Abstract
Human liver cancers, including hepatocellular carcinomas and intra-hepatic cholangiocarcinomas, are often diagnosed late with poor prognosis. A better understanding of cancer initiation could provide potential preventive therapies and increase survival. Models for studying human liver cancer initiation are largely missing. Here, using directly reprogrammed human hepatocytes (hiHeps) and inactivation of p53 and RB, we established organoids possessing liver architecture and function. HiHep organoids were genetically engineered to model the initial alterations in human liver cancers. Bona fide hepatocellular carcinomas were developed by overexpressing c-Myc. Excessive mitochondrion–endoplasmic reticulum coupling induced by c-Myc facilitated hepatocellular carcinoma initiation and seemed to be a target of preventive treatment. Furthermore, through the analysis of human intra-hepatic cholangiocarcinoma-enriched mutations, we demonstrate that the RAS-induced lineage conversion from hepatocytes to intra-hepatic cholangiocarcinoma cells can be prevented by the combined inhibition of Notch and JAK–STAT. Together, hiHep organoids represent a system that can be genetically manipulated to model cancer initiation and identify potential preventive therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The RNA sequencing data that support the findings of this study have been deposited in the Gene Expression Omnibus under accession codes GSE115853. Previously published RNA sequencing data that were re-analysed here are available under the accession codes GSE103078, GSE84073, GSE86007 and GSE61948. The human HCC and ICC data were derived from the TCGA Research Network (http://cancergenome.nih.gov/). The dataset derived from this resource supporting the findings of this study is available in Supplementary Table 3. The source data for the GSEA and gene ontology analyses in Figs. 2,4,5,7,8 and Supplementary Figs. 1, 3–5,7,8 have been provided in Supplementary Tables 3 and 4. The source data for the other quantitive analyses in Figs. 2,3,5,6,8 and Supplementary Fig. 1–5, 7–8 have been provided in Supplementary Table 5 (Statistics Source Data). All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Bruix, J., Han, K. H., Gores, G., Llovet, J. M. & Mazzaferro, V. Liver cancer: approaching a personalized care. J. Hepatol. 62, S144–S156 (2015).
Cancer Genome Atlas Research Network. Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell 169, 1327–1341 (2017).
Chaisaingmongkol, J. et al. Common molecular subtypes among Asian hepatocellular carcinoma and cholangiocarcinoma. Cancer Cell 32, 57–70 (2017).
Schulze, K. et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat. Genet. 47, 505–511 (2015).
Nakamura, H. et al. Genomic spectra of biliary tract cancer. Nat. Genet. 47, 1003–1010 (2015).
Zheng, K., Cubero, F. J. & Nevzorova, Y. A. c-MYC—making liver sick: role of c-MYC in hepatic cell function, homeostasis and disease. Genes 8, E123 (2017).
Marquardt, J. U., Andersen, J. B. & Thorgeirsson, S. S. Functional and genetic deconstruction of the cellular origin in liver cancer. Nat. Rev. Cancer 15, 653–667 (2015).
Fan, B. et al. Cholangiocarcinomas can originate from hepatocytes in mice. J. Clin. Invest. 122, 2911–2915 (2012).
Sekiya, S. & Suzuki, A. Intrahepatic cholangiocarcinoma can arise from Notch-mediated conversion of hepatocytes. J. Clin. Invest. 122, 3914–3918 (2012).
Odom, D. T. et al. Tissue-specific transcriptional regulation has diverged significantly between human and mouse. Nat. Genet. 39, 730–732 (2007).
Yu, Y. et al. A comparative analysis of liver transcriptome suggests divergent liver function among human, mouse and rat. Genomics 96, 281–289 (2010).
Qiu, Z. et al. Hepatocellular carcinoma cell lines retain the genomic and transcriptomic landscapes of primary human cancers. Sci. Rep. 6, 27411 (2016).
Broutier, L. et al. Human primary liver cancer-derived organoid cultures for disease modeling and drug screening. Nat. Med. 23, 1424–1435 (2017).
