Abstract
A robust nanopillar platform with increased spatial resolution reveals that perinuclear forces, originating from stress fibres spanning the nucleus of fibroblasts, are significantly higher on these nanostructured substrates than the forces acting on peripheral adhesions. Many perinuclear adhesions embrace several nanopillars at once, pulling them into β1-integrin- and zyxin-rich clusters, which are able to translocate in the direction of cell motion without losing their tensile strength. The high perinuclear forces are greatly reduced upon inhibition of cell contractility or actin polymerization and disruption of the actin cap by KASH dominant-negative mutant expression. LMNA null fibroblasts have higher peripheral versus perinuclear forces, impaired perinuclear β1-integrin recruitment, as well as YAP nuclear translocation, functional alterations that can be rescued by lamin A expression. These highly tensed actin-cap fibres are required for YAP nuclear signalling and thus play far more important roles in sensing nanotopographies and mechanochemical signal conversion than previously thought.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Vogel, V. & Sheetz, M. Local force and geometry sensing regulate cell functions. Nat. Rev. Mol. Cell Biol. 7, 265–275 (2006).
Fu, J. et al. Mechanical regulation of cell function with geometrically modulated elastomeric substrates. Nat. Methods 7, 733–736 (2010).
Dahl, K. N., Booth-Gauthier, E. A. & Ladoux, B. In the middle of it all: mutual mechanical regulation between the nucleus and the cytoskeleton. J. Biomech. 43, 2–8 (2010).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Trappmann, B. et al. Extracellular-matrix tethering regulates stem-cell fate. Nat. Mater. 11, 642–649 (2012).
Kim, D. H., Chambliss, A. B. & Wirtz, D. The multi-faceted role of the actin cap in cellular mechanosensation and mechanotransduction. Soft Matter 9, 5516–5523 (2013).
Piccolo, S. Role of the Hippo transducers YAP and TAZ in mechanotransduction and cancer stem cells. Eur. J. Cancer 50, S15–S15 (2014).
Iskratsch, T., Wolfenson, H. & Sheetz, M. P. Appreciating force and shape—the rise of mechanotransduction in cell biology. Nat. Rev. Mol. Cell Biol. 15, 825–833 (2014).
Humphrey, J. D., Dufresne, E. R. & Schwartz, M. A. Mechanotransduction and extracellular matrix homeostasis. Nat. Rev. Mol. Cell Biol. 15, 802–812 (2014).
Ihalainen, T. O. et al. Differential basal-to-apical accessibility of lamin A/C epitopes in the nuclear lamina regulated by changes in cytoskeletal tension. Nat. Mater. 14, 1252–1261 (2015).
Das, A., Fischer, R. S., Pan, D. & Waterman, C. M. YAP nuclear localization in the absence of cell–cell contact is mediated by a filamentous actin-dependent, myosin II- and phospho-YAP-independent pathway during extracellular matrix mechanosensing. J. Biol. Chem. 291, 6096–6110 (2016).
Elosegui-Artola, A. et al. Mechanical regulation of a molecular clutch defines force transmission and transduction in response to matrix rigidity. Nat. Cell Biol. 18, 540–548 (2016).
Discher, D. E. et al. Matrix mechanosensing: from scaling concepts in ‘omics data to mechanisms in the nucleus, regeneration, and cancer. Annu. Rev. Biophys. 46, 295–315 (2017).
Vogel, V. Unravelling the mechanobiology of extracellular matrix. Annu. Rev. Physiol. (in the press).
Roca-Cusachs, P., Gauthier, N. C., del Rio, A. & Sheetz, M. P. Clustering of α5β1 integrins determines adhesion strength whereas αvβ3 and talin enable mechanotransduction. Proc. Natl Acad. Sci. USA 106, 16245–16250 (2009).
Saltel, F. et al. New PI(4,5)P2- and membrane proximal integrin-binding motifs in the talin head control β3-integrin clustering. J. Cell Biol. 187, 715–731 (2009).
