Abstract
Pioneer transcription factors establish new cell-fate competence by triggering chromatin remodeling. However, many features of pioneer action, such as their kinetics and stability, remain poorly defined. Here, we show that Pax7, by opening a unique repertoire of enhancers, is necessary and sufficient for specification of one pituitary lineage. Pax7 binds its targeted enhancers rapidly, but chromatin remodeling and gene activation are slower. Enhancers opened by Pax7 show a loss of DNA methylation and acquire stable epigenetic memory, as evidenced by binding of nonpioneer factors after Pax7 withdrawal. This work shows that transient Pax7 expression is sufficient for stable specification of cell identity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Soufi, A., Donahue, G. & Zaret, K. S. Facilitators and impediments of the pluripotency reprogramming factors’ initial engagement with the genome. Cell 151, 994–1004 (2012).
Cirillo, L. A. et al. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol. Cell 9, 279–289 (2002).
Budry, L. et al. The selector gene Pax7 dictates alternate pituitary cell fates through its pioneer action on chromatin remodeling. Genes Dev. 26, 2299–2310 (2012).
van Oevelen, C. et al. C/EBPalpha activates pre-existing and de novo macrophage enhancers during induced pre-B cell transdifferentiation and myelopoiesis. Stem Cell Rep. 5, 232–247 (2015).
Boller, S. et al. Pioneering activity of the C-terminal domain of EBF1 shapes the chromatin landscape for B cell programming. Immunity 44, 527–541 (2016).
Wapinski, O. L. et al. Hierarchical mechanisms for direct reprogramming of fibroblasts to neurons. Cell 155, 621–635 (2013).
Pataskar, A. et al. NeuroD1 reprograms chromatin and transcription factor landscapes to induce the neuronal program. EMBO J. 35, 24–45 (2016).
Lamolet, B. et al. A pituitary cell-restricted T box factor, Tpit, activates POMC transcription in cooperation with Pitx homeoproteins. Cell 104, 849–859 (2001).
Pulichino, A. M. et al. Tpit determines alternate fates during pituitary cell differentiation. Genes Dev. 17, 738–747 (2003).
Lavoie, P. L., Budry, L., Balsalobre, A. & Drouin, J. Developmental dependence on NurRE and EboxNeuro for expression of pituitary proopiomelanocortin. Mol. Endocrinol. 22, 1647–1657 (2008).
Langlais, D., Couture, C., Kmita, M. & Drouin, J. Adult pituitary cell maintenance: lineage-specific contribution of self-duplication. Mol. Endocrinol. 27, 1103–1112 (2013).
Sheng, H. Z. & Westphal, H. Early steps in pituitary organogenesis. Trends Genet. 15, 236–240 (1999).
Charles, M. A. et al. PITX genes are required for cell survival and Lhx3 activation. Mol. Endocrinol. 19, 1893–1903 (2005).
Calo, E. & Wysocka, J. Modification of enhancer chromatin: what, how, and why? Mol. Cell 49, 825–837 (2013).
Langlais, D., Couture, C., Balsalobre, A. & Drouin, J. The Stat3/GR interaction code: predictive value of direct/indirect DNA recruitment for transcription outcome. Mol. Cell 47, 38–49 (2012).
Schuettengruber, B., Bourbon, H. M., Di Croce, L. & Cavalli, G. Genome regulation by Polycomb and Trithorax: 70 years and counting. Cell 171, 34–57 (2017).
Kawabe, Y., Wang, Y. X., McKinnell, I. W., Bedford, M. T. & Rudnicki, M. A. Carm1 regulates Pax7 transcriptional activity through MLL1/2 recruitment during asymmetric satellite stem cell divisions. Cell Stem Cell 11, 333–345 (2012).
Ghirlando, R. & Felsenfeld, G. CTCF: making the right connections. Genes Dev. 30, 881–891 (2016).
Liu, F., Wang, L., Perna, F. & Nimer, S. D. Beyond transcription factors: how oncogenic signalling reshapes the epigenetic landscape. Nat. Rev. Cancer 16, 359–372 (2016).
Drouin, J. Pituitary Development. In: S. Melmed ed. The Pituitary. 4th edn, 3–22. (Academic Press, Cambridge, MA, USA, 2017).
Rizzoti, K. Genetic regulation of murine pituitary development. J. Mol. Endocrinol. 54, R55–R73 (2015).
Zaret, K. S., Lerner, J. & Iwafuchi-Doi, M. Chromatin scanning by dynamic binding of pioneer factors. Mol. Cell 62, 665–667 (2016).
Voss, T. C. et al. Dynamic exchange at regulatory elements during chromatin remodeling underlies assisted loading mechanism. Cell 146, 544–554 (2011).
Soufi, A. et al. Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell 161, 555–568 (2015).
Swinstead, E. E. et al. Steroid receptors reprogram FoxA1 occupancy through dynamic chromatin transitions. Cell 165, 593–605 (2016).
