21, where 96.77% and 90.25% of the cDNAs had nearly perfect matches with the assemblies (with identity of >98%), respectively. The results from BACs and full-length cDNAs indicated that the de novo assemblies had both high accuracy and high genome coverage.
The 66 genome assemblies were anchored onto the Nipponbare reference genome to discover detailed sequence variations. We identified a total of 16,563,789 SNPs, 5,549,290 small insertions and deletions (indels) of ≤20 bp and 933,489 structural variants (SVs; which refer to large indels that range from 20 bp to 12 kb in this work). On the basis of the variants, the sequence diversity (π) of the O. sativa–O. rufipogon complex was calculated to be 0.018. Among the variants, 3.2% of SNPs, 2.5% of small indels and 2.0% of SVs were located in the coding regions of 27,655, 22,755 and 7,679 well-annotated genes, respectively (Supplementary Fig. 7). We investigated the allelic distributions for the ~23 million variants and found that most of the naturally occurring variations, including those with large effects on gene coding, were present in only one or a few accessions (Fig. 1a), similar to the characteristics of the populations among the Arabidopsis thaliana ecotypes22 and maize landraces23. In addition, we aligned the raw reads of each accession against the assembly of the same accession, and the sites with low read depth or abnormal distribution were masked. We found that, for each accession, ~2.1% of sequence variants were from the low-quality sites (Supplementary Fig. 8), implying that the variation calling was reliable in general. The ‘low-quality’ variants mostly resulted from assembly errors from multiple reads with ‘heterozygous genotypes’, especially in the simple-sequence repeat regions (see the examples in Supplementary Fig. 4d–f). We also identified a putative ‘identical-by-descent’ region on chromosome 1 (the interval of 0.0 to 5.0 Mb) between Tohoku IL9 and Daohuaxiang-2, on the basis of which the error rate of variant calling was estimated to be ~0.8%.
a, Allele frequency spectra for the indicated kinds of genetic variants. For each variant, we identified the minor allele across the 66 accessions and calculated the frequency of this allele. b, Neighbor-joining tree of the 66 rice accessions using whole-genome data. The accessions within different groups are indicated by different colors.
We used the genomic data to assess whether the 66 diverse accessions had a wide diversity. Previously, we resequenced a total of 1,529 accessions of O. sativa and O. rufipogon6. Among the common SNPs (minor allele frequency > 0.01) identified in the large population, 89.2% (1,405,349 of 1,575,718) were detected in the 66 genome assemblies as well, suggesting that the core collection captured a large proportion of common genetic variation in the O. sativa–O. rufipogon complex.
Domestication and introgression
We used the whole-genome variants to construct a phylogenetic tree for the 66 genomes (Fig. 1b), the pattern of which was generally consistent with that of 1,529 rice accessions6. Using the pan-genome-based variants, we performed a global analysis for the domestication selection scan (Supplementary Fig. 9). As expected, the results for the major domestication sweeps were almost the same as previous results from low-coverage resequencing of 1,529 accessions. There were six domestication sweeps identified using these pan-genome data that were missed in the previous results (Supplementary Data 3). We investigated estimates of sequence diversity (π) using the resequencing data and the pan-genome data. For five of the new domestication sweeps, the estimates of π in O. rufipgon using the pan-genome data were much higher than those using the resequencing data, as many more variants in O. rufipogon accessions were able to be discovered by the pan-genome approach. However, it was difficult for us to finely evaluate and conclude the effects of the pan-genome data in estimating genetic diversity because the two datasets had dramatically different population sizes (n = 60 and n = 1,529).
Beside indica and temperate japonica, there are three other groups in Asian cultivated rice—aus, aromatic and tropical japonica. Analysis at seven gene loci associated with rice domestication showed that aus accessions were not always included within the cultivated rice clade, for example, in analysis of aus rice at Bh4 (Os04g0460000)24 and An1 (Os04g0350700)25 (Supplementary Fig. 10). The results suggest that the aus group is under incomplete domestication selections, with some alleles associated with domestication not included in the genomes of aus rice.
Moreover, we found that there were potential clues for introgressions from indica into tropical japonica, both of which were cultivated in the same regions of tropical Asia. We identified 807,139 SNP sites with highly differentiated alleles between indica and temperate japonica and looked up their allelic information in each accession of tropical japonica (Supplementary Fig. 11). An average of ~16.0% of the whole rice genome in tropical japonica had evidence of introgression from indica. In particular, we identified nine loci with a clear introgression pattern; these included the thermotolerance allele of OsTT1 (Os03g0387100) and the large-grain allele of OsSPL13 (Os07g0505200), which have been reported to be introgressed from indica to tropical japonica26,27. The introgression probably contributes to the genetic composition of tropical japonica.
