Abstract
The first lineage specification of pluripotent mouse epiblast segregates neuroectoderm (NE) from mesoderm and definitive endoderm (ME) by mechanisms that are not well understood. Here we demonstrate that the induction of ME gene programs critically relies on the T-box transcription factors Eomesodermin (also known as Eomes) and Brachyury, which concomitantly repress pluripotency and NE gene programs. Cells deficient in these T-box transcription factors retain pluripotency and differentiate to NE lineages despite the presence of ME-inducing signals transforming growth factor β (TGF-β)/Nodal and Wnt. Pluripotency and NE gene networks are additionally repressed by ME factors downstream of T-box factor induction, demonstrating a redundancy in program regulation to safeguard mutually exclusive lineage specification. Analyses of chromatin revealed that accessibility of ME enhancers depends on T-box factor binding, whereas NE enhancers are accessible and already activation primed at pluripotency. This asymmetry of the chromatin landscape thus explains the default differentiation of pluripotent cells to NE in the absence of ME induction that depends on activating and repressive functions of Eomes and Brachyury.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
RNA-seq, ATAC-seq, and ChIP–seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE128466. Previously published sequencing data that were re-analysed here are available under accession numbers GSE99494 (RNA-seq data of mouse EpiSC32), GSE94250 (ATAC-seq of mESCs35) and GSE110164 (ATAC-seq of mouse EpiSCs36). Source data for Figs. 1, 2 and 6 and Extended Data Figs. 1, 2, 4, 6 and 7 are provided with the paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
All customized computational code used in this study is available from https://github.com/gwangjinkim/eomes-brachyury-rnaseq.
Change history
16 December 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Arnold, S. J. & Robertson, E. J. Making a commitment: cell lineage allocation and axis patterning in the early mouse embryo. Nat. Rev. Mol. Cell Biol. 10, 91–103 (2009).
Tesar, P. J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448, 196–199 (2007).
Brons, I. G. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007).
Nichols, J. & Smith, A. Naive and primed pluripotent states. Cell Stem Cell 4, 487–492 (2009).
Tam, P. P. L. & Loebel, D. A. F. Gene function in mouse embryogenesis: get set for gastrulation. Nature Rev. Genet. 8, 368–381 (2007).
Russ, A. P. et al. Eomesodermin is required for mouse trophoblast development and mesoderm formation. Nature 404, 95–99 (2000).
Arnold, S. J., Hofmann, U. K., Bikoff, E. K. & Robertson, E. J. Pivotal roles for eomesodermin during axis formation, epithelium-to-mesenchyme transition and endoderm specification in the mouse. Development 135, 501–511 (2008).
Costello, I. et al. The T-box transcription factor Eomesodermin acts upstream of Mesp1 to specify cardiac mesoderm during mouse gastrulation. Nat. Cell Biol. 13, 1084–1091 (2011).
Probst, S. & Arnold, S. J. Eomesodermin—at dawn of cell fate decisions during early embryogenesis. Curr. Top. Dev. Biol. 122, 93–115 (2017).
Herrmann, B. G., Labeit, S., Poustka, A., King, T. R. & Lehrach, H. Cloning of the T gene required in mesoderm formation in the mouse. Nature 343, 617–622 (1990).
Nelson, A. C. et al. Global identification of Smad2 and Eomesodermin targets in zebrafish identifies a conserved transcriptional network in mesendoderm and a novel role for Eomesodermin in repression of ectodermal gene expression. BMC biology 12, 1 (2014).
Evans, A. L. et al. Genomic targets of Brachyury (T) in differentiating mouse embryonic stem cells. PLoS ONE 7, e33346 (2012).
Faial, T. et al. Brachyury and SMAD signalling collaboratively orchestrate distinct mesoderm and endoderm gene regulatory networks in differentiating human embryonic stem cells. Development 142, 2121–2135 (2015).
Gentsch, G. E. et al. In vivo T-box transcription factor profiling reveals joint regulation of embryonic neuromesodermal bipotency. Cell Rep. 4, 1185–1196 (2013).
Teo, A. K. et al. Pluripotency factors regulate definitive endoderm specification through eomesodermin. Genes Dev. 25, 238–250 (2011).
Muñoz-Sanjuán, I. & Brivanlou, A. H. Neural induction, the default model and embryonic stem cells. Nat. Rev. Neurosci. 3, 271–280 (2002).