Ross, T. D. et al. Integrins in mechanotransduction. Curr. Opin. Cell Biol. 25, 613–618 (2013).
Rossier, O. et al. Integrins β1 and β3 exhibit distinct dynamic nanoscale organizations inside focal adhesions. Nat. Cell Biol. 14, 1057–1067 (2012).
Schiller, H. B. et al. β1- and αv-class integrins cooperate to regulate myosin II during rigidity sensing of fibronectin-based microenvironments. Nat. Cell Biol. 15, 625–636 (2013).
Sun, Z., Guo, S. S. & Fassler, R. Integrin-mediated mechanotransduction. J. Cell Biol. 215, 445–456 (2016).
Ungricht, R. & Kutay, U. Mechanisms and functions of nuclear envelope remodelling. Nat. Rev. Mol. Cell Biol. 18, 229–245 (2017).
Khatau, S. B. et al. A perinuclear actin cap regulates nuclear shape. Proc. Natl Acad. Sci. USA 106, 19017–19022 (2009).
Chancellor, T. J., Lee, J., Thodeti, C. K. & Lele, T. Actomyosin tension exerted on the nucleus through nesprin-1 connections influences endothelial cell adhesion, migration, and cyclic strain-induced reorientation. Biophys. J. 99, 115–123 (2010).
Chambliss, A. B. et al. The LINC-anchored actin cap connects the extracellular milieu to the nucleus for ultrafast mechanotransduction. Sci. Rep. 3, 1087 (2013).
Kim, D. H. & Wirtz, D. Cytoskeletal tension induces the polarized architecture of the nucleus. Biomaterials 48, 161–172 (2015).
Ramdas, N. M. & Shivashankar, G. V. Cytoskeletal control of nuclear morphology and chromatin organization. J. Mol. Biol. 427, 695–706 (2015).
Makhija, E., Jokhun, D. S. & Shivashankar, G. V. Nuclear deformability and telomere dynamics are regulated by cell geometric constraints. Proc. Natl Acad. Sci. USA 113, 32–40 (2016).
Polacheck, W. J. & Chen, C. S. Measuring cell-generated forces: a guide to the available tools. Nat. Methods 13, 415–423 (2016).
Sarangi, B. R. et al. Coordination between intra- and extracellular forces regulates focal adhesion dynamics. Nano Lett. 17, 399–406 (2017).
Lammerding, J. et al. Lamins A and C but not lamin B1 regulate nuclear mechanics. J. Biol. Chem. 281, 25768–25780 (2006).
Gay, O. et al. RefilinB (FAM101B) targets FilaminA to organize perinuclear actin networks and regulates nuclear shape. Proc. Natl Acad. Sci. USA 108, 11464–11469 (2011).
Falzone, T. T., Lenz, M., Kovar, D. R. & Gardel, M. L. Assembly kinetics determine the architecture of α-actinin crosslinked F-actin networks. Nat. Commun. 3, 861 (2012).
Roca-Cusachs, P. et al. Integrin-dependent force transmission to the extracellular matrix by α-actinin triggers adhesion maturation. Proc. Natl Acad. Sci. USA 110, E1361–E1370 (2013).
Ribeiro Ede, A. Jr et al. The structure and regulation of human muscle α-actinin. Cell 159, 1447–1460 (2014).
Meacci, G. et al. α-Actinin links extracellular matrix rigidity-sensing contractile units with periodic cell-edge retractions. Mol. Biol. Cell 27, 3471–3479 (2016).
Ylanne, J., Scheffzek, K., Young, P. & Saraste, M. Crystal structure of the α-actinin rod reveals an extensive torsional twist. Structure 9, 597–604 (2001).
Hirata, H., Tatsumi, H. & Sokabe, M. Mechanical forces facilitate actin polymerization at focal adhesions in a zyxin-dependent manner. J. Cell Sci. 121, 2795–2804 (2008).
Smith, M. A. et al. A zyxin-mediated mechanism for actin stress fiber maintenance and repair. Dev. Cell 19, 365–376 (2010).