Nuciforo, S. et al. Organoid models of human liver cancers derived from tumor needle biopsies. Cell Rep. 24, 1363–1376 (2018).
Matano, M. et al. Modeling colorectal cancer using CRISPR–Cas9-mediated engineering of human intestinal organoids. Nat. Med. 21, 256–262 (2015).
Li, X. et al. Oncogenic transformation of diverse gastrointestinal tissues in primary organoid culture. Nat. Med. 20, 769–777 (2014).
Huang, L. et al. Ductal pancreatic cancer modeling and drug screening using human pluripotent stem cell- and patient-derived tumor organoids. Nat. Med. 21, 1364–1371 (2015).
Crespo, M. et al. Colonic organoids derived from human induced pluripotent stem cells for modeling colorectal cancer and drug testing. Nat. Med. 23, 878–884 (2017).
Hu, H. L. et al. Long-term expansion of functional mouse and human hepatocytes as 3D organoids. Cell 175, 1591–1606 (2018).
Fu, G. B. et al. Expansion and differentiation of human hepatocyte-derived liver progenitor-like cells and their use for the study of hepatotropic pathogens. Cell Res. 29, 8–22 (2018).
Kim, Y. et al. Small molecule-mediated reprogramming of human hepatocytes into bipotent progenitor cells. J. Hepatol. 70, 97–107 (2019).
Zhang, K. et al. In vitro expansion of primary human hepatocytes with efficient liver repopulation capacity. Cell Stem Cell 23, 806–819 (2018).
Ji, S., Zhang, L. & Hui, L. Cell fate conversion: direct induction of hepatocyte-like cells from fibroblasts. J. Cell. Biochem. 114, 256–265 (2013).
Huang, P. Y. et al. Direct reprogramming of human fibroblasts to functional and expandable hepatocytes. Cell Stem Cell 14, 370–384 (2014).
Shi, X. L. et al. Improved survival of porcine acute liver failure by a bioartificial liver device implanted with induced human functional hepatocytes. Cell Res. 26, 206–216 (2016).
Shachaf, C. M. et al. MYC inactivation uncovers pluripotent differentiation and tumour dormancy in hepatocellular cancer. Nature 431, 1112–1117 (2004).
Dang, C. V. MYC on the path to cancer. Cell 149, 22–35 (2012).
Rowland, A. A. & Voeltz, G. K. Endoplasmic reticulum–mitochondria contacts: function of the junction. Nat. Rev. Mol. Cell Bio. 13, 607–615 (2012).
Sabharwal, S. S. & Schumacker, P. T. Mitochondrial ROS in cancer: initiators, amplifiers or an Achilles’ heel? Nat. Rev. Cancer 14, 709–721 (2014).
Cassidy-Stone, A. et al. Chemical inhibition of the mitochondrial division dynamin reveals its role in Bax/Bak-dependent mitochondrial outer membrane permeabilization. Dev. Cell 14, 193–204 (2008).
Kalia, R. et al. Structural basis of mitochondrial receptor binding and constriction by DRP1. Nature 558, 401–405 (2018).
Dolezal, J. M. et al. Sequential adaptive changes in a c-Myc-driven model of hepatocellular carcinoma. J. Biol. Chem. 292, 10068–10086 (2017).
Bridgewater, J. et al. Guidelines for the diagnosis and management of intrahepatic cholangiocarcinoma. J. Hepatol. 60, 1268–1289 (2014).
Park, S. Y. et al. Expression of MUC1, MUC2, MUC5AC and MUC6 in cholangiocarcinoma: prognostic impact. Oncol. Rep. 22, 649–657 (2009).
Reddy, K. P., Buschmann, R. J. & Chomet, B. Cholangiocarcinomas induced by feeding 3’-methyl-4-dimethylaminoazobenzene to rats. Histopathology and ultrastructure. Am. J. Pathol. 87, 189–204 (1977).