Hemmati-Brivanlou, A. & Melton, D. A. Inhibition of activin receptor signaling promotes neuralization in Xenopus. Cell 77, 273–281 (1994).
Tropepe, V. et al. Direct neural fate specification from embryonic stem cells: a primitive mammalian neural stem cell stage acquired through a default mechanism. Neuron 30, 65–78 (2001).
Perea-Gomez, A. et al. Nodal antagonists in the anterior visceral endoderm prevent the formation of multiple primitive streaks. Dev. Cell 3, 745–756 (2002).
Yamamoto, M. et al. Nodal antagonists regulate formation of the anteroposterior axis of the mouse embryo. Nature 428, 387–392 (2004).
Beddington, R. S. & Robertson, E. J. Axis development and early asymmetry in mammals. Cell 96, 195–209 (1999).
Iwafuchi-Doi, M. et al. Transcriptional regulatory networks in epiblast cells and during anterior neural plate development as modeled in epiblast stem cells. Development 139, 3926–3937 (2012).
Tang, K., Peng, G., Qiao, Y., Song, L. & Jing, N. Intrinsic regulations in neural fate commitment. Dev. Growth Differ. 57, 109–120 (2015).
Zhu, Q. et al. The transcription factor Pou3f1 promotes neural fate commitment via activation of neural lineage genes and inhibition of external signaling pathways. eLife 3, e02224 (2014).
Kubo, A. et al. Development of definitive endoderm from embryonic stem cells in culture. Development 131, 1651–1662 (2004).
Brennan, J. et al. Nodal signalling in the epiblast patterns the early mouse embryo. Nature 411, 965–969 (2001).
Arnold, S. J. et al. Brachyury is a target gene of the Wnt/β-catenin signaling pathway. Mech Dev 91, 249–258 (2000).
Yamaguchi, T. P., Takada, S., Yoshikawa, Y., Wu, N. & McMahon, A. P. T. T (Brachyury) is a direct target of Wnt3a during paraxial mesoderm specification. Genes Dev. 13, 3185–3190 (1999).
Wittler, L. et al. Expression of Msgn1 in the presomitic mesoderm is controlled by synergism of WNT signalling and Tbx6. EMBO Rep. 8, 784–789 (2007).
Nowotschin, S. et al. The T-box transcription factor Eomesodermin is essential for AVE induction in the mouse embryo. Genes Dev. 27, 997–1002 (2013).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Bao, S. et al. Derivation of hypermethylated pluripotent embryonic stem cells with high potency. Cell Res. 28, 22–34 (2018).
Pelton, T. A., Sharma, S., Schulz, T. C., Rathjen, J. & Rathjen, P. D. Transient pluripotent cell populations during primitive ectoderm formation: correlation of in vivo and in vitro pluripotent cell development. J. Cell Sci. 115, 329–339 (2002).
Fernando, R. I. et al. The T-box transcription factor Brachyury promotes epithelial–mesenchymal transition in human tumor cells. J. Clin. Invest. 120, 533–544 (2010).
Simon, C. S. et al. Functional characterisation of cis-regulatory elements governing dynamic Eomes expression in the early mouse embryo. Development 144, 1249–1260 (2017).
Senft, A. D. et al. Combinatorial Smad2/3 activities downstream of Nodal signaling maintain embryonic/extra-embryonic cell identities during lineage priming. Cell Rep. 24, 1977–1985 (2018).
Sadowski, I., Ma, J., Triezenberg, S. & Ptashne, M. GAL4–VP16 is an unusually potent transcriptional activator. Nature 335, 563–564 (1988).
Vickers, E. R. & Sharrocks, A. D. The use of inducible engrailed fusion proteins to study the cellular functions of eukaryotic transcription factors. Methods 26, 270–280 (2002).
Oliver, G. et al. Homeobox genes and connective tissue patterning. Development 121, 693–705 (1995).
Rivera-Pérez, J. A. & Magnuson, T. Primitive streak formation in mice is preceded by localized activation of Brachyury and Wnt3. Dev. Biol. 288, 363–371 (2005).
Izumi, N., Era, T., Akimaru, H., Yasunaga, M. & Nishikawa, S. Dissecting the molecular hierarchy for mesendoderm differentiation through a combination of embryonic stem cell culture and RNA interference. Stem Cells 25, 1664–1674 (2007).
Metzis, V. et al. Nervous system regionalization entails axial allocation before neural differentiation. Cell 175, 1105–1118 (2018).
