Abstract
Limited access to embryos has hampered the study of human embryogenesis and disorders that occur during early pregnancy. Human pluripotent stem cells provide an alternative means to study human development in a dish1,2,3,4,5,6,7. Recent advances in partial embryo models derived from human pluripotent stem cells have enabled human development to be examined at early post-implantation stages8,9,10,11,12,13,14. However, models of the pre-implantation human blastocyst are lacking. Starting from naive human pluripotent stem cells, here we developed an effective three-dimensional culture strategy with successive lineage differentiation and self-organization to generate blastocyst-like structures in vitro. These structures—which we term ‘human blastoids’—resemble human blastocysts in terms of their morphology, size, cell number, and composition and allocation of different cell lineages. Single-cell RNA-sequencing analyses also reveal the transcriptomic similarity of blastoids to blastocysts. Human blastoids are amenable to embryonic and extra-embryonic stem cell derivation and can further develop into peri-implantation embryo-like structures in vitro. Using chemical perturbations, we show that specific isozymes of protein kinase C have a critical function in the formation of the blastoid cavity. Human blastoids provide a readily accessible, scalable, versatile and perturbable alternative to blastocysts for studying early human development, understanding early pregnancy loss and gaining insights into early developmental defects.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
A total of nine single-cell RNA-seq datasets generated in this study have been deposited in the Gene Expression Omnibus (GEO) with accession code GSE150578. Source data are provided with this paper.
Code availability
Code used in this project is provided at https://github.com/jlduan/Human_blastoid.
Change history
02 August 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-03635-8
References
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Reubinoff, B. E., Pera, M. F., Fong, C. Y., Trounson, A. & Bongso, A. Embryonic stem cell lines from human blastocysts: somatic differentiation in vitro. Nat. Biotechnol. 18, 399–404 (2000).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Park, I. H. et al. Reprogramming of human somatic cells to pluripotency with defined factors. Nature 451, 141–146 (2008).
Wernig, M. et al. In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature 448, 318–324 (2007).
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007).
Aasen, T. et al. Efficient and rapid generation of induced pluripotent stem cells from human keratinocytes. Nat. Biotechnol. 26, 1276–1284 (2008).
Martyn, I., Kanno, T. Y., Ruzo, A., Siggia, E. D. & Brivanlou, A. H. Self-organization of a human organizer by combined Wnt and Nodal signalling. Nature 558, 132–135 (2018).
Simunovic, M. et al. A 3D model of a human epiblast reveals BMP4-driven symmetry breaking. Nat. Cell Biol. 21, 900–910 (2019).
Warmflash, A., Sorre, B., Etoc, F., Siggia, E. D. & Brivanlou, A. H. A method to recapitulate early embryonic spatial patterning in human embryonic stem cells. Nat. Methods 11, 847–854 (2014).
Shao, Y. et al. A pluripotent stem cell-based model for post-implantation human amniotic sac development. Nat. Commun. 8, 208 (2017).
Zheng, Y. et al. Controlled modelling of human epiblast and amnion development using stem cells. Nature 573, 421–425 (2019).
Xue, X. et al. Mechanics-guided embryonic patterning of neuroectoderm tissue from human pluripotent stem cells. Nat. Mater. 17, 633–641 (2018).
Moris, N. et al. An in vitro model of early anteroposterior organization during human development. Nature 582, 410–415 (2020).
Rivron, N. C. et al. Blastocyst-like structures generated solely from stem cells. Nature 557, 106–111 (2018).
Kime, C. et al. Induced 2C expression and implantation-competent blastocyst-like cysts from primed pluripotent stem cells. Stem Cell Rep. 13, 485–498 (2019).
Sozen, B. et al. Self-organization of mouse stem cells into an extended potential blastoid. Dev. Cell 51, 698–712 (2019).
Li, R. et al. Generation of blastocyst-like structures from mouse embryonic and adult cell cultures. Cell 179, 687–702 (2019).
Dong, C. et al. Derivation of trophoblast stem cells from naïve human pluripotent stem cells. eLife 9, e52504 (2020).
