Abstract
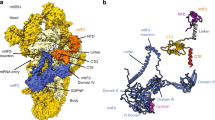
Mitochondria maintain their own specialized protein synthesis machinery, which in mammals is used exclusively for the synthesis of the membrane proteins responsible for oxidative phosphorylation1,2. The initiation of protein synthesis in mitochondria differs substantially from bacterial or cytosolic translation systems. Mitochondrial translation initiation lacks initiation factor 1, which is essential in all other translation systems from bacteria to mammals3,4. Furthermore, only one type of methionyl transfer RNA (tRNAMet) is used for both initiation and elongation4,5, necessitating that the initiation factor specifically recognizes the formylated version of tRNAMet (fMet–tRNAMet). Lastly, most mitochondrial mRNAs do not possess 5′ leader sequences to promote mRNA binding to the ribosome2. There is currently little mechanistic insight into mammalian mitochondrial translation initiation, and it is not clear how mRNA engagement, initiator-tRNA recruitment and start-codon selection occur. Here we determine the cryo-EM structure of the complete translation initiation complex from mammalian mitochondria at 3.2 Å. We describe the function of an additional domain insertion that is present in the mammalian mitochondrial initiation factor 2 (mtIF2). By closing the decoding centre, this insertion stabilizes the binding of leaderless mRNAs and induces conformational changes in the rRNA nucleotides involved in decoding. We identify unique features of mtIF2 that are required for specific recognition of fMet–tRNAMet and regulation of its GTPase activity. Finally, we observe that the ribosomal tunnel in the initiating ribosome is blocked by insertion of the N-terminal portion of mitochondrial protein mL45, which becomes exposed as the ribosome switches to elongation mode and may have an additional role in targeting of mitochondrial ribosomes to the protein-conducting pore in the inner mitochondrial membrane.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Greber, B. J. et al. The complete structure of the 55S mammalian mitochondrial ribosome. Science 348, 303–308 (2015).
Montoya, J., Ojala, D. & Attardi, G. Distinctive features of the 5′-terminal sequences of the human mitochondrial mRNAs. Nature 290, 465–470 (1981).
Atkinson, G. C. et al. Evolutionary and genetic analyses of mitochondrial translation initiation factors identify the missing mitochondrial IF3 in S. cerevisiae. Nucleic Acids Res. 40, 6122–6134 (2012).
Christian, B. E. & Spremulli, L. L. Mechanism of protein biosynthesis in mammalian mitochondria. Biochim. Biophys. Acta 1819, 1035–1054 (2012).
Kuzmenko, A. et al. Mitochondrial translation initiation machinery: conservation and diversification. Biochimie 100, 132–140 (2014).
Carter, A. P. et al. Crystal structure of an initiation factor bound to the 30S ribosomal subunit. Science 291, 498–501 (2001).
Weisser, M., Voigts-Hoffmann, F., Rabl, J., Leinundgut, M. & Ban, N. The crystal structure of the eukaryotic 40S ribosomal subunit in complex with eIF1 and eIF1A. Nat. Struct. Mol. Biol. 20, 1015–1017 (2013).
Weisser, M. et al. Structural and functional insights into human re-initiation complexes. Mol. Cell 67, 447–456 (2017).
Lomakin, I. B. et al. Crystal structure of the human ribosome in complex with DENR–MCT-1. Cell Reports 20, 521–528 (2017).
Gaur, R. et al. A single mammalian mitochondrial translation initiation factor functionally replaces two bacterial factors. Mol. Cell 29, 180–190 (2008).
Yassin, A. S. et al. Insertion domain within mammalian mitochondrial translation initiation factor 2 serves the role of eubacterial initiation factor 1. Proc. Natl Acad. Sci. USA 108, 3918–3923 (2011).
Shimizu, Y., Kanamori, T. & Ueda, T. Protein synthesis by pure translation system. Methods 36, 299–304 (2005).
Jones, C. N., Wilkinson, K. A., Hung, K. T., Weeks, K. M. & Spremulli, L. L. Lack of secondary structure characterizes the 5′ ends of mammalian mitochondrial mRNAs. RNA 14, 862–871 (2008).