Han, D. W. et al. Epiblast stem cell subpopulations represent mouse embryos of distinct pregastrulation stages. Cell 143, 617–627 (2010).
Lescroart, F. et al. Defining the earliest step of cardiovascular lineage segregation by single-cell RNA-seq. Science 359, 1177–1181 (2018).
Henrique, D., Abranches, E., Verrier, L. & Storey, K. G. Neuromesodermal progenitors and the making of the spinal cord. Development 142, 2864–2875 (2015).
Koch, F. et al. Antagonistic activities of Sox2 and Brachyury control the fate choice of neuro-mesodermal progenitors. Dev. Cell 42, 514–526 (2017).
Gouti, M. et al. A gene regulatory network balances neural and mesoderm specification during vertebrate trunk development. Dev. Cell 41, 243–261 (2017).
Chapman, D. L. & Papaioannou, V. E. Three neural tubes in mouse embryos with mutations in the T-box gene Tbx6. Nature 391, 695–697 (1998).
Takemoto, T. et al. Tbx6-dependent Sox2 regulation determines neural or mesodermal fate in axial stem cells. Nature 470, 394–398 (2011).
Arnold, S. J. et al. The T-box transcription factor Eomes/Tbr2 regulates neurogenesis in the cortical subventricular zone. Genes Dev. 22, 2479–2484 (2008).
Mihalas, A. B. et al. Intermediate progenitor cohorts differentially generate cortical layers and require Tbr2 for timely acquisition of neuronal subtype identity. Cell Rep. 16, 92–105 (2016).
Miller, S. A., Huang, A. C., Miazgowicz, M. M., Brassil, M. M. & Weinmann, A. S. Coordinated but physically separable interaction with H3K27-demethylase and H3K4-methyltransferase activities are required for T-box protein-mediated activation of developmental gene expression. Genes Dev. 22, 2980–2993 (2008).
Beisaw, A. et al. BRACHYURY directs histone acetylation to target loci during mesoderm development. EMBO Rep. 19, 118–134 (2018).
Guo, X. et al. A Linc1405/Eomes complex promotes cardiac mesoderm specification and cardiogenesis. Cell Stem Cell 22, 893–908 (2018).
Kartikasari, A. E. et al. The histone demethylase Jmjd3 sequentially associates with the transcription factors Tbx3 and Eomes to drive endoderm differentiation. EMBO J. 32, 1393–1408 (2013).
Schlesinger, S. & Meshorer, E. Open chromatin, epigenetic plasticity, and nuclear organization in pluripotency. Dev. Cell 48, 135–150 (2019).
Argelaguet, R. et al. Single cell multi-omics profiling reveals a hierarchical epigenetic landscape during mammalian germ layer specification. Preprint at bioRxiv https://doi.org/10.1101/519207 (2019).
Iwafuchi-Doi, M. & Zaret, K. S. Cell fate control by pioneer transcription factors. Development 143, 1833–1837 (2016).
Zhu, F. et al. The interaction landscape between transcription factors and the nucleosome. Nature 562, 76–81 (2018).
Zylicz, J. J. et al. G9a regulates temporal preimplantation developmental program and lineage segregation in blastocyst. 7, e33361 (2018).
Waldron, L. et al. The cardiac TBX5 interactome reveals a chromatin remodeling network essential for cardiac septation. Dev. Cell 36, 262–275 (2016).
Iacovino, M., Roth, M. E. & Kyba, M. Rapid genetic modification of mouse embryonic stem cells by Inducible Cassette Exchange recombination. Methods Mol. Biol. 1101, 339–351 (2014).
Beddington, R. S., Rashbass, P. & Wilson, V. Brachyury—a gene affecting mouse gastrulation and early organogenesis. Dev. Suppl. 1992, 157–165 (1992).
Cambray, S. et al. Activin induces cortical interneuron identity and differentiation in embryonic stem cell-derived telencephalic neural precursors. Nat. Commun. 3, 841 (2012).
Pierreux, C. E., Nicolas, F. J. & Hill, C. S. Transforming growth factor beta-independent shuttling of Smad4 between the cytoplasm and nucleus. Mol. Cell Biol. 20, 9041–9054 (2000).
Willert, K. et al. Wnt proteins are lipid-modified and can act as stem cell growth factors. Nature 423, 448–452 (2003).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Singh, A. A. et al. Optimized ChIP–seq method facilitates transcription factor profiling in human tumors. Life Sci. Alliance 2, e201800115 (2019).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108–e108 (2013).
