Abstract
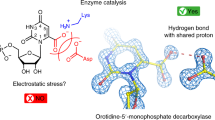
The UbiD enzyme plays an important role in bacterial ubiquinone (coenzyme Q) biosynthesis. It belongs to a family of reversible decarboxylases that interconvert propenoic or aromatic acids with the corresponding alkenes or aromatic compounds using a prenylated flavin mononucleotide cofactor. This cofactor is suggested to support (de)carboxylation through a reversible 1,3-dipolar cycloaddition process. Here, we report an atomic-level description of the reaction of the UbiD-related ferulic acid decarboxylase with substituted propenoic and propiolic acids (data ranging from 1.01–1.39 Å). The enzyme is only able to couple (de)carboxylation of cinnamic acid-type compounds to reversible 1,3-dipolar cycloaddition, while the formation of dead-end prenylated flavin mononucleotide cycloadducts occurs with distinct propenoic and propiolic acids. The active site imposes considerable strain on covalent intermediates formed with cinnamic and phenylpropiolic acids. Strain reduction through mutagenesis negatively affects catalytic rates with cinnamic acid, indicating a direct link between enzyme-induced strain and catalysis that is supported by computational studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data generated and analysed in this study, including the computational modelling data associated with all of the figures, are available from the corresponding authors on reasonable request. The diffraction data and corresponding atomic models have been deposited in the PDB under accession codes 6R3G, 6R3F, 6R3I, 6R2Z, 6R30, 6R3O, 6R2T, 6R2R, 6R3N, 6R3L, 6R3J, 6R2P, 6R34, 6R33 and 6R32.
References
Toney, M. D. Controlling reaction specificity in pyridoxal phosphate enzymes. Biochim. Biophys. Acta Proteins Proteom. 1814, 1407–1418 (2011).
Kluger, R. & Tittmann, K. Thiamin diphosphate catalysis: enzymic and nonenzymic covalent intermediates. Chem. Rev. 108, 1797–1833 (2008).
Piano, V., Palfey, B. A. & Mattevi, A. Flavins as covalent catalysts: new mechanisms emerge. Trends Biochem. Sci. 42, 457–469 (2017).
Marshall, S. A., Payne, K. A. P. & Leys, D. The UbiX-UbiD system: the biosynthesis and use of prenylated flavin (prFMN). Arch. Biochem. Biophys. 632, 209–221 (2017).
Pellissier, H. Asymmetric 1,3-dipolar cycloadditions. Tetrahedron 63, 3235–3285 (2007).
Jeon, B.-S. et al. Investigation of the mechanism of the SpnF-catalyzed [4+2]-cycloaddition reaction in the biosynthesis of spinosyn A. Proc. Natl Acad. Sci. USA 114, 10408–10413 (2017).
Meldal, M. & Tornøe, C. W. Cu-catalyzed azide−alkyne cycloaddition. Chem. Rev. 108, 2952–3015 (2008).
Jacobs, M. J., Schneider, G. & Blank, K. G. Mechanical reversibility of strain-promoted azide–alkyne cycloaddition reactions. Angew. Chem. Int. Ed. 55, 2899–2902 (2016).
Khanal, A., Long, F., Cao, B., Shahbazian-Yassar, R. & Fang, S. Evidence of splitting 1,2,3-triazole into an alkyne and azide by low mechanical force in the presence of other covalent bonds. Chemistry 22, 9760–9767 (2016).
Krupička, M., Dopieralski, P. & Marx, D. Unclicking the click: metal-assisted mechanochemical cycloreversion of triazoles Is possible. Angew. Chem. Int. Ed. 56, 7745–7749 (2017).
Stauch, T. & Dreuw, A. Force-induced retro-click reaction of triazoles competes with adjacent single-bond rupture. Chem. Sci. 8, 5567–5575 (2017).
Payne, K. A. et al. New cofactor supports α,β-unsaturated acid decarboxylation via 1,3-dipolar cycloaddition. Nature 522, 497–501 (2015).
Ferguson, K. L., Eschweiler, J. D., Ruotolo, B. T. & Marsh, E. N. G. Evidence for a 1,3-dipolar cyclo-addition mechanism in the decarboxylation of phenylacrylic acids catalyzed by ferulic acid decarboxylase. J. Am. Chem. Soc. 139, 10972–10975 (2017).
Bailey, S. S. et al. The role of conserved residues in UbiD/Fdc decarboxylase in oxidative maturation, isomerisation and catalysis of prenylated flavin mononucleotide. J. Biol. Chem. 293, 2272–2287 (2018).
Borden, W. T. Pyramidalized alkenes. Chem. Rev. 89, 1095–1109 (1989).
Okrasa, K. et al. Structure and mechanism of an unusual malonate decarboxylase and related racemases. Chemistry 14, 6609–6613 (2008).
