Abstract
Transcription factors (TFs) direct developmental transitions by binding to target DNA sequences, influencing gene expression and establishing complex gene-regultory networks. To systematically determine the molecular components that enable or constrain TF activity, we investigated the genomic occupancy of FOXA2, GATA4 and OCT4 in several cell types. Despite their classification as pioneer factors, all three TFs exhibit cell-type-specific binding, even when supraphysiologically and ectopically expressed. However, FOXA2 and GATA4 can be distinguished by low enrichment at loci that are highly occupied by these factors in alternative cell types. We find that expression of additional cofactors increases enrichment at a subset of these sites. Finally, FOXA2 occupancy and changes to DNA accessibility can occur in G1-arrested cells, but subsequent loss of DNA methylation requires DNA replication.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Spitz, F. & Furlong, E. E. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Cirillo, L. A. et al. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol. Cell 9, 279–289 (2002).
Iwafuchi-Doi, M. et al. The pioneer transcription factor FoxA maintains an accessible nucleosome configuration at enhancers for tissue-specific gene activation. Mol. Cell 62, 79–91 (2016).
Iwafuchi-Doi, M. & Zaret, K. S. Pioneer transcription factors in cell reprogramming. Genes Dev. 28, 2679–2692 (2014).
Soufi, A., Donahue, G. & Zaret, K. S. Facilitators and impediments of the pluripotency reprogramming factors’ initial engagement with the genome. Cell 151, 994–1004 (2012).
Soufi, A. et al. Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell 161, 555–568 (2015).
Zaret, K. S. & Carroll, J. S. Pioneer transcription factors: establishing competence for gene expression. Genes Dev. 25, 2227–2241 (2011).
Lupien, M. et al. FoxA1 translates epigenetic signatures into enhancer-driven lineage-specific transcription. Cell 132, 958–970 (2008).
Zaret, K. S. & Mango, S. E. Pioneer transcription factors, chromatin dynamics, and cell fate control. Curr. Opin. Genet. Dev. 37, 76–81 (2016).
Hurtado, A., Holmes, K. A., Ross-Innes, C. S., Schmidt, D. & Carroll, J. S. FOXA1 is a key determinant of estrogen receptor function and endocrine response. Nat. Genet. 43, 27–33 (2011).
Chen, J. et al. Single-molecule dynamics of enhanceosome assembly in embryonic stem cells. Cell 156, 1274–1285 (2014).
Swinstead, E. E. et al. Steroid receptors reprogram FoxA1 occupancy through dynamic chromatin transitions. Cell 165, 593–605 (2016).
Liu, Z. & Kraus, W. L. Catalytic-independent functions of PARP-1 determine Sox2 pioneer activity at intractable genomic loci. Mol. Cell 65, 589–603. e9 (2017).
Franco, H. L., Nagari, A. & Kraus, W. L. TNFα signaling exposes latent estrogen receptor binding sites to alter the breast cancer cell transcriptome. Mol. Cell 58, 21–34 (2015).
Tuteja, G., Jensen, S. T., White, P. & Kaestner, K. H. Cis-regulatory modules in the mammalian liver: composition depends on strength of Foxa2 consensus site. Nucleic Acids Res. 36, 4149–4157 (2008).
Li, Z., Schug, J., Tuteja, G., White, P. & Kaestner, K. H. The nucleosome map of the mammalian liver. Nat. Struct. Mol. Biol. 18, 742–746 (2011).
Cirillo, L. A. et al. Binding of the winged-helix transcription factor HNF3 to a linker histone site on the nucleosome. EMBO J. 17, 244–254 (1998).
Gifford, C. A. et al. Transcriptional and epigenetic dynamics during specification of human embryonic stem cells. Cell 153, 1149–1163 (2013).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Bailey, T. L. & Machanick, P. Inferring direct DNA binding from ChIP-seq. Nucleic Acids Res. 40, e128 (2012).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Voss, T. C. et al. Dynamic exchange at regulatory elements during chromatin remodeling underlies assisted loading mechanism. Cell 146, 544–554 (2011).
Gualdi, R. et al. Hepatic specification of the gut endoderm in vitro: cell signaling and transcriptional control. Genes Dev. 10, 1670–1682 (1996).