Giliberti, J., O’Donnell, S., Etten, W. J. & Janssen, G. R. A. 5′-terminal phosphate is required for stable ternary complex formation and translation of leaderless mRNA in Escherichia coli. RNA 18, 508–518 (2012).
Christian, B. E. & Spremulli, L. L. Preferential selection of the 5′-terminal start codon on leaderless mRNAs by mammalian mitochondrial ribosomes. J. Biol. Chem. 285, 28379–28386 (2010).
Voorhees, R. M., Schmeing, T. M., Kelley, A. C. & Ramakrishnan, V. The mechanism for activation of GTP hydrolysis on the ribosome. Science 330, 835–838 (2010).
Fernández, I. S. et al. Molecular architecture of a eukaryotic translation initiation complex. Science 342, (2013).
Szyrach, G., Ott, M., Bonnefoy, N., Neupert, W. & Herrmann, J. M. Ribosome binding to the Oxa1 complex facilitates co-translational protein insertion in mitochondria. EMBO 22, 6448–6457 (2003).
Keil, M. et al. Oxa1-ribosome complexes coordinate the assembly of cytochrome C oxidase in mitochondria. J. Biol. Chem. 287, 34484–34493 (2012).
Denks, K. et al. The signal recognition particle contacts uL23 and scans substrate translation inside the ribosomal tunnel. New Microbiol. 2, (2017).
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Greber, B. J. et al. Architecture of the large subunit of the mammalian mitochondrial ribosome. Nature 505, 515–519 (2014).
Spencer, A. C., Heck, A., Takeuchi, N., Watanabe, K. & Spremulli, L. L. Characterization of the human mitochondrial methionyl-tRNA synthetase. Biochemistry 43, 9743–9754 (2004).
Liao, H. X. & Spremulli, L. L. Effects of length and mRNA secondary structure on the interaction of bovine mitochondrial ribosomes with messenger RNA. J. Biol. Chem. 265, 11761–11765 (1990).
Blanquet, S., Dessen, P. & Kahn, D. Properties and specificity of methionyl-tRNAf Met formyltransferase from Escherichia coli. Methods Enzymol. 106, 141–152 (1984).
Koc, E. C. & Spremulli, L. L. Identification of mammalian mitochondrial translation initiation factor 3 and examination of its role in initiation complex formation with natural mRNAs. J. Biol. Chem. 277, 35541–35549 (2002).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, (2015).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Greber, B. J. et al. The complete structure of the large subunit of the mammalian mitochondrial ribosome. Nature 515, 283–286 (2014).
Jones, T. A. Interactive electron-density map interpretation: from INTER to O. Acta Crystallogr. D 60, 2115–2125 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Amunts, A., Brown, A., Toots, J., Scheres, S. H. W. & Ramakrishnan, V. The structure of the human mitochondrial ribosome. Science 348, 95–98 (2015).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015).
Diaconu, M. et al. Structural basis for the function of the ribosomal L7/12 stalk in factor binding and GTPase activation. Cell 121, 991–1004 (2005).
Leijonmarck, M. & Liljas, A. Structure of the C-terminal domain of ribosomal protein L7/L12 from Escherichia coli at 1.7 Å. J. Mol. Biol. 195, 555–579 (1987).
Roll-Mecak, A., Cao, C., Dever, T. E. & Burley, S. K. X-ray structures of the universal translation initiation factor IF2/eIF5B: conformational changes on GDP and GTP binding. Cell 103, 781–792 (2000).
Eiler, D., Lin, J., Simonetti, A., Klaholz, B. P. & Steitz, T. A. Initiation factor 2 crystal structure reveals a different domain organization from eukaryotic initiation factor 5B and mechanism among translational GTPases. Proc. Natl Acad. Sci. USA 110, 15662–15667 (2013).
Shi, H. & Moore, P. B. The crystal structure of yeast phenylalanine tRNA at 1.93 Å resolution: a classic structure revisited. RNA 6, 1091–1105 (2000).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Rackham, O. et al. Pentatricopeptide repeat domain protein 1 lowers the levels of mitochondrial leucine tRNAs in cells. Nucleic Acids Res. 37, 5859–5867 (2009).
