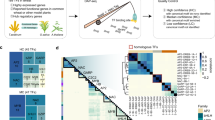
Abstract
Plants synthesize numerous ecologically specialized, lineage-specific metabolites through biosynthetic gene duplication and functional specialization. However, it remains unclear how duplicated genes are wired into existing regulatory networks. We show that the duplicated gene CYP82C2 has been recruited into the WRKY33 regulon and indole-3-carbonylnitrile (ICN) biosynthetic pathway through exaptation of a retroduplicated LINE retrotransposon (EPCOT3) into an enhancer. The stepwise development of a chromatin-accessible WRKY33-binding site on EPCOT3 has potentiated the regulatory neofunctionalization of CYP82C2 and the evolution of inducible defense metabolite 4-hydroxy-ICN in Arabidopsis thaliana. Although transposable elements (TEs) have long been recognized to have the potential to rewire regulatory networks, these results establish a more complete understanding of how duplicated genes and TEs contribute in concert to chemical diversity and pathogen defense.
Similar content being viewed by others
Introduction
Plant secondary or specialized metabolites are essential for plant survival in co-evolving biotic and fluctuating abiotic environments. The evolutionary process of chemical innovation resulted in the collective synthesis of hundreds of thousands of ecologically specialized, mostly lineage-specific metabolites1,2,3. Plant-specialized metabolic enzymes are ultimately produced from primary metabolic enzymes through gene duplication and subsequent functional divergence of one or both paralogs to produce enzymes with altered expression patterns and/or protein functions3,4,5. They are also often organized into transcription factor (TF) regulons of co-regulated genes for optimal timing, amplitude, and tissue-specific pathway gene expression and subsequent metabolite accumulation6,7.
Changes in cis-regulatory modules such as enhancers and promoters can accelerate the capture of duplicated genes into regulons, thus driving phenotypic diversity8,9,10. Enhancers consist of TF binding sites (TFBSs) and are derived either through mutation or co-option of a TFBS-carrying transposable element (TE)10,11. TE exaptations are hypothesized to be responsible for the rapid transcriptional rewiring of gene regulatory networks in ancient lineages of vertebrates12,13,14 and plants15, but general understandings of the physiological significance of this rewiring are greatly limited.
Bacteria elicit two primary immune defense modes in plants, pattern- and effector-triggered immunity (PTI and ETI)16. Pathogenic bacteria additionally compromise PTI via specific virulence effector proteins (effector-triggered susceptibility, ETS)16. PTI involves the extracellular perception of conserved molecules known as microbe-associated molecular patterns (MAMPs), whereas ETI involves the cytosolic perception of effectors. Although ETI results in the formation of more rapid and robust pathogen-specific responses including a form of programmed cell death known as the hypersensitive response (HR)16, both result in the ability of naive host cells to generate, through non-self perception and subsequent transcriptional reprogramming, pathogen-inducible specialized metabolites necessary for defense17,18,19.
Three pathogen-inducible tryptophan (Trp)-derived defense metabolites— 4-methoxyindol-3-ylmethylglucosinolate (4M-I3M)19,20, camalexin21,22, and 4-hydroxyindole-3-carbonylnitrile (4OH-ICN)23—are known to expand innate immunity in Arabidopsis thaliana. Their biosynthetic pathways share an early step, which is the conversion of Trp to indole-3-acetaldoxime (IAOx) via the genetically redundant P450 monooxygenases CYP79B2 and CYP79B323,24,25,26 (Fig. 1a). The camalexin and 4OH-ICN pathways additionally share the conversion of IAOx to indole-3-cyanohydrin (ICY) by partially redundant P450s CYP71A12 and CYP71A1323,27,28 (Fig. 1a). CYP71A13 and CYP71B15/PAD3 catalyze further reactions, leading to camalexin production28,29, whereas the flavin-dependent oxidase FOX1/AtBBE3 and P450 CYP82C2 convert ICY to 4OH-ICN (Fig. 1a)23. 4M-I3M is widely distributed across the mustard family (Brassicaceae), whereas camalexin is restricted to the Camelineae tribe of Brassicaceae30. The evolutionary conservation of 4OH-ICN has not yet been investigated.
4OH-ICN is synthesized under ETI-like responses. a Schematic of tryptophan (L-Trp)-derived specialized metabolism in A. thaliana. White arrows denote the presence of additional enzymes. ANI, aci-nitro indole; ICY, indole cyanohydrin. b. LC-DAD-FLD-MS analysis of camalexin (top), ICN (middle), and 4OH-ICN (bottom) in seedlings elicited with indicated MAMPs and bacterial strains for 27 h. Data represent mean ± SE of four replicates of 15 ± 2 seedlings each. Different letters denote statistically significant differences (P < 0.05, one-factor ANOVA coupled to Tukey’s test). ICN totals consist of the sum of ICN and methanolic degradation product ICA-ME. 4OH-ICN totals consist of the sum of aqueous and methanolic degradation products 4OH-ICA-ME and 4OH-ICA, respectively
The TF WRKY33 regulates the pathogen-inducible biosynthesis of camalexin in A. thaliana31,32 and its orthologs regulate numerous unrelated specialized metabolites in other flowering plant lineages33. The group I class of WRKYs to which WRKY33 belongs is an ancient clade of regulators; orthologs in the green alga Chlamydomonas reinhardtii may be ancestral to all higher plant WRKYs33,34. Although all WRKY TFs bind to the W-box core sequence [TTGAC(T/C)], WRKY33 preferentially binds W-boxes that are within 500 nt of the WRKY33-specific motif [(T/G)TTGAAT])35.
Here we show that a recent, lineage-specific TE exaptation has resulted in the expansion of a core regulon within the framework of Arabidopsis Trp-derived defense metabolism. Specifically, the LINE retrotransposon EPCOT3 has retroduplicated from a WRKY33-TFBS-carrying progenitor and inserted upstream of the newly duplicated gene CYP82C2. Subsequent chromatin remodeling in A. thaliana has led EPCOT3 to become a bona fide enhancer with demonstrated biochemical, regulatory, physiological, and fitness-promoting characteristics by way of WRKY33-binding and pathogen-responsive CYP82C2 transcription, 4OH-ICN biosynthesis, and antibacterial defense.
Results
4OH-ICN requires ETI-like responses
To identify the major Trp-derived specialized metabolites synthesized in ETI in A. thaliana, we compared host transcriptional and metabolic responses to the PTI-eliciting bacterial MAMPs flg22, elf26, and fungal MAMP chitosan; the PTI/ETS-eliciting pathogens Pseudomonas syringae pv. tomato DC3000 (Pto DC3000 or Pst); P. syringae pv. maculicola ES4326 (Pma); and the ETI-eliciting pathogens Pst avrRpm1 (Psta), Pst avrRpt2, Pst avrRps4, Pma M2, and Pma avrRpt2 under similar conditions as those of previous studies19,36. Psm M2 is an ETI-eliciting strain from which the avrRpm1 gene was originally isolated37. Both flg22 and Psta induced genes involved in camalexin, 4OH-ICN, and 4M-I3M biosynthesis, with camalexin and 4OH-ICN biosynthetic genes having a higher level of induction than those of 4M-I3M in Psta-inoculated plants36 (Supplementary Table 1). On the other hand, metabolite responses between PTI and ETI differed qualitatively. 4M-I3M and its immediate precursor 4-hydroxy-I3M (4OH-I3M) were present in uninfected plants and accumulated to modest levels at the expense of parent metabolite I3M in flg22- and Psta-inoculated plants19 (Supplementary Fig. 1a). By comparison, camalexin, ICN, and 4OH-ICN were absent in uninfected plants and accumulated to high levels upon inoculation with ETI-inducing pathogens (Fig. 1b and Supplementary Fig. 1b). Furthermore, camalexin, ICN, and 4OH-ICN metabolism was greatly diminished, and 4M-I3M, 4OH-I3M, and I3M levels were mostly unchanged in the rpm1 mutant (Supplementary Fig. 1), which is impaired in ETI recognition of Psta40. By contrast, camalexin and ICN were largely at low-to-undetectable levels in plants treated with saturating concentrations of the bacterial MAMPs flg22 and elf2638,39 and PTI/ETS-eliciting pathogens, with 4OH-ICN not detected in most cases (Fig. 1b). One exception was the fungal MAMP chitosan. Chitosan (150 μg/mL) induced high levels of camalexin and detectable levels of ICN (Fig. 1b), consistent with previous observations of camalexin biosynthetic gene upregulation41. Higher chitosan concentrations (≥ 200 μg/mL) have been shown to induce HR-like cell death in Arabidopsis42, a phenomenon commonly observed for ETI16. To our surprise, 300 μg/mL chitosan additionally induced detectable levels of 4OH-ICN (Fig. 1b). These results suggest that 4OH-I3M, 4M-I3M, camalexin, and ICN are synthesized in response to multiple PTI elicitors, whereas 4OH-ICN biosynthesis is specific to ETI-like responses.