Zaret, K. Developmental competence of the gut endoderm: genetic potentiation by GATA and HNF3/fork head proteins. Dev. Biol. 209, 1–10 (1999).
Bossard, P. & Zaret, K. S. GATA transcription factors as potentiators of gut endoderm differentiation. Development 125, 4909–4917 (1998).
Tsankov, A. M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349 (2015).
Jozwik, K. M., Chernukhin, I., Serandour, A. A., Nagarajan, S. & Carroll, J. S. FOXA1 directs H3K4 monomethylation at enhancers via recruitment of the methyltransferase MLL3. Cell Rep. 17, 2715–2723 (2016).
Sérandour, A. A. et al. Epigenetic switch involved in activation of pioneer factor FOXA1-dependent enhancers. Genome Res. 21, 555–565 (2011).
Wang, A. et al. Epigenetic priming of enhancers predicts developmental competence of hESC-derived endodermal lineage intermediates. Cell Stem Cell 16, 386–399 (2015).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Brinkman, A. B. et al. Sequential ChIP-bisulfite sequencing enables direct genome-scale investigation of chromatin and DNA methylation cross-talk. Genome Res. 22, 1128–1138 (2012).
Kohli, R. M. & Zhang, Y. TET enzymes, TDG and the dynamics of DNA demethylation. Nature 502, 472–479 (2013).
Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Smith, Z. D. & Meissner, A. The simplest explanation: passive DNA demethylation in PGCs. EMBO J. 32, 318–321 (2013).
Sakaue-Sawano, A. et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell 132, 487–498 (2008).
Sekiya, T., Muthurajan, U. M., Luger, K., Tulin, A. V. & Zaret, K. S. Nucleosome-binding affinity as a primary determinant of the nuclear mobility of the pioneer transcription factor FoxA. Genes Dev. 23, 804–809 (2009).
Chronis, C. et al. Cooperative binding of transcription factors orchestrates reprogramming. Cell 168, 442–459 (2017). e20.
Sekiya, T. & Zaret, K. S. Repression by Groucho/TLE/Grg proteins: genomic site recruitment generates compacted chromatin in vitro and impairs activator binding in vivo. Mol. Cell 28, 291–303 (2007).
Wang, J.-C. et al. Transducin-like enhancer of split proteins, the human homologs of Drosophila groucho, interact with hepatic nuclear factor 3b. J. Biol. Chem. 275, 18418–18423 (2000).
Zhang, Y. et al. Nucleation of DNA repair factors by FOXA1 links DNA demethylation to transcriptional pioneering. Nat. Genet. 48, 1003–1013 (2016).
Stark, R. & Brown, G. DiffBind: differential binding analysis of ChIP–seq peak data. http://bioconductor.org/packages/release/bioc/vignettes/DiffBind/inst/doc/DiffBind.pdf (2011).
Mikkelsen, T. S. et al. Comparative epigenomic analysis of murine and human adipogenesis. Cell 143, 156–169 (2010).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Landt, S. G. et al. ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813–1831 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Quinlan, A. R. BEDTools: The Swiss-Army Tool for Genome FeatureAnalysis. Curr. Protoc. Bioinformatics 47, 11–12 (2014).
Acknowledgements
We thank all members of the laboratory of A.M., specifically A. Tsankov for advice and experimental support. M.J.Z. is supported by BMBF grant 01ZX1504 and the Max-Planck-Society. A.M. is supported as a New York Stem Cell Foundation Robertson Investigator. This work was supported by the New York Stem Cell Foundation, NIH grant 1P50HG006193 and the Max Planck Society.