Tarlow, B. D. et al. Bipotential adult liver progenitors are derived from chronically injured mature hepatocytes. Cell Stem Cell 15, 605–618 (2014).
Raggi, C., Invernizzi, P. & Andersen, J. B. Impact of microenvironment and stem-like plasticity in cholangiocarcinoma: molecular networks and biological concepts. J. Hepatol. 62, 198–207 (2015).
Zong, Y. W. et al. Notch signaling controls liver development by regulating biliary differentiation. Development 136, 1727–1739 (2009).
Sia, D. et al. Integrative molecular analysis of intrahepatic cholangiocarcinoma reveals 2 classes that have different outcomes. Gastroenterology 144, 829–840 (2013).
Knudsen, E. S., Gopal, P. & Singal, A. G. The changing landscape of hepatocellular carcinoma etiology, genetics, and therapy. Am. J. Pathol. 184, 574–583 (2014).
Finkel, T. & Hwang, P. M. The Krebs cycle meets the cell cycle: mitochondria and the G1–S transition. Proc. Natl Acad. Sci. USA 106, 11825–11826 (2009).
Mitra, K., Wunder, C., Roysam, B., Lin, G. & Lippincott-Schwartz, J. A hyperfused mitochondrial state achieved at G1–S regulates cyclin E buildup and entry into S phase. Proc. Natl Acad. Sci. USA 106, 11960–11965 (2009).
Gao, Y. M. et al. Distinct gene expression and epigenetic signatures in hepatocyte-like cells produced by different strategies from the same donor. Stem Cell Rep. 9, 1813–1824 (2017).
Broutier, L. et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat. Protoc. 11, 1724–1743 (2016).
Arruda, A. P. et al. Chronic enrichment of hepatic endoplasmic reticulum-mitochondria contact leads to mitochondrial dysfunction in obesity. Nat. Med. 20, 1427–1435 (2014).
Tang, Z. F. et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45, W98–W102 (2017).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Tripathi, S. et al. Meta- and orthogonal integration of influenza “OMICs” data defines a role for UBR4 in virus budding. Cell Host Microbe 18, 723–735 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Jalan-Sakrikar, N. et al. Hedgehog signaling overcomes an EZH2-dependent epigenetic barrier to promote cholangiocyte expansion. Plos ONE 11, e0168266 (2016).
Shih, H. P. et al. A gene regulatory network cooperatively controlled by Pdx1 and Sox9 governs lineage allocation of foregut progenitor cells. Cell Rep. 13, 326–336 (2015).
Schuppan, D. & Kim, Y. O. Evolving therapies for liver fibrosis. J. Clin. Invest. 123, 1887–1901 (2013).
Michelotti, G. A., Machado, M. V. & Diehl, A. M. NAFLD, NASH and liver cancer. Nat. Rev. Gastroenterol. Hepat. 10, 656–665 (2013).
Friedman, S. L., Neuschwander-Tetri, B. A., Rinella, M. & Sanyal, A. J. Mechanisms of NAFLD development and therapeutic strategies. Nat. Med. 24, 908–922 (2018).
Friedman, S. L. Evolving challenges in hepatic fibrosis. Nat. Rev. Gastroenterol. Hepat. 7, 425–436 (2010).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Broad Institute TCGA Genome Data Analysis Center. Analysis overview for liver hepatocellular carcinoma (Primary solid tumor cohort). Broad Institute of MIT and Harvard, https://doi.org/10.7908/C1QV3KXG (2016).
Acknowledgements
We are thankful to X. He (Fudan University) and J. Hu (Interdisciplinary Research Center on Biology and Chemistry, CAS) for their critical reading of the manuscript, Q. Gao (Zhongshan Hospital) for providing the lentivirus vectors PTPN3, D. Li (East China Normal University) for providing the Tet-On 3G inducible system, D. Ye (Fudan University) for providing the IDH1 and IDH2 plasmids, the members of Cell Biology and Chemical Biology Core Facility of the SIBCB for technical support and the members of the L. Hui laboratory for their valuable suggestions. This study was supported by the Chinese Academy of Sciences (grant nos XDA16020201, XDA12050104, 153D31KYSB20160247), National Natural Science Foundation of China (grant nos 31630044, 31601186, 81703093 and 31801228), Shanghai Science and Technology Committee (16JC1400202), National Science and Technology Major Project (grant no. 2018ZX09711002-009) and National Special Support Plan for Top Talents.