Linneberg-Agerholm, M. et al. Naïve human pluripotent stem cells respond to Wnt, Nodal and LIF signalling to produce expandable naïve extra-embryonic endoderm. Development 146, dev180620 (2019).
Guo, G. et al. Trophectoderm potency is retained exclusively in human naïve cells. Preprint at https://doi.org/10.1101/2020.02.04.933812 (2020).
Theunissen, T. W. et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 15, 471–487 (2014).
Okae, H. et al. Derivation of human trophoblast stem cells. Cell Stem Cell 22, 50–63 (2018).
Wu, J. et al. Interspecies chimerism with mammalian pluripotent stem cells. Cell 168, 473–486 (2017).
Wu, J. et al. An alternative pluripotent state confers interspecies chimaeric competency. Nature 521, 316–321 (2015).
Deglincerti, A. et al. Self-organization of the in vitro attached human embryo. Nature 533, 251–254 (2016).
Moon, K. R. et al. Visualizing structure and transitions in high-dimensional biological data. Nat. Biotechnol. 37, 1482–1492 (2019).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Petropoulos, S. et al. Single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016).
Takashima, Y. et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 158, 1254–1269 (2014).
Zhou, F. et al. Reconstituting the transcriptome and DNA methylome landscapes of human implantation. Nature 572, 660–664 (2019).
Lv, B. et al. Single-cell RNA sequencing reveals regulatory mechanism for trophoblast cell-fate divergence in human peri-implantation conceptuses. PLoS Biol. 17, e3000187 (2019).
Xiang, L. et al. A developmental landscape of 3D-cultured human pre-gastrulation embryos. Nature 577, 537–542 (2020).
Cinkornpumin, J. K. et al. Naive human embryonic stem cells can give rise to cells with a trophoblast-like transcriptome and methylome. Stem Cell Rep. 15, 198–213 (2020).
Nakamura, T. et al. A developmental coordinate of pluripotency among mice, monkeys and humans. Nature 537, 57–62 (2016).
Blakeley, P. et al. Defining the three cell lineages of the human blastocyst by single-cell RNA-seq. Development 142, 3151–3165 (2015).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Niu, Y. et al. Dissecting primate early post-implantation development using long-term in vitro embryo culture. Science 366, eaaw5754 (2019).
Bedzhov, I., Leung, C. Y., Bialecka, M. & Zernicka-Goetz, M. In vitro culture of mouse blastocysts beyond the implantation stages. Nat. Protocols 9, 2732–2739 (2014).
Shahbazi, M. N. et al. Self-organization of the human embryo in the absence of maternal tissues. Nat. Cell Biol. 18, 700–708 (2016).
Bedzhov, I. & Zernicka-Goetz, M. Self-organizing properties of mouse pluripotent cells initiate morphogenesis upon implantation. Cell 156, 1032–1044 (2014).
Niimura, S. Time-lapse videomicrographic analyses of contractions in mouse blastocysts. J. Reprod. Dev. 49, 413–423 (2003).
Eckert, J. J. et al. PKC signalling regulates tight junction membrane assembly in the pre-implantation mouse embryo. Reproduction 127, 653–667 (2004).
Eckert, J. J. et al. Relative contribution of cell contact pattern, specific PKC isoforms and gap junctional communication in tight junction assembly in the mouse early embryo. Dev. Biol. 288, 234–247 (2005).
Yu, L. et al. A protocol for the generation of blastocyst-like structures from human pluripotent stem cells. Protocol Exchange https://doi.org/10.21203/rs.3.pex-1359/v1 (2021).
Bredenkamp, N., Stirparo, G. G., Nichols, J., Smith, A. & Guo, G. The cell-surface marker sushi containing domain 2 facilitates establishment of human naive pluripotent stem cells. Stem Cell Rep. 12, 1212–1222 (2019).