Identification of functionally diverse alleles
By using the genome assemblies, the fine-scale distribution of quantitative trait nucleotides (QTNs) underlying various agronomic traits could be explored and the demographic origins of these functionally important alleles could be traced. To demonstrate the evolutionary route, we chose five important quantitative trait loci (QTLs)—Hd3a (Os06g0157700), COLD1 (Os04g0600800), GW6a (Os06g0650300), TAC1 (Os09g0529300) and Sd1 (Os01g0883800), which are involved in flowering time, cold tolerance, grain weight, tiller angle and plant height, respectively (Fig. 2). All five genes have well-characterized causative variants28,29,30,31,32. For Hd3a, COLD1 and GW6a, the variation at the QTNs could be observed in the gene pools of wild rice, O. rufipogon, and the differentiated distribution within cultivated rice is probably due to founder effects. For TAC1, all O. rufipogon accessions in this collection contained the wild-type allele, whereas all accessions of japonica subspecies (including temperate japonica, tropical japonica and aromatic) had the mutated allele for a narrower tiller angle that enables more efficient plant architecture31, suggesting that the mutation may have been selected during japonica domestication. A similar situation was observed for sd1, the well-known Green Revolution target for modern breeding, in which a mutated semidwarf-1 allele (a 384-bp indel) was present mainly in indica cultivars. Moreover, the mutated allele of sd1 with two missense SNPs, which controls culm length, was found to be present in all accessions of japonica subspecies and may have been selected during japonica domestication as well32. As a rough estimation of the scenario, the variants that segregated in only temperate japonica or in only indica accounted for 5.8% and 11.0% of total variants, respectively.
We focused on the variants in gene-coding regions and attempted to predict their putative effects on protein coding. According to coding variants in this pan-genome dataset, each gene contained, on average, ten missense SNP sites and six polymorphic sites of relatively large effect (for example, see Supplementary Fig. 12), which often created multiple gene alleles. For example, we observed three missense SNP sites, one intron 1–exon 1 junction SNP site and one indel site in waxy (Os06g0133000; a major gene underlying grain quality)33 from seven representative haplotypes (Fig. 3a,b). As compared to the wild-type allele in most indica and aus cultivars, the T allele at the intron 1–exon 1 junction site led to lower amylase content (from 24.7% for the G allele to 14.6% for the T allele)5 by reducing the expression level of the waxy gene33, whereas the 23-bp duplication at the second exon was a frameshift mutation that resulted in no accumulation of amylase (that is, the phenotype of sticky rice for accessions GP551 and HP263)3. In Hd1 (Os06g0275000; a major gene underlying flowering time)34, there were a total of 22 SNP sites that resulted in missense mutations, 2 SNP sites that resulted in the formation of stop codons and 7 indel sites for seven representative haplotypes, where the 2-bp indel in the first exon (in Kasalath) and the 2-bp indel in the second exon (in HP263) resulted in lack of the CCT (CONSTANS, CO-like and TOC1) domain, which would cause a defect in the protein function of Hd1 (Fig. 3c,d)35. For a global picture of the potentially functional alleles, we further analyzed the variants in coding genes from 38 gene families in the rice genome (Supplementary Data 4). As expected, the gene families controlling basic biological processes (for example, the amino acid transporter family and peroxidase family) contained much fewer missense variants than those for plant immunology.
a, Neighbor-joining tree of 66 accessions using genetic variants in waxy. Seven diverse accessions representing different haplotypes were selected from the tree and are color-coded according to rice group. b, Allelic information of sequence variants in waxy for the seven accessions. c, Neighbor-joining tree using genetic variants in Hd1. d, Allelic information of sequence variants in Hd1 for the seven accessions selected in c.
Presence–absence variation of coding genes
Presence–absence varitaion (PAV) of genes, referring to the presence or absence of gene variability in diverse rice accessions, is one of the genetic factors underlying agronomic traits, and here the whole-genome de novo assemblies provided the opportunity to discover genes that are absent in the Nipponbare reference genome sequence and to explore the PAV information of all coding genes among the rice accessions. We performed genome annotations for all 67 assemblies (including that for Nipponbare). With the exclusion of repetitive sequences, we predicted all the non-transposable-element (non-TE) protein-coding genes for each genome. There were a total of 10,872 genes in the 67 rice accessions that were at least partially absent in the Nipponbare reference. These ‘newly identified’ genes were mostly due to large indels among accessions (for example, a large insertion relative to the Nipponbare variety; see Fig. 4a,b). A small fraction of the newly identified genes, however, should be located within the physical gaps of the Nipponbare reference genome sequence, because ~9.5% of the newly identified genes could be found in our Nipponbare assembly from whole-genome Illumina sequencing. To investigate whether the newly identified genes were expressed, we collected four tissues (young seedling, root, leaf and panicle) in two accessions (GLA4 and W1943) for RNA sequencing. We found transcripts for approximately 57.1% and 60.6% of the newly identified genes in GLA4 and W1943, respectively, although the expression levels of the newly identified genes (as measured by reads per kilobase per million reads (RPKM) value) were generally lower than those for the genes annotated in the Nipponbare reference (Fig. 4c and Supplementary Fig. 13). Moreover, previous studies had identified several genes that had not been observed in the Nipponbare reference, including Sub1A, SNORKEL1 and SNORKEL2, which control submergence tolerance11,12, and Pstol, which controls phosphorus-deficiency tolerance13. Sequence searching showed that all of these reported genes were among the newly identified genes found in the pan-genome (Fig. 5a). Taken together, these pieces of evidence suggest that at least some of the newly identified genes are functionally important.
