Abstract
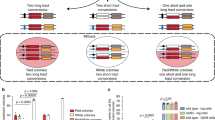
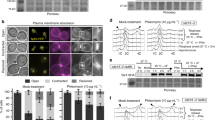
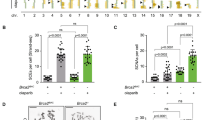
The resolution of joint molecules that link recombining sister chromatids is essential for chromosome segregation. Here, we determine the fate of unresolved recombination intermediates arising in cells lacking two nucleases required for resolution (GEN1 –/– knockout cells depleted of MUS81). We find that intermediates persist until mitosis and form a distinct class of anaphase bridges, which we term homologous recombination ultra-fine bridges (HR-UFBs). HR-UFBs are distinct from replication stress-associated UFBs, which arise at common fragile sites, and from centromeric UFBs. HR-UFBs are processed by BLM helicase to generate single-stranded RPA-coated bridges that are broken during mitosis. In the next cell cycle, DNA breaks activate the DNA damage checkpoint response, and chromosome fusions arise by non-homologous end joining. Consequently, the cells undergo cell cycle delay and massive cell death. These results lead us to present a model detailing how unresolved recombination intermediates can promote DNA damage and chromosomal instability.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Uhlmann, F. SMC complexes: from DNA to chromosomes. Nat. Rev. Mol. Cell Biol. 17, 399–412 (2016).
Chan, K. L. & Hickson, I. D. New insights into the formation and resolution of ultra-fine anaphase bridges. Sem. Cell Dev. Biol. 22, 906–912 (2011).
Baumann, C., Korner, R., Hofmann, K. & Nigg, E. A. PICH, a centromere-associated SNF2 family ATPase, is regulated by PLK1 and required for the spindle checkpoint. Cell 128, 101–114 (2007).
Chan, K. L., North, P. S. & Hickson, I. D. BLM is required for faithful chromosome segregation and its localization defines a class of ultrafine anaphase bridges. EMBO J. 26, 3397–3409 (2007).
Chan, K. L. & Hickson, I. D. On the origins of ultra-fine anaphase bridges. Cell Cycle 8, 3065–3066 (2009).
Germann, S. M. et al. TopBP1/Dpb11 binds DNA anaphase bridges to prevent genome instability. J. Cell Biol. 204, 45–59 (2014).
Liu, Y., Nielsen, C. F., Yao, Q. & Hickson, I. D. The origins and processing of ultra fine anaphase DNA bridges. Curr. Opin. Genet. Dev. 26, 1–5 (2014).
Nielsen, C. F. et al. PICH promotes sister chromatid disjunction and co-operates with topoisomerase II in mitosis. Nat. Commun. 6, 8962 (2015).
Wang, L. H., Schwarzbam, T., Speicher, M. R. & Nigg, E. A. Persistence of DNA threads in human anaphase cells suggests late completion of sister chromatid decatenation. Chromosoma 117, 123–135 (2008).
Chan, K. L., Palmai-Pallag, T., Ying, S. M. & Hickson, I. D. Replication stress induces sister-chromatid bridging at fragile site loci in mitosis. Nat. Cell Biol. 11, 753–760 (2009).
Naim, V. & Rosselli, F. The FANC pathway and BLM collaborate during mitosis to prevent micro-nucleation and chromosome abnormalities. Nat. Cell Biol. 11, 761–768 (2009).
Naim, V., Wilhelm, T., Debatisse, M. & Rosselli, F. ERCC1 and MUS81-EME1 promote sister chromatid separation by processing late replication intermediates at common fragile sites during mitosis. Nat. Cell Biol. 15, 1008–1015 (2013).
Ying, S. M. et al. MUS81 promotes common fragile site expression. Nat. Cell Biol. 15, 1001–1007 (2013).
Barefield, C. & Karlseder, J. The BLM helicase contributes to telomere maintenance through processing of late-replicating intermediate structures. Nucl. Acids Res. 40, 7358–7367 (2012).
Maciejowski, J., Li, Y., Bosco, N., Campbell, P. J. & de Lange, T. Chromothripsis and kataegis induced by telomere crisis. Cell 163, 1641–1654 (2015).
Nera, B., Huang, H.-S., Lai, T. & Xu, L. X. Elevated levels of TRF2 induce telomeric ultrafine anaphase bridges and rapid telomere deletions. Nat. Commun. 6, 10132 (2015).
Sarbajna, S., Davies, D. & West, S. C. Roles of SLX1-SLX4, MUS81-EME1 and GEN1 in avoiding genome instability and mitotic catastrophe. Genes Dev. 28, 1124–1136 (2014).
Wu, L. & Hickson, I. D. The Bloom’s syndrome helicase suppresses crossing over during homologous recombination. Nature 426, 870–874 (2003).
Chen, X. B. et al. Human MUS81-associated endonuclease cleaves Holliday junctions in vitro. Mol. Cell 8, 1117–1127 (2001).