Ma, B. et al. Zyxin-Siah2-Lats2 axis mediates cooperation between Hippo and TGF-β signalling pathways. Nat. Commun. 7, 11123 (2016).
Lord, M. Cytoskeletal regulation: sorting out stress fibers with tropomyosin. Curr. Biol. 21, R255–R257 (2011).
Wolfenson, H. et al. Tropomyosin controls sarcomere-like contractions for rigidity sensing and suppressing growth on soft matrices. Nat. Cell Biol. 18, 33–42 (2016).
Aragona, M. et al. A mechanical checkpoint controls multicellular growth through YAP/TAZ regulation by actin-processing factors. Cell 154, 1047–1059 (2013).
Liu, M., Sun, J. R., Sun, Y., Bock, C. & Chen, Q. F. Thickness-dependent mechanical properties of polydimethylsiloxane membranes. J. Micromech. Microeng. 19, 035028 (2009).
Ghassemi, S. et al. Cells test substrate rigidity by local contractions on submicrometer pillars. Proc. Natl Acad. Sci. USA 109, 5328–5333 (2012).
Kuo, C. W. et al. Polymeric nanopillar arrays for cell traction force measurements. Electrophoresis 31, 3152–3158 (2010).
Zamir, E. et al. Dynamics and segregation of cell-matrix adhesions in cultured fibroblasts. Nat. Cell Biol. 2, 191–196 (2000).
Mould, A. P. et al. Integrin activation involves a conformational change in the α1 helix of the β subunit A-domain. J. Biol. Chem. 277, 19800–19805 (2002).
Kim, D. H., Cho, S. & Wirtz, D. Tight coupling between nucleus and cell migration through the perinuclear actin cap. J. Cell Sci. 127, 2528–2541 (2014).
Dalby, M. J., Gadegaard, N. & Oreffo, R. O. Harnessing nanotopography and integrin–matrix interactions to influence stem cell fate. Nat. Mater. 13, 558–569 (2014).
Liang, E. I., Mah, E. J., Yee, A. F. & Digman, M. A. Correlation of focal adhesion assembly and disassembly with cell migration on nanotopography. Integr. Biol. 9, 145–155 (2017).
Jiguet, S., Bertsch, A., Judelewicz, M., Hofmann, H. & Renaud, P. SU-8 nanocomposite photoresist with low stress properties for microfabrication applications. Microelectron. Eng. 83, 1966–1970 (2006).
Gere, J. M. & Timoshenko, S. P. Mechanics of Materials. (PWS, Boston, 1997).
Schoen, I., Hu, W., Klotzsch, E. & Vogel, V. Probing cellular traction forces by micropillar arrays: contribution of substrate warping to pillar deflection. Nano Lett. 10, 1823–1830 (2010).
Vallotton, P. et al. Diatrack particle tracking software: review of applications and performance evaluation. Traffic 18, 840–852 (2017).
Acknowledgements
The authors thank M. Vihinen-Ranta and D. Hodzic, who provided the eGFP-lamin A and KASH2/ext_eGFP constructs, R. Foisner and R. Fässler for the LMNA null and WT MEF cells, B. Wehrle-Haller (Geneva) for the REFs expressing β3-integrin-eGFP, and K. Maniura (EMPA, St. Gallen) for use of the Amaxa Nucleofector II system. This work was made possible by financial support from the Portuguese Foundation for Science and Technology, doctoral grant SFRH/BD/42019/2007 (to L.A.), an ERC Advanced Grant GA 233157 (to V.V.), the Swiss National Science Foundation (SNF 310030B_133122/1 to V.V.), as well as the SNF NCCR ‘Molecular Systems Engineering’ and the Nano-Tera.ch ‘LiveSense’ RTD: 20NAN1_123593. The authors acknowledge access to the ETH FIRST User Facility for micro- and nanofabrication.