WRKY33 is required to activate 4OH-ICN in response to Psta
4OH-ICN biosynthetic genes are highly co-expressed with each other23 and with camalexin biosynthetic genes (Supplementary Table 2), which are in the WRKY33 regulon31,43. To determine whether 4OH-ICN biosynthetic genes are also in the WRKY33 regulon, we compared camalexin, ICN, and 4OH-ICN levels between wild-type and a wrky33 loss-of-function mutant that encodes two differently truncated proteins44 (Fig. 2a). Consistent with a previous report31, wrky33 was impaired in camalexin biosynthesis in response to Psta and Pst avrRps4 (Fig. 2b and Supplementary Fig. 2a). The wrky33 mutant was similarly impaired in 4OH-ICN biosynthesis (Fig. 2b and Supplementary Fig. 2a). These results indicate that WRKY33 is required for camalexin and 4OH-ICN biosynthesis in response to multiple ETI elicitors.
Intraspecific variation in WRKY33 affects 4OH-ICN and immunity. a Schematic of WRKY33 proteins in Col-0, Col-0 wrky33, Ler-1, and Di-G. Black boxes denote WRKY domains (W), nuclear localization signal (NLS), or C-terminal domain (CTD). b LC-DAD-MS analysis of camalexin, ICN, and 4OH-ICN in seedlings inoculated with Psta for 24 h. Data represent mean ± SE of four replicates of 15 ± 2 seedlings each. c Bacterial growth analysis of Pst in surface-inoculated leaves. Middle and right panels were pre-treated with 20 μM dex for 6–8 h. Data represent mean ± SE of 4 (left); 11, 6, 7, 7, 7 (middle); 6, 6, 8 (right) replicates of 15 ± 2 seedlings each. CFU, colony-forming units. Different letters in b, c denote statistically significant differences (P < 0.05, one-factor ANOVA coupled to Tukey’s test). Source data of Figs. 2b and 2c are provided as a Source Data file
To confirm that WRKY33 is required to activate the 4OH-ICN pathway, we used a two-component glucocorticoid-inducible system to generate wrky33 plants that in the presence of the glucocorticoid hormone dexamethasone (dex) express a wild-type copy of WRKY33 with a C-terminal fusion to 1× flag epitope (wrky33/DEX:WRKY33-flag; Supplementary Fig. 2b–c). Induced expression of WRKY33-flag restored camalexin and 4OH-ICN biosynthesis in Psta-challenged wrky33 plants to greater than wild-type levels (Supplementary Fig. 2d). These results indicate that WRKY33 is required to activate camalexin and 4OH-ICN biosynthesis in response to Psta.
Natural variation in WRKY33 affects metabolism and defense
Intraspecific variation in TFs can contribute to gain or loss of phenotypes, such as branching in maize45 or pelvic loss in three-spined stickleback fish46. In addition, the wide variation in camalexin biosynthesis reported among natural accessions of A. thaliana47 suggests that a similar variation in 4OH-ICN biosynthesis may exist. To identify additional transcriptional activators of 4OH-ICN biosynthesis that otherwise might be refractory to traditional genetic approaches, we compared intraspecific variation in Psta-induced camalexin, ICN, and 4OH-ICN among 35 re-sequenced accessions and wrky33 (Col-0 accession). We found camalexin and 4OH-ICN levels to be positively correlated among accessions (R2 = 0.37; Supplementary Fig. 3a), lending further support to their co-regulation by WRKY33. Accession Dijon-G (Di-G) was identified to produce less camalexin and 4OH-ICN and more ICN than its near-isogenic relatives, the Landsberg accessions Ler-0 and Ler-1 (Fig. 2b and Supplementary Fig. 3a–b). In addition, differences observed in the metabolite response between Landsberg accessions and Di-G most closely resembled those between Col-0 and wrky33 mutant (Fig. 2b and Supplementary Fig. 3a). These results led us to hypothesize that genetic variation in a regulatory gene, as opposed to an immune signaling gene, is responsible for the metabolite phenotypes observed in Di-G. To test this hypothesis, genetic variation between Di-G and three sequenced Landsberg accessions (La-0, Ler-0, and Ler-1) were used to identify 354 genes that were differentially mutated to high effect in Di-G (Supplementary Fig. 3c). Twenty-eight of these mutated Di-G genes were annotated by Gene Ontology to have roles in defense, including WRKY33 (Supplementary Table 3). We confirmed by Sanger sequencing that Di-G WRKY33 harbors a nonsense mutation early in the N-terminal DNA-binding motif (Fig. 2a), likely abolishing protein function. Our findings indicate that camalexin and 4OH-ICN are sensitive to intraspecific variation in WRKY33.
Camalexin and 4OH-ICN promote plant fitness by contributing non-redundantly to pathogen defense against the fitness-reducing Pst23. To confirm that disease resistance to Pst is also sensitive to intraspecific variation in WRKY33, we measured bacterial growth in adult leaves of wrky33, Di-G, and their respective (near-)isogenic accessions Col-0 and Ler-1. wrky33 and Di-G were more susceptible to Pst than their (near-)isogenic relatives and comparable to the 4OH-ICN biosynthetic mutant cyp82C223 (Fig. 2c)
We additionally generated wrky33 plants that in the presence of dex express a wild-type copy of WRKY33 with a C-terminal fusion to a larger 6× myc epitope (wrky33/DEX:WRKY33-myc; Supplementary Fig. 4a–c). Induced expression of WRKY33-myc in wrky33 and Di-G plants restored and/or exceeded Col-0 and Ler-1 levels of resistance to Pst (Fig. 2c) and of Psta-induced camalexin and ICN, respectively (Supplementary Fig. 4d–e). Together, our results support a role of WRKY33 in pathogen defense as an activator of Trp-derived specialized metabolism.
WRKY33 activates 4OH-ICN biosynthesis
To confirm that the 4OH-ICN biosynthetic pathway is in the WRKY33 regulon, we first compared WRKY33, CYP71A13, CYP71B15, FOX1, and CYP82C2 transcript levels among WT, wrky33, wrky33/DEX:WRKY33-flag, and wrky33/DEX:WRKY33-myc. Consistent with previous reports31, CYP71A13, CYP71B15, and FOX1 expression was downregulated in wrky33 plants in response to Psta and upregulated in both wrky33/DEX:WRKY33-flag and wrky33/DEX:WRKY33-myc (Fig. 3a) (Supplementary Figs. 4f and 5a). Interestingly, CYP82C2 expression and 4OH-ICN production were restored in wrky33/DEX:WRKY33-flag but not wrky33/DEX:WRKY33-myc or Di-G/DEX:WRKY33-myc plants (Figs. 2d and 3a, and Supplementary Fig. 4d–f), likely due to the interference of the larger myc tag with the WRKY33 C-terminus, a region previously linked with transactivation activity48. These transcriptional and metabolic findings indicate that WRKY33 mediates camalexin and 4OH-ICN biosynthesis in response to pathogen effectors.
WRKY33 directly activates 4OH-ICN biosynthetic genes. a qPCR analysis of 4OH-ICN regulatory and biosynthetic genes in seedlings co-treated with 20 μM dex and Psta for 9 and 12 h. Different letters denote statistically significant differences (P < 0.05, one-factor ANOVA coupled to Tukey’s test). Lowercase and uppercase letters denote comparisons across 9 and 12 h time points, respectively. Data represent mean ± SE of 4, 5, 4, 5 (9 h) and 6, 6, 6, 5 (12 h) replicates of 15 ± 2 seedlings each. b Schematic of FOX1 and CYP82C2 loci, indicating nt positions of W-box-containing regions (W). c ChIP-PCR analysis of W-box-containing regions upstream of FOX1 and CYP82C2 in wrky33/DEX:WRKY33-flag plants co-treated with 20 μM dex (D) or mock solution (M) and Psta for 9 h. Dashed line represents the fivefold cutoff between weak and strong TF-DNA interactions. Data represent median ± SE of four replicates of 15 ± 2 seedlings each. Source data of Figs. 3a and 3c are provided as a Source Data file
We then tested for WRKY33-binding to W-box-containing regions upstream of camalexin and 4OH-ICN biosynthetic genes in dex-treated and Psta-infected wrky33/DEX:WRKY33-flag seedlings by chromatin immunoprecipitation (ChIP)-PCR. WRKY33 reportedly binds upstream of 4OH-ICN biosynthetic gene CYP71A12 to a W-box region that also contains three WRKY33-specific motifs49. We additionally observed that Psta-induced WRKY33 bound strongly (greater than fivefold enrichment) upstream of 4OH-ICN biosynthetic genes FOX1 and CYP82C2 to W-box regions that also contain one to three WRKY33-specific motifs (W2 and W4, respectively; Fig. 3b, c and Supplementary Fig. 5b). Together with our expression analysis, our findings indicate that WRKY33 uses preferred WRKY33-binding sites to directly activate 4OH-ICN biosynthetic genes in response to pathogen effectors.
Interestingly, Psta-induced WRKY33 did not bind to the W5 region upstream of CYP82C2 (Fig. 3b, c), a W-box region that does not contain any WRKY33-specific motifs and is just upstream of neighboring gene of unknown function At4g31960. WRKY33 reportedly binds to W5 in response to flg2249 and Botrytis cinerea35. By contrast, Psta-induced WRKY33 bound strongly to the W1 region upstream of CYP71B15 (Supplementary Fig. 5c–d), a W-box region that also does not contain any WRKY33-specific motifs. WRKY33 reportedly binds to a region encompassing W1 in response to flg2231,49 and Psta31. These findings suggest that WRKY33 may use W-box extended motifs or alternative specificity motifs to target camalexin biosynthetic genes in response to pathogen effectors, or 4OH-ICN biosynthetic genes in response to MAMPs or fungal pathogens.