Author information
Authors and Affiliations
Contributions
L.S. and L.H. designed the project. L.S. performed most of the experiments with support from Y.W. Y.W. and L.C. constructed the lentiviral vectors and performed the RT–qPCR assays and virus packaging. J.C. performed the orthotopic transplantations. X.M., H.L. and Z.Q. performed the bioinformatics analyses. Z.Z. helped with the drug screening. R.-Z.Y. and J.-S.K. helped with the mitochondrial-related analyses. C.W. and X.C. helped with the western blots and cloning. Y.Y. and Y.J. performed the histological analyses and provided the human ICC samples. L.W. and G.P. characterized the CYP metabolism. H.Z. provided the human HCC samples. Y.J. supervised all histological analyses and conceptually contributed to phenotype characterization. Y.-W.Z. supervised L.S. in the three-dimensional culture and functional analyses of organoids and conceptually contributed to the experimental design. S.Z. supervised Y.W. and contributed to the experimental design. L.S. and L.H. analysed the data and wrote the paper with suggestions from other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Characterization of hiHep organoids.
(a) Immunofluorescent staining of MRP2, an organic anion transporter located in bile canaliculi between adjacent cells. Scale bars: 100 μm. (b) Immunofluorescent staining of mature hepatic proteins ALB and AAT in hiHep organoids. Scale bars: 100 μm. (c) Immunofluorescent staining of ALB and fetal hepatocyte protein AFP in hiHep and HepG2 organoids. Scale bars: 50 μm. (d-e) Enriched gene ontology (GO) terms in PHHs and hiHep organoids by analyses of the differentially expressed genes (FDR q-value <5%, n=2 biologically independent samples for each group). (f-i) 2D_hiHep cells and hiHep organoids showed diverse expression levels of hepatic genes as determined by q-PCR. The hepatic genes included the genes of liver-specific secretory proteins (f), glycogen synthesis (g), lipid metabolism (h) and CYP450 family (i). Cryopreserved PHHs cultured for 2 days after thaw were used as positive control. HepG2 cells were also used as a control. Data were normalized to PHHs (n=4 independent hiHep samples were collected). (j) The expression levels of cell-cycle related genes were detected by q-PCR. All data were normalized to those of 2D_hiHeps (n=3 independent experiments). (k) Cell cycle analysis of 2D_hiHeps and hiHep organoids by DAPI staining and FACS (n=4 independent experiments). (l) Transmission electron microscopy image for glycogen granules (arrows) in hiHep organoids. Scale bar: 0.5 μm. a-c, l were repeated 3 times independently, with similar results. Data in f-k were presented as mean ± SEM. All P values were calculated using two-tailed unpaired Student’s t-test. The gating strategy of flow cell analyses was shown in Supplementary Table 2. Statistics Source Data were provided in Supplementary Table 5.
Supplementary Figure 2 Characterization of c-MYC organoids and the derived HCCs.