a, Detection of an expressed gene on chromosome 6 of the W1943 genome assembly in a 3.7-kb insertion. The black lines and green boxes indicate genome seqeunces and gene-coding regions, respectively. Domain information from InterPro scans is indicated. b, Detection of an expressed gene on chromosome 11 of the W1943 assembly in an insertion of 3.6 kb. c, Comparison of the expression levels of genes annotated in the Nipponbare reference and the newly identified genes in the W1943 genome in four tissues of W1943.
a, PAV of six functionally characterized genes in the 67 genomes. The accessions within different groups are color-coded as in Fig. 1b. The absence of a gene in the genome is indicated by a blank box. b, A 67 × 67 matrix comparing the coding genes of the accessions by pairs. For each rice accession, we searched for the genes it shared with each of the 67 accessions. The color index corresponds to the number of shared coding genes. c, Presence and absence information of 42,580 genes in the 67 rice accessions. The order of the 67 accessions (from “GLA4” on the left to “W3095-2” on the right) is the same as in a. The core genome set and the dispensable genome set refer to coding genes present in ≥90% of rice accessions and genes present in <90% of accessions, respectively. Presence is color-coded as in a, and the absence of a gene is indicated by white.
We observed that, even for the genes annotated in the Nipponbare reference, there were extensive PAVs among diverse rice accessions, for example, Ghd7 (Os07g0261200; which controls flowering time) and OsFBX310 (Os09g0292900; which controls hull color) (Fig. 5a)36,37. Hence, to obtain a clear picture of PAV in rice, we compiled a list of all of the coding genes in the 67 genomes together and excluded any redundancies (see the RicePanGenome database). There were a total of 42,580 non-TE genes annotated in at least one of the 67 rice accessions. We further tried to estimate the total gene number of the rice species using the approach from the study of the maize pan-genome and pan-transcriptome38. Stepwise addition of rice accessions from n = 2 to n = 67 showed that the number of coding genes (42,580) at n = 67 was close to a plateau (Supplementary Fig. 14). Further sampling of more diverse rice accessions will likely result in limited gene discoveries for the dispensable genome set. We searched the orthologs of the gene set against each of the 67 rice genomes (see the number of shared genes between two accessions in Fig. 5b) and generated a list of one-to-one correspondences and their presence-or-absence information in different accessions (Fig. 5c). According to PAV of the genes, there were 26,372 and 16,208 genes present in ≥60 rice accessions (90% of the collection) and present in <60 accessions, respectively, and these were defined as the core genome set and the dispensable genome set of coding genes in rice. Among the dispensable genome set, there were 285 group-specific genes (Supplementary Fig. 15), whereas most of the genes were present in only a few accessions. We screened InterPro domains (from a database of protein families, domains and functional sites) for coding genes in the core genome set and those in the dispensable genome set and compared the functional classifications of the coding genes from the two sets (Supplementary Fig. 16). As expected, the genes of the dispensable genome set were enriched for abiotic and biotic response genes, especially for NBS-LRR (nucleotide-binding site–leucine-rich repeat) and NB-ARC (nucleotide-binding adaptor shared by APAF-1, R proteins and CED-4) genes, which control disease resistance in rice. Furthermore, in the core genome set, ~77.6% of the coding genes contained InterPro domains, a much higher proportion than that in the dispensable genome set (~35.8%), implying that a portion of the PAV genes in the dispensable genome set may be just artifacts or pseudogenes.
Discussion
We have generated a pan-genome dataset for the O. sativa–O. rufipogon species complex, a resource for in-depth functional genomics studies and molecular breeding that should be useful in future. Using the pan-genome dataset, genome-wide comparisons of the assemblies enabled the characterization of numerous complex variants, including many large-effect coding variants and many coding genes that were absent in the rice reference genome sequence, which should be helpful in pinpointing the causal variation in QTL cloning and in genome-wide association studies (GWAS)5,7,9,27.