Rackham, O. et al. Hierarchical RNA processing is required for mitochondrial ribosome assembly. Cell Reports 16, 1874–1890 (2016).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Fernandez-Leiro, R. & Scheres, S. H. W. A pipeline approach to single-particle processing in RELION. Acta Crystallogr. D. 73, 496–502 (2017).
Wahl, M. C. & Möller, W. Structure and function of the acidic ribosomal stalk proteins. Curr. Protein Pept. Sci. 3, 93–106 (2002).
Helgstrand, M. et al. The ribosomal stalk binds to translation factors IF2, EF-Tu, EF-G and RF3 via a conserved region of the L12 C-terminal domain. J. Mol. Biol. 365, 468–479 (2007).
Tourigny, D. S., Fernández, I. S., Kelley, A. C. & Ramakrishnan, V. Elongation factor G bound to the ribosome in an intermediate state of translocation. Science 340, (2013).
Pallesen, J. et al. Cryo-EM visualization of the ribosome in termination complex with apo-RF3 and RF1. eLife 2, (2013).
Kuhle, B. & Ficner, R. A monovalent cation acts as a structural and catalytic cofactor in translational GTPases. EMBO J. 33, 2547–2563 (2014).
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006).
Dolinsky, T. J., Nielsen, J. E., McCammon, J. A. & Baker, N. A. PDB2PQR: an automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acid Res. 32, W665–W667.
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001).
Li, L. et al. DelPhi: a comprehensive suite for DelPhi software and associated resources. BMC Biophys. 4, (2012).
Englmeier, R., Pfeffer, S. & Förster, F. Structure of the human mitochondrial ribosome studied in situ by cryoelectron tomography. Structure 25, 1574–1581 (2017).
Acknowledgements
We thank T. Schönhut for help with purification of mtIF2 variants; A. Scaiola for support with computational problems and data analysis; V. Godinic for advice on tRNA charging; T. Schönhut, P. Bieri and M. Saurer for help with mitochondria preparation; K. Warinner for help during cloning and for creating a great working atmosphere. We would like to thank the ETH Zürich scientific centre for optical and electron microscopy (ScopeM) for access to electron microscopy equipment and P. Tittmann for technical support. E.K. was supported by an EMBO long-term fellowship (1196-2014). This work was supported by the Swiss National Science Foundation grant (310030B_163478) and via the National Centre of Excellence in RNA and Disease and project funding 138262 to N.B. We thank the NHMRC, ARC and Cancer Council of WA for grants and fellowships to A.F. and O.R. and UWA for a postgraduate scholarship to R.G.L.
Reviewer information
Nature thanks L. Sazanov and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
E.K. prepared initiation complex components, ribosomal subunits and cryo-EM samples. D.B. and E.K. acquired the cryo-EM data. E.K. calculated the cryo-EM reconstructions with support from D.B. M.L. built the atomic model and performed coordinate refinement of the atomic model. E.K., M.L. and N.B. interpreted the structure. R.G.L., O.R. and A.F. performed in vivo experiments in HEK cells. E.K. performed in vitro translation assays and size-exclusion chromatography for ternary complex formation. All authors contributed to the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
: The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Classification scheme.
a, Schematic representation of how the different cryo-EM maps have been calculated in RELION31,47. Classification yielded three maps used for model building. Maps 1 and 2 were calculated from identical particles with map 1 being refined over the entire particle volume whereas map 2 refinement was focused on the 28S subunit of the ribosome. Map 3 derived from a different particle subset and was refined over the entire particle volume. Resolutions have been estimated in RELION by post-processing the entire ribosome (map 1), 28S including mtIF2 and tRNA (map 2, indicated in red) or 39S including mtIF2 and tRNA (map 3, indicated in red). b, Depiction of masks that have been applied for focused 3D classification or 3D refinement in RELION. Mask I encompasses the mtIF2 G domain, domain II and domain III. Mask II includes the 28S A site and the mtIF2 insert. Mask III focuses on fMet–tRNAMet and domain IV of mtIF2. Mask IV includes the 28S subunit.