CYP82C2 underwent regulatory neofunctionalization
CYP82C2 catalyzes the last step in 4OH-ICN biosynthesis, hydroxylating ICN to form 4OH-ICN23, and likely was the last 4OH-ICN pathway gene to be recruited to the WRKY33 regulon in A. thaliana. To explore the phylogenetic distribution pattern of 4OH-ICN biosynthesis, we profiled ICN and 4OH-ICN metabolites in close and distant relatives of A. thaliana in response to Psta. Although ICN biosynthesis was observed across multiple close relatives, 4OH-ICN was only detected in A. thaliana (Fig. 4a and Supplementary Fig. 6a). This result suggests that 4OH-ICN manifests a species-specific diversification of pathogen-inducible Trp-derived metabolism in the mustard family.
Regulatory neofunctionalization of CYP82C2. a (Left) Phylogenetic species tree. (Right) HPLC-DAD analysis of 4OH-ICN in seedlings inoculated with Psta for 30 h. Data in box plots represent median (center line), 25th percentile (lower box limit), 75th percentile (upper box limit), and full range of variation (whiskers) for n = 13, 9, 3, 6, 6, 6, 6, 3, 3, 6 replicates of 15 ± 2 seedlings each. Data were pooled from several independent experiments with A. thaliana as the positive control. 4OH-ICA and 4OH-ICA-ME are aqueous and methanolic degradation products of 4OH-ICN, respectively. DW, dry weight; n.d., not detected. b (Left) Phylogenetic species tree. (Right) Synteny map of CYP82C genes. Gray arrows or rectangles represent non-CYP82C genes. Gray dotted lines represent large ( > 500 nt) sequence gaps. c, d qPCR analysis of 4OH-ICN and sideretin biosynthetic genes in seedlings inoculated with Psta (c) or grown in iron-deficient medium (d). Data in c represent mean ± SE of 3 (12 h Arabidopsis lyrata) or 4 (all other) replicates of 15 ± 2 seedlings each. Data in d represent mean ± SE of 3 replicates of 15 ± 2 seedlings each. Asterisks denote statistically significant differences of stress-treated relative to untreated samples (P < 0.05, two-tailed t-test). Source data of Figs. 4a, 4c, and 4d are provided as a Source Data file
In A. thaliana, CYP82C2 resides in a near-tandem cluster with paralogs CYP82C3 and CYP82C4 (Fig. 4b). We performed phylogenetic and syntenic analyses to identify putative CYP82C2 orthologs in a clade inclusive of ICN-synthesizing species. All identified homologs are syntenic to CYP82C2 or CYP82C4, and encode proteins with > 88% identity to one another (Fig. 4b and Supplementary Fig. 6b–c). CYP82C3 is present only in A. thaliana and, although more similar to CYP82C2 than CYP82C4 in sequence (Fig. 4b and Supplementary Fig. 6b), it is not functionally redundant with CYP82C223. CYP82C4 is required for the biosynthesis of sideretin, a widely conserved, phenylalanine-derived metabolite required for iron acquisition50. CYP82C4 has syntenic orthologs in the mustard family (Fig. 4b and Supplementary Fig. 6b), correlating with the distribution of sideretin biosynthesis50. By contrast, CYP82C2 has syntenic orthologs only within the Arabidopsis genus (Fig. 4b and Supplementary Fig. 6b). These results suggest that CYP82C2 duplicated from CYP82C4 prior to the formation of the Arabidopsis genus and then acquired a new expression pattern and/or catalytic function prior to A. thaliana speciation ~2 million years later51,52.
CYP82C2 and CYP82C4 were previously characterized to 5-hydroxylate with equal efficiency the specialized metabolite 8-methoxypsoralen, a molecule structurally reminiscent of ICN and sideretin53. The apparent similarities in substrate specificity and catalytic function suggest that CYP82C2 may have diverged from CYP82C4 in expression but not protein function. To test this, we first compared the expression of CYP82C2 and CYP82C4 in A. lyrata and A. thaliana in response to Psta. 4OH-ICN biosynthetic genes CYP79B2, CYP71A12, and FOX1 were upregulated in both species, consistent with the common presence of ICN (Fig. 4a, c). By contrast, CYP82C2 levels were respectively upregulated and unchanged in A. thaliana and A. lyrata, correlating with the distribution of 4OH-ICN in these species (Fig. 4a, c). CYP82C4 expression was unchanged in both species (Fig. 4c). These results indicate that 4OH-ICN biosynthesis is linked with pathogen-induced expression of CYP82C2.
We then compared the aligned upstream sequences of CYP82C2 and CYP82C4 in A. lyrata and A. thaliana, and observed good sequence conservation among orthologs but poor conservation among paralogs (Supplementary Fig. 6d), indicating that sequences upstream of CYP82C4 and CYP82C2 were independently derived. We performed expression analysis in A. thaliana to confirm that CYP82C2 and CYP82C4 have different expression patterns. Consistent with previous reports23,50, CYP82C2 expression is upregulated in response to Psta and unchanged under iron deficiency, whereas CYP82C4 is upregulated under iron deficiency and unchanged in response to Psta (Fig. 4c, d and Supplementary Table 1). Finally, CYP82C4 expression is unchanged in Psta-challenged, dex-induced wrky33 and wrky33/DEX:WRKY33-flag (Supplementary Fig. 6e). Our findings suggest that CYP82C2 diverged from CYP82C4 by acquiring WRKY33 regulation for its pathogen-induced expression.
We next assessed dN/dS ratios along branches of the CYP82C phylogenetic tree (Supplementary Fig. 6b) and found good support for purifying selection acting on CYP82C enzymes (ω = 0.21) and no support for positive selection acting on CYP82C2/3 enzymes (Supplementary Data 1). Lastly, we identified non-conserved amino acid residues among CYP82C homologs and mapped this information onto a homology model of CYP82C2. The protein inner core, which encompasses the active site and substrate channel, is highly conserved among CYP82C homologs (Supplementary Fig. 6f), and is consistent with CYP82C2 and CYP82C4’s reportedly redundant catalytic functions53. Altogether, our findings suggest that CYP82C2 underwent regulatory neofunctionalization, diverging from CYP82C4 in expression but not protein function.
TE EPCOT3 is a CYP82C2 enhancer
WRKY33 regulation of CYP82C2 is mediated by a WRKY33-TFBS in the W4 region (Figs. 3 and 5a; Supplementary Fig. 5c). Preferential WRKY33-binding at this region should also be influenced by chromatin features associated with cis-regulatory elements such as enhancers and basal promoters54. To investigate how CYP82C2 acquired WRKY33-binding for its pathogen-induced expression, we compared the aligned upstream sequences of CYP82C homologs within a clade inclusive of ICN-synthesizing species. We observed three large upstream sequences specific to A. thaliana CYP82C2, hereafter named Eighty-two-C2 Promoter Contained Only in A. Thaliana1-3 (EPCOT1–3; Fig. 5a). EPCOT3 in particular is a 240 nt region that completely encompasses W4 (Fig. 5a), indicating that WRKY33’s regulation of CYP82C2 in response to Psta may be species-specific. Further bioinformatics analysis revealed that EPCOT3 is enriched with the activating histone mark H3K4me2 and lacks the repressive histone mark H3K27me3 (Fig. 5b)55,56, which are epigenetic signatures of an active enhancer57,58,59. Our findings suggest that EPCOT3 functions as an enhancer that mediates WRKY33-binding and activation of CYP82C2 in response to pathogen effectors.
