Abstract
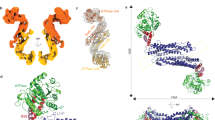
Mitochondrial inheritance, genome maintenance and metabolic adaptation depend on organelle fission by dynamin-related protein 1 (DRP1) and its mitochondrial receptors. DRP1 receptors include the paralogues mitochondrial dynamics proteins of 49 and 51 kDa (MID49 and MID51) and mitochondrial fission factor (MFF); however, the mechanisms by which these proteins recruit and regulate DRP1 are unknown. Here we present a cryo-electron microscopy structure of full-length human DRP1 co-assembled with MID49 and an analysis of structure- and disease-based mutations. We report that GTP induces a marked elongation and rotation of the GTPase domain, bundle-signalling element and connecting hinge loops of DRP1. In this conformation, a network of multivalent interactions promotes the polymerization of a linear DRP1 filament with MID49 or MID51. After co-assembly, GTP hydrolysis and exchange lead to MID receptor dissociation, filament shortening and curling of DRP1 oligomers into constricted and closed rings. Together, these views of full-length, receptor- and nucleotide-bound conformations reveal how DRP1 performs mechanical work through nucleotide-driven allostery.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mishra, P. & Chan, D. C. Mitochondrial dynamics and inheritance during cell division, development and disease. Nat. Rev. Mol. Cell Biol. 15, 634–646 (2014).
Bleazard, W. et al. The dynamin-related GTPase Dnm1 regulates mitochondrial fission in yeast. Nat. Cell Biol. 1, 298–304 (1999).
Toyama, E. Q. et al. AMP-activated protein kinase mediates mitochondrial fission in response to energy stress. Science 351, 275–281 (2016).
Roy, M., Reddy, P. H., Iijima, M. & Sesaki, H. Mitochondrial division and fusion in metabolism. Curr. Opin. Cell Biol. 33, 111–118 (2015).
Twig, G. et al. Fission and selective fusion govern mitochondrial segregation and elimination by autophagy. EMBO J. 27, 433–446 (2008).
Mao, K., Wang, K., Liu, X. & Klionsky, D. J. The scaffold protein Atg11 recruits fission machinery to drive selective mitochondria degradation by autophagy. Dev. Cell 26, 9–18 (2013).
Chan, D. C. Fusion and fission: interlinked processes critical for mitochondrial health. Annu. Rev. Genet. 46, 265–287 (2012).
van der Bliek, A. M., Shen, Q. & Kawajiri, S. Mechanisms of mitochondrial fission and fusion. Cold Spring Harb. Perspect. Biol. 5, a011072 (2013).
Frank, S. et al. The role of dynamin-related protein 1, a mediator of mitochondrial fission, in apoptosis. Dev. Cell 1, 515–525 (2001).
Ishihara, N. et al. Mitochondrial fission factor Drp1 is essential for embryonic development and synapse formation in mice. Nat. Cell Biol. 11, 958–966 (2009).
Wakabayashi, J. et al. The dynamin-related GTPase Drp1 is required for embryonic and brain development in mice. J. Cell Biol. 186, 805–816 (2009).
Chatel-Chaix, L. et al. Dengue virus perturbs mitochondrial morphodynamics to dampen innate immune responses. Cell Host Microbe 20, 342–356 (2016).
Kim, S. J. et al. Hepatitis B virus disrupts mitochondrial dynamics: induces fission and mitophagy to attenuate apoptosis. PLoS Pathog. 9, e1003722 (2013).
Friedman, J. R. et al. ER tubules mark sites of mitochondrial division. Science 334, 358–362 (2011).
Murley, A. et al. ER-associated mitochondrial division links the distribution of mitochondria and mitochondrial DNA in yeast. eLife 2, e00422 (2013).
Lewis, S. C., Uchiyama, L. F. & Nunnari, J. ER-mitochondria contacts couple mtDNA synthesis with mitochondrial division in human cells. Science 353, aaf5549 (2016).
Osman, C., Noriega, T. R., Okreglak, V., Fung, J. C. & Walter, P. Integrity of the yeast mitochondrial genome, but not its distribution and inheritance, relies on mitochondrial fission and fusion. Proc. Natl Acad. Sci. USA 112, E947–E956 (2015).