Wysocka, J., Myers, M. P., Laherty, C. D., Eisenman, R. N. & Herr, W. Human Sin3 deacetylase and trithorax-related Set1/Ash2 histone H3-K4 methyltransferase are tethered together selectively by the cell-proliferation factor HCF-1. Genes Dev. 17, 896–911 (2003).
Laganière, J. et al. Location analysis of estrogen receptor α target promoters reveals that FOXA1 defines a domain of the estrogen response. Proc. Natl. Acad. Sci. USA 102, 11651–11656 (2005).
Wang, Q. et al. A hierarchical network of transcription factors governs androgen receptor-dependent prostate cancer growth. Mol. Cell 27, 380–392 (2007).
Carroll, J. S. et al. Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the forkhead protein FoxA1. Cell 122, 33–43 (2005).
Sherwood, R. I. et al. Discovery of directional and nondirectional pioneer transcription factors by modeling DNase profile magnitude and shape. Nat. Biotechnol. 32, 171–178 (2014).
Almouzni, G. & Cedar, H. Maintenance of epigenetic information. Cold Spring Harb. Perspect. Biol. 8, a019372 (2016).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Rhee, H. S. & Pugh, B. F. Comprehensive genome-wide protein-DNA interactions detected at single-nucleotide resolution. Cell 147, 1408–1419 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Akalin, A. et al. methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 13, R87 (2012).
Lerdrup, M., Johansen, J. V., Agrawal-Singh, S. & Hansen, K. An interactive environment for agile analysis and visualization of ChIP-sequencing data. Nat. Struct. Mol. Biol. 23, 349–357 (2016).
Acknowledgements
We are grateful to our colleagues N. Francis for critical comments on the manuscript; L. Budry and S. Nemec for profiling data and Pitx1 ChIP–seq, respectively; O. Neyret for NGS analyses; E. Massicotte for FACS sorting; A. Blanchet-Cohen for WGBS data analysis; and E. Joyal for expert secretarial assistance. A.M. was supported by an IRCM Challenge fellowship. This work was supported by grants to J.D. from the Canadian Institutes of Health Research.
Author information
Authors and Affiliations
Contributions
A.M., K.K., F.H. and Y.G. performed experiments. A.M., T.P. and J.D. conceived and designed the experiments. A.M., A.B. and J.D. analyzed the data. A.M. and J.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 High degree of correlation between RNA-seq and ATAC–seq replicates of FACS-enriched pituitary cells
(a) Dispersion plots showing correlation between RNA-seq values for all expressed genes (FPKM >1 in at least one sample) of two independent primary cell replicates of pituitary corticotropes and melanotropes. (b) Density plots showing correlation between two independent primary cell replicates of ATAC-seq′s performed in corticotropes and melanotropes. Only 200-bp bins with more than 50 reads in one sample were used for calculating the Pearson coefficient. Bins with more than 200 reads are shown on the plot as 201 reads. (c) Bar graphs showing the proportion of ATAC-seq peaks with P values (derived from MACS) between 10−5 and 10−20 (grey) and less than 10−20 (green). (d) Cross comparisons of ATAC-seq peaks in corticotropes (blue) and melanotropes (red) between the different replicates with the indicated P-value (derived from MACS) thresholds.
Supplementary Figure 2 Lineage-specific transcriptional program controlled by cell-specific DARs
(a,b) Expression levels of the 20 most highly expressed genes in corticotropes (a) and melanotropes (b). (c) Boxplot showing the distances between cortico- (n = 558) or melano- (n = 2,891) specific DARs and the closest differentially expressed genes. Center lines show medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th to 75th percentiles; outliers are represented by dots. P values are assessed by unpaired two-sided t test. (d) Boxplot showing the level of conservation (PhastCons) of lineage specific DARs compared to random control regions. Center lines show medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th to 75th percentiles; outliers are represented by dots. P values assessed by unpaired two-sided t test. (e, f) Genome browser views of lineage-specific genes for two melanotrope (e) and two corticotrope markers (f).
Supplementary Figure 3 DNA-sequence-motif searches at DARs identify Pax7 as a cell-fate regulator
(a, b) Motif enrichments obtained using HOMER on corticotrope (a) and melanotrope (b) DARs within a 200-bp window around DARs summit. The top three de novo and known motifs are shown. The bZIP motif identified in the melano-specific DARs (b) appears unique to this subset. However, this bZIP motif is not unique to melanotrope DARs since it is also a major motif extracted from DARs shared between the two POMC lineages and it is also found in gonadotrope DARs (not shown). (c) Distribution of changes (P <0.05) in cortico- and melano-specific gene expression in Pax7-/- neuro-intermediate lobe3.