Extended Data Fig. 2 Map evaluation.
a, Local resolution estimation performed in RELION yielded maps that were filtered according to the local resolution estimate47. Displayed are a front view and a slabbed view at a position indicated by the arrow on the left. Colour keys indicate the local resolution in Å. In contrast to the FSC curves in Extended Data Fig. 2b, local resolution has been estimated and depicted for the entire volume of map 1, map 2 and map 3. Map 2 indicates that the ribosomal subunits exhibit a substantial rotational freedom in the initiation complex leading to a poorly resolved 39S if the alignment is focused on the 28S during refinement. b, FSC curves calculated from the two particle half sets from gold-standard 3D refinement (black) or from model versus map (blue) for all three deposited maps and their corresponding PDBs. c, A representative micrograph shows the particle distribution of the 55S initiation complex on cryo-EM grids. d, Euler angle distribution of particles included in the final 3D reconstructions are shown using the .bild file generated in Relion. Distributions for map 1 and map 3 are displayed. Because map 1 and map 2 were generated from the same particle subset, map 2 distribution is expected to be very similar to the one from map 1 and is therefore not shown.
Extended Data Fig. 3 Quality of the cryo-EM maps around mtIF2 and the role of C-terminal domain of bL12m in subunit joining.
a, Sliced representation of the ternary complex (red) bound between the ribosomal subunits. All domains of mtIF2 are clearly resolved (maps at 2.2 and 4σ). b–e, Magnified views of different areas of mtIF2. b, The contact site of the G domain with domains II and III (maps at 3 and 6σ). c, GTPγS coordinated by the mtIF2 P loop and switch regions 1 and 2 (maps at 2.75 and 5.5σ). d, Domain IV (maps at 2.75 and 5.5σ). e, Map filtered to 5 Å showing bL12m-CTD bound to the solvent-side of the mtIF2 G-domain and opposite to GTPγS. The ribosomal L7–L12 stalk forms part of the conserved GTPase activating centre of the ribosomal large subunit in all kingdoms of life48. It enhances recruitment of translational GTPases to cytoplasmic ribosomes via its flexibly attached C-terminal domain (CTD), which has been observed to bind the G′ domains in EF-G and RF349,50,51. However, G′ does not exist in mtIF2. In the initiation complex, we show that bL12m-CTD recognizes the G-domain of mtIF2 on the surface-exposed side opposite the catalytic centre. Since initiation complex formation involves binding of mtIF2 to the small ribosomal subunit before the large subunit joins, the observed interactions of mtIF2 with bL12m may be important to promote subunit joining to form the 55S mitoribosomal initiation complex rather than recruitment of mtIF2 to the assembled ribosome. The bL12m-CTD was modelled with PHYRE236 using PDB 1CTF38 as a template and docked as a rigid body. Experimental densities are shown at two different contour levels (1.5 and 3σ).
Extended Data Fig. 4 Sequence alignments for different functionally important regions of mtIF2.
Boxes indicate where the aligned sequences are located. The alignments contain sequences from mammals as well as other vertebrates to depict a more general conservation. The alignment for the fMet interaction site also contains eIF5B homologues for which structures have already been published (Saccharomyces cerevisiae17, Methanothermobacter thermautotrophicus39, Chaetomium thermophilum52). The alignment for the C-terminal extension of mtIF2 also contains S. cerevisiae mtIF2 and cytosolic IF2 from E. coli, both of which lack the extension.
Extended Data Fig. 5 The mtIF2 insert contacts the decoding centre and closes the mRNA channel.
a, Two views of the α-helical element of the mtIF2 insert occupying the A site with experimental maps at two different contour levels (maps at 3 and 5σ). W486 stacks on top of decoding nucleotide G256 and F494 contacts the flipped out A919. mRNA bases are numbered according to their position relative to the 5′ end of the mRNA. b, The mtIF2 insert substantially extends α-helix 8 of the mitochondrial IF2 homologue and then enters the A site. A stable contact is established by a number of conserved positively charged residues facing the 12S rRNA (maps at 3 and 5σ). c, The fMet–tRNAMet anticodon stem loop (ASL) that is in contact with the MT-CO3 AUG start codon is stabilized in the P site by numerous conserved interactions with 12S rRNA53. The anticodon fully base-pairs with the start codon, with the tRNA wobble base of the anticodon stacking on top of C844 (C1400 in T. thermophilus) and its ribose against A571 (G966). G782/A783 (G1338/A1339) protrude from the 28S head to form A-minor interactions with fMet–tRNAMet specific G–C pairs 26:38 and 27:37. A430 (A790) stacks onto the ribose of tRNA nucleotide 35 to stabilize tRNA binding from the opposite side of the ASL. Protein uS13 is not present in mitoribosomes und thus uS9m is the only ribosomal protein to contact tRNA in the P site. Its C-terminal tail (comprising residues K396 and R397) reaches into a cavity formed by phosphates 30, 31 and 31 of the ASL backbone (map at 5σ).