Ciccia, A., Constantinou, A. & West, S. C. Identification and characterization of the human MUS81/EME1 endonuclease. J. Biol. Chem. 278, 25172–25178 (2003).
Fekairi, S. et al. Human SLX4 is a Holliday junction resolvase subunit that binds multiple DNA repair/recombination endonucleases. Cell 138, 78–89 (2009).
Munoz, I. M. et al. Coordination of structure-specific nucleases by human SLX4/BTBD12 is required for DNA repair. Mol. Cell 35, 116–127 (2009).
Svendsen, J. M. et al. Mammalian BTBD12/SLX4 assembles a Holliday junction resolvase and is required for DNA repair. Cell 138, 63–77 (2009).
Ip, S. C. Y. et al. Identification of Holliday junction resolvases from humans and yeast. Nature 456, 357–361 (2008).
Rass, U. et al. Mechanism of Holliday junction resolution by the human GEN1 protein. Genes Dev. 24, 1559–1569 (2010).
Chan, Y. W. & West, S. C. GEN1 promotes Holliday junction resolution by a coordinated nick and counter-nick mechanism. Nucl. Acids Res. 43, 10882–10892 (2015).
Wyatt, H. D. M., Laister, R. C., Martin, S. R., Arrowsmith, C. H. & West, S. C. The SMX DNA repair tri-nuclease. Mol. Cell 65, 848–860 (2017).
Wyatt, H. D. M., Sarbajna, S., Matos, J. & West, S. C. Coordinated actions of SLX1-SLX4 and MUS81-EME1 for Holliday junction resolution in human cells. Mol. Cell 52, 234–247 (2013).
Wechsler, T., Newman, S. & West, S. C. Aberrant chromosome morphology in human cells defective for Holliday junction resolution. Nature 471, 642–646 (2011).
Castor, D. et al. Cooperative control of Holliday junction resolution and DNA repair by the SLX1 and MUS81-EME1 nucleases. Mol. Cell 52, 221–233 (2013).
Garner, E., Kim, Y., Lach, F. P., Kottemann, M. C. & Smogorzewska, A. Human GEN1 and the SLX4-associated nucleases MUS81 and SLX1 are essential for the resolution of replication-induced Holliday junctions. Cell Rep. 5, 207–215 (2013).
Duda, H. et al. A mechanism for controlled breakage of under-replicated chromosomes during mitosis. Dev. Cell 39, 740–755 (2016).
Matos, J., Blanco, M. G., Maslen, S. L., Skehel, J. M. & West, S. C. Regulatory control of the resolution of DNA recombination intermediates during meiosis and mitosis. Cell 147, 158–172 (2011).
Chan, Y. W. & West, S. C. Spatial control of the GEN1 Holliday junction resolvase ensures genome stability. Nat. Commun. 5, 5844 (2014).
Nair, N., Castor, D., Macartney, T. & Rouse, J. Identification and characterization of MUS81 point mutations that abolish interaction with the SLX4 scaffold protein. DNA Rep. 24, 131–137 (2014).
Minocherhomji, S. et al. Replication stress activates DNA repair synthesis in mitosis. Nature 528, 286–290 (2015).
Schlacher, K. et al. Double-strand break repair-independent role for BRCA2 in blocking stalled replication fork degradation by MRE11. Cell 145, 529–542 (2011).
Lukas, C. et al. 53BP1 nuclear bodies form around DNA lesions generated by mitotic transmission of chromosomes under replication stress. Nat. Cell Biol. 13, 243–253 (2011).
Harrigan, J. A. et al. Replication stress induces 53BP1-containing OPT domains in G1 cells. J. Cell Biol. 193, 97–108 (2011).
Klein Douwel, D. et al. XPF-ERCC1 acts in unhooking DNA interstrand crosslinks in cooperation with FANCD2 and FANCP/SLX4. Mol. Cell 54, 460–471 (2014).
Hodskinson, M. R. G. et al. Mouse SLX4 is a tumour suppressor that stimulates the activity of the nuclease XPF-ERCC1 in DNA crosslink repair. Mol. Cell 54, 472–484 (2014).
Burrell, R. A. et al. Replication stress links structural and numerical cancer chromosomal instability. Nature 494, 492–496 (2013).
Ke, Y. et al. PICH and BLM limit histone association with anaphase centromeric threads and promote their resolution. EMBO J. 30, 3309–3321 (2011).
Hengeveld, R. C. et al. Rif1 Is required for resolution of ultrafine DNA bridges in anaphase to ensure genomic stability. Dev. Cell 34, 466–474 (2015).
Houchmandzadeh, B., Marko, J. F., Chatenay, D. & Libchaber, A. Elasticity and structure of eukaryote chromosomes studied by micromanipulation and micropipette aspiration. J. Cell Biol. 139, 1–12 (1997).
Alexander, S. P. & Rieder, C. L. Chromosome motion during attachment to the vertebrate spindle: initial saltatory-like behavior of chromosomes and quantitative analysis of force production by nascent kinetochore fibers. J. Cell Biol. 113, 805–815 (1991).