Author information
Authors and Affiliations
Contributions
J.D. and A.M. conceived and designed the experimental study. J.D. and J.S.C. performed all experiments. S.T. and J.C. performed analysis of next-generation sequencing data. J.D., S.T. and J.C. performed all data interpretation and analysis with guidance from A.M., Z.D.S., and M.J.Z. D.C., C.A.G. and Z.D.S. provided experimental support. Initial data processing and analyses were performed by K.C., R.K., D.R.K. and M.J.Z. with the guidance of A.M. and J.L.R. H.G. and E.K.S. generated WGBS libraries under the guidance of A.G. R.P. assisted with flow cytometry analysis. J.D. and A.M. wrote the manuscript with help from S.T., Z.D.S. and J.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Pioneer-TF systems
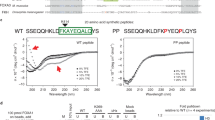
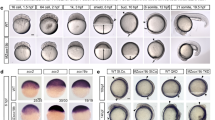
(a) FOXA2, part of the Forkhead box TF family, was first characterized as a pioneer TF for its ability to remodel nucleosomes at the repressed enhancers of the Albumin locus during endoderm development17,23. Ablation of FOXA2 in mice is embryonic lethal due to defects in early developmental structures, pointing to a critical role in lineage specification. Interestingly, however, after early development, FOXA2 is widely expressed across most endodermal and some ectodermal cell types, suggesting the need for specificity in its regulation29. Likewise, studies looking at FOXA1 occupancy across similar breast cancer cell types noted evidence of cell type specific binding8,9,10. Taken together, this has suggested that FOXA’s specific activity is likely not directed solely by the presence of its cognate DNA motif sequence and that there are perhaps additional features guiding even pioneer factor occupancy. To dissect this we used different Motif logo’s of the PWMs (shown in Fig. 1a) for identifying genome-wide occurrence of selected motifs throughout Hg19 using FIMO. (b) Chart displaying name of PWM used in each motif analysis, number of times the PWM mapped across the genome, the number of motifs within potentially ‘active’ regulatory regions, motifs in ‘active’ regions bound by FOXA and the calculated percentage of bound motifs. Potentially active regulatory regions were identified by utilizing all DNAse-seq, H3K27ac, and H3K4me3 data from the ENCODE project (utilizing Irreproducibility Discovery Rate (IDR) peak calling on ChIP-seq experiments; see Online Methods). (c) Peak saturation analysis of new FOXA2 peak calls obtained with FOXA2 ChIP-sequencing experiments in additional cell types. (d) Expression bar plot generated in CummeRbund displaying log2 FPKM values for FOXA family members: FOXA2, FOXA1 and FOXA3 in human hepatocytes (positive control) BJ fibroblasts, BJ fibroblasts infected with control RFP virus (negative control). Error bars represent a 95% confidence interval around the average values. (e) Immunostaining for FOXA2 in the JD1 BJFOXA2 line (10x magnification shown). White scale bar is equal to 345 nm. (f) qRT-PCR measurements of FOXA2 transcript level at four time points over a 10 day time course. No expression is measured on day 0. Stable FOXA2 transcript level is observed across days 1, 4 and 10 following induction. (g) Bright-field images show morphological change in JD1 BJFOXA2 cells after 3 days of doxycycline. White scale bar is equal to 345 nm. (h) Venn diagram displays the strong overlap and similar number of MACS peak calls for FOXA2 ChIP-sequencing after 4 and 10 days of FOXA2 induction. (i) Venn diagram demonstrating the overlap of the intersection in MACS peak calls between the BJFOXA2 day 4/day 10 time points (combined n = 73,827) and FOXA2 ChIP-sequencing after 1 day of FOXA2 induction. (j) Centrimo (4.10.2) analysis displaying top three motifs located at the summit of BJFOXA2 peaks and p-values associated with motifs.
Supplementary Figure 2 Pioneer-TF sampling
a) Read density heat maps of FOXA2 enrichment in BJFOXA2 ChIP-seq data at FOXA2 endogenous peaks from HepG2 (n = 34,595) and A549 (n = 33,041) cells. Black bars indicate peak calls in common between ectopic FOXA2 ChIP-sequencing data and endogenous (HepG2 or A549) FOXA2 ChIP-sequencing. Dashed lines represent the start and end of FOXA2 peaks. Similar to the dEN results, most HepG2 and A549 sites still show some level of enrichment of FOXA2 in BJs that are however not called as significantly enriched by our peak calling. (b) Schematic of OCT4 and GATA4 ectopic systems with corresponding cropped western blots demonstrating protein levels and immunostaining. Inf, infection (see Online Methods). White scale bar is equal to 345 nm. (c) Read density heat maps of OCT4 enrichment in BJ cells infected with OCT4, SOX2, KLF4 and cMYC5 at OCT4 bound regions in human ESCs. Dashed lines represent the start and end of OCT4 peaks. (d) Endogenous sampling demonstrated by read density heat maps of FOXA2 enrichment in HepG2 and dEN at A549 bound FOXA2 sites. Bar indicates peak calls in common between HepG2 and dEN FOXA2 ChIP-seq data and A549 FOXA2 ChIP-seq. Dashed lines mark the start and end of FOXA2 peaks. (e) Density plots displaying FOXA2 log2 RPKM enrichment in A549, HepG2, dEN and/or BJ cells, at endogenous peak sets (A549, HepG2 and dEN, respectively). Dashed lines demarcate regions within the background distribution, regions called as sampled sites and regions that were called as peaks.