Jez, J. M., Ferrer, J. L., Bowman, M. E., Dixon, R. A. & Noel, J. P. Dissection of malonyl-coenzyme A decarboxylation from polyketide formation in the reaction mechanism of a plant polyketide synthase. Biochemistry 39, 890–902 (2000).
Blake, C. C. et al. Crystallographic studies of the activity of hen egg-white lysozyme. Proc. R. Soc. Lond. B Biol. Sci. 167, 378–388 (1967).
Vocadlo, D. J., Davies, G. J., Laine, R. & Withers, S. G. Catalysis by hen egg-white lysozyme proceeds via a covalent intermediate. Nature 412, 835–838 (2001).
Anderson, V. E. Quantifying energetic contributions to ground state destabilization. Arch. Biochem. Biophys. 433, 27–33 (2005).
Zhang, Y. & Schramm, V. L. Ground-state destabilization in orotate phosphoribosyltransferases by binding isotope effects. Biochemistry 50, 4813–4818 (2011).
Lehwess-Litzmann, A. et al. Twisted Schiff base intermediates and substrate locale revise transaldolase mechanism. Nat. Chem. Biol. 7, 678–684 (2011).
Milić, D. et al. Crystallographic snapshots of tyrosine phenol-lyase show that substrate strain plays a role in C–C bond cleavage. J. Am. Chem. Soc. 133, 16468–16476 (2011).
Vladimirova, A. et al. Substrate distortion and the catalytic reaction mechanism of 5-carboxyvanillate decarboxylase. J. Am. Chem. Soc. 138, 826–836 (2016).
Lüdtke, S. et al. Sub-ångström-resolution crystallography reveals physical distortions that enhance reactivity of a covalent enzymatic intermediate. Nat. Chem. 5, 762–767 (2013).
Albery, W. J. & Knowles, J. R. Efficiency and evolution of enzyme catalysis. Angew. Chem. Int. Ed. 16, 285–293 (1977).
Abu Laban, N., Selesi, D., Rattei, T., Tischler, P. & Meckenstock, R. U. Identification of enzymes involved in anaerobic benzene degradation by a strictly anaerobic iron-reducing enrichment culture. Environ. Microbiol. 12, 2783–2796 (2010).
Luo, F. et al. Metatranscriptome of an anaerobic benzene-degrading, nitrate-reducing enrichment culture reveals involvement of carboxylation in benzene ring activation. Appl. Environ. Microbiol. 80, 4095–4107 (2014).
Baunach, M. & Hertweck, C. Natural 1,3-dipolar cycloadditions. Angew. Chem. Int. Ed. 54, 12550–12552 (2015).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Grimme, S., Ehrlich, S. & Goerigk, L. Effect of the damping function in dispersion corrected density functional theory. J. Comp. Chem. 32, 1456–1465 (2011).
Acknowledgements
This work was supported by the grants BBSRC BB/K017802 and ERC pre-FAB 695013. We acknowledge assistance via use of the Manchester Protein Structure Facility and Diamond Light Source for access (proposal numbers MX12788 and MX17773), which contributed to the results presented here. We also acknowledge the assistance given by IT Services and the use of the Computational Shared Facility at The University of Manchester. D.L. is a Royal Society Wolfson Merit Award holder.
Author information
Authors and Affiliations
Contributions
S.S.B. cloned, expressed and purified AnFdc1 and various variants. S.S.B. determined all crystal structures, with guidance from D.L. K.A.P.P. generated various AnFdc1 variants and collected solution data with cinnamic acid. A.S. assisted with crystallization of the Int3crotonic state. S.A.M. assisted with AnFdc1 reconstitution. I.G. assisted with crystallization of FMN-substituted AnFdc1 and the Int3butynoic state (with help from I.K.). K.F. assisted with protein purification and solution data collection and analysis. S.H. performed all of the computational studies. All authors discussed the results and participated in writing the manuscript. D.L. initiated and directed the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Supplementary Table 1.
Calculations archive file
Density functional theory coordinates used in this study.
Rights and permissions
About this article
Cite this article
Bailey, S.S., Payne, K.A.P., Saaret, A. et al. Enzymatic control of cycloadduct conformation ensures reversible 1,3-dipolar cycloaddition in a prFMN-dependent decarboxylase. Nat. Chem. 11, 1049–1057 (2019). https://doi.org/10.1038/s41557-019-0324-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-019-0324-8
This article is cited by
-
UbiD domain dynamics underpins aromatic decarboxylation
Nature Communications (2021)
-
Direct 1,3-butadiene biosynthesis in Escherichia coli via a tailored ferulic acid decarboxylase mutant
Nature Communications (2021)
-
Directed evolution of prenylated FMN-dependent Fdc supports efficient in vivo isobutene production
Nature Communications (2021)
-
Structural basis for antibiotic action of the B1 antivitamin 2′-methoxy-thiamine
Nature Chemical Biology (2020)
-
Enzymatic C–H activation of aromatic compounds through CO2 fixation
Nature Chemical Biology (2020)