Labbé, K., Murley, A. & Nunnari, J. Determinants and functions of mitochondrial behavior. Annu. Rev. Cell Dev. Biol. 30, 357–391 (2014).
Kraus, F. & Ryan, M. T. The constriction and scission machineries involved in mitochondrial fission. J. Cell Sci. 130, 2953–2960 (2017).
Osellame, L. D. et al. Cooperative and independent roles of the Drp1 adaptors Mff, MiD49 and MiD51 in mitochondrial fission. J. Cell Sci. 129, 2170–2181 (2016).
Palmer, C. S. et al. Adaptor proteins MiD49 and MiD51 can act independently of Mff and Fis1 in Drp1 recruitment and are specific for mitochondrial fission. J. Biol. Chem. 288, 27584–27593 (2013).
Palmer, C. S. et al. MiD49 and MiD51, new components of the mitochondrial fission machinery. EMBO Rep. 12, 565–573 (2011).
Koirala, S. et al. Interchangeable adaptors regulate mitochondrial dynamin assembly for membrane scission. Proc. Natl Acad. Sci. USA 110, E1342–E1351 (2013).
Gandre-Babbe, S. & van der Bliek, A. M. The novel tail-anchored membrane protein Mff controls mitochondrial and peroxisomal fission in mammalian cells. Mol. Biol. Cell 19, 2402–2412 (2008).
Fröhlich, C. et al. Structural insights into oligomerization and mitochondrial remodelling of dynamin 1-like protein. EMBO J. 32, 1280–1292 (2013).
Mears, J. A. et al. Conformational changes in Dnm1 support a contractile mechanism for mitochondrial fission. Nat. Struct. Mol. Biol. 18, 20–26 (2011).
Ingerman, E. et al. Dnm1 forms spirals that are structurally tailored to fit mitochondria. J. Cell Biol. 170, 1021–1027 (2005).
Daumke, O. & Praefcke, G. J. Mechanisms of GTP hydrolysis and conformational transitions in the dynamin superfamily. Biopolymers 105, 580–593 (2016).
Lee, J. E., Westrate, L. M., Wu, H., Page, C. & Voeltz, G. K. Multiple dynamin family members collaborate to drive mitochondrial division. Nature 540, 139–143 (2016).
Losón, O. C., Song, Z., Chen, H. & Chan, D. C. Fis1, Mff, MiD49, and MiD51 mediate Drp1 recruitment in mitochondrial fission. Mol. Biol. Cell 24, 659–667 (2013).
Lackner, L. L., Horner, J. S. & Nunnari, J. Mechanistic analysis of a dynamin effector. Science 325, 874–877 (2009).
Faelber, K. et al. Crystal structure of nucleotide-free dynamin. Nature 477, 556–560 (2011).
Ford, M. G., Jenni, S. & Nunnari, J. The crystal structure of dynamin. Nature 477, 561–566 (2011).
Richter, V. et al. Structural and functional analysis of MiD51, a dynamin receptor required for mitochondrial fission. J. Cell Biol. 204, 477–486 (2014).
Losón, O. C. et al. The mitochondrial fission receptor MiD51 requires ADP as a cofactor. Structure 22, 367–377 (2014).
Liu, R. & Chan, D. C. The mitochondrial fission receptor Mff selectively recruits oligomerized Drp1. Mol. Biol. Cell 26, 4466–4477 (2015).
Losón, O. C. et al. Crystal structure and functional analysis of MiD49, a receptor for the mitochondrial fission protein Drp1. Protein Sci. 24, 386–394 (2015).
Gao, S. et al. Structural basis of oligomerization in the stalk region of dynamin-like MxA. Nature 465, 502–506 (2010).
Haller, O., Gao, S., von der Malsburg, A., Daumke, O. & Kochs, G. Dynamin-like MxA GTPase: structural insights into oligomerization and implications for antiviral activity. J. Biol. Chem. 285, 28419–28424 (2010).
Reubold, T. F. et al. Crystal structure of the dynamin tetramer. Nature 525, 404–408 (2015).