Supplementary Figure 3 Influence of epigenome on pioneer-TF occupancy
a) IGV browser tracks displaying FOXA2 binding at each chromatin state we defined (coordinates from left to right: chr6:109,366,481–109,381,042; chr14:75,743,837–75,747,300; chr6:108,485,215–108,512,013; chr6:108,213,193–108,245,723; chr20:43,024,520–43,048,327). Classification was defined and employed hierarchically. (b) Chromatin state map defining percentages of dEN FOXA2 bound regions using hESC chromatin data. Spearman correlation with dEN FOXA2 peaks and human ESC chromatin. FOXA2 ESC 5.6 FPKM, and dEN FPKM 20.1. (c) Left: Stacked bar plots display FOXA2, GATA4, and OCT4 closed chromatin bound regions and levels of H3K27me3. Right: Stacked bar plot displays levels of DNAme at FOXA2 and GATA4 bound regions in closed chromatin. Pie chart displaying BJGATA4 peak overlap with CpG Islands (CGIs). (d) Schematic representation of system used to generate BJHNF1A cells. (e) IGV browser shots displaying a 400-kb genomic region in HNF1A (using V5 antibody) ChIP experiments. Top three experiments are distinct biological replicate experiments in BJHNF1A cells. In contrast, the bottom track represents HNF1A binding when FOXA2 is co-expressed. (f) Cropped western blot analysis of HNF1A (V5) protein levels in soluble nuclear, chromatin bound and whole cell lysates in BJHNF1A cells compared to BJFOXA2-HNF1A cells. Control blots in BJFOXA2 cells demonstrate difference in chromatin bound protein fraction of the two factors assessed. Full western blot is shown in Supplementary Figure 9. (g) Read density heat map displaying enrichment of BJ H3K9me3 ChIP-sequencing (REMC) at heterochromatin domains (n = 256) defined in ref.5. (h) Read density heat map displaying FOXA2 enrichment of BJ FOXA2 ChIP-sequencing (REMC) at heterochromatin domains defined in ref.5. (i) Representative IGV browser tracks showing a zoomed out view on chromosome 8 (305,736- 42,374,902) that visualizes the general depletion of FOXA2 binding within H3K9me3 marked regions. (j) Percentage of exclusively bound endogenous sites that are found in K9-domains.
Supplementary Figure 4 Cofactor influence on pioneer-TF occupancy
(a) Differential motif analysis displaying –log10 P-value of enriched motifs in BJ exclusive sites versus dEN exclusive sites with the most significant motifs on the left. Expression (log2 FPKM) of the TF associated with the motif in both BJ and dEN is shown on the bottom. (b) Venn diagram showing the overlap between IDR peak calls that are co-bound by FOXA2 and GATA4 in dEN and dEN exclusive targets as compared to BJs. (c) Box plots displaying the RPKM of GATA4 enrichment in BJGATA4 and BJFOXA2-GATA4 at the subset of regions that are GATA4 stabilized compared to the non-enriched subset. Boxes indicate interquartile range, and whiskers show maximum and minimum values. Outliers are removed. (d) Box plots displaying RPKM of ATAC-seq enrichment in uninduced BJFOXA2 versus BJFOXA2-GATA4 at GATA4 stabilized bites. Boxes indicate interquartile range and whiskers show maximum and minimum values. Outliers are removed.