Yu, L. et al. Derivation of intermediate pluripotent stem cells amenable to primordial germ cell specification. Cell Stem Cell https://doi.org/10.1016/j.stem.2020.11.003 (2020).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in unique molecular identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Linderman, G. C., Rachh, M., Hoskins, J. G., Steinerberger, S. & Kluger, Y. Fast interpolation-based t-SNE for improved visualization of single-cell RNA-seq data. Nat. Methods 16, 243–245 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Acknowledgements
We thank R. Jaenisch and T. Theunissen for providing the naive WIBR3 (OCT4-2A-GFP) cells; C. Zheng for help with cell culture; L. Zhang for technical support; and all other members of the Wu laboratory for discussion and suggestions. We acknowledge the BioHPC computational infrastructure at UT Southwestern for providing high-performance computing (HPC) and storage resources that have contributed to the research results reported within this paper. We also acknowledge UT Southwestern Genomics and Microarray Core for providing next-generation sequencing services. J.W. is a Virginia Murchison Linthicum Scholar in Medical Research and funded by CPRIT (RR170076) and Hamon Center for Regenerative Science and Medicine. G.H. is supported by CPRIT (RP190451), the Welch Foundation (I-1926-20170325), the Burroughs Wellcome Fund (1019804) and the Green Center for Reproductive Biology.
Author information
Authors and Affiliations
Contributions
J.W. and L.Y. conceptualized the idea, designed, analysed and interpreted the experimental results. L.Y. and Y.W. performed most of the human blastoid experiments. M.S. helped with immunofluorescence analysis. D.A.S. designed and prepared DNA constructs. J.D. and G.H. performed scRNA-seq analyses. L.W. performed scRNA-seq experiments. Y.W. and M.S. performed mouse embryo microinjection experiments. K.W. and S.Z. performed all human blastocyst experiments including human blastocyst thaw and culture, immunostaining and cell counting, and performed the grading of human blastocysts and blastoids. J.W. and G.H. supervised the study. L.Y., J.D., G.H. and J.W. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Jianping Fu, Josephine Johnston and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Generation of human blastoids from naive human PSCs.
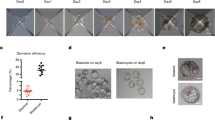
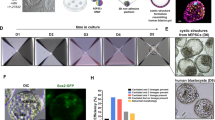
a, Representative immunofluorescence co-staining images of SOX2, GATA3 and GATA6 showing differentiation of naive human PSCs in human TSM (left) and HDM (right). Scale bars, 100 μm. b, Representative immunofluorescence co-staining images of SOX2, GATA3 and GATA6 showing differentiation of naive human PSCs in TSM followed by HDM (top) and HDM followed by TSM (bottom). Scale bars, 100 μm. c, A representative phase-contrast image showing the presence of ELCs, HLCs and TLCs in a day-7 codifferentiation culture. Green, purple and red dashed lines encircle a typical colony of ELCs, TLCs and HLCs, respectively. Scale bar, 100 μm. d, Representative phase-contrast images showing cell aggregates formed in media with different TSM and 5i/L/A ratios. Red arrowheads indicate blastocyst-like structures. Scale bars, 100 μm. e, Schematic of human blastoid formation from naive human PSCs using the TH method. f, Representative phase-contrast images of human blastoids (TH method). Scale bars, 100 μm. g, Representative phase-contrast images showing human blastoids generated from 5i/L/A-cultured HFF and BJ human iPS cells and PXGL-cultured WIBR3 human ES cells (HT method). Scale bar, 100 μm. h, Human blastoid formation efficiencies from 5i/L/A-cultured HFF human iPS cells (HT method, n = 6 biological replicates; TH method, n = 3 biological replicates), 5i/L/A-cultured BJ human iPS cells (HT method, n = 3 biological replicates) and PXGL-cultured WIBR3 human ES cells (HT method, n = 3 biological replicates). Data are mean ± s.d. i, Lengths and widths of human blastocysts (n = 13 biological replicates) and human blastoids (TH method, n = 36 biological replicates; HT method, n = 110 biological replicates). j, Total number of cells per human blastocyst (n = 9 biological replicates) and human blastoid (HT method, n = 52 biological replicates; TH method, n = 16 biological replicates). Box plots show median (centre line), 25th and 75th percentiles (bottom and top of box, respectively), and minimum and maximum values (bottom and top whisker, respectively). k, Morphological grading of human blastocysts (n = 11 biological replicates) and human blastoids (HT method, n = 97 biological replicates; TH method, n = 21 biological replicates). TE, trophectoderm. Images in a–d, f, g are representative of at least three independent experiments.