Love, M. I., Huber, W. & Anders, S. et al. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res. 44, W3–W10 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Metsalu, T. & Vilo, J. ClustVis: a web tool for visualizing clustering of multivariate data using Principal Component Analysis and heatmap. Nucleic Acids Res. 43, W566–W570 (2015).
Zhang, Y. et al. Model-based analysis of ChIP–Seq (MACS). Genome Biol. 9, R137 (2008).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Yu, G., Wang, L.-G. & He, Q.-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Acknowledgements
We thank T. Bass and C. Domisch for technical assistance; B. G. Herrmann for providing T2J mice and M. Kyba for A2lox-Cre ES cells; the Freiburg Galaxy Team, especially B. Grüning and computing support by the state of Baden-Württemberg through bwHPC, University of Tübingen; C. Schwan for imaging support; S. Prekovic for valuable advice concerning ChIP; S. Nothjunge and R. Gilsbach for technical advice; M. Timmers for advising on GFP-fusion constructs; C. Hill and T. Gaarenstroom for the pGL2-6xARE-Lux reporter; G. Schmidt and S. Kowarschick for supporting luciferase assays; D. Onichtchouk, M.A. Morgan and P. Walentek for critical reading and discussion; the staff of the Life Imaging Centre (Albert-Ludwigs-University Freiburg) for help with confocal microscopy and the Genomics Core Facility at EMBL (Heidelberg) for sequencing. This study was supported by the German Research Foundation (DFG) through the Emmy Noether and Heisenberg Programs (AR 732/1-1/2/3 and AR 732/3-1), project grant (AR 732/2-1), project B07 of SFB 1140 (project ID 246781735), project A03 of SFB 850 (project ID 89986987), Germany’s Excellence Strategy (CIBSS–EXC-2189–Project ID 390939984) to S.J.A., TRR 152 (project ID 239283807) project P03, Germany’s Excellence Strategy (CIBSS–EXC-2189–Project ID 390939984) to M.K., and project B03 of SFB 992 (project ID 192904750) to L.H.
Author information
Authors and Affiliations
Contributions
J.T., M.P., C.M.S., S.-L.M., M.B., S.P. and S.J.A. performed experiments. J.T. and S.J.A. conceived and analysed experiments. J.T. and G.-J.K. performed bioinformatics data analysis. A.H. and M.K. designed TALENs. L.H. analysed data and edited the manuscript. J.T. and S.J.A. prepared figures and wrote and edited the manuscript with input from all authors. S.J.A. conceived the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterisation of EOMES and BRACHYURY dKO cells.
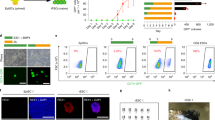
a, Schematic of EomesGfp/∆ (EoKO) and BrachyurytdTomato/∆ (BraKO) alleles. b, Relative mRNA expression of mesoderm and endoderm (ME) genes alongside with Eomes and Brachyury over 5 days of differentiation of WT cells. n=3 biological replicates. c, Expression levels of Eomes and Brachyury transcripts during 5 days of differentiation of WT, BraKO, EoKO and dKO cells measured by RT-qPCR. n=3 biological replicates. P-values for differences of mean expression between WT, EoKO, BraKO and dKO samples were calculated using two-tailed Student’s t-test. NS, not significant (P > 0.05), *:p≤0.05, **:p≤0.01, ***:p≤0.001. Exact P-values are provided in Supplementary Table 10. d, Representative immunofluorescence (IF) staining at day 4 of differentiation demonstrating the absence of EOMES and BRACHYURY in indicated loss-of-function cells of 3 biological replicates. e, EomesGfp and BratdTomato reporter activation in EomesGfp/+ and dKO EBs at day 4 of differentiation. Representative images of 10 biological replicates are shown. Maximum intensity projections of z-stacks. f, Protein levels of non-phosphorylated (active) β-CATENIN and total β-CATENIN in WT and dKO cells show responsiveness to WNT. β-ACTIN served as loading control. 2 independent experiments. g, Protein levels of phosphorylated SMAD2 and total SMAD2 in WT and dKO cells show responsiveness to ActA. ɑ-TUBULIN served as loading control. 2 independent experiments. h,i, Super 8x TOPflash and 6xARE luciferase reporter assays demonstrate responsiveness of WT and dKO cells to WNT and TGF-β/Nodal signals. n=3 biological replicates. P-values for differences of mean expression between treated and untreated samples were calculated using two-tailed Student’s t-test. *:p≤0.05, **:p≤0.01. Exact P-values are provided in Supplementary Table 10. j, Representative IF staining of EBs at day 4 and 7 of differentiation show the absence of endoderm (SOX17 and FOXA2) and mesoderm (FN1 and FOXC2) markers in dKO cells of 3 biological replicates. Scale bars in d,e, and j: 100 μm. Error bars represent s.e.m. in b,c,h, i.