In rice, hundreds of genes have been functionally investigated through mutagenesis-based approaches or through transgenic methods (for example, overexpression or RNAi), some of which were later found to be the causative genes underlying complex traits using QTL cloning—for example, the cases for LAX1 (Os01g0831000) and NAL1 (Os04g0615200)39,40. Hence, integration of the information from studies of gene function and the natural variation in the genome assemblies could provide a complementary approach to forward genetic studies. Among the functionally characterized genes in the rice genome (according to the information in the RiceData database), a total of 867 genes were found to contain important coding variation in at least one rice accession. For instance, the sd-g (Os05g0407500) gene was cloned from a semidwarf mutant that was insensitive to gibberellin41, and a total of five frameshift indels located within the coding region of this gene were detected in four rice accessions, which probably result in plant height variation (Supplementary Fig. 12).
In particular, our study demonstrated that most of the naturally occurring variants in rice are of low frequency (Fig. 1a). A small fraction of these low-frequency alleles disrupt gene coding and might have important biological functions underlying the variation of complex traits. However, in conventional GWAS, it is very difficult to identify associations from rare alleles by statistical methods unless extremely large sample sizes are used42,43. In human genetic studies, the existence of numerous rare variants with large effects is regarded as one of the major causes of the 'missing heritability' problem44 (for example, in human adult height45). Functional genomics methodologies, such as genome editing technology, coupled with in-depth annotations for the genetic variants could be used to verify the functional effects of these rare alleles in rice.
To date, few studies using multiple collaborative populations in rice for joint analyses have been reported. Considering that there are only weak reproductive isolations within the O. sativa–O. rufipogon species complex, the divergent accessions in this rice pan-genome can be crossed with a couple of common reference parents in each group (for example, Nipponbare in the temperate japonica group, GLA4 in the indica group and Kasalath in the aus group) to generate backcross inbred line (BIL) populations, similar to the designs in maize46 and in Arabidopsis47,48. Such a panel with multiple BIL populations collectively will be useful for both breeding and mapping of complex traits. In particular, it is not possible to perform GWAS in combined populations of cultivated rice and wild rice species owing to large genomic and phenotypic divergence. When numerous ‘novel’ alleles from diverse genetic backgrounds that underlie specific agronomic traits are introduced into common reference parents through the BIL approach, large-scale genetic mapping will become feasible and the rich gene resources can be used efficiently. To further improve the assembly quality of the pan-genome data and to compensate for the limitation of assembly from short reads, we will utilize new sequencing technology to build pseudomolecules, especially for the common reference parents in each rice group. These genome assemblies, coupled with genetic populations and transcriptome and epigenomics data generated in future work, will facilitate the mining of natural variation for genetic studies and breeding.
Methods
Sampling and sequencing
The initial set of 1,529 accessions was selected from a collection of ~50,000 rice accessions that are preserved at the China National Rice Research Institute in China and the National Institute of Genetics in Japan6. From database records of the phenotypic variation and geographic origins of the germplasm, we generated a data matrix and conducted a cluster analysis. On the basis of the resulting tree, we sampled 1,083 O. sativa accessions and 446 O. rufipogon accessions to represent the entire range of phenotypic diversity and geographic distribution and sequenced them with twofold genome coverage. Using the whole-genome resequencing data, we constructed neighbor-joining trees for O. sativa and O. rufipogon. According to the two phylogenetic trees, several divergent accessions were selected for each clade in the trees. A total of 57 representative accessions were selected in the O. sativa–O. rufipogon species complex. Moreover, nine widely used modern cultivars in China, Japan and India were also included in the representative collection. Genomic DNA from the resulting 66 accessions was prepared from the fresh leaf tissue of a single plant of each accession using the DNeasy Plant Mini Kit (Qiagen). A sequencing library was constructed with an insert size of ~400 bp or ~700 bp on an Illumina HiSeq 2500 system using the manufacturer’s protocol, and an amplification-free method of library preparation49 was used to reduce the incidence of duplicate sequences, thus facilitating genome assembly. This study generated a total of 3.1 Tb of raw data of 100-bp and 150-bp paired-end reads, with an average of 115-fold coverage for each accession. For quality control, we also sequenced the Nipponbare genome using the same method with 91× coverage.