Author information
Authors and Affiliations
Contributions
J.S., L.A. and V.V. developed the story line, experimental design and wrote the manuscript. J.S. and L.A. carried out the experimental work and Z.L. contributed to the YAP1 translocation studies. J.S. fabricated the nanopillar arrays and performed all data analyses.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Production and characterization of roboust nanopillar arrays for cellular traction force measurement.
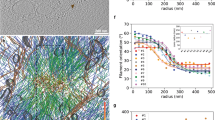
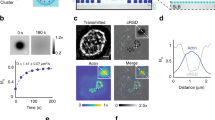
a, Scanning electron microscopy images of the nanopillar arrays with the dimensions 1.5 μm height, 0.25μm width and center-to-center pitch 0.8 μm (upper, scale bar 1μm) and of adhering fibroblasts (scale bars 5 μm and 2 μm, bottom left and right, respectively) b, Schematic of the colloidal templating process used to fabricate the SU-8 nanopillar arrays. The single layer of polystyrene beads was spin-coated on a silicon substrate to create close-packed sphere nanostructures. Oxygen plasma treatment reduced the size of the polystyrene beads from 800 nm to 250 nm at time T0 and T, respectively. Plasma etching was used to create the silicon nanopillar array (nanopillars: 1.5 μm height with a 800 nm pitch) whereby the polystyrene beads served as mask. PDMS was filled into the silicon nanopillar structure to create a mold. The PDMS mold was peeled off from the silicon nanopillar array. The SU-8 photoresist solution was then filled into the PDMS mold, topped with a glass coverslip followed by degasing under vacuum and finally turned upside down and cured by UV exposure (2 minutes). After curing, the PDMS mold was peeled off and could be reused multiple time.The SU-8 nanopillar arrays were coated with 25μg/μl Fn by contact printing (room temperature for 30 minutes) and washed with PBS. Finally, the nanopillar arrays were immersed in cell culture medium followed by cell seeding. c, Spring constant of nanopillars (diameter = 250 nm, length = 1.5 μm) calculated conventionally using Hook’s law (kbend), corrected considering substrate warping (kbend)1, and calculated from AFM measurements (kAFM) (for additional information see Supporting Information). Quantification of the spring constant of nanopillars measured by AFM (circles) and calculated theoretically (rectangular and triangle) (additional supplemental information). The different colors of the circles represented the distinct positions where the AFM tip (spring constant 62 nN/μm) applied force on a nanopillar (the different lengths from pillar base to top were: 1.45 μm (red), 1.25 μm (blue) and 1 μm (green)). Each position was loaded and unloaded for 10 times. Inner cross-section image showing the nanopillar array that was scanned by AFM (using tapping mode). Scale bar is 1 μm. L refers to the length of the nanopillar and D to its diameter. Real-time SEM images showing the ability of a SU-8 nanopillar to swing back to the original position upon cutting the filopodium using an electron beam. Scale bar 200 nm. d, XY-confocal confocal images of a membrane stained fibroblast (Vybrant/Dil, red) on Fn-coated (Alexa 488 labeled, green) nanopillar array (top), 30 min after seeding. Zoom-in from boxed-in white region: fluorescence (left, membrane and middle, Fn) and DIC (right) images from nanopillars taken at the top (T) and bottom (B) planes, respectively as indicated (n=285 pillars). Scale bars are 5 μm. Together with respective quantification of pillar displacements using the top and bottom planes of the nanopillar array. P values were accessed with two-sided Student’s T-test (*p<0.05, **p<0.01, ***p<0.001), the differences between these pairwise displacements (T vs. B and T vs. T’) are less than 1%. NS, not significantly different. f, Histrogram for pillar displacement comparing thermal noise as measured on nanopillars by cell contact (REF, red) and not (black), respectively. Pairwise pillar displacement measurements by comparing DIC images taken at the pillar tops (T) and pillar bottoms (B), respectively, as well as at the pillar tops before (T) and after (T’) cell detachment (post-trypsination, 2 min). Each experiment was repeated independently 3 times.