Extended Data Fig. 2 Optimization of blastoid formation from naive human PSCs.
a, Ratio of blastoids to all aggregates generated from different starting cell numbers (HT method, n = 3 biological replicates; mean ± s.d.). b, Ratio of empty wells or dead wells (wells containing only dead cells) to all wells using different starting cell numbers (HT method, n = 3 biological replicates; mean ± s.d.). c, The effect of each TDM component on blastoid formation efficiency (HT method).
Extended Data Fig. 3 Cell-lineage composition of human blastoids.
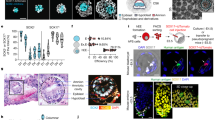
a, Representative immunofluorescence co-staining images of CDX2, GATA3 and SOX2 in a human blastoid. A higher-magnification image of the boxed area is shown on the right. Yellow arrowheads indicate CDX2+GATA3+ cells. Scale bar, 100 μm. b, Representative immunofluorescence co-staining images of TFAP2A, TFAP2C and SOX2 in a human blastoid. Scale bar, 100 μm. c, Representative immunofluorescence co-staining images of TEAD4 and SOX2 in a human blastoid. Scale bar, 100 μm. d, Representative immunofluorescence co-staining images of OCT4, GATA6 and SOX17 in a human blastoid. Left, maximum intensity projection of z-stack images; right, higher-magnification single-plane images of the boxed area. Yellow arrowheads indicate GATA6+SOX17+ cells. Scale bar, 100 μm. e, Representative immunofluorescence co-staining images of TFCP2L1, SUSD2 and KLF4 in a human blastoid. Left, maximum intensity projection of z-stack images; right, higher-magnification single-plane images of the boxed area. Scale bar, 100 μm. f, Representative immunofluorescence co-staining images of KLF17, CD24 and OTX2 in a human blastoid. Scale bar, 100 μm. g, Representative immunofluorescence co-staining images of SOX2, EOMES and T in a human blastoid. Scale bar, 100 μm. h, Representative immunofluorescence co-staining images of GATA3 and OCT4 (top) and GATA6 and SOX2 (bottom) in human blastoids generated from 5i/L/A-cultured HFF iPS cells. Scale bars, 100 μm. i, Representative immunofluorescence co-staining images of SOX2, SOX17 and GATA3 in a human blastoid (TH method). Scale bar, 100 μm. j, Quantification of the number of SOX2+, SOX17+ and GATA3+ cells in human blastocysts (n = 9 biological replicates) and human blastoids (TH method, n = 16 biological replicates; HT method, n = 30 biological replicates). Box plots show median (centre line), 25th and 75th percentiles (bottom and top of box, respectively), and minimum and maximum values (bottom and top whisker, respectively). k, Pie charts showing the frequencies of different expression patterns of SOX2, SOX17 and GATA3 in human blastocysts (left) and human blastoids (TH method, middle; HT method, right). l, m, Left, representative immunofluorescence co-staining images of OCT4 and GATA3 in cell aggregates generated by the HT (l) and TH (m) method. Scale bars, 100 μm. Right, percentages of cavityless structures, trophoblast spheroids and blastoids among all aggregates generated by the HT (l) and TH (m) method. Images in a–i, l, m are representative of at least three independent experiments.
Extended Data Fig. 4 Time-course studies of lineage differentiation dynamics during human blastoid formation and extended culture of human blastoids.
a, Representative immunofluorescence co-staining images of SOX2, GATA6 and GATA3 in cell aggregates generated at different time points during blastoid formation. Scale bars, 100 μm. b, Representative phase-contrast images showing extended culture of human blastoids in TDM. Scale bar, 100 μm. c, Percentages of EdU+ or Ki67+ cells in human blastoids after extended culture in TDM (day 9, n = 5; day 10, n = 6; day 11, n = 4 (biological replicates); mean ± s.d.). d, Representative immunofluorescence co-staining images of EdU and Ki67 in human blastoids after extended culture in TDM. Higher-magnification images of the boxed areas are shown on the right. Scale bars, 100 μm. Images in a, b, d are representative of at least three independent experiments.