TE EPCOT3 is a CYP82C2 enhancer. a mVISTA plot of CYP82C2 upstream sequence, indicating nt positions of unique (EPCOT1–3; gray boxes) and conserved regions (≥ 70% identity, pink) among homologous sequences. Also indicated are positions of W-boxes (green) and WRKY33-specific motifs (blue) present (solid lines) or absent (dashed lines) in each homologous sequence, known WRKY33 TFBSs (diamonds) and ChIP-tested regions (W1–5). Al, A. lyrata; Ah, Arabidopsis halleri; Bs, Boechera stricta; Cg, Capsella grandiflora; Cr, Capsella rubella; TSS, transcriptional start site. b Epigenetic map of CYP82C2 upstream sequence, indicating positions of significant H3K4me2 (blue–gray bars) and H3K27me3 (purple bars). c (Left) Schematic of EPCOT3 and related LINE retrotransposons in A. thaliana, indicating positions of CYP82C2 and reverse-transcriptase (RT) domains. See also Supplementary Note 1. Dashed box outlines W-boxes (green lines) and/or WRKY33-binding motifs (blue lines) within EPCOT3/EPLs. (Right) Phylogenetic maximum likelihood tree. d (Upper left) Schematic of CYP82C2 and AlCYP82C2 transgenic loci used for WRKY33 transactivation experiments. (Lower left) RT-PCR images of CYP82C2, AlCYP82C2, and NbACTIN1 in N. benthamiana leaves co-transfected with DEX:WRKY33-flag and CYP82C2 or AlCYP82C2 locus, and incubated with 1 μM flg22 and mock solution (0.5% DMSO) or 20 μM dex for 30 h (CYP82C2/AlCYP82C2) or 24 hr (NbACTIN1). Data represent five replicates (three leaf discs each). (Lower right) RT-PCR images of CYP82C2, AlCYP82C2, and EIF4A1 in A. thaliana cyp82C2 protoplasts transfected with CYP82C2 or AlCYP82C2 locus and elicited 6 h with 1 μM flg22. As original CYP82C2 primers detect endogenous transcription downstream of the cyp82C2 T-DNA insertion (see CYP82C2 + cyp82C2-2, second row), a second set of primers (CYP82C2*, Supplementary Data 2) flanking the insertion was used to test WRKY33 transactivation (see CYP82C2, first row). Data represent four replicates of 2.5 × 105 protoplasts each. e ChIP-PCR analysis of W-box-containing regions (W) within EPLs in wrky33/DEX:WRKY33-flag plants co-treated 9 h with 20 μM dex or mock solution and Psta. Data represent median ± SE of four replicates (~210 seedlings each). Dashed line represents fivefold cutoff between weak and strong TF-DNA interactions. Source data of Figs. 5d and 5e are provided as a Source Data file
EPCOT3 contains a 3′-poly-A tail and is flanked by variable-length target site duplications (Fig. 5c and Supplementary Fig. 7a), which are hallmarks of eukaryotic LINE retrotransposons60. LINE retrotransposition (reverse transcription and integration) results in frequent 5′-truncation of retrocopies61. We identified 11 variably truncated retrocopies similar to EPCOT3 throughout the genome (Fig. 5c, Supplementary Fig. 7a–b, and Supplementary Table 4), including Ta22, one of the first LINEs characterized in A. thaliana62. EPCOT3-related LINEs were sorted into two groups roughly correspondent to their phylogenetic placement: EPCOT3-LIKE (EPL) for those with high identity (> 65%) to EPCOT3, and Ta22 or Ta22-LIKE (Ta22L) for the remainder (Supplementary Fig. 7a and Supplementary Table 4). Only Ta22 and Ta22L1 are full-length LINEs (Fig. 5c), presumably encoding the proteins necessary for their own transposition and for the transposition of non-autonomous family members such as EPCOT3. Through synteny analysis, we also identified two species-specific Ta22Ls, but no EPLs, in A. lyrata (Supplementary Table 4). To confirm the involvement of EPCOT3 in species-specific expression of CYP82C2, we introduced WRKY33 into Nicotiana benthamiana (tobacco) leaves and A. thaliana cyp82C2 protoplasts transfected with either the A. thaliana or A. lyrata CYP82C2 locus (coding and 3000 nt upstream sequences, Fig. 5d). We observed transactivation by WRKY33 of the A. thaliana gene, but not that of A. lyrata (Fig. 5d and Supplementary Fig. 7d). Altogether, these data indicate that EPCOT3 and EPLs arose from retrotransposition following the speciation of A. thaliana, and that the EPCOT3-containing A. thaliana CYP82C2 promoter is sufficient to confer WRKY33-mediated transcription of CYP82C2.
Of the EPL retrocopies, EPL1 is most similar to EPCOT3 (85.4% identity), sharing the W-box and WRKY33-specific motif, whereas EPL2 is less similar (67%) and lacks the WRKY33-specific motif (Fig. 5c, Supplementary Fig. 7a, and Supplementary Table 4). EPL1 and EPL2 are much less truncated than EPCOT3 (Fig. 5c) and lack epigenetic signatures typical of cis-regulatory sequences55,56 (Supplementary Fig. 7c). To investigate whether the sequences and chromatin features associated with EPLs are sufficient for WRKY33 binding, we tested for WRKY33 binding to EPL sequences homologous to the W4 region of EPCOT3 in dex-treated, Psta-infected wrky33/DEX:WRKY33-flag plants by ChIP-(q)PCR. Compared with EPCOT3 (Fig. 3c), WRKY33 bound weakly or not at all to EPL1 and EPL2, respectively (Fig. 5e, and Supplementary Fig. 7e). Our findings suggest the following history: (1) EPL1 likely retroduplicated from EPL2 or its progenitor, which already contained a W-box; (2) EPL1 then acquired a WRKY33-specific motif by mutation; and (3) EPCOT3 retroduplicated from EPL1 and then acquired epigenetic signatures of an enhancer, thereby allowing selection to act on standing variation rather than de novo mutation for CYP82C2 recruitment into the 4OH-ICN biosynthetic pathway.
Discussion
TEs were originally conceived to act as controlling elements of several loci in the genome63, and exaptation of TEs into cis-regulatory modules has been hypothesized to be responsible for the rapid transcriptional rewiring in more ancient lineages of vertebrates12,13,14. However, few (if any) evolutionarily recent TE exaptation events in vertebrates and higher plants have been demonstrated to have biochemical, regulatory, physiological, and fitness-promoting functions14. With well over a dozen genomes available including the genetic model A. thaliana, the mustard family presents an excellent system for examining such events. In this study, we show that EPCOT3 is a TE-derived enhancer that mediates WRKY33 binding, pathogen-responsive transcription of CYP82C2, synthesis of the species-specific metabolite 4OH-ICN, and pathogen defense (Fig. 6). These results demonstrate how a recent TE exaptation can wire a new gene into an ancient regulon, ultimately leading to a positive effect on fitness.
Model of regulatory neofunctionalization of CYP82C2. An ancestral gene with roles in iron-stress responses (CYP82C4) underwent gene duplication in a progenitor species to A. thaliana and A. lyrata, leading to ancestral CYP82C2. Subsequent speciation led to ancestral A. thaliana and A. lyrata. In the former species, a significant degree of retroduplication, mutagenesis, and transposition events occurred, culminating with the formation of W-box and WRKY33-specific sequences in the ancestral EPCOT3 and its integration upstream of CYP82C2. Subsequent epigenetic modifications in A. thaliana were necessary to permit WRKY33 binding and CYP82C2 activation. Features in black have a hypothesized function, whereas features in gray have no known function. Double-dashed line indicates features omitted from view (e.g., CYP82C3)
Although the EPL1/EPCOT3 progenitor retrotransposed a preferred WRKY33-TFBS in the form of EPCOT3 upstream of CYP82C2, a further series of epigenetic modifications were needed to facilitate optimal access of EPCOT3 by WRKY33 (Fig. 6). EPL1 exists in a silenced heterochromatin state55,56 (Supplementary Fig. 7c), typical for TEs64, and is bound weakly by WRKY33 (Fig. 5e), whereas EPCOT3 is in an open chromatin state55,56 (Fig. 5b) and bound relatively strongly by WRKY33 (Fig. 3c). The more severe 5′-truncation of EPCOT3 could account for its release from TE-silencing mechanisms and the initially weak WRKY33 binding could provide a seed for chromatin remodelers to drive the exaptation of newly retrotransposed EPCOT3 into a bona fide enhancer. Further epigenomic sampling within Arabidopsis is needed to better clarify the epigenetic transformations underlying the EPCOT3 exaptation event.
Compared with closely related Landsberg accessions (Supplementary Fig. 3), Di-G synthesizes less camalexin and 4OH-ICN47 (Fig. 2b), and is more susceptible to a range of bacterial and fungal pathogens47,65 (Fig. 2c). WRKY33 has been implicated in camalexin biosynthesis31 and antifungal defense44. We identified WRKY33 as causal for some if not all of these phenotypes in Di-G. In addition, WRKY33’s involvement in antibacterial defense is consistent with the contribution of camalexin and 4OH-ICN toward antibacterial defense23.
WRKY33 is an ancient TF responsible for many fitness-promoting traits in plants; thus, it is unexpected that an A. thaliana accession would have a naturally occurring wrky33 mutation (C536T transversion). Di-G is the sole member of 1,135 sequenced accessions to have a high-effect single-nucleotide polymorphism (SNP) in WRKY3366, and may have originated from a Ler-0 ethyl methanesulfonate (EMS) mutagenesis screen, whose mutagenesis rate67 is well within the range of ~25,000 SNPs that are not concordant between Di-G and Ler-066 (Supplementary Fig. 2f). However, features of EMS mutations (i.e., transversion mutations) or X-ray mutations (i.e., indels) are not enriched in the Di-G pseudogenome relative to related pseudogenomes (Supplementary Table 5). These findings suggest that the wrky33 Di-G mutation is naturally derived.