Chappie, J. S., Acharya, S., Leonard, M., Schmid, S. L. & Dyda, F. G domain dimerization controls dynamin’s assembly-stimulated GTPase activity. Nature 465, 435–440 (2010).
Vanstone, J. R. et al. DNM1L-related mitochondrial fission defect presenting as refractory epilepsy. Eur. J. Hum. Genet. 24, 1084–1088 (2016).
Sheffer, R. et al. Postnatal microcephaly and pain insensitivity due to a de novo heterozygous DNM1L mutation causing impaired mitochondrial fission and function. Am. J. Med. Genet. A. 170, 1603–1607 (2016).
Chang, C. R. et al. A lethal de novo mutation in the middle domain of the dynamin-related GTPase Drp1 impairs higher order assembly and mitochondrial division. J. Biol. Chem. 285, 32494–32503 (2010).
Chang, C. R. & Blackstone, C. Dynamic regulation of mitochondrial fission through modification of the dynamin-related protein Drp1. Ann. NY Acad. Sci. 1201, 34–39 (2010).
Chang, C. R. & Blackstone, C. Cyclic AMP-dependent protein kinase phosphorylation of Drp1 regulates its GTPase activity and mitochondrial morphology. J. Biol. Chem. 282, 21583–21587 (2007).
Cribbs, J. T. & Strack, S. Reversible phosphorylation of Drp1 by cyclic AMP-dependent protein kinase and calcineurin regulates mitochondrial fission and cell death. EMBO Rep. 8, 939–944 (2007).
Cereghetti, G. M. et al. Dephosphorylation by calcineurin regulates translocation of Drp1 to mitochondria. Proc. Natl Acad. Sci. USA 105, 15803–15808 (2008
Chappie, J. S. et al. A pseudoatomic model of the dynamin polymer identifies a hydrolysis-dependent powerstroke. Cell 147, 209–222 (2011).
Ugarte-Uribe, B., Prevost, C., Das, K. K., Bassereau, P. & Garcia-Saez, A. J. Drp1 polymerization stabilizes curved tubular membranes similar to those of constricted mitochondria. J. Cell Sci. 132, jcs208603 (2019).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Kalia, R., Talledge, N. & Frost, A. Structural and functional studies of membrane remodeling machines. Methods Cell Biol. 128, 165–200 (2015).
Blommel, P. G., Becker, K. J., Duvnjak, P. & Fox, B. G. Enhanced bacterial protein expression during auto-induction obtained by alteration of lac repressor dosage and medium composition. Biotechnol. Prog. 23, 585–598 (2007).
Studier, F. W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q., Palovcak, E., Armache, J.-P., Cheng, Y. & Agard, D. A. MotionCor2: Anisotropic correction of beam-induced motion for improved single- particle electron cryo-microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Bell, J. M., Chen, M., Baldwin, P. R. & Ludtke, S. J. High resolution single particle refinement in EMAN2.1. Methods 100, 25–34 (2016).
Scheres, S. H. W. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015).
Egelman, E. H. Reconstruction of helical filaments and tubes. Methods Enzymol. 482, 167–183 (2010).
Ge, P. et al. Cryo-EM model of the bullet-shaped vesicular stomatitis virus. Science 327, 689–693 (2010).
Pintilie, G. D., Zhang, J., Goddard, T. D., Chiu, W. & Gossard, D. C. Quantitative analysis of cryo-EM density map segmentation by watershed and scale-space filtering, and fitting of structures by alignment to regions. J. Struct. Biol. 170, 427–438 (2010).
Goddard, T. D., Huang, C. C. & Ferrin, T. E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007).
Goddard, T. D., Huang, C. C. & Ferrin, T. E. Software extensions to UCSF chimera for interactive visualization of large molecular assemblies. Structure 13, 473–482 (2005).
Meng, E. C., Pettersen, E. F., Couch, G. S., Huang, C. C. & Ferrin, T. E. Tools for integrated sequence-structure analysis with UCSF Chimera. BMC Bioinformatics 7, 339 (2006).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Wang, R. Y. R. et al. Automated structure refinement of macromolecular assemblies from cryo-EM maps using Rosetta. eLife 5, 1–22 (2016).