Extended Data Fig. 2 Eomes- and Brachyury-deficient cells retain pluripotency and express neuroectoderm markers during differentiation.
a, Relative mRNA expression of pluripotency genes alongside with Eomes and Brachyury expression over 5 days of differentiation of WT cells. n=3 biological replicates. b, Representative immunofluorescence staining for NANOG, SOX2 and OCT4 of plated EBs at day 7 of differentiation show maintained expression of pluripotency markers in dKO cells, that are absent in WT, BraKO and EoKO cells. 3 biological replicates. c, Relative mRNA expression of neuroectoderm (NE) genes alongside with Eomes and Brachyury expression over 5 days of differentiation of WT cells. n=3 biological replicates. d, Co-immunofluorescence staining in dKO EBs at day 7 of differentiation show OCT4 and SOX1 co-expression in a small proportion of cells (arrowheads), few cells express only OCT4 (dashed line), and most cells express only SOX1. Yellow rectangles are shown at higher magnification to the right. 2 biological replicates in d and e. e, Co-immunofluorescence staining in dKO EBs at day 7 of differentiation show NANOG and SOX1 co-expression in few cells indicated by arrowheads. Most cells show only SOX1 staining (dashed line). Single NANOG positive cells are not detected. Yellow rectangles are shown at higher magnification to the right. Scale bars in b,d, and e: 100 μm. Error bars represent s.e.m. in a,c.
Extended Data Fig. 3 EOMES and BRACHYURY co-expression in the posterior epiblast and nascent mesoderm, and embryonic dKO phenotype.
a, Representative immunofluorescence staining of transverse sections of E6.5, E7.0 and E7.5 WT embryos showing EOMES and BRACHYURY co-expression in posterior-proximal epiblast and nascent mesoderm at E6.5. At E7.0 and E7.5 EOMES expression extends more anteriorly in the epiblast than BRACHYURY. Double positive cells are found in the epiblast and in nascent mesoderm. 3 biological replicates. b, mRNA in situ hybridization analysis of transversal sections of E7.5 embryos of indicated genotypes for Snai1 and Sox17 show the absence of both markers in the T2J/2J;EomesΔEpi embryos. Arrowheads indicate sites of staining for Snai1 and Sox17 expression. 2 biological replicates. Scale bars in a: 50 μm; in b: 100 μm.
Extended Data Fig. 4 Expression of Eomes- and Brachyury- GFP and FL constructs restores ME differentiation.
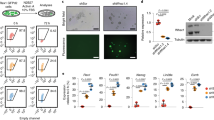
a, Schematic of Eo-GFP and Bra-GFP constructs for ChIP-seq. GFP was fused to the C-terminus of full-length Eomes or Brachyury and introduced into the doxycycline-inducible locus (TRE) of dKO mESCs. b,c, Protein levels of BRACHYURY (b) and EOMES (c) in WT cells compared to dKO+Bra-GFP and dKO+Eo-GFP cells after 4 and 5 days of differentiation. β-ACTIN served as loading control. 2 biological replicates. d, ME differentiation protocol of dKO+Eo-GFP and dKO+Bra-GFP cells by administration of doxycycline (dox) from days 3 -5 of differentiation. e, Principal component (PC) analysis of RNA-seq data of indicated cell lines at day 5 of differentiation. dKO+Eo-GFP and dKO+Eo-FL cells cluster more closely to BraKO cells, and dKO+Bra-GFP and dKO+Bra-FL cluster more closely to EoKO cells, indicating functional rescues by GFP fusion constructs. n=3 biological replicates. f-i, Expression levels indicated by centred scaled counts of mesoderm and endoderm (ME) and pluripotency and neuroectoderm (EPI and NE) genes that are rescued after induced expression of Eomes- (f, h) or Brachyury- (g, i) FL and GFP constructs. Bars represent centred scaled counts from n=3 biologically independent RNA-seq experiments. j, Representative immunofluorescence staining for α-ACTININ and SOX17 of plated EBs at day 8 of differentiation demonstrating cardiomyocyte and endoderm differentiation of WT and dKO+Eo-GFP cells, but not of dKO cells of 2 biological replicates. Scale bars 100 μm. Error bars indicate s.e.m. in f-i.