Grandbois, M., Beyer, M., Rief, M., Clausen-Schaumann, H. & Gaub, H. E. How strong is a covalent bond? Science 283, 1727–1730 (1999).
Bustamante, C., Smith, S. B., Liphardt, J. & Smith, D. R. Single-molecule studies of DNA molecules. Curr. Opin. Struct. Biol. 10, 279–285 (2000).
Mullins, J. M. & Biesele, J. J. Terminal phase of cytokinesis in D-98s cells. J. Cell Biol. 73, 672–684 (1977).
Steigemann, P. et al. Aurora B-mediated abscission checkpoint protects against tetraploidization. Cell 136, 473–484 (2009).
Janssen, A., van der Burg, M., Szuhai, K., Kops, G. J. & Medema, R. H. Chromosome segregation errors as a cause of DNA damage and structural chromosome aberrations. Science 333, 1895–1898 (2011).
Ly, P. et al. Selective Y centromere inactivation triggers chromosome shattering in micronuclei and repair by non-homologous end joining. Nat. Cell Biol. 19, 68–75 (2017).
Thompson, S. L. & Compton, D. A. Examining the link between chromosomal instability and aneuploidy in human cells. J. Cell Biol. 180, 665–672 (2008).
Tanno, Y. et al. The inner centromere-shugoshin network prevents chromosomal instability. Science 349, 1237–1240 (2015).
Raderschall, E. et al. Formation of higher-order nuclear RAD51 structures is functionally linked to p21 expression and protection from DNA damage-induced apoptosis. J. Cell Sci. 115, 153–164 (2002).
Klein, H. L. The consequences of RAD51 overexpression for normal and tumor cells. DNA Repair 7, 686–693 (2008).
Marsden, C. G. et al. The tumor-associated variant RAD51 G151D induces a hyper-recombination phenotype. PLoS Genet. 12, e1006208 (2016).
Rodriguez-Lopez, A. M., Whitby, M. C., Borer, C. M., Bachler, M. A. & Cox, L. S. Correction of proliferation and drug sensitivity defects in the progeriod Werner’s Syndrome by Holliday junction resolution. Rejuvenation Res. 10, 27–40 (2007).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Essletzbichler, P. et al. Megabase-scale deletion using CRISPR/Cas9 to generate a fully haploid human cell line. Genome Res. 24, 2059–2065 (2014).
Paliwal, S., Kanagaraj, R., Sturzenegger, A., Burdova, K. & Janscak, P. Human RECQ5 helicase promotes repair of DNA double-strand breaks by synthesis-dependent strand annealing. Nucl. Acids Res. 42, 2380–2390 (2013).
Chowdhury, D. et al. The exonuclease TREX1 is in the SET complex and acts in concert with NM23-H1 to degrade DNA during granzyme A-mediated cell death. Mol. Cell 23, 133–142 (2006).
Fugger, K. et al. Human FBH1 helicase contributes to genome maintenance via pro- and anti-recombinase activities. J. Cell Biol. 186, 655–663 (2009).
Petermann, E., Orta, M. L., Issaeva, N., Schultz, N. & Helleday, T. Hydroxyurea-stalled replication forks become progressively inactivated and require two different RAD51-mediated pathways for restart and repair. Mol. Cell 37, 492–502 (2010).
Acknowledgements
We thank members of the West Lab for their help and encouragement, and K. L. Chan (University of Sussex, UK) for sharing unpublished data. This work was supported by the Francis Crick Institute (FC10212), the European Research Council (ERC-ADG-249145 and ERC-ADG-666400) and the Louis-Jeantet Foundation. The Francis Crick Institute receives core funding from Cancer Research UK, the Medical Research Council and the Wellcome Trust. K.F. is the recipient of a fellowship from the Lundbeck Foundation.
Author information
Authors and Affiliations
Contributions
Y.W.C. carried out the experiments and was assisted by K.F with the fibre assays and some of the immunofluorescence experiments. Y.W.C. and S.C.W. designed the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and Table 1 Legend.
Supplementary Table 1
Statistics Source Data
Rights and permissions
About this article
Cite this article
Chan, Y.W., Fugger, K. & West, S.C. Unresolved recombination intermediates lead to ultra-fine anaphase bridges, chromosome breaks and aberrations. Nat Cell Biol 20, 92–103 (2018). https://doi.org/10.1038/s41556-017-0011-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-017-0011-1
This article is cited by
-
Histone demethylase KDM2A is a selective vulnerability of cancers relying on alternative telomere maintenance
Nature Communications (2023)
-
Mitotic DNA synthesis in response to replication stress requires the sequential action of DNA polymerases zeta and delta in human cells
Nature Communications (2023)
-
The SUMO–NIP45 pathway processes toxic DNA catenanes to prevent mitotic failure
Nature Structural & Molecular Biology (2023)
-
An Aurora B-RPA signaling axis secures chromosome segregation fidelity
Nature Communications (2023)
-
Effects of aneuploidy on cell behaviour and function
Nature Reviews Molecular Cell Biology (2022)