Extended Data Fig. 5 Single-cell RNA-seq analyses of human blastoids.
a, PHATE embedding of blastoid cells. Shown is the expression of three lineage markers: SOX2 (epiblast), COL4A1 (hypoblast) and GATA2 (trophectoderm) b, Left, t-SNE plots comparing the transcriptomes of blastoid cells from TH and HT derivation protocols. Cluster labels are indicated. Middle, cells are coloured by derivation protocol. 5i/L/A and PXGL indicate two culture conditions for naive human PSCs before blastoid derivation. Right, quantification of the cellular composition of each cluster. c, t-SNE plots comparing the transcriptomes of blastoid cells from TH and HT derivation protocols. Shown is the expression of three lineage markers: SOX2 (epiblast), COL4A1 (hypoblast) and GATA2 (trophectoderm). d, Joint UMAP embedding of single-cell transcriptomes from blastoids and human pre-implantation embryos (Fig. 2a). Shown is the expression of lineage markers. To emphasize blastoid cells, cells derived from human pre-implantation embryos are coloured grey. e, Violin plots illustrating the expression of selected lineage markers from single-cell expression data. Quantified cells are split between blastoids (left) and pre-implantation human embryos (right). f, Dot plot indicating the expression of well-known markers of epiblast (EPI), hypoblast (HYP) and trophectoderm (TE). Clusters contain only blastoid cells, and do not contain cells from human embryos. g, PCA of blastoid-derived HLCs (left) and TLCs (right) relative to published maps from human embryos and established stem cell lines19,20,29,31,32,33,34. Embryonic time points and cell line names are indicated (labelled as in each previous study), and the dotted grey lines indicate developmental time axes estimated from human embryo datasets. h, Enrichment of GO biological process terms for cluster-specific genes. In e–h, clusters contain only blastoid cells, and do not contain cells from human embryos.
Extended Data Fig. 6 Additional single-cell RNA-seq analyses of human blastoids.
a, Correlation analysis of blastoid clusters with cells from in vivo cynomolgus monkey embryos (gast, gastrulating cells; postE-EPI, post-implantation early epiblast; postL-EPI, post-implantation late epiblast; post-paTE, post-implantation parietal trophectoderm; pre-EPI, pre-implantation epiblast; preE-TE, pre-implantation early trophectoderm; preL-TE, pre-implantation late trophoectoderm). b, Single-cell RNA-seq was performed at days 3, 6 and 9 during blastoid derivation (n = 3). t-SNE plot of sequenced cells, coloured by cluster assignment. c, Quantification of the cellular composition of clusters in b. d, Expression of lineage markers from the t-SNE analysis in b. e, Pseudotime trajectories for the ELCs across a time course of blastoid derivation. Epiblast cells from human pre-implantation embryos are included as a reference. Plots show pseudotime (top), sample time point (middle) and cell identity (bottom). The variability of U6 cells across time points may be due to cellular heterogeneity and unsynchronized differentiation of naive human PSCs during blastoid formation. f, g, Pseudotime-dependent gene expression signatures for the ELCs. This analysis only includes epiblast-specific genes defined in Zhou et al.31. h, Representative immunofluorescence and phase-contrast images showing the expression of CCR7 in a human blastoid. Images are representative of at least three independent experiments. Scale bar, 100 μm. i, t-SNE plot from Fig. 2h, without velocity analysis.
Extended Data Fig. 7 Derivation of stem cells from human blastoids.