Song, Y. et al. High-resolution comparative modeling with RosettaCM. Structure 21, 1735–1742 (2013).
DiMaio, F., Leaver-Fay, A., Bradley, P., Baker, D. & André, I. Modeling symmetric macromolecular structures in Rosetta3. PLoS ONE 6, e20450 (2011).
DiMaio, F. et al. Atomic-accuracy models from 4.5-Å cryo-electron microscopy data with density-guided iterative local refinement. Nat. Methods 12, 361–365 (2015).
Jakobi, A. J., Wilmanns, M. & Sachse, C. Model-based local density sharpening of cryo-EM maps. eLife 6, 1–26 (2017).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Acknowledgements
We thank M. Braunfeld, C. Kennedy, D. Bulkley, and A. Myasnikov and the University of California San Francisco (UCSF) Center for Advanced Cryo-EM, which is supported in part from National Institutes of Health (NIH) grants S10OD020054 and 1S10OD021741 and the Howard Hughes Medical Institute (HHMI). We thank the QB3 shared cluster and NIH grant 1S10OD021596-01, J.-P. Armache, N. Talledge for microscopy advice, C. Greenberg for consulting on structural modelling, D. Winge for discussions and M. Gu for facilitating mass spectrometry of proteins. This work was further supported by a Faculty Scholar grant from the HHMI (A.F.), the Searle Scholars Program (A.F.), NIH grant 1DP2GM110772-01 (A.F.), NIH grants GM53466 and GM84970 (J.M.S.), the Sandler Family Foundation through the UCSF Program for Breakthrough Biomedical Research and the American Asthma Foundation, and the HHMI (R.Y.-R.W., J.M.S. and D.A.A.). R.Y.-R.W. is an HHMI Fellow of the Life Sciences Research Foundation. A.F. is a Chan Zuckerberg Biohub investigator.
Reviewer information
Nature thanks M. Ryan, H. Zhou and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
R.K., J.M.S. and A.F. conceived the study. R.K., A.Y. and P.V.T. performed all experiments. R.K., R.Y.-R.W. and A.F. performed the computational analyses. D.A.A. advised R.Y.-R.W. and R.K. on model building. All authors evaluated the results and edited the manuscript. R.K. and A.F. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
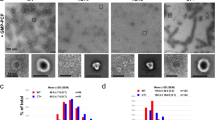
Extended Data Fig. 1 DRP1 and MID49 assembly states.
a–j, DRP1 assembly states visualized with negative-stain electron microscopy in the presence of different guanine nucleotides and MID49(126–454). Both proteins were incubated at concentrations of 2 μM. Scale bars, 100 nm. k–m, MID49(126–454) and MID51(132–463) form indistinguishable assemblies with DRP1: DRP1 + MID49(126–454) and GMPPCP (k), DRP1 + MID51(132–463) and GMPPCP (l), DRP1 + both equimolar MID49 and MID51 (m). Scale bars, 100 nm.
Extended Data Fig. 2 Cryo-EM and 3D reconstruction.
a, A cryo-EM micrograph of DRP1–MID49(126–454) filaments formed with GMPPCP. Scale bar, 100 nm. Inset, a representative 2D class average. Scale bar, 10 nm. b, Cross-section of the 3D reconstruction of the filament and the distribution of views determined during helical reconstruction. The length of the cylinders and the colour code correspond to the number of particles for that viewing direction (from few to many, blue to red). The 3D structure has been segmented and coloured with DRP1 in grey and MID49 in golden yellow. c, Particle numbers and workflow for the reconstruction protocol. DRP1 density is shown in grey and MID49 is in golden yellow.