Methods
Plant materials and growth
For quantitative PCR (qPCR) and high-performance liquid chromatography coupled with diode array detection and fluorescence detection (HPLC-DAD-FLD) analyses, surface-sterilized A. thaliana accession Columbia-0 (Col-0) seeds were sown in 12-well microtiter plates sealed with Micropore tape (3 M, St. Paul, MN), each well containing ~15 ± 2 seeds and 1 mL of either filter-sterilized 1× Murashige and Skoog media (pH 5.7–5.8) (4.43 g/L Murashige and Skoog basal medium with vitamins [Phytotechnology Laboratories, Shawnee Missions, KS], 0.05% MES hydrate, 0.5% sucrose) or iron-deficient media (amounts per liter): sucrose, 5.0 g; potassium nitrate, 1.9 g; ammonium nitrate, 1.65 g; MES monohydrate, 0.5 g; calcium chloride dihydrate, 0.44 g; magnesium sulfate heptahydrate, 0.37 g; monopotassium phosphate, 0.17 g; myo-inositol, 0.1 g; disodium EDTA, 29.2 mg; manganese sulfate monohydrate, 16.9 mg; zinc sulfate heptahydrate, 8.6 mg; boric acid, 6.2 mg; glycine, 2.0 mg; potassium iodide, 0.83 mg; nicotinic acid, 0.5 mg; pyridoxine hydrochloride, 0.5 mg; sodium molybdate dihydrate, 0.25 mg; thiamine hydrochloride, 0.1 mg; cobalt chloride hexahydrate, 25.0 μg; and copper sulfate pentahydrate, 25.0 μg. On day 9, seedlings were transferred to 6-well microtiter plates, each well containing ~15 seeds and 3 mL Murashige and Skoog or iron-deficient media. For Polyctenium fremontii, surface-sterilized seeds were sown on Murashige and Skoog agar plates. For all other species, surface-sterilized seeds were sown in 6-well plates, each well containing ~15 seeds and 3 mL Murashige and Skoog media. On day 9, media were refreshed prior to bacterial elicitation. Microtiter plates were placed on grid-like shelves over water-filled trays on a Floralight cart (Toronto, Canada) and plants were grown at 21 °C with 60% humidity under a 16 h light cycle (70–80 μE m−2 s−1 light intensity). For ChIP analyses, ~200 surface-sterilized seeds were sown in a 100 × 15 mm petri plate containing 20 mL of 1× Murashige and Skoog media. Media were exchanged for fresh media on day 9. For bacterial infection assays, plants were grown on soil (3:1 mix of Farfard Growing Mix 2 [Sun Gro Horticulture, Vancouver, Canada] to vermiculite [Scotts, Marysville, OH]) at 22 °C daytime/18 °C nighttime with 60% humidity under a 12 h light cycle [50 (dawn/dusk) and 100 (midday) μE m−2 s−1 light intensity]. Seed stock information is shown in Supplementary Table 6.
Vector construction and transformation
To generate the DEX:WRKY33-flag construct, WRKY33 was PCR-amplified from genomic DNA using the primers WRKY33gXhoF (5′-AACTCGAGAAGAACAAGAACCATCAC-3′) and W33flagSpeR (5′-CGACTAGTCTACTTGTCGTCATCGTCTTTGTAGTCGGGCATAAACGAATCGAAA-3′), and subcloned into the XhoI/SpeI sites of pTA7002 vector68. To generate the DEX:WRKY33-myc construct, WRKY33 was PCR-amplified using the primers WRKY33gXhoF and WRKY33gStuR (5′-AAGGCCTGGCATAAACGAATCGAAAAATG-3′), and subcloned into the XhoI/StuI sites of pTA7002-6x c-Myc vector69. Constructs were introduced into wrky33 and Di-G plants via Agrobacterium tumefaciens-mediated floral dip method70 and transformants were selected on agar media containing 15 μg/mL hygromycin B (Invitrogen, Carlsbad, CA).
To generate the CYP82C2 locus construct, the CYP82C2 upstream and coding sequences were PCR-amplified from A. thaliana genomic DNA using the primers At82C2proXbaF (5′-GCTCTAGAAGCTTCCAATAAAACATTC-3′) and At82C2proBamR (5′-GCGGATCCAGTGGTTTGAGCGTGCAAA-3′), and At82C2geneBamF (5′-GCGGATCCATGGATACTTCCCTCTTTTC-3′) and At82C2geneSmaR (5′-TTCCCGGGCTACTTGTCGTCATCGTCTTTGTAGTCCACATAAAGCCCTTCCTTAAG-3′). Sequences were subcloned into the XbaI/SmaI sites of pBI101 vector71. To generate the AlCYP82C2 locus construct, the AlCYP82C2 upstream and coding sequences were PCR-amplified from A. lyrata genomic DNA using the primers Al82C2proSalF (5′-CGGTCGACTATTCCAGGAGCATACAA-3′) and Al82C2proBglIIR (5′-GGAGATCTAATGTTTTAAAAGTGCAAAAGAG-3′), and Al82C2geneBamHF (5′-GCGGATCCATGGATACATCCCTCTTTTC-3′) and Al82C2geneSmaR (5′-TTCCCGGGCTACTTGTCGTCATCGTCTTTGTAGTCCACAAAAAGTTCTTCCTTAAGAC-3′), and subcloned into the SalI/SmaI sites of pBI101 vector. DEX:WRKY33-flag, CYP82C2, and AlCYP82C2 constructs were introduced into N. benthamiana leaves as previously described23 with the following modifications: leaves were infiltrated with transformed Agrobacterium strains that were grown in lysogeny broth (LB) medium supplemented with 30 μg/mL gentamycin and 50 μg/mL kanamycin to an OD600 of 0.7. Sixteen hours post-Agro-infiltration, leaves were sprayed with 20 μM dex, 0.1% Tween-20, and 1 µM flg22, and incubated for 24 and 30 h. Three 8 mm leaf discs were pooled per sample and snap-frozen for reverse-transcriptase PCR (RT-PCR) analyses. CYP82C2 and AlCYP82C2 constructs were introduced into A. thaliana cyp82C2 via PEG-mediated protoplast transformation72 with the following modifications: 2.5 × 105 protoplasts were transfected with 3 μg of construct for 20 min, recovered in 2.5× volume of W5 solution, elicited with 1 μM flg22 in 1 mL W5 solution for 6 h, and snap-frozen for RT-PCR analyses.
Bacterial infection and MAMP elicitation
A single colony of P. syringae pv. maculicola (Pma) M2 (containing avrRpm1, but not avrRps4 or avrRpt2), Pma ES4326 (containing no aforementioned effectors), Pma ES4326 avrRpt2, P. syringae pv. tomato DC3000 (Pst, containing no aforementioned effectors), Pst avrRpm1, Pst avrRps4, and Pst avrRpt2 from a freshly streaked 3-day-old agar plate was used to inoculate 2 mL of LB medium containing appropriate antibiotics. Strains were grown to log phase, washed in sterile water twice, resuspended in sterile water to OD600 of 0.2, and incubated at room temperature with no agitation for 3 to 6 h, prior to infection. Chitosan (90% deacetylated chitin; Spectrum Chemical Mfg Corp, New Brunswich, NJ) was prepared in 0.1 N acetic acid and neutralized with 0.1 N NaOH to pH 5.8, to a stock concentration of 1.2 mg/mL.
Hydroponically grown 9-day-old seedlings were inoculated with bacterial strains to OD600 of 0.013 or treated with 10 μM flg22 (QRLSTGSRINSAKDDAAGLQIA; Genscript, Nanjing, China, 10 μM elf26 (ac–SKEKFERTKPHVNVGTIGHVDHGKTT; Genscript), and 150 or 300 μg/mL chitosan. Seedlings were snap-frozen 9 h post infection for ChIP analyses, 12 h post infection for qPCR analyses, and 24–28 h post infection for HPLC-DAD analyses.
Four- to-five-week-old adult leaves were treated with 0.0125% Silwet (Phytotechnology Laboratories) or 0.0125% Silwet and 20 μM dex for 20 s, and incubated on 0.8% (w/v) tissue-culture agar plates on a light cart at 21 °C for 6–8 h. Leaves were then surface-inoculated with Pst (OD600 = 0.002 or 106 colony-forming units (cfu)/cm2 leaf area) in the presence of 0.01% (v/v) Silwet L-77 for 15 s and incubated on 0.8% (w/v) tissue-culture agar plates at 21 °C under a 16 h light cycle (70–80 μE m−2 s−1 light intensity) for 3 days. Leaves were then surface-sterilized in 70% ethanol for 10 s, rinsed in sterile water, surface-dried on paper towels, and the bacteria were extracted into water, using an 8 mm stainless steel bead and a ball mill (20 Hz for 3 min). Serial dilutions of the extracted bacteria were plated on LB agar plates for cfu counting.
RNA isolation and RT-PCR
Total RNA was extracted from snap-frozen seedlings using TRIzol reagent (Invitrogen) and 2.5 µg was reverse-transcribed with 3.75 µM random hexamers (Qiagen), 20 units of M-MuLV (New England Biolabs), and 20 units of ProtoScript II (New England Biolabs) for 1 h at 42 °C and then for 15 min at 70 °C. Resulting cDNA:RNA hybrids were treated with 10 units of DNase I (Roche) for 30 min at 37 °C and purified on PCR clean-up columns (Macherey–Nagel). qPCR was performed with Kapa SYBR Fast qPCR master mix (Kapa Biosystems) and CFX96 real-time PCR machine (Bio-Rad). Biological and technical replicates were performed on the same 384-well PCR plate, and EIF4A1 (AT3G13920) and AlEIF4A1 (AL3G26100) housekeeping genes were used to normalize mRNA levels between different samples. Primer sequences and efficiencies are listed in Supplementary Data 2. Total RNA from tobacco leaf discs and A. thaliana protoplasts was extracted from snap-frozen samples using 300 µL of Cell Lysis Solution (2% (w/v) SDS, 63 mM sodium citrate, 132 mM citric acid, 1 mM EDTA), 100 µL of Protein-DNA Precipitation Solution (4 M NaCl, 16 mM sodium citrate, 32 mM citric acid), and 300 µL of isopropanol. 2 μg RNA was then reverse-transcribed and complementary DNA was diluted 7.5-fold. Four microliters of cDNA was used in 20 μL PCR reactions and resulting PCR products were separated on 2% agarose gels. PCR was performed on C1000 thermal cycler (Bio-Rad) with the following thermal cycling program: 95 °C for 3 min; 40 cycles of 95 °C for 10 s, 53 °C for 15 s, and 72 °C for 7 s (WRKY33, CYP82C2, AlCYP82C2), 15 s (NbACTIN1), or 21 s (CYP82C2*). Primer sequences are listed in Supplementary Data 2.