Extended Data Fig. 6 mRNA binding and start codon selection on the mitochondrial ribosome.
a, Map 2 (classified for as described in Extended Data Fig. 1 and displayed without post-processing) shown colour coded according to the underlying atomic coordinates (grey, the small ribosomal subunit and mtIF2; yellow, uS5m). A lower contour level of map 2 is shown in transparency and locally filtered in Relion47. Density that cannot be assigned to the underlying atomic coordinates reaches from mS39 towards the mRNA entry site (magenta). We believe that this density contains mostly mRNA but it may also include 6 unassigned amino acids from mS39 and possibly part of 21 unassigned amino acids from the N terminus of mS35. b, The mRNA entry is surrounded by uS5m. mRNA (magenta) follows the positively charged surface of the mitochondria-specific uS5m extension towards the A site. The surface potential for uS5m was calculated using PDB 2PQR54 and visualized with the APBS tool55 from PyMOL (M. Lerner and H. Carlson, University of Michigan). ± 5 kT/e electrostatic potential of uS5m have been plotted. c, Although resolved to atomic resolution only in the area of the start codon–anticodon interaction, cryo-EM density for the mRNA can be assigned along its entire path through the mRNA channel, reaching from the P site—where the AUG start codon is located—into the A site, which is shielded by the mtIF2 insert. Subsequently, density nestles alongside protein uS5m, which substantially restricts the diameter of the mRNA channel and places a delineation that may prevent mRNA from slipping out of the mRNA channel. Map 2 is shown unfiltered (blue) and filtered to 5 Å (grey) at two contour levels. For clarity, cryo-EM density for the entire ribosome and mtIF2 has been subtracted from map 2 in Chimera and the difference density is carved 10 Å around our modelled mRNA (contour levels are 10 and 15σ). The mRNA occupies a similar position as in the elongation complex1. d, Alignment of the first 70 nucleotides of the 11 mRNA 5′ ends in human mitochondria (MT-ND1, MT-ND2, MT-CO1, MT-CO2, MT-ATP8, MT-CO3, MT-ND3, MT-ND4L, MT-ND5, MT-CYB, MT-ND6), starting precisely at the start codon. Codons are indicated by bars and numbered. Alignments were generated using the weblogo server (https://weblogo.berkeley.edu).
Extended Data Fig. 7 mL45 serves as ribosomal membrane anchor and is crucial for insertion of oxidative phosphorylation proteins.
a, Sequence alignment of the mL45 N-terminal extension in vertebrates shows strong sites of conservation—predominantly in mammals. b, The surface potential of mL45 (calculated using DELPHI implemented in MOLDRAW56 and visualized with the APBS tool of PyMOL) shows that the membrane-facing side of mL45 contains a large positively charged patch that may mediate association to the negatively charged inner mitochondrial membrane57. For comparison, the structure is shown as a cartoon in the right panel, with positions of positively charged residues in the putative membrane interaction area as blue spheres. c, Polypeptide synthesis necessitates displacement of the NTE at a hinge region around G64. The mL45 NTE was truncated at positions G64 and K71 to study its role in vivo. Locations of positively charged residues in mL45 α-helices proposed to mediate membrane association of the ribosome are indicated as blue spheres. d, Left, cell lysates (25 µg) from HEK293T cells co-transfected with control or mL45 CRISPR/Cas9 plasmids and MRPL45 wild type or deletion mutant expressing plasmids were resolved on 4–20% SDS–PAGE gels and immunoblotted to investigate the steady-state levels of nuclear- and mitochondrial-encoded oxidative phosphorylation (OXPHOS) and ribosomal proteins. β-actin was used as a loading control. The data are representative of at least three independent biological experiments. Right, quantification of the relative abundance of the OXPHOS polypeptides relative to control and normalized to the β-actin loading control. Error bars indicate standard error of the mean. **P < 0.01, ***P < 0.001, Student’s t-test. e, A continuous 10–30% sucrose gradient was used to determine the distribution of the small and large ribosomal subunit and polysomes in mitochondria isolated from cells expressing wild-type or truncated mL45. Mitochondrial ribosomal protein markers of the small (bS16m) and large (mL45) ribosomal subunits were detected by immunoblotting. The input, mitochondrial lysate, was used as a positive control. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 8 Additional information on in vitro translation assays.