Supplementary Figure 2 Characterization of fibroblasts adhering on fibronectin coated nanopillars by identifying peripheral β3-integrin rich region.
a, Prior to the formation of an actin cap, no pillar clusters can be observed in the perinuclear region. z-maximum intensity projection from confocal image of a REF immunostained for actin (red) (top left), together with correspondent XZ-cross section below (bottom left). The cell on a Fn-coated nanopillar array were fixed 10 min after cell seeding (top right). Vectoral directions of pillar deflections are indicated for inward (red) and outward (green), respectively. At the cell periphery, the outer line of pillars is deflected inwards, while the next row of pillars are pulled in opposite direction as reported previously.3, 4 At the perinuclear region, the pillars are mostly deflected outwards, but no nanopillar cluster formation is yet seen so early in the spreading phase since the stress fibers spanning the nucleus have not yet been assembled (bottom left). Scale bar 5 μm. b, Fluorescent confocal images of single fibroblasts 30 min after seeding on different SU-8 substrates, stained with a membrane dye (red, Dil) on flat substrates (left), as well as on and nanopillars (middle) and microposts (right), together with a respective comparison of cell spreading areas (histogram, n=20 cells per condition). The largest cell spreading areas were measured on flat SU-8 surfaces (1673 ± 87 μm2), followed by the cell spreading areas on SU-8 nanopillar arrays (1454 ± 77 μm2), and finally on SU-8 micropost arrays (520 ± 66 μm2). Scale bar is 5 μm. c, REFs were seeded for 2h on Fn coated nanopillar substrates. Percentage of β3-integrin clusters were grouped into three distinct categories: peripheral rim (5 μm wide peripheral rim at the cell edge, green), perinuclear region (red) and the region underneath the cell nucleus (blue). Scale bar is 10 μm, n = 20 cells. d, Typical approach to categorize and quantify the three distinct regions for region-specific nanopillar displacement quantifications. Absolute fractions of color coded cell areas occupied by these three regions (periphery, perinuclear and nuclear) (n=20 cells). The data present here is average plus and minus the standard deviation, P values were accessed with two-sided Student’s T-test (*p<0.05, **p<0.01, ***p<0.001). Each experiment was repeated independently 3 times.
Supplementary Figure 3 Correlative nanopillar displacement measurements from DIC vs fluorescence confocal images.
a, Z-maximum intensity projection from confocal image of a wt MEF immunostained for actin (green) and the nucleus (DAPI, blue) which adhered for 1 h to fluorescently labeled fibronectin (Alexa 633, red) coated nanopillars, together with zoom-in view from yellow rectangle and respective nanopillar images obtained from DIC and fluorescence microscopy. Scale bar 10 μm. b, Color mapping of pillar displacements from the boxed-in region (yellow rectangle) and respective quantification from DIC and fluorescence images (c&d). The data present here is average plus and minus the standard deviation (Box:SE whisker:SD). The same results for different cell types or treatments are obtained either from DIC or fluorescence images of nanopillar arrays. e-h, Similar data representation for LMNA MEF. No difference was observed between correlative quantifications of pillar displacements using DIC versus fluorescence images. i, Average pillar displacement quantification of DIC and fluorescence images for wt MEFs upon LatB treatment (n=20 cells). The data present here is average plus and minus the standard deviation. The same results for different cell types or treatments are obtained either from DIC or fluorescence images of nanopillar arrays. Each experiment was repeated independently 3 times.
Supplementary Figure 4 Total horizontal force measurements, actin cap formation and YAP nuclear translocation for different fibroblasts on nanopillar arrays.
