Abstract
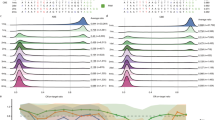
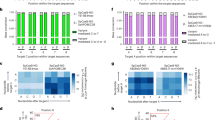
Base editors, including adenine base editors (ABEs)1 and cytosine base editors (CBEs)2,3, are widely used to induce point mutations. However, determining whether a specific nucleotide in its genomic context can be edited requires time-consuming experiments. Furthermore, when the editable window contains multiple target nucleotides, various genotypic products can be generated. To develop computational tools to predict base-editing efficiency and outcome product frequencies, we first evaluated the efficiencies of an ABE and a CBE and the outcome product frequencies at 13,504 and 14,157 target sequences, respectively, in human cells. We found that there were only modest asymmetric correlations between the activities of the base editors and Cas9 at the same targets. Using deep-learning-based computational modeling, we built tools to predict the efficiencies and outcome frequencies of ABE- and CBE-directed editing at any target sequence, with Pearson correlations ranging from 0.50 to 0.95. These tools and results will facilitate modeling and therapeutic correction of genetic diseases by base editing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The deep sequencing data from this study have been submitted to the National Center for Biotechnology Information (NCBI) Sequence Read Archive under accession number SRP150719 (PRJNA476544). The data sets used in this study are provided as Supplementary Tables 2, 3 and 4.
Code availability
Source code for DeepABE, DeepCBE, DeepCBE-CA and custom Python scripts used for the base-editing frequency and outcome calculations are available on github (at https://github.com/MyungjaeSong/Paired-Library and https://github.com/CRISPRJWCHOI/BaseEditing_tool).
References
Gaudelli, N. M. et al. Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, pii: aaf8729 (2016).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Doench, J. G. et al. Rational design of highly active sgRNAs for CRISPR–Cas9-mediated gene inactivation. Nat. Biotechnol. 32, 1262–1267 (2014).
Wong, N., Liu, W. & Wang, X. WU-CRISPR: characteristics of functional guide RNAs for the CRISPR/Cas9 system. Genome Biol. 16, 218 (2015).
Xu, H. et al. Sequence determinants of improved CRISPR sgRNA design. Genome Res. 25, 1147–1157 (2015).
Moreno-Mateos, M. A. et al. CRISPRscan: designing highly efficient sgRNAs for CRISPR-–Cas9 targeting in vivo. Nat. Methods 12, 982–988 (2015).
Chari, R., Mali, P., Moosburner, M. & Church, G. M. Unraveling CRISPR–Cas9 genome engineering parameters via a library-on-library approach. Nat. Methods 12, 823–826 (2015).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Kim, H. K. et al. In vivo high-throughput profiling of CRISPR-Cpf1 activity. Nat. Methods 14, 153–159 (2017).
Kuan, P. F. et al. A systematic evaluation of nucleotide properties for CRISPR sgRNA design. BMC Bioinformatics 18, 297 (2017).
Chari, R., Yeo, N. C., Chavez, A. & Church, G. M. sgRNA Scorer 2.0: a species-independent model to predict CRISPR/Cas9 activity. ACS Synth. Biol. 6, 902–904 (2017).
Labuhn, M. et al. Refined sgRNA efficacy prediction improves large- and small-scale CRISPR-Cas9 applications. Nucleic Acids Res. 46, 1375–1385 (2018).
Peng, H., Zheng, Y., Blumenstein, M., Tao, D. & Li, J. CRISPR/Cas9 cleavage efficiency regression through boosting algorithms and Markov sequence profiling. Bioinformatics 34, 3069–3077 (2018).
Wilson, L. O. W., Reti, D., O’Brien, A. R., Dunne, R. A. & Bauer, D. C. High activity target-site identification using phenotypic independent CRISPR-Cas9 core functionality. CRISPR J. 1, 182–190 (2018).