Whole-genome de novo assembly and validation
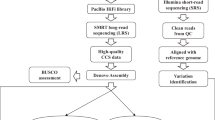
The 66 rice genomes were de novo assembled by using a pipeline that combined both the SOAPdenovo2 package (version 2.23)50 and the Fermi package (version 1.1)51. Briefly, raw reads were assembled in parallel into contigs by Fermi (run-fermi.pl -Pe) and SOAPdenovo2, and the software GapCloser (version 1.12-r6) was used to fill gaps in the draft assembly results from SOAPdenovo2. All of the contigs derived from Fermi and SOAPdenovo2 were merged to form draft contigs using a C program REPLACE. The N50 length of the final genome assemblies was evaluated, and all small contigs of <200 bp were excluded. To check the quality of the assembly, the genome assemblies of Nipponbare and GLA4 were aligned against the Nipponbare reference and GLA4 BAC-based sequences (with a total size of ~22 Mb) using the software MUMmer52 (with the parameters ‘show-coords -rcl; delta-filter -q; show-coords -rcl’) and ClustalW53 (with default parameters). The number of errors per base was estimated according to the sequence variants between them. On rice chromosome 4, a total of 273 BACs of indica GLA4 were sequenced and assembled using the Sanger-based method (Supplementary Fig. 4a). The sequences of 273 BACs were merged into 87 contigs (with an average size of ~250 kb). We compared the 87 BAC-based contigs with the GLA4 assemblies in this pan-genome, and there were a total of 4,353 substitutions, 4,283 small indels and 40 relatively large-scale variants. Among the 22-Mb BAC-based regions, the GLA4 assemblies in the pan-genome contained a total of 979 gaps. Detailed statistics for each of the 87 BAC-based contigs are provided in Supplementary Data 2. All of the full-length cDNA sequences of accession GLA4 (n = 10,082) and accession W1943 (n = 2,045) were aligned against the genome assemblies using the software BLASTN54 with the parameters ‘-e 1e-10’ and ‘-F F’ and a sequence identity of >98%.
Identification of genomic variation
The contig sequences of the whole-genome assemblies were anchored to the rice reference genome sequence (IRGSP build 4 version) using the software package MUMmer. According to the results from MUMmer, one-to-one alignment blocks (that is, each contig of the genome assemblies and its corresponding local sequence in the Nipponbare reference) were generated, and sequence variants were further called using the diffseq program in the EMBOSS package55 (version 4.0) with the parameter ‘-wordsize 10’. SVs of large size were called based on the alignment results from MUMmer. At the site of each sequence variant, the genotypic information (that is, the reference allele or the alternative allele) for all of the 66 rice accessions was called according to the results of the one-to-one alignments. The potential effects of the variants were predicted based on GFF files from RAP-DB (release 2). In addition to variants in well-annotated genes, a total of 1,171,090 variants were found to be located in the coding regions of 21,644 predicted genes (without cDNA or EST support in RAP-DB). The software programs ClustalW and BLASTN were used for detailed haplotype analyses for the well-characterized genes in rice. Moreover, we aligned the raw reads of each accession against the genome assembly of the same accession using the software Bowtie2 (version 2.2.6) and default parameters to generate BAM files. With the sorted BAM result for each genome, pileup results were generated using the SAMtools package (version 0.1.19). According to these results, we identified ‘low-quality’ SNPs and small indels (1–3 bp) using the parameter ‘varFilter -D200’ and a mapping quality of ≥30.
Evolutionary analysis
Simple matching coefficients were calculated from whole-genome SNPs or the SNPs at the local regions (including Bh4, PROG1, An1 and An2) of the 66 rice accessions. The 66 × 66 matrix of simple matching coefficients was used to construct phylogenetic trees through the ‘neighbor’ software in PHYLIP56, and the package MEGA5 was used to display the phylogenetic trees. In the analysis of introgression events, we first identified SNP sites with highly differentiated alleles between indica and temperate japonica, requiring that the SNP site have an allele frequency of >0.95 in indica and an allele frequency of <0.05 in temperate japonica. At these SNP sites, the allele information (indica-specific type or temperate-japonica-specific type) in each accession of tropical japonica was called across the rice genome. For each tropical japonica accession, the sizes of the introgression segments in its genome were determined to estimate the proportion of the potential introgression events in tropical japonica. Information for functionally characterized genes in the rice genome was based on the database in the China Rice Data Center, with all redundancies removed.
Presence–absence variation analysis of coding genes
For each genome assembly, the package RepeatMasker (version 4.0.6; with parameter ‘-species rice -nolow’) was used first to annotate and mask the repetitive sequences, including simple-sequence repeats and kinds of TEs. Sequences from microbial genomes (including those from pathogen infection of rice plants), which had no homologs with the rice genome, were masked as well. The software FGeneSH (Softberry)57 was used for gene structure prediction in the 67 rice genomes (Nipponbare included) with the parameters trained on monocotyledons. The predicted genes were searched against the annotated coding genes of the Nipponbare reference (RAP-DB on IRGSP-1.0 and RGAP 7) using BLASTN (with the parameters '-e 1e-10 –F F'). Genes that showed no hits with the Nipponbare reference genes or only partial sequence matches (coverage <50%) were regarded to be ‘newly identified genes’ that were absent in the Nipponbare reference.
