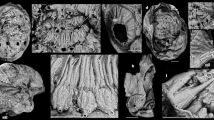
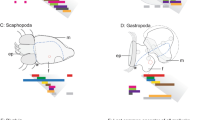
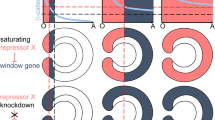
Abstract
The mouth opening of bilaterian animals develops either separate from (deuterostomy) or connected to (protostomy) the embryonic blastopore, the site of endomesoderm internalization. Although this distinction preluded the classification of bilaterian animals in Deuterostomia and Protostomia, and has influenced major scenarios of bilaterian evolution, the developmental basis for the appearance of these different embryonic patterns remains unclear. To identify the underlying mechanisms, we compared the development of two brachiopod species that show deuterostomy (Novocrania anomala) and protostomy (Terebratalia transversa), respectively. We show that the differential activity of Wnt signalling, together with the timing and location of mesoderm formation, correlate with the differential behaviour and fate of the blastopore. We further assess these principles in the spiral-cleaving group Annelida, and propose that the developmental relationships of mouth and blastoporal openings are secondary by-products of variations in axial and mesoderm development. This challenges the previous evolutionary emphasis on extant blastoporal behaviours to explain the origin and diversification of bilaterian animals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Gilbert, S. F. & Raunio, A. M. (eds) Embryology, Constructing the Organism (Sinauer Associates, 1997).
Grobben, K. Die systematische Einteilung des Tierreichs. Verh. Zool. Bot. Ges. Wien 58, 491–511 (1908).
Dunn, C. W., Giribet, G., Edgecombe, G. D. & Hejnol, A. Animal phylogeny and its evolutionary implications. Annu. Rev. Ecol. Evol. Syst. 45, 371–395 (2014).
Cannon, J. T. et al. Xenacoelomorpha is the sister group to Nephrozoa. Nature 530, 89–93 (2016).
Martín-Durán, J. M., Janssen, R., Wennberg, S., Budd, G. E. & Hejnol, A. Deuterostomic development in the protostome Priapulus caudatus . Curr. Biol. 22, 2161–2166 (2012).
Hejnol, A. & Martindale, M. Q. in Animal Evolution: Genes, Genomes, Fossils and Trees (eds Telford M. J. & Littlewood D. T. J. ) 33–40 (Oxford Univ. Press, 2009).
Sedgwick, A. On the origin of metameric segmentation and some other morphological questions. Q. J. Microsc. Sci. 24, 43–82 (1884).
Jägersten, G. On the early phylogeny of the Metazoa: the bilatero-gastrea theory. Zool. Bidr. Uppsala 30, 321–354 (1955).
Remane, A. Die entstehung der metamerie der wirbellosen. Zool. Anz. 14, 18–23 (1950).
Arendt, D. & Nübler-Jung, K. Dorsal or ventral: similarities in fate maps and gastrulation patterns in annelids, arthropods and chordates. Mech. Dev. 61, 7–21 (1997).
Steinmetz, P. R. H., Zelada-Gonzales, F., Burgtorf, C., Wittbrodt, J. & Arendt, D. Polychaete trunk neuroectoderm converges and extends by mediolateral cell intercalation. Proc. Natl Acad. Sci. USA 104, 2727–2732 (2007).
Arendt, D., Technau, U. & Wittbrodt, J. Evolution of the bilaterian larval foregut. Nature 409, 81–85 (2001).
Nielsen, C. Evolution of deuterostomy – and origin of the chordates. Biol. Rev. Camb. Philos. Soc. (2015).
Malakhov, V. V. New ideas on the origin of bilateral animals. Russ. J. Mar. Biol. 30, S22–S33 (2004).
von Graff, L. Die Organisation der Turbellaria Acoela (von Wilhelm Engelmann, 1891).
Hyman, L. H. The Invertebrates. Vol II: Platyhelminthes and Rhynchocoela. (McGraw-Hill, 1951).
Salvini-Plawen, L. On the origin and evolution of the lower Metazoa. Zeitschr. Zool. Syst. Evol.-Forsch. 16, 40–88 (1978).
Beklemishev, V. N. Principles of Comparative Anatomy of Invertebrates (Univ. Chicago Press, 1969).
Salvini-Plawen, L. v. Phylogenetischer status und bedeutung der mesenchymaten Bilateria. Zool. Jahrb. Anat. 103, 354–373 (1980).
Lankester, E. R. Notes on the embryology and classification of the animal kingdom: comprising a revision of speculations relative to the origin and significance of the germ-layers. Q. J. Microsc. Sci. s2-s17, 399–454 (1877).