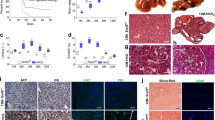
(a) List of c-MYC alterations in human HCCs (see Methods). 348 HCC samples in TCGA were analysed with 12 HCCs showing both c-MYC amplification and up-regulation, 27 HCCs showing amplification and 21 HCCs showing up-regulation. (b) Western blot of c-MYC protein in GFP and c-MYC organoids. (c) FACS analysis of c-MYC over-expression. (d) Immunofluorescent staining of ALB and AFP in c-MYC organoids. HiHep organoids and HepG2 were used as controls. Scale bars: 50 μm. (e) Representative images of c-MYC organoid-derived cancers in NOD/SCID mice. No cancer was detected from GFP organoids. (f-g) AFP levels were measured in supernatants of cultured c-MYC organoids (f) and in sera from mice with c-MYC organoid cancers (g) by ELISA (n=3 independent experiments, data were presented as mean ± SEM). (h) The expression levels of cholangiocyte genes in c-MYC transfected cholangiocyte organoids were detected by q-PCR. Data were normalized to GFP transfected cholangiocyte organoids (n=4 biologically independent samples, data were presented as mean ± SD). (i) Morphologies of constitutively activated β-catenin (β-catenin∆ex3) and β-catenin∆ex3+c-MYC organoids during culture. Scale bars: 100 μm. (j) Representative H&E staining of β-catenin and β-catenin∆ex3+c-MYC organoids. Scale bars: 50 μm. (k) Representative images of β-catenin∆ex3+c-MYC organoid-derived cancers in NOD/SCID mice. No cancer was detected from GFP and β-catenin∆ex3 organoids. (l) Representative H&E staining of β-catenin∆ex3+c-MYC organoid-derived HCCs. Arrowheads indicate eosinophilic nucleoli. Arrows indicate trabecular structures. Scale bar: 50 μm. b-c, e, i-j were repeated three times independently, d, k-l were repeated twice independently, with similar results. The gating strategy of flow cell analyses was shown in Supplementary Table 2. Scanned images of unprocessed blots were shown in Supplementary Figure 9. Statistics Source Data were provided in Supplementary Table 5.
Supplementary Figure 3 Mitochondria-associated functions in c-MYC organoids and c-MYC-positive HCCs.
(a) GSEA analyses for Reactome S-phase geneset and Reactome translation genesets. FDR q<25%. (b) The expression levels of cell cycle-related genes were detected by q-PCR. Data were normalized to GFP organoids (n=3 independent experiments). (c) Cell cycle analysis of c-MYC organoids by DAPI staining and FACS (n=4 independent experiments). (d-e) Enriched gene ontology (GO) terms in c-MYC and GFP organoids by analysis of the differentially expressed genes (DEG, see Methods). P <0.01. (f) Statistical quantification of the numbers of mitochondria in 2D-cultured c-MYC cells and c-MYC organoids (n=12 biologically independent samples for 2D-cultured c-MYC cells; n=18 biologically independent samples for c-MYC organoids). (g-i) The mitochondrion organization and biogenesis pathway and the mitochondrial fission pathway were enriched in HCCs positive in c-MYC alone (g) or in combination with P53 (h) and RB mutations (i). Data were retrieved from TCGA database (see Methods). FDR q<25%. (j) Cellular response to oxidative stress pathway was reduced in HCCs positive in c-MYC alone or in combination with P53 and RB mutations. FDR q<25%. n=2 biologically independent samples for a, d-e. n=50 biologically independent samples for c-MYC-positive HCCs (including n=18 biologically independent samples for P53-mutated ones and n=4 biologically independent samples for RB-mutated ones) and Human_HCC-normals in g-j. Data in b, c, f were presented as mean ± SEM. P values were calculated using two-tailed unpaired Student’s t-test. The gating strategy of flow cell analyses was shown in Supplementary Table 2. Statistics Source Data were provided in Supplementary Table 5.
Supplementary Figure 4 Analyses of ICC-enriched mutations in hiHep organoids.
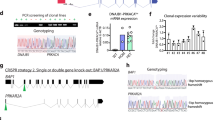
(a) List of frequencies of ICC-enriched mutations using ICC database from cBioPortal. (b) The over-expression levels of ICC-enriched mutations were detected by q-PCR analysis (n=4 biologically independent samples. Data were presented as mean ± SD). GFP organoids were used as controls. (c) Alcian blue staining of hiHep organoids transfected with ICC-enriched mutations. Scale bars: 50 μm. (d) Heatmap of ICC-related genes and hepatocyte genes in c-MYC organoids, RAS organoids, TCGA-ICCs and TCGA-HCCs, and published human HCC and CC organoids1. c was performed once. Statistics Source Data were provided in Supplementary Table 5.