Extended Data Fig. 3 Intra- and inter-filament interactions.
a, The triangular structure seen in cross section. Side 1 of the triangular structure has the atomic model placed in the density. The G-domain-to-G-domain contact between adjacent sides is circled. b, The sum of the three sides with the model fit in density. c, Ribbon diagram of the same atomic model as in b. The rotated view shows eight chains each of DRP1 and MID49. The chains further from the reader are rendered transparent. d, An isolated tetramer of DRP1 from the filament, rendered to highlight the stalk interfaces 1, 2 and 3 observed for DRP1 and other GTPases of the dynamin family. e, Expanded view of the circular region in a illustrating a salt bridge between adjacent G domains. f–i, Negative-stain micrographs of: DRP1-only wild-type polymers incubated with GMPPCP (f), DRP1 co-assembly with wild-type MID49(126–454) and GMPPCP (g; inset, high-magnification view), DRP1(E116R) mutant polymers (h), DRP1(E116R) mutant co-assembly with MiD49(126–454) (i). Shorter, single-sided filaments predominate. Disordered ‘triangular assemblies’ were also observed, but were much shorter and infrequent compared with the wild-type proteins. Scale bars, 50 nm.
Extended Data Fig. 4 Resolution estimates.
a, b, Local resolution estimates computed by Resmap74. Histogram of voxel values (a), and results in a depicted as a heat map of a cross-section through the reconstruction (b). c, d, Fourier shell correlation plots for the half-maps with and without symmetry (c) and model-to-map correlations for each sub-region of the structure (d).
Extended Data Fig. 5 Rosetta-based model refinement.
a, Complete assembly used for Rosetta-based model building with the asymmetric unit shown in green. b, Atomic B-factors for one asymmetric unit, DRP1 versus MID49 models (ribbon) and bound GMPPCP (space filling). c, Sequence alignment between human and mouse MID49 sequences. d, Overlay of the homology model of human MID49(126–454) (blue, with DRR in yellow, ribbon) modelled within the cryo-EM density overlaid with the mouse MID49 crystal structure (PDB: 4WOY, grey ribbon)37 which was used as a constraint. No density attributable to ADP within the nucleotidyltransferase domain was observed.
Extended Data Fig. 6 Map-to-model fits and role of the L1NS and L2S loops.
a–d, Examples of models fit within B-factor-sharpened cryo-EM density for a helix from the DRP1 stalk (a), the backbone of the L1NS loop from the stalk (b), an elongated helix and turn found in MID49 (c) and GMPPCP and Mg within the G domain (d). e, Roles of L1NS and L2S in linear filament formation. Top, an isolated Drp1 tetramer from the cryo-EM model. The circled region is expanded in the lower panel. Bottom, interactions that the conserved loop L1Ns makes within the assembly. G362 is highlighted with an arrow. f, DRP1 stalk and MID49 at receptor interface-3. g, Dyn3 stalk and pleckstrin homology domain from PDB 5A3F40. h, Overlay of f and g.
Extended Data Fig. 7 Sequence conservation and key interaction sites.
a, Multiple sequence alignment of the regions near and including the DRP1 residues mutated in this study: D190, D221 and G362, G363. The residue numbers apply to human DRP1, isoform 2 (UNIPROT identifier: O00429-3 which is also known as DLP1a). b, Sequence alignment of MID49 and MID51 at the region around residue R235 of MID49. R235 of MID49 corresponds to R243 of MID51. c, Sequence alignment of DRP1 isoform 1 and isoform 2 showing the correspondence of S637 (isoform 1) and S611 (isoform 2).
Extended Data Fig. 8 Biochemical and structural characterization of mutants.
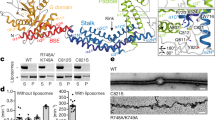
a–c, Size-exclusion chromatography traces for DRP1 wild-type and mutants used in the study. Comparison between wild-type (WT) and G362D (a), wild-type and D221A (b), and wild-type and D190A (c). d, Gel filtration standards with annotated molecular weights from the same column and chromatography system. e–h, DRP1 assembly and co-assembly reactions with GMPPCP for DRP1(D190A) alone (e), DRP1(D190A) + MID49(126–454) (f), DRP1(D221A) alone (g) and DRP1(D221A) + MID49(126–454) (h). Scale bars, 100 nm. i–l, DRP1(G362D) assembly and co-assembly reactions with GMPPCP or GTP. DRP1(G362D) forms rings but not linear filaments without MID49 (i, k), with MID49 (j, l), with GMPPCP (i, j) or with GTP (k, l). Scale bars, 100 nm.