Extended Data Fig. 5 EOMES and BRACHYURY bind to regulatory regions of ME genes, and of EPI and NE genes, and regulate chromatin accessibility.
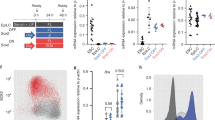
a, Overlap of genes associated to regions bound by EOMES or BRACHYURY shows that the majority of genes contain ChIP-seq peaks for both T-box factors. b, Genomic distribution of EOMES- and BRACHYURY-bound sites (ChIP-seq) showing predominant binding to regions in the proximity to gene bodies. c, RNA-seq, ATAC-seq and ChIP-seq coverage tracks of differentiated WT and dKO cells, ESCs35 and EpiSCs36 at loci of proposed T-box factors-activated target genes. ATAC peaks at regulatory sites of ME genes that are bound by EOMES and BRACHYURY are not present in pluripotent cells (ESCs and EpiSCs) and become established during differentiation to ME in a T-box-dependent manner. Counts normalized to RPKM are indicated in c and d. d, RNA-seq, ATAC-seq and ChIP-seq coverage tracks of T-box factors-repressed pluripotency (Nanog, Sox2, Lefty2) and NE (Sox1, Pou3f1, Neurog2) target genes show that regulatory sites are already accessible in pluripotent cells (ESCs and EpiSCs).
Extended Data Fig. 6 Eomes- and Brachyury-rescue constructs restore ME program activation and repression of pluripotency and NE programs.
a, ChIP-seq coverage tracks of Eo-GFP, Eo-FL, and Eo-VP16 for indicated genes show identical binding of GFP and VP16 fusion constructs to the full-length (FL) EOMES. Counts normalized to RPKM are indicated in a-c. b, RNA-seq coverage tracks of mesoderm (Mesp1, Msgn1, Mixl1, Myocd and Nkx2-5) and definitive endoderm (Foxa2 and Hhex) markers that are rescued by FL and VP16 activator constructs (Eo-FL, Bra-FL, Eo-VP16 and Bra-VP16) but not by repressor constructs (Eo-EnR and Bra-EnR) or Eo-TBX. c, RNA-seq coverage tracks of pluripotency (Pou5f1, Sox2, Lefty2 and Nkx1-2) and NE (Sox3, Brsk2 and Nefm) markers show selectivity of transcriptional repression by Eomes- or Brachyury- FL, VP16, and EnR constructs. Pluripotency genes are predominantly repressed by Eomes, and NE markers by Brachyury in a direct manner as demonstrated by EnR-mediated repression, and indirectly by VP16 activator-mediated mechanisms. Expression of Eo-TBX does not rescue the expression of EPI genes. d, e, Expression levels indicated by centred scaled counts of mesoderm (Mes) and definitive endoderm (DE) marker genes downregulated in dKO cells are rescued after induced expression of Eomes- (d) or Brachyury- (e) FL, VP16-activator, but not of EnR-repressor constructs. f, g, Expression levels indicated by scaled centred counts of EPI and NE marker genes after induced expression of Eomes- (f) or Brachyury- (g) rescue constructs show reduced expression by FL, VP16 activator, and EnR repressor constructs, but not by Eo-TBX. Bars represent centred scaled counts from n=3 biologically independent RNA-seq experiments in d-g. Error bars indicate s.e.m.
Extended Data Fig. 7 Expression of Eomes or Brachyury downstream targets Mesp1, Msgn1, Mixl1, Foxa2 and Six2 in dKO cells activates specific ME gene programs.
a, Expression levels of Mesp1, Msgn1, Mixl1 and Foxa2 in WT, dKO and dKO cells induced with Eomes- or Brachyury- FL, VP16 and EnR rescue constructs. Expression levels are shown as log10 RPKM from n=3 biologically independent RNA-seq experiments. b-f, Expression levels of ME genes after induced expression of Mesp1 (b), Msgn1 (c), Mixl1 (d), Foxa2 (e) or Six2 (f) in dKO cells. Bars represent centred scaled counts from n=3 biologically independent RNA-seq experiments. Error bars indicate s.e.m.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5 provide additional analysis of the ChIP-seq and ATAC-seq results shown in Figs. 4 and 5.