Camalexin and 4OH-ICN extraction and LC-DAD-MS
Snap-frozen seedlings were lyophilized, weighed, and homogenized using a 5 mm stainless steel bead and ball mill (20 Hz, 4 min). For phytoalexin analysis, homogenate was extracted with 300 μL 80% (v/v) aqueous methanol containing 0.08% (v/v) formate and 2.5 μL internal standard (200 μM 4-methoxyindole/4M-I [Sigma-Aldrich] in 100% methanol) per mg sample dry weight. Extracts were sonicated for 5 min and centrifuged at 16,000 × g for 2 min. The supernatant was filtered using a 0.45 μm polypropylene filter plate (GE Healthcare, Chicago, IL).
Samples were separated on an Ultimate 3000 HPLC (Dionex, Sunnyvale, CA) system, using a 3.5 μm, 3 × 150 mm Zorbax SB-Aq column (Agilent, Santa Clara, CA) with the gradient shown in Supplementary Table 7. A coupled DAD-3000RS diode array detector (Dionex), FLD-311 fluorescence detector (Dionex), and MSQPlus mass spectrometer (MS) (Dionex) collected UV absorption spectra in the range of 190–560 nm, a collected fluorescence data at 275/350 nm (ex/em), and collected electrospray ionization (ESI) mass spectra in positive and negative ion modes in the range of 100–1000 m/z, respectively. Total ICN, 4OH-ICN, and camalexin amounts were quantified using standard curves of standards prepared in cyp79B2 cyp79B3 seedling extract and integrated areas in the UV chromatographs at 260 nm for 4M-I (retention time [RT] = 7.7 min); 340 nm for ICN (RT = 11.5 min); 280 nm for ICN degradation product ICA-ME (RT = 9.5 min), and co-eluting 4OH-ICN degradation products 4OH-ICA and 4OH-ICA-ME (RT = 10.1 min); and 320 nm for camalexin (RT = 12.1 min). For Fig. 1b, total camalexin amounts were quantified using integrated areas in the FLD chromatograph. For some experiments, 2.5 μL 200 μM indole butrytic acid (IBA; RT = 10.1 min) was added per mg sample dry weight instead of 4M-I. Relative amounts of ICN, 4OH-ICN, and amounts were quantified by dividing the peak areas at m/z 169 [M-H]− (ICN), 174 [M-H]− (ICA-ME), 176 [M-H]− (4OH-ICA), 190 [M-H]− (4OH-ICME), and 201 [M + H]+ (camalexin), by the peak area at m/z 202 [M-H]− (IBA).
Glucosinolate extraction and LC-DAD-FLD-MS
For glucosinolate extraction, a 96-well 0.45 μm polyvinylidene fluoride (PVDF) filter plate (EMD Millipore, Billerica, MA) was charged with 45 mg DEAE Sephadex A25 (GE Heathcare) and 300 μL of water per well, and equilibrated at room temp for 2 h. Prior to sample homogenization, the plate was centrifuged at 400 × g for 1 min to remove the water. The homogenate was extracted with 500 μL 70% (v/v) aqueous methanol at 67.5 °C for 10 min and centrifuged at 16,000 × g for 2 min. Added to the supernatant was 3 μL of IS (1.25 mM sinigrin (Sigma-Aldrich) in 80% (v/v) ethanol) per mg sample dry weight. Extract was applied to and incubated on the ion exchanger for 10 min. The sephadex resin was washed three times with 70% (v/v) methanol, three times with distilled deionized water (ddH2O), and two times with 20 mM sodium acetate (pH 5). Twenty microliters of 25 mg/mL aryl sulfatase (Type H1 from Helix pomatia, Sigma-Aldrich) was applied to and incubated on the sephadex resin at RT overnight. The plate was centrifuged at 400 × g for 1 min and desulfoglucosinolates were eluted from the sephadex resin by two 100 μL washes with 60% (v/v) methanol and two 100 μL washes with ddH2O. Eluate volume was reduced to 250–350 μL using an evaporator.
Samples were separated on an Ultimate 3000 HPLC system, using a 3.5 μm, 3 × 150 mm Zorbax SB-Aq column with the gradient shown in Supplementary Table 7. A coupled DAD-3000RS diode array detector, FLD-311 fluorescence detector, and MSQPlus mass spectrometer collected UV absorption spectra at 229 nm, fluorescence spectra at 275/350 nm (ex/em), and ESI mass spectra in positive/negative ion modes at 100–1000 m/z, respectively. Glucosinolates were quantified using integrated areas of desulfoglucosinolates in the UV chromatographs at 229 nm and published response factors73.
ChIP and PCR
For ChIP experiments on wrky33/DEX:WRKY33-flag nuclear extracts, approximately two hundred and ten 9-day-old seedlings were inoculated with Psta to OD600 of 0.013 and co-treated with mock solution of dimethyl sulfoxide (M) or 20 μM dex (D) for 9 h. Total protein was extracted in 25 mL of Extraction buffer 1 (0.4 M sucrose, 10 mM Tris-Cl [pH 8], 10 mM MgCl2, 5 mM 2-mercaptoethanol, 0.1 mM AEBSF, Complete EDTA-free protease inhibitor cocktail [Roche]. After a 10 min fixing step with 1% (v/v) formaldehyde solution and a 5 min quenching step with 2 M glycine, seedlings were washed three times with deionized water, vacuum-dried, and snap-frozen with liquid nitrogen. Following frozen homogenization, the homogenate was filtered once through a 70 µm mesh (Carolina Biological) and a 0.45 µm filter (EMD Millipore). Filtered homogenate was then washed once in 500 µL of Extraction buffer 2 (0.25 M sucrose, 10 mM Tris-Cl [pH 8], 10 mM MgCl2, 1% [v/v] Triton X-100, 5 mM 2-mercaptoethanol, 0.1 mM AEBSF, Complete EDTA-free protease inhibitor cocktail) and resuspended in 300 µL of Extraction buffer 3 (1.7 M sucrose, 10 mM Tris-Cl [pH 8], 0.15% [v/v] Triton X-100, 2 mM MgCl2, 5 mM 2-mercaptoethanol, 0.1 mM AEBSF, Complete-Mini EDTA-free protease inhibitor cocktail) prior to sucrose centrifugation. Following nuclear extraction, samples were resuspended in 125 µL of Nuclei Lysis buffer (50 mM Tris-Cl [pH 8], 10 mM EDTA, 1% [v/v] SDS, 0.1 mM AEBSF, Complete-Mini EDTA-free protease inhibitor cocktail), and 250 µL of ChIP dilution buffer (1% [v/v] Triton X-100, 1.2 mM EDTA, 16.7 mM Tris-Cl [pH 8], 167 mM NaCl, Complete EDTA-free protease inhibitor cocktail), sonicated in a Covaris S2 sonicator (Covaris, Woburn, MA) using 10% duty, 7% intensity, 200 cycles per burst for a total time of 11 min, and centrifuged at 16,000 × g for 10 min at 4 °C to precipitate SDS. ChIP was performed using Anti-FLAG M2 Affinity Gel (Sigma-Aldrich). Beads were pre-treated with 0.1% (w/v) non-fat milk in 1× phosphate-buffered saline (PBS) and 0.5 mg/mL sheared salmon sperm DNA (Invitrogen). Following de-crosslinking, DNA samples were phenol-chloroform-extracted, diluted to the same OD260 concentration, and 1.5 μL was used in a 15 μL PCR reaction.
PCR analysis was performed on nuclear extracts prior to antibody incubation (input) and after ChIP. PCR conditions were as follows: 95 °C for 3 min; 40 cycles of 95 °C for 15 s, 53 °C for 15 s, and 72 °C for 1 min; 72 °C for 5 min. Densitometric determination of signal intensity in each ChIP and input sample was calculated using ImageJ. Fold enrichment was determined by calculating the ratio of PCR product intensities in ChIP D/M to Input D/M. In cases where amplicons were absent, an arbitrary value of 10 was assigned. For EPL2, qPCR analysis was additionally performed to confirm absence of amplicons in ChIP samples. RLU counts at the 25th cycle were used for quantification. Primer sequences are listed in Supplementary Data 2.
SDS-PAGE and western blotting
Total protein was extracted from snap-frozen seedlings into 80 µL of extraction buffer (50 mM Tris-Cl [pH 7.5], 50 mM dithiothreitol, 4% [w/v] SDS, 10% [v/v] glycerol) using a 5 mm stainless steel bead and ball mill (20 Hz for 3 min). Samples were centrifuged briefly, incubated at 95 °C for 10 min, and centrifuged at 12,000 × g for 8 min to precipitate insoluble material.