a, Derivation efficiency for blastoid naive ES cells, blastoid nEND, and blastoid TSCs. Thirty blastoids were used for each. b, Growth curves of blastoid naive ES cells, blastoid nEND and blastoid TSCs (n = 3 biological replicates; mean ± s.d.). c, Representative immunofluorescence images of blastoid naive ES cells (OCT4, KLF17), blastoid nEND (SOX17, FOXA2) and blastoid TSCs (CDH1, TEAD4). Scale bars, 100 μm. d, RT–qPCR results showing the relative expression levels of epiblast (POU5F1, SOX2, KLF17), hypoblast (GATA4, GATA6, SOX17) and trophectoderm (GATA3, TFAP2C) markers in blastoid naive ES cells, blastoid nEND and blastoid TSCs (n = 3 biological replicates; mean ± s.d.). e, t-SNE of naive ES cells, blastoid naive ES cells, blastoid nEND and blastoid TSCs. Plots are coloured according to sample (top left), SOX2 expression (top right), GATA6 expression (bottom left) and GATA3 expression (bottom right). f, Correlation analysis of blastoid naive ES cells, blastoid nEND and blastoid TSCs compared to published transcriptomes19,20,29,31,32,33,34 from established human ES cells, human TSCs and nEND. Each indicated sample represents merged cells (scRNA-seq datasets) or replicates (bulk RNA-seq datasets). The gene sets used in this analysis were defined in Zhou et al.31. g, Left, representative phase-contrast images showing manual cutting (red dashed line) and outgrowths (days 1 and 7) during the derivation of blastoid TSCs. Right, a representative phase-contrast image showing the morphology of blastoid TSCs (passage 5) derived from manually cut TLCs. Scale bars, 100 μm. h, A representative immunofluorescence image showing that human TSCs derived from cut TLCs express GATA3. Scale bar, 100 μm. i, Schematic showing the strategy to derive blastoid nEND and blastoid TSCs from OCT4-GFP− and SUSD2− double-negative single cells (DNSCs) isolated from human blastoids. j, FACS gating and sorting of OCT4-GFP− and SUSD2− DNSCs from dissociated human blastoids. k, l, Representative phase-contrast and immunofluorescence images showing that blastoid TSCs (k) and blastoid nEND (l) derived from FACS-sorted DNSCs express GATA3 and GATA6, respectively. Scale bars, 100 μm. m–o, Representative phase-contrast (left) and immunofluorescence (right) images showing that monomeric Kusabira-Orange (mKO)-labelled blastoid TSCs (m), blastoid nEND (n) and blastoid naive ES cells (o) contribute to the extraembryonic ectoderm, visceral endoderm and epiblast compartments of day-6 IVC mouse embryos after blastocyst injections. Higher-magnification images of the boxed areas are shown on the right. Scale bars, 100 μm. Images in c, g, h, k–o are representative of at least three independent experiments.
Extended Data Fig. 8 In vitro differentiation of stem cells derived from human blastoids.
a, Representative phase-contrast images showing blastoid TSCs, as well as EVT-like cells and SCT-like cells differentiated from blastoid TSCs. Scale bars, 100 μm. b, Representative immunofluorescence images of GATA3, CK7 and CDH1 (trophectoderm markers), SDC1 and CGB (SCT markers) and HLA-G (an EVT marker) in EVT-like cells and SCT-like cells differentiated from blastoid TSCs. Scale bars, 100 μm. c, RT–qPCR results showing the relative expression levels of EVT (FN1, HLA-G, ITGA5, ITGA1) and SCT (CGA, CGB (detects all 6 CGB genes), PSG1, CSH1, CYP19A1) markers in blastoid TSCs, as well as in EVT-like and SCT-like cells differentiated from blastoid TSCs (n = 3 biological replicates, mean ± s.d.). d, A representative phase-contrast image showing a day-9 differentiation culture of blastoid nEND cells. Scale bar, 100 μm. e, f, Representative immunofluorescence co-staining images of COL6A1, FOXA1 and GATA6 in a day-9 differentiation culture of blastoid nEND cells. White dashed lines indicate FOXA1−COL6A1+GATA6− cells (e) and FOXA1+COLA6A1−GATA6− cells (f). Scale bars, 100 μm. g, RT–qPCR results showing relative expression levels of VE/YE and EXMC markers in day-4 and day-9 differentiation cultures of blastoid nEND cells. (n = 3 biological replicates; mean ± s.d.). h, A representative phase-contrast image showing the colony morphology of primed human ES cells converted from blastoid naive ES cells. Scale bar, 100 μm. i, Representative immunofluorescence images showing that primed human ES cells converted from blastoid naive ES cells stained positive for SOX2 and CD24. Scale bars, 100 μm. j, Representative H&E staining images showing that the teratomas formed by primed human ES cells contain tissues from all three embryonic germ layers. Scale bar, 100 μm. Images in a, b, d–f, h–j are representative of at least three independent experiments.