Four tissues (young seedling, root, leaf and panicle) were collected from O. rufipogon W1943 and O. sativa indica GLA4 plants to perform RNA sequencing (RNA-seq) experiments. Paired-end cDNA libraries were constructed by using the RNA-seq Library Preparation Kit (Gnomegen, cat no. K02421T-L). cDNA fragments of ~300 bp in size were excised, followed by enrichment using PCR amplification for ~15 cycles. The resulting paired-end cDNA libraries were sequenced using the Illumina HiSeq 2500 system to generate 100-bp paired-end reads (29.1 Gb and 35.1 Gb of raw data for W1943 and GLA4, respectively). RNA-seq reads were aligned against the annotated genes in Nipponbare and the newly identified genes using the software SMALT (version 0.5.7) with the parameters ‘map -i 700 -j 50 -m 30’. The numbers of uniquely mapped reads (mapping score ≥50) were converted to quantify the transcript levels of genes from the two gene sets using RPKM values58.
To explore the PAV information of all coding genes among the rice accessions, we integrated the DNA sequences of all non-TE genes annotated in at least one of the 67 rice accessions. We searched the sequence of each gene against those of the annotated genes in other genome assemblies through BLASTN and generated a list of one-to-one correspondences. For the whole set of coding genes in the pan-genome data, the protein sequences were searched for protein domain information and protein function classification using the software InterProScan59 (version 5.7-48.0) with the parameters ‘-f TSV -iprlookup -goterms’.
URLs
SOAPdenovo2, https://sourceforge.net/projects/soapdenovo2/files/SOAPdenovo2/; GapCloser, http://soap.genomics.org.cn/about.html; Fermi package, https://github.com/lh3/fermi; REPLACE, ftp://ftp.sanger.ac.uk/pub/users/zn1/merge/replace/; MUMmer, http://mummer.sourceforge.net/; ClustalW, http://www.clustal.org/; IRGSP build 4, http://rapdb.dna.affrc.go.jp/download/build4.html; EMBOSS, http://emboss.sourceforge.net; Bowtie2, http://bowtie-bio.sourceforge.net/bowtie2/index.shtml; SAMtools, http://samtools.sourceforge.net/; PHYLIP, http://evolution.genetics.washington.edu/phylip.html; MEGA5, http://www.megasoftware.net/index.php; China Rice Data Center, http://www.ricedata.cn/gene/; RepeatMasker, http://www.repeatmasker.org/; RAP-DB on IRGSP4, http://rapdb.dna.affrc.go.jp/download/build4.html; RAP-DB on IRGSP-1.0, http://rapdb.dna.affrc.go.jp/download/irgsp1.html; RGAP 7, http://rice.plantbiology.msu.edu/pub/data/Eukaryotic_Projects/o_sativa/annotation_dbs/pseudomolecules/version_7.0/; SMALT, http://www.sanger.ac.uk/science/tools/smalt-0; InterProScan, http://www.ebi.ac.uk/interpro/download.html.
Life Sciences Reporting Summary
Further information on experimental design is available in the Life Sciences Reporting Summary.
Data availability
The DNA sequencing data are deposited in the European Nucleotide Archive under accession numbers PRJEB19404. The 67 genome assemblies, the BLAST searches and related information are available at the RicePanGenome database (http://www.ncgr.ac.cn/RicePanGenome).
Change history
01 June 2018
When published, this article did not initially appear open access. This error has been corrected, and the open access status of the paper is noted in all versions of the paper.
References
Khush, G. S. Origin, dispersal, cultivation and variation of rice. Plant Mol. Biol. 35, 25–34 (1997).
Huang, X. et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 42, 961–967 (2010).
Huang, X. et al. Genome-wide association study of flowering time and grain-yield traits in a worldwide collection of rice germplasm. Nat. Genet. 44, 32–39 (2011).
Xu, X. et al. Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat. Biotechnol. 30, 105–111 (2011).
Huang, X. et al. Genomic analysis of hybrid rice varieties reveals numerous superior alleles that contribute to heterosis. Nat. Commun. 6, 6258 (2015).
Huang, X. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501 (2012).
Chen, W. et al. Genome-wide association analyses provide genetic and biochemical insights into natural variation in rice metabolism. Nat. Genet. 46, 714–721 (2014).
3,000 rice genomes project. The 3,000 rice genomes project. Gigascience 3, 7 (2014).