Extended Data Fig. 9 DRP1(G362D) rings on model membranes.
a, DOPS liposomes used in the study. b, DOPS liposomes after incubation with DRP1(G362D) showing ring-like assemblies on the membrane and in the background. c Lipid nanotubes incubated with DRP1(G362D). d, Power spectrum of the area shown within the dashed square in b. e, Power spectrum of the area shown within the dashed square in c. In both d and e, layer lines indicative of helical geometry are not detectable. Scale bars, 50 nm.
Extended Data Fig. 10 DRP1(G362D) forms 12-dimer closed rings.
a, 2D class average of the rings. The red dashed circle indicates density that may be attributable to the variable domain. b, 2D class average of infrequent, orthogonal or side views used as a constraint during model building. c–f, Top (c) and side (d) projections of the model; top (e) and side (f) views of the final model rendered as ribbons. Scale bars, 100 Å. Green, G domain; red, BSE; purple, stalk.
Supplementary information
Supplementary Table 1
Cryo-EM data collection, refinement and validation statistics.
Video 1: Nucleotide-induced opening of hinge 2 between the G-domain and Bundle Signaling.
Element (BSE): Green: G-domain, Red: BSE. G2/Switch1 is indicated by an arrow and label. GTP/GMPPCP indicated as labeled. When the nucleotide binds, the G2/Switch1 loop of the G-domain closes like a lid over the nucleotide. This motion propagates through the associated beta sheet to push open hinge 2 of the BSE.
Video 2: Nucleotide-induced opening of hinges 1 and 2, receptor binding versus curling into rings.
Green: G-domain, Red: BSE, Blue: MID49, Yellow on MID49: Dynamin Recruitment Region (DRR), Orange on G-domains: Exposed receptor interfaces. As the G-domains bind a GTP molecule (not shown), an allosteric relay propagates through hinge 1 (between the BSE and the stalk) and hinge 2 (between the G-domain and the BSE) until the G-domain turns ~90degrees and moves ~40Å relative to the stalk. In this nucleotide-bound state, linear copolymers are stabilized by MID49/51 receptor binding. GTP hydrolysis and exchange returns the hinges to the apo conformation and promotes receptor dissociation. Upon nucleotide rebinding in the absence of MID49/51 receptors, the hinges can access a second GTP-dependent conformation that promotes curling by inducing a ~30 degree bending between adjacent interface-2 DRP1 dimers. This is the conformation that leads to closed ring formation.
Video 3: A hypothetical path-dependent allosteric curling reaction to constrict mitochondria.
DRP1: Green, MID49: Blue, white dots: GTP, pink dots: GDP. MID49 recruits tetramers of DRP1GTP, stabilizing a ~linear arrangement of DRP1 stalks and copolymerization of a filament on the outer mitochondrial membrane (OMM). The variable domain, which is rendered as a loop-like region, may also interact with the lipids of the OMM. GTP hydrolysis and exchange returns the hinges to the apo conformation and promotes receptor dissociation. Subsequent rounds of GTP hydrolysis and exchange progressively shorten the filament and enable DRP1’s hinges to access a second, GTP-dependent conformation that promotes curling by inducing a ~30 degree bend between adjacent interface-2 DRP1 dimers. This is the conformation that promotes curling into closed rings as the filaments shorten to an average length of ~12 dimers.
Rights and permissions
About this article
Cite this article
Kalia, R., Wang, R.YR., Yusuf, A. et al. Structural basis of mitochondrial receptor binding and constriction by DRP1. Nature 558, 401–405 (2018). https://doi.org/10.1038/s41586-018-0211-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0211-2
This article is cited by
-
Mitochondrial dynamics and colorectal cancer biology: mechanisms and potential targets
Cell Communication and Signaling (2024)
-
A partial Drp1 knockout improves autophagy flux independent of mitochondrial function
Molecular Neurodegeneration (2024)
-
Structural basis for regulated assembly of the mitochondrial fission GTPase Drp1
Nature Communications (2024)
-
Allosteric control of dynamin-related protein 1 through a disordered C-terminal Short Linear Motif
Nature Communications (2024)
-
Fatty acyl-coenzyme A activates mitochondrial division through oligomerization of MiD49 and MiD51
Nature Cell Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.