Supplementary Tables
Supplementary Table 1: Related to Figure 1 a, Clustered groups of scaled counts normalized by library size and gene-wise dispersion of downregulated genes (adjusted P-value≤0.05, log2(FC)≤-2.5) in dKO compared to WT cells at day 5 of differentiation analysed by RNA-seq. n=3 biological replicates. Statistical significance of differentially expressed genes was assessed using DEseq2 based on two-sided Wald test with Benjamini-Hochberg adjusted P-values. b, Enriched GO-terms of downregulated genes in dKO compared to WT cells. n=3 biological replicates. Statistical significance was assessed using standard hypergeometric test with FDR adjusted P-values. Supplementary Table 2: Related to Figure 2 a, Clustered groups of scaled counts normalized by library size and gene-wise dispersion of upregulated genes (adjusted P-value≤0.05, log2(FC)≥ 2.5) in dKO compared to WT cells at day 5 of differentiation analysed by RNA-seq. n=3 biological replicates. Statistical significance of differentially expressed genes was assessed using DEseq2 based on two-sided Wald test with Benjamini-Hochberg adjusted P-values. b, Enriched GO-terms of upregulated genes in dKO compared to WT cells. n=3 biological replicates. Statistical significance was assessed using standard hypergeometric test with FDR adjusted P-values. c, Clustered groups of scaled counts normalized by library size and gene-wise dispersion of upregulated genes (adjusted P-value≤0.05, log2(FC)≥ 2.5) with RNA-seq profiles of ESCs differentiated to NE (NE Ctrl) and epiblast stem cells (EpiSCs)37. n=3 biological replicates. Statistical significance of differentially expressed genes was assessed using DEseq2 based on two-sided Wald test with Benjamini-Hochberg adjusted P-values. Supplementary Table 3: Related to Figure 3 a, Clustered groups of scaled counts normalized by library size and gene-wise dispersion of differentially expressed genes (adjusted P-value≤0.05, log2(FC) of +/-1.0) of T2J/2J, Eomes∆Epi, and dKO (T2J/2J;Eomes∆Epi) embryos compared to WT assayed by RNA-seq at E7.5. n=3 biological replicates. Statistical significance of differentially expressed genes in was assessed using DEseq2 based on two-sided Wald test with Benjamini-Hochberg adjusted P-values. b, Enriched GO-terms of downregulated genes in dKO compared to WT embryos. n=3 biological replicates. Statistical significance was assessed using standard hypergeometric test with FDR adjusted P-values. c, Enriched GO-terms of upregulated genes in dKO compared to WT embryos. n=3 biological replicates. Statistical significance was assessed using standard hypergeometric test with FDR adjusted P-values. Supplementary Table 4: Related to Figure 4 and Extended data 4 a, List of genes associated to EOMES and BRACHYURY ChIP-seq peaks. n=2 biological replicates. b, List of genes bound by EOMES+BRACHYURY (ChIP-seq) that are downregulated in dKO cells compared to WT cells obtained by RNA-seq analysis at day 5 of differentiation. c, List of genes bound by EOMES+BRACHYRY (ChIP-seq) that are upregulated in dKO cells compared to WT cells obtained by RNA-seq analysis at day 5 of differentiation. Supplementary Table 5: Related to Figure 4 a, Transcription factor-binding motifs enriched within accessible chromatin in WT over dKO cells (ATAC-seq). n=2 biological replicates in a and b. b, Transcription factor-binding motifs enriched within ATAC peaks of dKO cells compared to WT cells. Supplementary Table 6: Related to Figure 5 a,b, List of genes downregulated in dKO cells that are rescued by (a) Eo-FL and Eo-VP16, and (b) Bra-FL and Bra-VP16 when analysed by RNA-seq. Adjusted P-value≤0.05, log2(FC) of +/-1.5 was used for rescue constructs in a-d. n=3 biological replicates in a-d. Statistical significance of differentially expressed genes in a-d was assessed using DEseq2 based on two-sided Wald test with Benjamini-Hochberg adjusted P-values. c, d, List of genes upregulated in dKO cells that are reduced upon expression of (c) Eo-FL and Eo-EnR, and (d) Bra-FL and Bra-EnR. Supplementary Table 7: Related to Figure 5 and Extended data 5 a, Centred scaled counts of differentially expressed genes between WT and dKO cells and after the induced expression of Eo-FL, Eo-VP16, Eo-EnR, and Eo-TBX constructs, with standard deviations and s.e.m. n=3 biological replicates