Martindale, M. Q. & Hejnol, A. A developmental perspective: changes in the position of the blastopore during bilaterian evolution. Dev. Cell 17, 162–174 (2009).
Freeman, G. Regional specification during embryogenesis in the craniiform brachiopod Crania anomala. Dev. Biol. 227, 219–238 (2000).
Freeman, G. Regional specification during embryogenesis in the articulate brachiopod Terebratalia. Dev. Biol. 160, 196–213 (1993).
Nielsen, C. The development of the brachiopod Crania (Neocrania) anomala (O. F. Müller) and its phylogenetic significance. Acta Zool. 72, 7–28 (1991).
Freeman, G. Regional specification during embryogenesis in Rhynchonelliform brachiopods. Dev. Biol. 261, 268–287 (2003).
Long, J. A. & Stricker, S. A. in Reproduction of Marine Invertebrates (eds Giese, A. C., Pearse, J. S. & Pearse V. B. ) 47–84 (Boxwood Press, 1991).
Petersen, C. P. & Reddien, P. W. Wnt signaling and the polarity of the primary body axis. Cell 139, 1056–1068 (2009).
Kunick, C., Lauenroth, K., Leost, M., Meijer, L. & Lemcke, T. 1-Azakenpaullone is a selective inhibitor of glycogen synthase kinase-3 beta. Bioorg. Med. Chem. Lett. 14, 413–416 (2004).
Jho, E. H. et al. Wnt/beta-catenin/Tcf signaling induces the transcription of Axin2, a negative regulator of the signaling pathway. Mol. Cell. Biol. 22, 1172–1183 (2002).
De Robertis, E. M. Evo–devo: variations on ancestral themes. Cell 132, 185–195 (2008).
Hao, J. et al. In vivo structure–activity relationship study of dorsomorphin analogues identifies selective VEGF and BMP inhibitors. ACS Chem. Biol. 5, 245–253 (2010).
Fuentealba, L. C. et al. Integrating patterning signals: Wnt/GSK3 regulates the duration of the BMP/Smad1 signal. Cell 131, 980–993 (2007).
Hashiguchi, M. & Mullins, M. C. Anteroposterior and dorsoventral patterning are coordinated by an identical patterning clock. Development 140, 1970–1980 (2013).
Wei, Z., Range, R., Angerer, R. & Angerer, L. Axial patterning interactions in the sea urchin embryo: suppression of nodal by Wnt1 signaling. Development 139, 1662–1669 (2012).
Genikhovich, G. et al. Axis patterning by BMPs: cnidarian network reveals evolutionary constraints. Cell Rep. 10, 1646–1654 (2015).
Leclère, L. & Rentzsch, F. RGM regulates BMP-mediated secondary axis formation in the sea anemone Nematostella vectensis . Cell Rep. 9, 1921–1930 (2014).
Kraus, Y., Aman, A., Technau, U. & Genikhovich, G. Pre-bilaterian origin of the blastoporal axial organizer. Nat. Commun. 7, 11694 (2016).
Weigert, A. et al. Illuminating the base of the annelid tree using transcriptomics. Mol. Biol. Evol. 31, 1391–1401 (2014).
Seaver, E. C. Variation in spiralian development: insights from polychaetes. Int. J. Dev. Biol. 58, 457–467 (2014).
Smart, T. I. & Von Dassow, G. Unusual development of the mitraria larva in the polychaete Owenia collaris . Biol. Bull. 217, 253–268 (2009).
Wilson, D. P. On the mitraria larva of Owenia fusiformis Delle Chiaje. Phil. Trans. R. Soc. Lond. B 221, 231–334 (1932).
Eisig, H. Zur entwicklungsgeschichte der Capitelliden. Mitt. Aus Der Zool. Station Zu Neapel 13, 1–292 (1899).
Seaver, E. C., Yamaguchi, E., Richards, G. S. & Meyer, N. P. Expression of the pair-rule gene homologs runt, Pax3/7, even-skipped-1 and even-skipped-2 during larval and juvenile development of the polychaete annelid Capitella teleta does not support a role in segmentation. EvoDevo 3, 8 (2012).
Boyle, M. J., Yamaguchi, E. & Seaver, E. Molecular conservation of metazoan gut formation: evidence from expression of endomesoderm genes in Capitella teleta (Annelida). EvoDevo 5, 39 (2014).
Fröbius, A. C. & Seaver, E. C. ParaHox gene expression in the polychaete annelid Capitella sp. I. Dev. Genes Evol. 216, 81–88 (2006).
Amiel, A. R., Henry, J. Q. & Seaver, E. C. An organizing activity is required for head patterning and cell fate specification in the polychaete annelid Capitella teleta: new insights into cell-cell signaling in Lophotrochozoa. Dev. Biol. 379, 107–122 (2013).