Supplementary Figure 5 Transcriptional and epigenetic effect of pioneer-TF occupancy
(a) Volcano plot of differentially expressed genes on day 4 of FOXA2 induction in the BJFOXA2 line compared to the uninduced control. Differentially expressed genes are identified using cufflinks. y-axis represents –log10 of P-value while x-axis shows fold change in log2 scale. (b) Scatter plots of H3K4me1/2 and H3K27ac signal at pre- versus post-FOXA2 induction. Dots highlighted in red are at least 2-fold upregulated and reach at least 1 RPKM of post-FOXA2 induction. Ellipses roughly highlight de novo gained versus enhanced changes. De novo regions have minimal levels of either modification prior to FOXA2 occupancy, gain at least 2-fold signal as well as become enriched above RPKM = 1 whereas enhanced regions have prior enrichment for either mark and gain at least 2-fold more enrichment upon occupancy. (c) Composite plots of ATAC-seq signal pre- and post-FOXA2 induction as well as after 10 days of DOX followed by 2 days withdrawal at regions that become accessible (left) and remain inaccessible (right) in BJFOXA2. (d) Differential motif analysis displayed as a bar plot using Homer for regions that become accessible versus regions that remain closed. (e) Mean enrichment of FOXA2 (RPKM) at regions that remain inaccessible versus those becoming fully accessible. Boxes indicate interquartile range and whiskers show maximum and minimum values. Outliers are removed. (f) Composite line plot of FOXA motif frequency across peak regions in those that become accessible (black) compared to the inaccessible set (grey). (g) Mean enrichment (RPKM) of pre-existing ATAC-seq signal at FOXA2 target site that remain closed and become open. Boxes indicate interquartile range and whiskers show maximum and minimum values. Outliers are removed. (h) Binned scatter plot for BJ peaks is pre-existing closed chromatin (ATAC RPKM >1) comparing ATAC-seq and H3K4me1, H3K4me2 or H3K27ac signal post- FOXA2 induction. Regions were binned based on ATAC signal and then mean ATAC and chromatin mark signal for each bin is plotted as dots. (i) Composite plot of H3K4me2 (black line) and H3K27ac (gray line) signal at all active promoter regions as defined by RNA-seq FPKM >3. (j) Cropped western blot analysis of FOXA2 and H3 protein as loading control after FOXA2 induction for 1 day and 10 days followed by 2 and 4 days of doxycycline withdrawal. Full western blot is shown in Supplementary Figure 9.
Supplementary Figure 6 FOXA2-deletion constructs
a) Schematic representation of the FOXA2 deletion constructs. (b) Cropped western blots corresponding to FOXA2 deletion constructs using V5 antibody and FOXA2 antibody. Full western blot is shown in Supplementary Figure 9. (c) Violin plots displaying mean methylation (of regions covered at least 3x in deletion construct data) in WGBS, FOXA2 wild-type or FOXA2 deletion conditions.
Supplementary Figure 7 FOXA2 occupancy and DNAme
(a) Oligomer probes were designed for electrophoretic mobility shift assay (EMSA) at the FOXA2 binding sites in the AFM genes as shown in the IGV browser track (chr4:74,263,092–74,395,230). Two oligo versions were synthesized for AFM (with and without a methylated CpG). Motif sequence is highlighted in red and CpG in blue. (b) EMSA using purified Halo-tagged FOXA2 protein demonstrates FOXA2 interacts equally with methylated, hemi-methylated and non-methylated oligomers. Competition experiments were performed with non-biotinylated oligomers at 10x and 100x the concentration of the biotinlyated oligomers. (c) Scatter plot of FOXA2 enrichment at class 3-1 regions compared to their change in DNAme. (d) Density plot capturing distance to nearest CpG from the summit of FOXA2 ChIP-sequencing peaks. Class 2, black. Class 3-1, blue. (e) Density plot capturing the percent methylation of the nearest CpG from the summit of FOXA2 ChIP-sequencing peaks. Class 2, black. Class 3-1, blue. (f) Average distance and methylation status to the nearest CpG from the peak summit. Statistical significance shown by Welch t-test. (g) Box plot shows the percent methylation of CpGs within 20bp windows from the summit of the peak extended 200 bp. Methylation measurements were taken from WGBS data prior to FOXA2 induction. Class 2, black. Class 3-1, blue. Boxes indicate interquartile range and whiskers show maximum and minimum values. Outliers are removed. (h) Density plot of ATAC-seq coverage 2 days after FOXA2 induction for class 2 (black) and class 3-1 (blue) target sites (left). Accompanying browser tracks on the right display FOXA2 ChIP-sequencing, BJ WGBS, FOXA2 ChIP-BS, and ATAC-seq prior to FOXA2 induction as well as 2 days following the induction. Class 2 region shown is chr12:54,011,044–54,012,658, and class 3-1 region shown is chr1:28,720,983–28,722,960. CpGs included in the analysis are shown in red and highlighted by a gray box. (i) Box plots displaying mean RPKM values of H3K4me2 (left) and H3K27ac (right) at class 3-1 compared to class 2 targets in pre- and post- FOXA2 induction conditions.