Supplementary Figure 5 ICC-related features in RAS organoids and the derived cancers.
Supplementary Figure 5 (a) Protein levels of RAS and the active forms of its downstream molecules ERK and AKT were detected by Western blot. (b) FACS analysis of RAS organoids by RAS staining. (c) Representative images of RAS organoid-derived ICCs. (d) Gene expression levels of RAS organoid-derived ICCs. C-MYC organoid-derived HCCs were used as controls (n=4 biologically independent samples. Data were presented as mean ± SD). (e) Immunostaining images of human-specific antibodies on mouse livers, including Ki67, CK19, MUC5 and CK7. Scale bars: 50 μm. (f) P-AKT and p-ERK staining of RAS organoid-derived ICCs. Scale bars: 50 μm. (g) The expression levels of cell cycle-related genes were detected by q-PCR (n=3 independent experiments and data were normalized to GFP organoids). (h) Cell cycle analysis of RAS organoids by DAPI staining and FACS (n=4 independent experiments). (i) Quantitation of cleaved-caspases 3-positive cells in GFP and RAS organoids (n=3 independent experiments). (j) Compared with 2D-cultured RAS cells, RAS organoids were enriched in gland development pathway, bile acid and bile salt metabolism pathway, and the reported CC-related geneset (FDR q<25%, n=2 biologically independent samples for each group). (k) Typical glandular structures were formed in RAS organoid-derived ICC (arrows), but not in 2D-cultured RAS cells-derived cancers. Scale bars: 50 μm. a-c, k were repeated three times independently, with similar results, e-f were performed once. Data in g-i were presented as mean ± SEM. The gating strategy of flow cell analyses was shown in Supplementary Table 2. Scanned images of unprocessed blots were shown in Supplementary Figure 9. Statistics Source Data were provided in Supplementary Table 5.
Supplementary Figure 6 Cancers from transplantation of organoids transfected with ICC oncogenes.
(a) Representative images of YAP and IDH2 organoid-derived cancers in NOD/SCID mice. No cancer was detected from NICD, IDH1 and AKT organoids. (b-c) Representative H&E, CK19, SOX9 and Ki67 staining of YAP (b) and ΙDΗ2 (c) organoid-derived cancers. Scale bars: 50 μm. a, c was performed once. b was repeated twice independently, with similar results.
Supplementary Figure 7 In vitro lineage conversion in RAS organoids.
(a-b) The expression levels of cholangiocyte markers (a) and hepatic markers (b) were detected by q-PCR analysis. All the q-PCR data were normalized to those of GFP groups. (c-d) Immunofluorescent staining and q-PCR analyses of cholangiocyte markers CK19 (c) and SOX9 (d). Scale bars: 50 μm. (e) The expression levels of ICC-related genes were detected by q-PCR in RAS organoids which were cultured for 20, 30 and 60 days (n=4 independent experiments, and data were presented as mean ± SEM). (f) ALB and AFP staining of RAS organoids. Scale bars: 50 μm. (g) Venn diagram for iPSC-derived cholangiocyte (CH)- and iPSC-derived LPC-enriched genes (see Methods). (h-i) GSEA analyses of CH- and LPC-enriched genes in human ICCs. All the human samples were from TCGA dataset (see Methods, n=25 biologically independent samples for No RAS-mutated ICCs; n=5 biologically independent samples for RAS-mutated ICCs; n=9 biologically independent samples for Human-ICC_Normals). FDR q<25%, P<0.05. (j) K-means clustering of RAS organoids at indicated days using the whole transcriptome. (k-l) Enriched gene ontology (GO) terms in Cluster 4 (k) and Cluster 3 (l). Note the Notch signalling pathway, JAK-STAT and cytokine related pathways were included. Images in c-d were performed three times, f was performed twice independently, with similar results. n=2 biologically independent samples for g, j-l. 4 biologically independent samples were detected by q-PCR in a-d, and data were presented as mean ± SD. Statistics Source Data were provided in Supplementary Table 5.