Total horizontal force divided by the cell spreading area. Histograms of horizontal force per cell area in a, Different pan-integrin-null fibroblasts (pKO-αv, pKO-β1 and pKO-αv/β1). b, MEFs treated with Mn2+ (0.5 mM for 30min) and one of the inhibitors: ML-7 (25 μM), LatB (50nM) or Y27632 (10 μM) for 2h. c, Different MEFs, wild type (wt), LMNA null and LMNA null transiently transfected with lamin A tagged with eGFP. d, Quantification of average nanopillar displacements from under the whole cell. The data presented here are averages plus and minus the standard deviation (n=10 cells). P values were accessed with one-way analysis of variance (ANOVA) Tukey’s post test (*p<0.05, ** p<0.01, ***p<0.001). Each experiment was repeated independently 3 times. e, REFs pre-adhered for 2 h on a Fn coated glass-bottom dish. XY-confocal images of actin immunostaining (grey) from apical and basal planes of a control (no drug treatment) (top) versus LatB treated2 (50 nM, 2 h) fibroblast. Scale bars 20 μm. Together withg respective quantification of the percentage of fibroblasts having an actin cap from control versus LatB treated cells (based on the presence or absence of apical actin fibers) The data presented here are averages plus and minus the standard deviation (n=50 cells). f, Fibroblasts were seeded overnight on Fn coated nanopillar arrays. z-maximum intensity projections from confocal images of the indicated cell types immunostained for actin (gray) merged with cell nucleus (DAPI, blue) (left) and YAP (green) (right). Enlargements of the apical actin filaments (center) from the boxed-in region (yellow dash square) of the left side images. Scale bar 20 μm. Together with quantification of the percentage of fibroblasts having an actin cap (based on the acquired apical actin confocal images), from wt versus LMNA null MEFs for different conditions: control (normal and overnight starved) versus RhoA activator (CN03) treated fibroblasts (n=10 cells). LMNA null MEFs treated with CN03 significantly increased actin cap formation, but still below the wt levels. The data present here is average plus and minus the standard deviation, P values were accessed with two-sided Student T-test (*p<0.05, **p<0.01, ***p<0.001). Each experiment was repeated independently 3 times.
Supplementary Figure 5 Perinuclear adhesion are significantly enriched β1-integrin, α-actinin and zyxin and show high nanopillar deflections.
a, XY-confocal image from the basal plane of LMNA null MEFs immunostained for integrin β1 (red), cell nucleus (DAPI, blue), lamin A (eGFP tagged, green) (top) and actin (Phalloidin, yellow) (bottom). Scale bars 10 μm. Together with respective intensity of line profile along the white dash line from LMNA fibroblast not expressing (no rescue) versus expressing lamin A eGFP. Upon transient lamin A rescue, there is an increase of integrin β1 recruitment to the perinuclear region even though the apical actin is still not fully reorganized. b,αvβ3-integrin distribution and clustering for the same REF adhering simultaneously to a nanopillar array and flat surface. XY-confocal image of the basal plane of a single fibroblast (cell edge marked in blue dashed line) expressing eGFP tagged αvβ3-integrin (green) adhering to a nanopillar array (inside of white dash-line) and flat surface (DIC image, grey) (left). Zoom-in from boxed-in regions (yellow squares), showing clusters of eGFP tagged αvβ3-integrin (green) formed in a region of the cell adhering to nanopillars (top right) or to the flat surface (bottom right). Scale bars are 20 and 10 µm. c, Confocal image of a MEF on a flat (left) versus nanopillar surface (right), immunostained for actin (red), zyxin (green) and the cell nucleus (DAPI, blue) (n=10 cells).. Scale bar 20 μm. wt and LMNA nul MEFs were cultured on the nanopillar arrays for 12h. d, XY-confocal images of the basal plane of the indicated cell type immunostained for α-actinin (purple), actin (yellow) and nucleus (DAPI, blue) (left). Together with the normalized intensity of α-actinin as function of distance along the white dotted arrow shown in the respective immunostained images. Scale bar 10 μm. e. Hyperstack images from α-actinin immunostainings color-coded according to depth (from bottom (pillar bottom) to top (of the cell) of the indicated cell types (left). Together with the average signal intensity (grey) of α-actinin (each image over a total of 2 μm Z-stack, boxed in with respective colour code according to z- position from violet, to red and finally yellow), from the bottom (violet box) to the top (yellow box) (left). Scale bar 10 μm. (n=10 cells) f, XY-confocal image of the basal plane of the indicated cell type immunostained for zyxin (cyan), actin (yellow) and nucleus (DAPI, blue) (left). Together with normalized intensity of zyxin as function of distance along the white dotted arrow shown in the respective immunostained images. Scale bar 10 μm. g, Similar representation for zyxin immunostaining as in e (n=10 cells). Each experiment was repeated independently 3 times.