in a and f. Statistical significance of differentially expressed genes in a and f was assessed using DEseq2 based on two-sided Wald test with Benjamini-Hochberg adjusted P-values. b, Centred scaled counts of downregulated genes in dKO cells, dKO-dox cells and after induction of Eo-FL, Eo-VP16, Eo-EnR, and Eo-TBX rescue constructs. c, Enriched GO-terms of genes listed in b. Statistical significance was assessed using standard hypergeometric test with FDR adjusted P-values in c,e,h, and j. d, Centred scaled counts of upregulated genes in dKO, dKO-dox cells and after induction of Eo-FL, Eo-VP16, Eo-EnR, and Eo-TBX rescue constructs. e, Enriched GO-terms of genes listed in c. f, Centred scaled counts of differentially expressed genes between WT and dKO cells and after the induced expression of Bra-FL, Bra-VP16, and Bra-EnR constructs, with standard deviations and s.e.m. g, Centred scaled counts of downregulated genes in dKO, dKO-dox cells and after induction of Bra-FL, Bra-VP16, and Bra-EnR rescue constructs. h, Enriched GO-terms of genes listed in g. i, Centred scaled counts of upregulated genes in dKO, dKO-dox cells and after induction of Bra-FL, Bra-VP16, and Bra-EnR rescue constructs. j, Enriched GO-terms analyses of genes listed in i. Supplementary Table 8: Related to Figure 6 a, Centred scaled counts of differentially expressed genes between WT and dKO cells and after the induced expression of Mesp1, Msgn1 and Six2 constructs, with standard deviations and s.e.m. n=3 biological replicates in a-c. Statistical significance of differentially expressed genes in a-c was assessed using DEseq2 based on two-sided Wald test with Benjamini-Hochberg adjusted P-values. b, Centred scaled counts of differentially expressed genes between WT and dKO cells and after the induced expression of Mixl1, Foxa2 and Six2 constructs, with standard deviations and s.e.m. c, Centred scaled counts of differentially expressed genes between WT and dKO cells and after the induced expression of Six2 control construct, with standard deviations and s.e.m. Supplementary Table 9: Key resources a, List of antibodies used for immunofluorescence staining, ChIP-seq, and Western blot experiments. b, List of oligos used for the RT-qPCR analysis. Supplementary Table 10: P-values a, List of exact P-values for indicated figures.
Supplementary Video
Supplementary Video 1: Related to Extended data 4 Beating cardiomyocytes obtained from dKO cells after induced expression of Eo-GFP construct, demonstrating cardiac mesoderm differentiation.
Source data
Source Data Fig. 1
Statistical source data for Figure 1
Source Data Fig. 2
Statistical source data for Figure 2
Source Data Fig. 6
Statistical source data for Figure 6
Source Data Extended Data Fig. 1
Statistical source data for Extended Data Fig. 1
Source Data Extended Data Fig. 1
Unprocessed western blots related to Extended Data Figs. 1f and 1g
Source Data Extended Data Fig. 2
Statistical source data for Extended Data Fig. 2
Source Data Extended Data Fig. 4
Statistical source data for Extended Data Fig. 4
Source Data Extended Data Fig. 4
Unprocessed western blots related to Extended Data Fig. 4b and 4c
Source Data Extended Data Fig. 6
Statistical source data for Extended Data Fig. 6
Source Data Extended Data Fig. 7
Statistical source data for Extended Data Fig. 7
Rights and permissions
About this article
Cite this article
Tosic, J., Kim, GJ., Pavlovic, M. et al. Eomes and Brachyury control pluripotency exit and germ-layer segregation by changing the chromatin state. Nat Cell Biol 21, 1518–1531 (2019). https://doi.org/10.1038/s41556-019-0423-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0423-1
This article is cited by
-
Single-cell transcriptome analyses reveal critical regulators of spermatogonial stem cell fate transitions
BMC Genomics (2024)
-
On the genetic basis of tail-loss evolution in humans and apes
Nature (2024)
-
DELVE: feature selection for preserving biological trajectories in single-cell data
Nature Communications (2024)
-
Comparison of Four Protocols for In Vitro Differentiation of Human Embryonic Stem Cells into Trophoblast Lineages by BMP4 and Dual Inhibition of Activin/Nodal and FGF2 Signaling
Reproductive Sciences (2024)
-
Statistical learning quantifies transposable element-mediated cis-regulation
Genome Biology (2023)