Yano, K. et al. Genome-wide association study using whole-genome sequencing rapidly identifies new genes influencing agronomic traits in rice. Nat. Genet. 48, 927–934 (2016).
International Rice Genome Sequencing Project. The map-based sequence of the rice genome. Nature 436, 793–800 (2005).
Xu, K. et al. Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice. Nature 442, 705–708 (2006).
Hattori, Y. et al. The ethylene response factors SNORKEL1 and SNORKEL2 allow rice to adapt to deep water. Nature 460, 1026–1030 (2009).
Gamuyao, R. et al. The protein kinase Pstol1 from traditional rice confers tolerance of phosphorus deficiency. Nature 488, 535–539 (2012).
Gan, X. et al. Multiple reference genomes and transcriptomes for Arabidopsis thaliana. Nature 477, 419–423 (2011).
Li, Y. H. et al. De novo assembly of soybean wild relatives for pan-genome analysis of diversity and agronomic traits. Nat. Biotechnol. 32, 1045–1052 (2014).
Sakai, H. et al. Construction of pseudomolecule sequences of the aus rice cultivar Kasalath for comparative genomics of Asian cultivated rice. DNA Res. 21, 397–405 (2014).
Zhang, J. et al. Extensive sequence divergence between the reference genomes of two elite indica rice varieties Zhenshan 97 and Minghui 63. Proc. Natl Acad. Sci. USA 113, E5163–E5171 (2016).
Dong, J. et al. Analysis of tandem gene copies in maize chromosomal regions reconstructed from long sequence reads. Proc. Natl Acad. Sci. USA 113, 7949–7956 (2016).
Schatz, M. C. et al. Whole-genome de novo assemblies of three divergent strains of rice, Oryza sativa, document novel gene space of aus and indica. Genome Biol. 15, 506 (2014).
Lu, T. et al. Collection and comparative analysis of 1,888 full-length cDNAs from wild rice Oryza rufipogon Griff. W1943. DNA Res. 15, 285–295 (2008).
Liu, X. et al. A collection of 10,096 indica rice full-length cDNAs reveals highly expressed sequence divergence between Oryza sativa indica and japonica subspecies. Plant Mol. Biol. 65, 403–415 (2007).
Cao, J. et al. Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nat. Genet. 43, 956–963 (2011).
Navarro, J. A. R. et al. A study of allelic diversity underlying flowering-time adaptation in maize landraces. Nat. Genet. 49, 476–490 (2017).
Zhu, B. F. et al. Genetic control of a transition from black to straw-white seed hull in rice domestication. Plant Physiol. 155, 1301–1311 (2011).
Luo, J. et al. An-1 encodes a basic helix-loop-helix protein that regulates awn development, grain size and grain number in rice. Plant Cell 25, 3360–3376 (2013).
Li, X. M. et al. Natural alleles of a proteasome α2 subunit gene contribute to thermotolerance and adaptation of African rice. Nat. Genet. 47, 827–833 (2015).
Si, L. et al. OsSPL13 controls grain size in cultivated rice. Nat. Genet. 48, 447–456 (2016).
Kojima, S. et al. Hd3a, a rice ortholog of the Arabidopsis FT gene, promotes transition to flowering downstream of Hd1 under short-day conditions. Plant Cell Physiol. 43, 1096–1105 (2002).
Ma, Y. et al. COLD1 confers chilling tolerance in rice. Cell 160, 1209–1221 (2015).
Song, X. J. et al. Rare allele of a previously unidentified histone H4 acetyltransferase enhances grain weight, yield and plant biomass in rice. Proc. Natl Acad. Sci. USA 112, 76–81 (2015).
Yu, B. et al. TAC1, a major quantitative trait locus controlling tiller angle in rice. Plant J. 52, 891–898 (2007).
Asano, K. et al. Artificial selection for a green revolution gene during japonica rice domestication. Proc. Natl Acad. Sci. USA 108, 11034–11039 (2011).
Wang, Z. Y. et al. The amylose content in rice endosperm is related to the post-transcriptional regulation of the waxy gene. Plant J. 7, 613–622 (1995).
Yano, M. et al. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12, 2473–2484 (2000).
Takahashi, Y., Teshima, K. M., Yokoi, S., Innan, H. & Shimamoto, K. Variations in Hd1 proteins, Hd3a promoters, and Ehd1 expression levels contribute to diversity of flowering time in cultivated rice. Proc. Natl Acad. Sci. USA 106, 4555–4560 (2009).
Xue, W. et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 40, 761–767 (2008).
Cui, J. et al. Characterization and fine mapping of the ibf mutant in rice. J. Integr. Plant Biol. 49, 678–685 (2007).
Hirsch, C. N. et al. Insights into the maize pan-genome and pan-transcriptome. Plant Cell 26, 121–135 (2014).