a, Depiction of the construct used for in vitro translation assays. Human DHFR was fused to streptactin binding protein (SBP) via a linker encoding the amino acid sequence GSSGS. Ribosome binding site (RBS), linker regions (nt, nucleotides) as well as T7 promoter and terminator have been copied from the PURE Express template plasmid provided by the manufacturer (New England Biolabs). The XhoI cleavage site was used to generate DNA templates for run-off transcription. DHFR–SPB was efficiently expressed either upon addition of DNA as template or after supplying DHFR–SBP mRNA directly (data not shown). We decided to perform all subsequent experiments providing equal amounts of mRNA to ensure that every sample contains the same concentration of template in order to make translation yields comparable. b, Production of DHFR–SBP was monitored after 2 h at 37 °C at different concentrations of mtIF2(WT). The positive control contained E. coli IF1, IF2 and IF3 but no mtIF2. The negative control contained all E. coli initiation factors but lacked mRNA. mtIF2 was tested at the given concentrations and in the presence of E. coli IF3. Immunoblots (left) were quantified using the gel analysis routine in ImageJ (right). The sample containing 4 µM mtIF2 was excluded from quantification because it was only partially transferred onto the nitrocellulose membrane during blotting. c, SDS–PAGE with 2 μg protein loaded for each mtIF2 variant to show that protein concentrations have been estimated correctly for all variants before in vitro translation was performed (data for mtIF2(H678A) not shown). d, Since experiments to determine the translation activity of mtIF2(H678A) were performed at a later time point than for other variants, samples were analysed on separate immunoblots. The immunoblot shows samples from 4 independent experiments for mtIF2(H678A) and 2 independent experiments for mtIF2(WT). Bands have been quantified as for other mtIF2 variants using the gel analysis routine in ImageJ and activity was normalized to mtIF2(WT) (see Fig. 2).
Supplementary information
Supplementary Figure 1
Images contained in this study that have been cropped are shown as full scanned images with boxes indicating how gels and immunoblots were trimmed for the final figures. The respective figure numbers for the cropped images are provided together with molecular weight markers.
Supplementary Table 1
Components of the mitochondrial 55S initiation complex. Table containing all components of the mammalian mitochondrial 55S initiation complex with their respective chain IDs, an indication for which residues of the full-length molecule have been build and their accession codes.* The individually deposited 28S and 39S ribosomal subunits carry the same chain IDs as the 55S assembly. Nomenclature according to Ban et al.58.† Nomenclature according to ribosomal protein gene database60.‡ Full-length protein sequence including putative mitochondrial targeting peptides.§ Unassigned residues were modeled as poly-alanine and deposited as UNK.
Rights and permissions
About this article
Cite this article
Kummer, E., Leibundgut, M., Rackham, O. et al. Unique features of mammalian mitochondrial translation initiation revealed by cryo-EM. Nature 560, 263–267 (2018). https://doi.org/10.1038/s41586-018-0373-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0373-y
This article is cited by
-
Structural insights into the role of GTPBP10 in the RNA maturation of the mitoribosome
Nature Communications (2023)
-
The P-type pentatricopeptide repeat protein DWEORG1 is a non-previously reported rPPR protein of Arabidopsis mitochondria
Scientific Reports (2022)
-
Organization and expression of the mammalian mitochondrial genome
Nature Reviews Genetics (2022)
-
Mechanism of mitoribosomal small subunit biogenesis and preinitiation
Nature (2022)
-
Distinct mechanisms of the human mitoribosome recycling and antibiotic resistance
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.