Supplementary Figure 4 Analysis of subsets of Pax7 genomic targets
(a) Scheme for clustering subsets of Pax7 binding sites identified (P < 10−5, derived using MACS) by ChIP-seq (n = 89,206). Pax7 peaks were associated with the presence of H3K4me1 and p300 peaks (P < 10−5) before or after Pax7 expression (summit of H3K4me1 peak +/− 1 kb from Pax7 summit, +/− 500 bp for p300 peaks). The subsets with H3K4me1 before and after Pax7 were defined as Constitutively Active (p300 present before and after Pax7) and Pax7 Activated putative enhancers (p300 present only after Pax7). The subset that gained H3K4me1 after Pax7 binding was deemed to include putative Pioneer sites, and was further subdivided into Primed or Activated pioneer subsets depending on the accompanying gain of p300. Finally, pioneer targets being mostly at intergenic and intronic regions, we extracted for all subsets the intergenic and intronic peaks to define putative enhancers for further analyses. The number of peaks in each subset is indicated. (b) P value (derived from MACS) distribution of the four subsets (n indicated above in a) of Pax7 targets used for analyses. (c) Average DNAse hypersensitivity (GEO SRX034837) profiles for four subsets of peaks defined in a. (d) Average H3K4me1 profiles before (blue) or after (red) Pax7 at the four subsets described in Fig. 3. (e) Heatmaps of Tpit and Stat3 ChIP-seq signals at the four subsets of Pax7 peaks described in Fig. 3. (f) GR binding changes, measured by ChIP-seq (P < 10−5, derived from MACS), before or after Pax7 action at the indicated subsets (n indicated above in a) of Pax7 peaks. (g) Genome browser views of the four loci used for qPCR validation in h. H3K4me1 ChIP-seq data are shown in AtT-20 cells with/without Pax7 and replicate 1 of ATAC-seq data from corticotropes (C) and melanotropes (M). (h) ChIP-qPCR for H3K4me1 at two Pax7 pioneer targets in the Kif21b and Pcsk2 loci, one putative enhancer of the GR locus open in both lineages (C and M), and another GR locus putative enhancer specifically opened in corticotropes (C) as indicated. The data shown are means +/- s.e.m. of tissue triplicates assessed by duplicate qPCR measurements. The negative control sites (Neg1 and Neg2) do not show any H3K4me1 enrichment.
Supplementary Figure 5 DNA sequence motifs identified in Pax7 subsets
(a) De novo motif enrichments identified using HOMER at the indicated subsets (n indicated in Supplementary Fig. 4a) of Pax7 targets. The top three identified motifs are shown. (b) De novo motif enrichments identified using HOMER at the indicated subsets of Pax7 targets using the indicated subsets as background to identify sequences associated with specific Pax7 subsets. The top three identified motifs are shown.
Supplementary Figure 6 Validation of chromatin-mark data produced in this study
(a) Heatmaps of indicated ChIP-seq data around 24,061 RefSeq TSS in AtT-20 cells before or after Pax7 ranked according to expression levels derived from RNA-seq data shown at right. (b) Genome-wide Pearson correlations of ChIP-seq signals of all chromatin marks used in this study. Genomic windows of 500 bp with ≥ 20 reads were used to calculate correlations; windows with more than 500 reads were downscaled to 501 reads. The repressive histone marks (H3K9me3, H3K9me2 and H3K27me3) cluster together, while the active mark H3K27ac correlates with both H3K4me1 and H3K4me3. In all cases, the strongest correlation is obtained when comparing each dataset with or without Pax7 (>0.89).
Supplementary Figure 7 Characterization of inducible ER-Pax7 cells
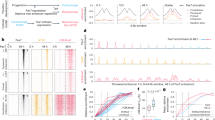
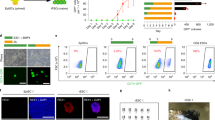
(a) Western blot showing Pax7 protein expression in Tam-induced ER-Pax7 AtT-20 cells compared to stable Pax7-expressing AtT-20 cells. A representative of two independent experiments is shown. (b) Gene induction curves of ER-Pax7 gene targets assessed by RNA-seq in cells treated with Tam for 12 h, 24 h and 96 h in comparison with non-treated cells. Gene targets were separated into two induction dynamics matching those observed in Fig. 5e. 100% corresponds to the maximum level of gene induction for each gene, and 0% representing the lowest level of the four time points measured. All target genes were induced at least 2-fold in stable Pax7-expressing and in ER-Pax7-expressing cells. Late targets show lower than 40% gene induction at 24 h, while early targets show more than 80% induction at 24 h.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7
Supplementary Tables
Supplementary Tables 1 and 2
Rights and permissions
About this article
Cite this article
Mayran, A., Khetchoumian, K., Hariri, F. et al. Pioneer factor Pax7 deploys a stable enhancer repertoire for specification of cell fate. Nat Genet 50, 259–269 (2018). https://doi.org/10.1038/s41588-017-0035-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-017-0035-2
This article is cited by
-
Pioneer factor Pax7 initiates two-step cell-cycle-dependent chromatin opening
Nature Structural & Molecular Biology (2024)
-
Protein-intrinsic properties and context-dependent effects regulate pioneer factor binding and function
Nature Structural & Molecular Biology (2024)
-
Pioneer factors — key regulators of chromatin and gene expression
Nature Reviews Genetics (2023)
-
PAX3-FOXO1 dictates myogenic reprogramming and rhabdomyosarcoma identity in endothelial progenitors
Nature Communications (2023)
-
The identification of PAX7 variants and a potential role of muscle development dysfunction in congenital scoliosis
Cell Regeneration (2022)