Five (for WRKY33-flag) or 15 µL (for WRKY33-myc) of extract was loaded onto a 10% SDS-PAGE gel and the separated proteins were transferred to PVDF membrane (Millipore, Billerica, MA), stained with Ponceau S for labeling of total protein, and probed with either FLAG M2 (Sigma-Aldrich, cat# F1804) or c-Myc 9E10 (Santa Cruz Biotechnology, cat# sc-40) antibodies diluted 1:1000 or 1:750, respectively, in 1× PBS containing 5% (w/v) non-fat milk.
Comparative genomics
All phylogenetic species trees were adapted from published data74,75. To generate phylogenetic maximum likelihood (ML) trees, sequences were aligned using MUSCLE in MEGA776 and the JTT model (for CYP82C and LINE alignments) or Tamura-Nei model (for the EPCOT3 alignment). Sequences for all genes with the description “non-LTR retrotransposon family (LINE)” (n = 263) were batch-downloaded from TAIR (https://arabidopsis.org). Of these, sequences containing intact reverse-transcriptase domains (PGPDG, LIPK, FRPISL, or FADD sequences; n = 126) were used for subsequent phylogenetic analysis (Supplementary Notes 1 and 2). Gaps were removed from the CYP82C alignment, leaving a total of 480 codons. Information on genomes used for synteny analysis is shown in Supplementary Table 8.
Selection estimates based on nonsynonymous-to-synonymous substitution ratios were calculated from the CYP82C ML tree. A Newick tree file was generated from this ML tree (Supplementary Fig. 4b and Supplementary Data 1) and for Branch site models, branches were pre-defined. CodeML analysis in PAML77 was then conducted with the following modified parameters: ncatG = 8, CodonFreq = 3. The M0 test was performed with model = 0 and NSsites = 0. The M1a-null test was performed with model = 0 and NSsites = 1. A more stringent null test (fixed omega) was performed for each Branch site model to be tested (model = 2 and NSsites = 2), where omega was fixed to 1. Branch site models were then tested with unfixed omega. Likelihood ratio tests were performed by comparing critical values and degrees of freedom between each unfixed Branch site test and either the M1a test or the corresponding fixed-omega test. Pre-defined branches with P-values < 0.05 for both tests were regarded as under positive selection (Supplementary Data 1).
Bioinformatics
Epigenetics data were obtained from published work55,56. Percent identity matrices were constructed from Clustal Omega Multiple Sequence Alignments (https://www.ebi.ac.uk/Tools/msa/clustalo/). Promoter alignment plots were generated using mVISTA (http://genome.lbl.gov/vista/mvista/submit.shtml)78.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The authors declare that all data supporting the findings of this study are available within the paper and the Supplementary Information or are available from the corresponding authors upon request. A reporting summary for this article is available as a Supplementary Information file. The source data underlying Figs. 1b, 2b, 2c, 3a, 3c, 4a, 4c, 4d, 5d, and 5e, as well as Supplementary Figures 1b, 1c, 2a, 2c, 2d, 3a, 4b-f, 5a, 5b, 5d, 5e, 6a, 6c, 6e, 7d, and 7e are provided as a Source Data file.
Code availability
Custom codes to visualize the CYP82C2 homology model in MacPyMOL and conduct statistics in R are available at https://github.com/brendenbarco.
References
Dixon, R. A. & Strack, D. Phytochemistry meets genome analysis and beyond. Phytochemistry 62, 815–816 (2003).
Chae, L., Kim, T., Nilo-Poyanco, R. & Rhee, S. Y. Genomic signatures of specialized metabolism in plants. Science 344, 510–513 (2014).
Weng, J. K., Philippe, R. N. & Noel, J. P. The rise of chemodiversity in plants. Science 336, 1667–1670 (2012).
Ohno, S. Evolution by Gene Duplication (Springer-Verlag, Berlin, 1970).
Force, A. et al. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151, 1531–1545 (1999).
Tohge, T. & Fernie, A. R. Co-expression and co-responses: within and beyond transcription. Front. Plant Sci. 3, 248 (2012).
Omranian, N. et al. Differential metabolic and coexpression networks of plant metabolism. Trends Plant Sci. 20, 266–268 (2015).
Prud’homme, B., Gompel, N. & Carroll, S. B. Emerging principles of regulatory evolution. Proc. Natl Acad. Sci. USA 104, 8605–8612 (2007).
Wray, G. A. The evolutionary significance of cis-regulatory mutations. Nat. Rev. Genet. 8, 206–216 (2007).
Wittkopp, P. J. & Kalay, G. Cis-regulatory elements: molecular mechanisms and evolutionary processes underlying divergence. Nat. Rev. Genet. 13, 59–69 (2012).
Spitz, F. & Furlong, E. E. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Feschotte, C. Transposable elements and the evolution of regulatory networks. Nat. Rev. Genet. 9, 397–405 (2008).
Bourque, G. Transposable elements in gene regulation and in the evolution of vertebrate genomes. Curr. Opin. Genet. Dev. 19, 607–612 (2009).
de Souza, F. S., Franchini, L. F. & Rubinstein, M. Exaptation of transposable elements into novel cis-regulatory elements: is the evidence always strong? Mol. Biol. Evol. 30, 1239–1251 (2013).
Hénaff, E. et al. Extensive amplification of the E2F transcription factor binding sites by transposons during evolution of Brassica species. Plant J. 77, 852–862 (2014).
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Hammerschmidt, R. PHYTOALEXINS: What have we learned after 60 years? Annu. Rev. Phytopathol. 37, 285–306 (1999).
Mansfield, J. W. in Mechanisms of Resistance to Plant Diseases (eds Slusarenko, A. J., Fraser, R. S., van Loon, L. C.) 325–370 (Springer, The Netherlands, 2000).
Clay, N. K., Adio, A. M., Denoux, C., Jander, G. & Ausubel, F. M. Glucosinolate metabolites required for an Arabidopsis innate immune response. Science 323, 95–101 (2009).
Bednarek, P. et al. A glucosinolate metabolism pathway in living plant cells mediates broad-spectrum antifungal defense. Science 323, 101–106 (2009).
Tsuji, J., Jackson, E. P., Gage, D. A., Hammerschmidt, R. & Somerville, S. C. (1992) Phytoalexin accumulation in Arabidopsis thaliana during the hypersensitive reaction to Pseudomonas syringae pv syringae. Plant Physiol. 98, 1304–1309 (1992).
Thomma, B. P., Nelissen, I., Eggermont, K. & Broekaert, W. F. Deficiency in phytoalexin production causes enhanced susceptibility of Arabidopsis thaliana to the fungus Alternaria brassicicola. Plant J. 19, 163–171 (1999).
Rajniak, J., Barco, B., Clay, N. K. & Sattely, E. S. A new cyanogenic metabolite in Arabidopsis required for inducible pathogen defence. Nature 525, 376–379 (2015).
Hull, A. K., Vij, R. & Celenza, J. L. Arabidopsis cytochrome P450s that catalyze the first step of tryptophan-dependent indole-3-acetic acid biosynthesis. Proc. Natl Acad. Sci. USA 97, 2379–2384 (2000).
Mikkelsen, M. D., Hansen, C. H., Wittstock, U. & Halkier, B. A. Cytochrome P450 CYP79B2 from Arabidopsis catalyzes the conversion of tryptophan to indole-3-acetaldoxime, a precursor of indole glucosinolates and indole-3-acetic acid. J. Biol. Chem. 275, 33712–33717 (2000).
Glawischnig, E., Hansen, B. G., Olsen, C. E. & Halkier, B. A. Camalexin is synthesized from indole-3-acetaldoxime, a key branching point between primary and secondary metabolism in Arabidopsis. Proc. Natl Acad. Sci. USA 101, 8245–8250 (2004).
Klein, A. P., Anarat-Cappillino, G. & Sattely, E. S. Minimum set of cytochromes P450 for reconstituting the biosynthesis of camalexin, a major Arabidopsis antibiotic. Angew. Chem. Int. Ed. Engl. 52, 13625–13628 (2013).
Nafisi, M. et al. Arabidopsis cytochrome P450 monooxygenase 71A13 catalyzes the conversion of indole-3-acetaldoxime in camalexin synthesis. Plant Cell 19, 2039–2052 (2007).
Böttcher, C. et al. The multifunctional enzyme CYP71B15 (PHYTOALEXIN DEFICIENT3) converts cysteine-indole-3-acetonitrile to camalexin in the indole-3-acetonitrile metabolic network of Arabidopsis thaliana. Plant Cell 21, 1830–1845 (2009).
Bednarek, P. et al. Conservation and clade-specific diversification of pathogen-inducible tryptophan and indole glucosinolate metabolism in Arabidopsis thaliana relatives. New Phytol. 192, 713–726 (2011).
Qiu, J. L. et al. Arabidopsis MAP kinase 4 regulates gene expression through transcription factor release in the nucleus. EMBO J. 27, 2214–2221 (2008).
Mao, G. et al. Phosphorylation of a WRKY transcription factor by two pathogen-responsive MAPKs drives phytoalexin biosynthesis in Arabidopsis. Plant Cell 23, 1639–1653 (2011).
Schluttenhofer, C. & Yuan, L. Regulation of specialized metabolism by WRKY transcription factors. Plant Physiol. 167, 295–306 (2015).
Rinerson, C. I., Rabara, R. C., Tripathi, P., Shen, Q. J. & Rushton, P. J. The evolution of WRKY transcription factors. BMC Plant Biol. 15, 66 (2015).