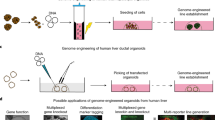
Supplementary Figure 8 Characterization of hiHep organoids with inducible SV40LT and ProliHH organoids upon c-MYC and RAS over-expression.
(a) Representative images of hihep organoids cultured for 10 days with or without dox-induced SV40LT expression. Scale bars: 50 μm. (b) The expression levels of SV40LT in hihep organoids with or without dox were detected by q-PCR (n=4 biologically independent samples). Wild-type fibroblasts-derived hiHeps were used as a control. (c) Q-PCR analyses of the expression levels of c-MYC-related genes in c-MYC organoids with or without dox-induced SV40LT expression (n=4 biologically independent samples). (d) The percentage of ERs adjacent to mitochondria as determined by electronic microscope. Pictures of 27 biologically independent samples with same magnification (21000X) were quantitated. (e) Q-PCR analysis of the expression of ICC-related genes in RAS organoids with or without dox-induced SV40LT expression (n=4 biologically independent samples). (f) Principal-component analysis (PCA) was performed to compare gene expression profiles of ProliHH organoids (n=1), human EpCAM+ liver organoids (n=3 biologically independent samples), hiHep organoids (n=2 biologically independent samples) and PHHs (n=3 biologically independent samples). Data of human EpCAM+ liver organoids were downloaded from the GEO database, GSE84073. (g) Enrichment of hepatic pathways were analysed using GSEA by pairwise comparison of ProliHH organoids, PHHs, hiHep organoids to human fibroblasts. (h) Heatmap of key liver progenitor genes in ProliHH organoids, PHHs and hiHep organoids. (i) ProliHH organoids transfected with c-MYC were enriched in c-MYC signature genes. FDR q<25%. (j) The percentage of ERs adjacent to mitochondria as determined by electronic microscope. Pictures of 24 biologically independent samples with same magnification (21000X) were quantitated. (k) ProliHH organoids transfected with RAS were enriched in RAS signature genes. FDR q<25%. (l) Q-PCR analyses of the expression of ICC-related genes in ProliHH organoids transfected with RAS. a was repeated three times independently, with similar results. g, h, i, k were performed with the same samples as listed in f. Data were shown as mean ± SD in b, c and e; data were shown as mean ± SEM in d and j. P values were calculated using two-tailed unpaired Student’s t-test. Statistics Source Data were provided in Supplementary Table 5.
Supplementary Figure 9
Unprocessed original scans of blots.
Supplementary information
Supplementary Information
Supplementary Figs 1–9, Supplementary Table titles/legends.
Supplementary Table 1
Summary of orthotopic transplantation and cancer formation
Supplementary Table 2
Summary of primers, kits, antibodies, HMM medium components and the work flow of flow cytometry analyses.
Supplementary Table 3
List of raw RNA sequencing data and mutation information of TCGA samples
Supplementary Table 4
Raw data for the RNA analyses
Supplementary Table 5
Statistics source data for this paper
Rights and permissions
About this article
Cite this article
Sun, L., Wang, Y., Cen, J. et al. Modelling liver cancer initiation with organoids derived from directly reprogrammed human hepatocytes. Nat Cell Biol 21, 1015–1026 (2019). https://doi.org/10.1038/s41556-019-0359-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0359-5
This article is cited by
-
Progress, application and challenges of liver organoids
Clinical Cancer Bulletin (2024)
-
Evolving cognition of the JAK-STAT signaling pathway: autoimmune disorders and cancer
Signal Transduction and Targeted Therapy (2023)
-
MYC in liver cancer: mechanisms and targeted therapy opportunities
Oncogene (2023)
-
Liver cell therapies: cellular sources and grafting strategies
Frontiers of Medicine (2023)
-
Criteria for preclinical models of cholangiocarcinoma: scientific and medical relevance
Nature Reviews Gastroenterology & Hepatology (2023)