Supplementary Figure 8 Loss of DNAme is dependent on DNA replication
(a) CFSE time course signal for samples after 24 h and 48 h labeling plus/minus FOXA2 induction overlaid on Day 0 labeling time point. Bar plot shows the median CFSE signal for cells plus/minus Dox induction of FOXA2 over 4 days. (b) Venn diagram of the overlap shows high similarity in called peaks between the two samples. (c) IGV browser shot of a 589-kb genomic region (chr 20:52,141,443-52,731,602) in BJFOXA2-CDT1 mimosine-treated compared to two replicates of BJFOXA2 FOXA2 ChIP-seq during mimosine-halted and released conditions. (d) Scatter plot displaying FOXA2 enrichment value at BJ FOXA2 peak set for mimosine-halted BJFOXA2-CDT1 compared to mimosine-halted BJFOXA2 FOXA2 ChIP-seq. (e) Box plots show average methylation of all Class 3 in BJ WGBS, BJFOXA2 ChIP-BS and BJFOXA2-CDT1 ChIP-BS data. Regions shown had at least 10x coverage. Boxes indicate interquartile range and whiskers show maximum and minimum values. Outliers are removed.
Supplementary Figure 9 Full western blots
a) Full western blot of BJFOXA2 clonal cell lines from Fig. 1c. Part of the blot included in the main figure is boxed. (b) Full western blot of BJOCT4 and BJGATA4 lines from Supplementary Figure 2b. Part of the blot included in the main figure is boxed. (c) Full western blot of HNF1A soluble nuclear/ chromatin bound protein extracts from Supplementary Figure 3f. Part of the blot included in the main figure is boxed. (d) Full western blot of BJFOXA2 withdrawal conditions from Supplementary Figure 5h. Part of the blot included in the main figure is boxed. (e) Full western blot of FOXA2 deletion constructs from Supplementary Figure 6. Part of the blot included in the main figure is boxed. (f) Full western blot of FOXA2 and H3 in BJFOXA2-CDT1 in wild-type conditions, G1 block and G2/M block from Fig. 6e. Part of the blot included in the main figure is boxed.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9.
Supplementary Table 1
FOXA2 enrichment.
Supplementary Table 2
GATA4 enrichment.
Supplementary Table 3
OCT4 enrichment
Supplementary Table 4
Alignment of data.
Supplementary Table 5
Expression analysis uninduced versus FOXA2 induced.
Supplementary Table 6
FOXA2 ChIP-BS-seq.
Supplementary Table 7
FOXA2, G1 block ChIP-BS-seq.
Supplementary Table 8
FOXA2, replicating ChIP-BS-seq.
Rights and permissions
About this article
Cite this article
Donaghey, J., Thakurela, S., Charlton, J. et al. Genetic determinants and epigenetic effects of pioneer-factor occupancy. Nat Genet 50, 250–258 (2018). https://doi.org/10.1038/s41588-017-0034-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-017-0034-3
This article is cited by
-
Pioneer factor Pax7 initiates two-step cell-cycle-dependent chromatin opening
Nature Structural & Molecular Biology (2024)
-
Protein-intrinsic properties and context-dependent effects regulate pioneer factor binding and function
Nature Structural & Molecular Biology (2024)
-
Pioneer factors — key regulators of chromatin and gene expression
Nature Reviews Genetics (2023)
-
Activator-blocker model of transcriptional regulation by pioneer-like factors
Nature Communications (2023)
-
Histone modifications regulate pioneer transcription factor cooperativity
Nature (2023)