Meyer, N. P., Boyle, M. J., Martindale, M. Q. & Seaver, E. C. A comprehensive fate map by intracellular injection of identified blastomeres in the marine polychaete Capitella teleta . EvoDevo 1, 8 (2010).
Kraus, Y., Fritzenwanker, J. H., Genikhovich, G. & Technau, U. The blastoporal organiser of a sea anemone. Curr. Biol. 17, R874–R876 (2007).
Gould, S. J. & Lewontin, R. C. The spandrels of San Marco and the Panglossian paradigm: a critique of the adaptationist programme. Proc. R. Soc. Lond. B 205, 581–598 (1979).
Christiaen, L. et al. Evolutionary modification of mouth position in deuterostomes. Semin. Cell Dev. Biol. 18, 502–511 (2007).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Hejnol, A. & Martindale, M. Q. Acoel development indicates the independent evolution of the bilaterian mouth and anus. Nature 456, 382–386 (2008).
Hejnol, A. & Schnabel, R. What a couple of dimensions can do for you: Comparative developmental studies using 4D microscopy—examples from tardigrade development. Integr. Comp. Biol. 46, 151–161 (2006).
Acknowledgements
We thank H. Hausen and O. Voecking for sharing the RNAseq data of O. fusiformis and expertise with the spawnings, B. C. Vellutini for help with collections and drug treatments, and G. S. Richards, F. Rentzsch, M. Iglesias and the members of the Hejnol laboratory for their comments on the manuscript. We also thank the staff at Friday Harbor Laboratories, Espeland Marine Biological Station and Station Biologique de Roscoff for assistance with animal collections. The study was funded by the core budget of the Sars Centre and supported by The European Research Council Community’s Framework Program Horizon 2020 (2014–2020) ERC grant agreement 648861 and an L. Meltzers Høyskolefond grant to A.H. J.M.M.-D. was supported by Marie Curie fellowship IEF 329024.
Author information
Authors and Affiliations
Contributions
J.M.M.-D. and A.H. conceived the project. J.M.M.-D., Y.J.P. and A.H. performed animal collections and cloned genes, J.M.M.-D conducted the experiments, and J.M.M.-D. and A.H. performed the four-dimensional recordings. Y.J.P. carried out the EdU analysis in T. transversa. J.M.M.-D. and A.H. analysed the data and wrote the manuscript, and Y.J.P. and M.Q.M. edited the paper. All authors discussed and commented on the data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary figures 1–12; Supplementary Tables 1–9; Supplementary References and legends for Supplementary Videos (PDF 4910 kb)
Supplementary Animation
Animated summary of the main findings of the manuscript regarding brachiopod gastrulation. (HTML 204 kb)
Supplementary Video 1
Description: Time-lapse recording of an embryo of N. anomala from the 5 early blastula stage to gastrulation, viewed from the animal hemisphere. (MP4 2036 kb)
Supplementary Video 2
Time-lapse recording of N. anomala from the early gastrula stage to the onset of axial elongation (shift of the blastopore to a ventral-posterior position), viewed from the vegetal pole. (MP4 12442 kb)
Supplementary Video 3
Time-lapse recording of N. anomala during early axial elongation, viewed from the ventral side. (MP4 2468 kb)
Supplementary Video 4
Time-lapse recording of N. anomala during axial elongation and blastopore closure, viewed from the ventral side. (MP4 10529 kb)
Supplementary Video 5
Time-lapse recording of N. anomala during late axial elongation and early larva differentiation (apical lobe-mantle lobe boundary formation; closure of the blastopore), viewed from the ventral side. (MP4 3006 kb)
Rights and permissions
About this article
Cite this article
Martín-Durán, J., Passamaneck, Y., Martindale, M. et al. The developmental basis for the recurrent evolution of deuterostomy and protostomy. Nat Ecol Evol 1, 0005 (2017). https://doi.org/10.1038/s41559-016-0005
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41559-016-0005
This article is cited by
-
The development of the adult nervous system in the annelid Owenia fusiformis
Neural Development (2024)
-
Highly conserved and extremely evolvable: BMP signalling in secondary axis patterning of Cnidaria and Bilateria
Development Genes and Evolution (2024)
-
Annelid functional genomics reveal the origins of bilaterian life cycles
Nature (2023)
-
Brachiopod and mollusc biomineralisation is a conserved process that was lost in the phoronid–bryozoan stem lineage
EvoDevo (2022)
-
ERK1/2 is an ancestral organising signal in spiral cleavage
Nature Communications (2022)