Chuai, G. et al. DeepCRISPR: optimized CRISPR guide RNA design by deep learning. Genome Biol. 19, 80 (2018).
Kim, H. K. et al. Deep learning improves prediction of CRISPR-Cpf1 guide RNA activity. Nat. Biotechnol. 36, 239–241 (2018).
Kim, H. K. et al. SpCas9 activity prediction by DeepSpCas9, a deep learning-based model with high generalization performance. Sci. Adv. 5, eaax9249 (2019).
Bae, S., Kweon, J., Kim, H. S. & Kim, J. S. Microhomology-based choice of Cas9 nuclease target sites. Nat. Methods 11, 705–706 (2014).
Shen, M. W. et al. Predictable and precise template-free CRISPR editing of pathogenic variants. Nature 563, 646–651 (2018).
Chakrabarti, A. M. et al. Target-specific precision of CRISPR-mediated genome editing. Mol. Cell 73, 699–713 (2019). e696.
Allen, F. et al. Predicting the mutations generated by repair of Cas9-induced double-strand breaks. Nat. Biotechnol. 37, 64–72 (2018).
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017).
Truong, D. J. et al. Development of an intein-mediated split-Cas9 system for gene therapy. Nucleic Acids Res. 43, 6450–6458 (2015).
Lai, Y. et al. Efficient in vivo gene expression by trans-splicing adeno-associated viral vectors. Nat. Biotechnol. 23, 1435–1439 (2005).
Ryu, S. M. et al. Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy. Nat. Biotechnol. 36, 536–539 (2018).
Villiger, L. et al. Treatment of a metabolic liver disease by in vivo genome base editing in adult mice. Nat. Med. 24, 1519–1525 (2018).
Zhou, C. et al. Off-target RNA mutation induced by DNA base editing and its elimination by mutagenesis. Nature 571, 275–278 (2019).
Koblan, L. W. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36, 843–846 (2018).
Gehrke, J. M. et al. An APOBEC3A-Cas9 base editor with minimized bystander and off-target activities. Nat. Biotechnol. 36, 977–982 (2018).
Thuronyi, B. W. et al. Continuous evolution of base editors with expanded target compatibility and improved activity. Nat. Biotechnol. 37, 1070–1079 (2019).
Grunewald, J. et al. CRISPR DNA base editors with reduced RNA off-target and self-editing activities. Nat. Biotechnol. 37, 1041–1048 (2019).
Doman, J. L., Raguram, A., Newby, G. A. & Liu, D. R. Evaluation and minimization of Cas9-independent off-target DNA editing by cytosine base editors. Nat. Biotechnol. 38, 620–628 (2020).
Zuo, E. et al. A rationally engineered cytosine base editor retains high on-target activity while reducing both DNA and RNA off-target effects. Nat. Methods 17, 600–604 (2020).
Gaudelli, N. M. et al. Directed evolution of adenine base editors with increased activity and therapeutic application. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0491-6 (2020).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0453-z (2020).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Shen, J. P. et al. Combinatorial CRISPR–Cas9 screens for de novo mapping of genetic interactions. Nat. Methods 14, 573–576 (2017).
Kim, H. W. et al. Differential effects on sodium current impairments by distinct SCN1A mutations in GABAergic neurons derived from Dravet syndrome patients. Brain Dev. 40, 287–298 (2018).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Lecun, Y., Bottou, L., Bengio, Y. & Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 86, 2278–2324 (1998).
Alipanahi, B., Delong, A., Weirauch, M. T. & Frey, B. J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Kelley, D. R., Snoek, J. & Rinn, J. L. Basset: learning the regulatory code of the accessible genome with deep convolutional neural networks. Genome Res. 26, 990–999 (2016).
Mnih, V. et al. Human-level control through deep reinforcement learning. Nature 518, 529–533 (2015).