Komatsu, M., Maekawa, M., Shimamoto, K. & Kyozuka, J. The LAX1 and FRIZZY PANICLE 2 genes determine the inflorescence architecture of rice by controlling rachis-branch and spikelet development. Dev. Biol. 231, 364–373 (2001).
Fujita, D. et al. NAL1 allele from a rice landrace greatly increases yield in modern indica cultivars. Proc. Natl Acad. Sci. USA 110, 20431–20436 (2013).
Sui, J. et al. A new GA-insensitive semidwarf mutant of rice (Oryza sativa L.) with a missense mutation in the SDG gene. Plant Mol. Biol. Report. 30, 187–194 (2012).
Nordborg, M. & Weigel, D. Next-generation genetics in plants. Nature 456, 720–723 (2008).
Myles, S. et al. Association mapping: critical considerations shift from genotyping to experimental design. Plant Cell 21, 2194–2202 (2009).
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Marouli, E. et al. Rare and low-frequency coding variants alter human adult height. Nature 542, 186–190 (2017).
McMullen, M. D. et al. Genetic properties of the maize nested-association mapping population. Science 325, 737–740 (2009).
Kover, P. X. et al. A multiparent advanced generation inter-cross to fine-map quantitative traits in Arabidopsis thaliana. PLoS Genet. 5, e1000551 (2009).
Huang, X. et al. Analysis of natural allelic variation in Arabidopsis using a multiparent recombinant inbred line population. Proc. Natl Acad. Sci. USA 108, 4488–4493 (2011).
Kozarewa, I. et al. Amplification-free Illumina sequencing-library preparation facilitates improved mapping and assembly of (G+C)-biased genomes. Nat. Methods 6, 291–295 (2009).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Li, H. Exploring single-sample SNP and indel calling with whole-genome de novo assembly. Bioinformatics 28, 1838–1844 (2012).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet. 16, 276–277 (2000).
Felsenstein, J. PHYLIP: phylogeny inference package (version 3.2). Cladistics 5, 164–166 (1989).
Salamov, A. A. & Solovyev, V. V. Ab initio gene finding in Drosophila genomic DNA. Genome Res. 10, 516–522 (2000).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-seq. Nat. Methods 5, 621–628 (2008).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Acknowledgements
We thank the China National Rice Research Institute for providing the cultivated rice germplasm and the Chinese wild rice accessions. The rest of the wild rice accessions were distributed from the National Institute of Genetics, which is supported by the National Bioresource Project, MEXT, Japan. We thank Z. Ning for assistance with genome assembly. This work was funded by the National Natural Science Foundation of China (91535202 and 91635302; to X.H.), the Ministry of Science and Technology of China (2016YFD0100902; to X.H.) and the Chinese Academy of Sciences (XDA08020101 and XDPB0401; to B.H.).
Author information
Authors and Affiliations
Contributions
X.H. and B.H. designed studies and contributed to the original concept of the project; A.W., Q. Zhan, Y.W., Z.W., Z.-X.W., Q.X. and X.W. contributed to the collection and planting of the rice accessions; Q.T., Y. Lu, D.F., C. Zhou, W.L., J.C., Q.W. and Q.F. performed the genome sequencing; Q. Zhao, H.L., Y. Li, L.Z., Y.Z., T.H. and C. Zhu performed genome data analysis; and X.H., Q. Zhao and B.H. analyzed whole data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Figures
Supplementary Figures 1–16.
Supplementary Data 1
Summary of 67 representative rice accessions for deep sequencing and de novo assembly.
Supplementary Data 2
Detailed statistics of the comparison between the GLA4 assembly and 87 BAC-based contigs.
Supplementary Data 3
List of six new domestication peaks identified using the pan-genome data.
Supplementary Data 4
Effects of the variants for the coding genes from 38 gene families.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, Q., Feng, Q., Lu, H. et al. Pan-genome analysis highlights the extent of genomic variation in cultivated and wild rice. Nat Genet 50, 278–284 (2018). https://doi.org/10.1038/s41588-018-0041-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0041-z
This article is cited by
-
Plant pangenomes for crop improvement, biodiversity and evolution
Nature Reviews Genetics (2024)
-
Dynamics of extrachromosomal circular DNA in rice
Nature Communications (2024)
-
A combination of conserved and diverged responses underlies Theobroma cacao’s defense response to Phytophthora palmivora
BMC Biology (2024)
-
A syntelog-based pan-genome provides insights into rice domestication and de-domestication
Genome Biology (2023)
-
Long-read, chromosome-scale assembly of Vitis rotundifolia cv. Carlos and its unique resistance to Xylella fastidiosa subsp. fastidiosa
BMC Genomics (2023)