Supplementary Figure 6 Phospho-Myosin Light Chain (P-MLC) and tropomyosin (Tpm) are significantly enriched in the peripheral region.
wt and LMNA null MEFs were cultured on the nanopillar arrays for 12h a, Single XY-confocal image of the basal plane of the indicated cell type immunostained for phosphorylated myosin (grey) and nucleus (DAPI, blue) (left). b, Hyperstack images of phosphorylated myosin color-coded according to depth (from bottom (pillar bottom) to top (of the cell) (right). Scale bar 20 μm. c, Normalized intensity of tropomyosin as function of distance along the white dotted arrow shown in the respective immunostained images (right) (n=10 cells). c, Single XY-confocal image of the basal plane of the indicated cell type immunostained for trompomyosin (grey) and nucleus (DAPI, blue) (left) d, Similar representation for trompomyosin immunostainings as in b. Both phosphorylated myosin and tropomyosin signals are mostly enriched at the periphery. For trompomyosin no significant difference is observed between wt and LMNA MEFs (n=10 cells). Each experiment was repeated independently 3 times.
Supplementary information
Supplementary Information
Supplementary Figures 1–6, Supplementary Video legends.
Videos
Supplementary Video 1
REF migration on Fn-coated nanopillar array.
Supplementary Video 2
Fibroblast on Fn-coated nanopillar versus micropost arrays.
Supplementary Video 3
Colourimetric displacement map of perinuclear nanopillars underneath a fibroblast.
Supplementary Video 4
Colourimetric displacement map of perinuclear nanopillars underneath a fibroblast.
Supplementary Video 5
Zoomed-in views of perinuclear versus peripheral nanopillar remodelling.
Supplementary Video 6
Disruption of the actin cap by Latrunculin B treatment.
Supplementary Video 7
Nanopillar clustering induced by stress fibres that form the actin cap.
Supplementary Video 8
Actin-cap-associated perinuclear nanopillar displacements during cell migration.
Supplementary Video 9
Live imaging of WT MEFs expressing RFP-tagged zyxin on a nanopillar array.
Supplementary Video 10
Zoomed-in views of the dynamic recruitment of zyxin to sites of perinuclear nanopillar cluster remodelling.
Supplementary Video 11
3D reconstruction of the actin cap of a WT MEF cultured on Fn-coated nanopillar array for 90 min.
Supplementary Video 12
3D reconstruction of a REF cultured on Fn-coated nanopillar array for 10 min and immunostained for actin (red).
Rights and permissions
About this article
Cite this article
Shiu, JY., Aires, L., Lin, Z. et al. Nanopillar force measurements reveal actin-cap-mediated YAP mechanotransduction. Nat Cell Biol 20, 262–271 (2018). https://doi.org/10.1038/s41556-017-0030-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-017-0030-y
This article is cited by
-
From gene to mechanics: a comprehensive insight into the mechanobiology of LMNA mutations in cardiomyopathy
Cell Communication and Signaling (2024)
-
Retinoic acid and proteotoxic stress induce AML cell death overcoming stromal cell protection
Journal of Experimental & Clinical Cancer Research (2023)
-
Nanotube patterning reduces macrophage inflammatory response via nuclear mechanotransduction
Journal of Nanobiotechnology (2023)
-
Aberrant MET activation impairs perinuclear actin cap organization with YAP1 cytosolic relocation
Communications Biology (2023)
-
Biophysics in tumor growth and progression: from single mechano-sensitive molecules to mechanomedicine
Oncogene (2023)