Caruana, R., Lawrence, S. & Giles, C. L. Overfitting in neural nets: backpropagation, conjugate gradient, and early stopping. In Proc. Advances in Neural Information Processing Systems Vol. 13 (eds Leen, T. K. et al.) 402 (MIT Press, 2000); https://dl.acm.org/doi/proceedings/10.5555/3008751
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Machine Learn. Res. 15, 1929–1958 (2014).
Kingma, D. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2014).
Wolfe, J., Jin, X., Bahr, T. & Holzer, N. Application of Softmax regression and its validation for spectral-based land cover mapping. Int. Arch. Photogramm. Remote Sens. Spatial Inf. Sci. XLII-1/W1, 455–459 (2017).
Acknowledgements
We are grateful to W. Wurst and O. Ortiz for providing the split-Cas9 plasmids. We thank J. Lee, J. Kweon, Y. Kim and G. Yu for technical assistance. We are also grateful to S. Park and Y. Kim for assisting with the experiments. This work was supported in part by the National Research Foundation of Korea (grant nos. 2017R1A2B3004198 (H.K.), 2017M3A9B4062403 (H.K.) and 2018R1A5A2025079 (H.K.)), Brain Korea 21 Plus Project (Yonsei University College of Medicine), Yonsei University Future-leading Research Initiative of 2015 (Challenge; grant no. RMS2 2015-22-0092) and the Korean Health Technology R&D Project, Ministry of Health and Welfare, Republic of Korea (grant nos. HI17C0676 (H.K.) and HI16C1012 (H.K.)).
Author information
Authors and Affiliations
Contributions
M.S. performed experiments to build data sets of efficiencies and outcomes of base editing at integrated and endogenous sites. Y.K. and S.-Y.S. critically assisted the base-editing data generation by M.S., S.L., S.M., S.Y., M.S. and H.K.K. developed the framework and carried out the model training, computational validation and development of the web tools. J.P. and J.W.C. contributed substantially to bioinformatics analyses. Z.Q., J.H.K. and H.C.K. contributed to experiments using human iPSCs. H.H.K. conceived and designed the study. H.K.K., M.S., S.L. and H.H.K. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Yonsei University has filed a patent application based on this work, in which M.S., H.K.K., S.L. and H.H.K. are coinventors (patent no. PCT/KR2019/011166).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Text, Figs. 1–12 and Table 1.
Supplementary Table 2
Data sets of ABE- and CBE-directed target conversion efficiencies and edited sequence outcomes at integrated sequences.
Supplementary Table 3
Data sets of ABE- and CBE-directed target conversion efficiencies and edited sequence outcomes at endogenous sites.
Supplementary Table 4
Data subsets of Endo_ABE and Endo_CBE obtained by stratified random sampling and the exact P value for Fig. 2e.
Supplementary Table 5
The results of computational prediction of the base-editing efficiencies and outcomes for the ABE- and CBE-directed generation and correction of pathogenic/likely pathogenic point mutations.
Supplementary Table 6
Oligonucleotides used in this study.
Rights and permissions
About this article
Cite this article
Song, M., Kim, H.K., Lee, S. et al. Sequence-specific prediction of the efficiencies of adenine and cytosine base editors. Nat Biotechnol 38, 1037–1043 (2020). https://doi.org/10.1038/s41587-020-0573-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0573-5
This article is cited by
-
Deep learning models to predict the editing efficiencies and outcomes of diverse base editors
Nature Biotechnology (2024)
-
Deep learning models incorporating endogenous factors beyond DNA sequences improve the prediction accuracy of base editing outcomes
Cell Discovery (2024)
-
An adenine base editor variant expands context compatibility
Nature Biotechnology (2024)
-
Recent advances in CRISPR-based functional genomics for the study of disease-associated genetic variants
Experimental & Molecular Medicine (2024)
-
Cytosine base editors (CBEs) for inducing targeted DNA base editing in Nicotiana benthamiana
BMC Plant Biology (2023)