Liu, S., Kracher, B., Ziegler, J., Birkenbihl, R. P. & Somssich, I. E. Negative regulation of ABA signaling by WRKY33 is critical for Arabidopsis immunity towards Botrytis cinerea 2100. eLife 4, e07295 (2015).
Denoux, C. et al. Activation of defense response pathways by OGs and Flg22 elicitors in Arabidopsis seedlings. Mol. Plant 1, 423–445 (2008).
Debener, T., Lehnackers, H., Arnold, M. & Dangl, J. L. Identification and molecular mapping of a single Arabidopsis thaliana locus determining resistance to a phytopathogenic Pseudomonas syringae isolate. Plant J. 1, 289–302 (1991).
Felix, G., Duran, J. D., Volko, S. & Boller, T. Plants have a sensitive perception system for the most conserved domain of bacterial flagellin. Plant J. 18, 265–276 (1999).
Zipfel, C. et al. Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125, 749–760 (2006).
Bisgrove, S. R., Simonich, M. T., Smith, N. M., Sattler, A. & Innes, R. W. A disease resistance gene in Arabidopsis with specificity for two different pathogen avirulence genes. Plant Cell 6, 927–933 (1994).
Povero, G. et al. Transcript profiling of chitosan-treated Arabidopsis seedlings. J. Plant Res. 124, 619–629 (2011).
Cabrera, J. C., Messiaen, J., Cambier, P. & Van Cutsem, P. Size, acetylation and concentration of chitooligosaccharide elicitors determine the switch from defence involving PAL activation to cell death and water peroxide production in Arabidopsis cell suspensions. Physiol. Plant 127, 44–56 (2006).
Birkenbihl, R. P., Diezel, C. & Somssich, I. E. Arabidopsis WRKY33 is a key transcriptional regulator of hormonal and metabolic responses toward Botrytis cinerea infection. Plant Physiol. 159, 266–285 (2012).
Zheng, Z., Qamar, S. A., Chen, Z. & Mengiste, T. Arabidopsis WRKY33 transcription factor is required for resistance to necrotrophic fungal pathogens. Plant J. 48, 592–605 (2006).
Studer, A., Zhao, Q., Ross-Ibarra, J. & Doebley, J. Identification of a functional transposon insertion in the maize domestication gene tb1. Nat. Genet. 43, 1160–1163 (2011).
Chan, Y. F. et al. Adaptive evolution of pelvic reduction in sticklebacks by recurrent deletion of a Pitx1 enhancer. Science 237, 302–305 (2010).
Kagan, I. A. & Hammerschmidt, R. Arabidopsis ecotype variability in camalexin production and reaction to infection by Alternaria brassicicola. J. Chem. Ecol. 28, 2121–2140 (2002).
Zhou, J. et al. Characterization of the promoter and extended C-terminal domain of Arabidopsis WRKY33 and functional analysis of tomato WRKY33 homologues in plant stress responses. J. Exp. Bot. 66, 4567–4583 (2015).
Birkenbihl, R. P., Kracher, B. & Somssich, I. E. Induced genome-wide binding of three Arabidopsis WRKY transcription factors during early MAMP-triggered immunity. Plant Cell 29, 20–38 (2017).
Rajniak, J. et al. Biosynthesis of redox-active metabolites in response to iron deficiency in plants. Nat. Chem. Biol. 14, 442–450 (2018).
Hu, T. T. et al. The Arabidopsis lyrata genome sequence and the basis of rapid genome size change. Nat. Genet. 43, 476–481 (2011).
Hohmann, N., Wolf, E. M., Lysak, M. A. & Koch, M. A. A time-calibrated road map of Brassicaceae species radiation and evolutionary history. Plant Cell 27, 2770–2784 (2015).
Kruse, T. et al. In planta biocatalysis screen of P450s identifies 8-methoxypsoralen as a substrate for the CYP82C subfamily yielding original chemical structures. Chem. Biol. 15, 149–156 (2008).
Slattery, M. et al. Absence of a simple code: how transcription factors read the genome. Trends Biochem. Sci. 39, 381–399 (2014).
Roudier, F. et al. Integrative epigenomic mapping defines four main chromatin states in Arabidopsis. EMBO J. 30, 1928–1938 (2011).
Liu, Y. et al. PCSD: a plant chromatin state database. Nucleic Acids Res 46, D1157–D1167 (2018).
Wang, Y., Li, X. & Hu, H. H3K4me2 reliably defines transcription factor binding regions in different cells. Genomics 103, 222–228 (2014).
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
He, H. H. et al. Positioned nucleosomes flanking a labile nucleosome characterize transcriptional enhancers. Nat. Genet. 42, 343–347 (2010).
Malik, H. S., Burke, W. D. & Eickbush, T. H. The age and evolution of non-LTR retrotransposable elements. Mol. Biol. Evol. 16, 793–805 (1999).
Luan, D. D., Korman, M. H., Jakubczak, J. L. & Eickbush, T. H. Reverse transcription of R2Bm RNA is primed by a nick at the chromosomal target site: a mechanism for non-LTR retrotransposition. Cell 72, 595–605 (1993).
Wright, D. A. et al. Multiple non-LTR retrotransposons in the genome of Arabidopsis thaliana. Genetics 142, 569–578 (1996).
McClintock, B. Controlling elements and the gene. Cold Spring Harb. Symp. Quant. Biol. 21, 197–216 (1956).
Slotkin, R. K. & Martienssen, R. Transposable elements and the epigenetic regulation of the genome. Nat. Rev. Genet. 8, 272–285 (2007).
Mukherjee, A. K., Lev, S., Gepstein, S. & Horwitz, B. B. A compatible interaction of Alternaria brassicicola with Arabidopsis thaliana ecotype DiG: evidence for a specific transcriptional signature. BMC Plant Biol. 9, 31 (2009).
1001 Genomes Consortium. 1,135 genomes reveal the global pattern of polymorphism in Arabidopsis thaliana. Cell 166, 481–491 (2016).
Rédei, G. P. & Koncz, C. in Methods in Arabidopsis Research (eds Koncz, C., Chua, N.-H., Schell, J.) 16–82 (World Scientific, 1993).
Aoyama, T. & Chua, N.-H. A glucocorticoid-mediated transcriptional induction system in transgenic plants. Plant J. 11, 605–612 (1997).
Chezem, W. R., Memon, A., Li, F. S., Weng, J. K. & Clay, N. K. SG2-type R2R3-MYB transcription factor MYB15 controls defense-induced lignification and basal immunity in Arabidopsis. Plant Cell 29, 1907–1926 (2017).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Jefferson, R. A., Kavanagh, T. A. & Bevan, M. W. GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 6, 3901–3907 (1987).
Sheen J. A transient expression assay using Arabidopsis mesophyll protoplasts. http://genetics.mgh.harvard.edu/sheenweb/ (2002).
Clarke, D. B. Glucosinolates, structures and analysis in food. Anal. Methods 2, 301–416 (2010).
Koch, M. A. & Kiefer, M. Genome evolution among cruciferous plants: a lecture from the comparison of the genetic maps of three diploid species—Capsella rubella, Arabidopsis lyrata subsp. petraea, and A. thaliana. Am. J. Bot. 92, 761–767 (2005).
Couvreur, T. L. et al. Molecular phylogenetics, temporal diversification, and principles of evolution in the mustard family (Brassicaceae). Mol. Biol. Evol. 27, 55–71 (2009).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Yang, Z. PAML 4: Phylogenetic Analysis by Maximum Likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Frazer, K. A., Pachter, L., Poliakov, A., Rubin, E. M. & Dubchak, I. VISTA: computational tools for comparative genomics. Nucleic Acids Res. 32, W273–W279 (2004).
Acknowledgements
We thank J.L. Celenza for the cyp79B2 cyp79B3 mutant. We thank E.S. Sattely for ICN/ICN-ME, 4OH-ICA/4OH-ICA-ME, and camalexin standards. This work was supported by T32-GM007499 (to B.B.) and Elsevier/Phytochemistry Young Investigator Award (to N.K.C.).
Author information
Authors and Affiliations
Contributions
B.B. and N.K.C. performed pathogen assays, immunoblotting, and ChIP-PCR experiments. N.K.C. performed RT-PCR experiments. B.B. and Y.K. profiled accessions and species. B.B. performed all other experiments. B.B. and N.K.C. interpreted the results and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barco, B., Kim, Y. & Clay, N.K. Expansion of a core regulon by transposable elements promotes Arabidopsis chemical diversity and pathogen defense. Nat Commun 10, 3444 (2019). https://doi.org/10.1038/s41467-019-11406-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-11406-3
This article is cited by
-
Dynamic enhancer transcription associates with reprogramming of immune genes during pattern triggered immunity in Arabidopsis
BMC Biology (2022)
-
Nearby transposable elements impact plant stress gene regulatory networks: a meta-analysis in A. thaliana and S. lycopersicum
BMC Genomics (2022)
-
Emerging mechanistic insights into the regulation of specialized metabolism in plants
Nature Plants (2022)
-
Regulatory regions in natural transposable element insertions drive interindividual differences in response to immune challenges in Drosophila
Genome Biology (2021)
-
Genome-wide (ChIP-seq) identification of target genes regulated by WRKY33 during submergence stress in Arabidopsis
BMC Genomic Data (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.