Extended Data Fig. 9 Culturing human blastoids beyond implantation stages.
a, Representative immunofluorescence co-staining images of SOX2, GATA6 and GATA3 in selected day-4 IVC human blastoids, showing abnormal growth of ELCs, TLCs and/or HLCs. Scale bars, 100 μm. b, Representative immunofluorescence co-staining images of phalloidin and OCT4 showing a rosette-like structure in a day-4 IVC human blastoid. A higher-magnification image of the boxed area is shown on the right. Green arrowhead indicates the apical domain of the rosette-like structure. Scale bar, 100 μm. c, Representative immunofluorescence co-staining images of OTX2 and laminin in a day-4 IVC human blastoid. Side views of the yellow dashed line are shown on the right. Scale bar, 100 μm. d Representative immunofluorescence co-staining images of OCT4 and CK7 in a day-4 IVC human blastoid. Higher-magnification images of the boxed area are shown on the right. Scale bar, 100 μm. e, Representative immunofluorescence co-staining images of HCGB and SOX2 in a day-4 IVC human blastoid. A higher-magnification image of the boxed area is shown on the right. White dotted line encircles multinucleated cells. Scale bar, 100 μm. f, Representative results using a human pregnancy test kit on medium collected from day-4 IVC human blastoids and IVC medium control (Ct.). Images in a–e are representative of at least three independent experiments.
Extended Data Fig. 10 PKC isozymes regulate cavity formation in human blastoids.
a, Representative immunofluorescence co-staining images of phalloidin and ZO1 in cell aggregates generated at different time points during blastoid formation. Scale bars, 100 μm. b, Representative phase-contrast images of cell aggregates after treatments with DMSO control (top), Gö6976 (middle) or Gö6983 (bottom). Red arrowheads indicate blastoids. Scale bar, 100 μm. c, Counts per million (CPM) values calculated from a published scRNA-seq dataset29, showing the expression levels of different PKC isozymes at different stages during human pre-implantation development. E, early stage; L, late stage. d, e, Representative immunofluorescence co-staining images of ZO1, phalloidin and GATA3 in cell aggregates after treatments with inhibitors of PKCα (d) and PKCζ (e). Higher-magnification images of the boxed areas are shown on the left. Scale bars, 100 μm. Images in a, b, d, e are representative of at least three independent experiments.
Supplementary information
Supplementary Table 1
Sequencing statistics and differential gene expression.
Supplementary Table 2
GO enrichment for blastoid cells - full list of Gene Ontology terms enriched for ELCs, HLCs and TLCs specific genes.
Supplementary Table 3
Summary of ex vivo human-mouse chimera experiments. Monomeric Kusabira-Orange (mKO) labeled blastoid-hTSCs, blastoid-nEND, and blastoid-nESCs were injected into E3.5 mouse blastocysts followed by 3 (IVC3) and 5/6 (IVC5/6) days in vitro embryo culture.
Supplementary Table 4
A list of antibodies and primers used in this study.
Video 1
: Cavity expansion in human blastoids Time-lapse imaging shows human blastoids undergo cavity expansion.
Rights and permissions
About this article
Cite this article
Yu, L., Wei, Y., Duan, J. et al. Blastocyst-like structures generated from human pluripotent stem cells. Nature 591, 620–626 (2021). https://doi.org/10.1038/s41586-021-03356-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03356-y
This article is cited by
-
Self-renewing human naïve pluripotent stem cells dedifferentiate in 3D culture and form blastoids spontaneously
Nature Communications (2024)
-
VGLL1 cooperates with TEAD4 to control human trophectoderm lineage specification
Nature Communications (2024)
-
Reprogramming fibroblast into human iBlastoids
Nature Protocols (2024)
-
Modelling post-implantation human development to yolk sac blood emergence
Nature (2024)
-
Hypoblast from human pluripotent stem cells regulates epiblast development
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.