Abstract
Phloem plays a fundamental role in plants by transporting hormones, nutrients, proteins, RNAs, and carbohydrates essential for plant growth and development. However, the identity of the underlying phloem genes and pathways remain enigmatic especially in agriculturally important perennial crops, in part, due to the technical difficulty of phloem sampling. Here, we used two phloem-specific promoters and a translating ribosome affinity purification (TRAP) strategy to characterize the phloem translatome during leaf development at 2, 4, and 6 weeks post vernalization in plum (Prunus domestica L.). Results provide insight into the changing phloem processes that occur during leaf development. These processes included the early activation of DNA replication genes that are likely involved in phloem cell division during leaf expansion, as well as the upregulation of phloem genes associated with sink to source conversion, induction of defense processes, and signaling for reproduction. Combined these results reveal the dynamics of phloem gene expression during leaf development and establish the TRAP system as a powerful tool for studying phloem-specific functions and responses in trees.
Similar content being viewed by others
Introduction
In plants, the phloem is the major conduit for the long-distance transport of photoassimilates, phytohormones, small molecules, and macromolecules including RNAs and proteins. This long-distance transport system is vital for plant development and physiology and allows the plant to respond to a diverse array of abiotic and biotic stresses1,2,3. Plant pathogens, such as viruses and some bacteria, can also utilize the phloem to spread systemically throughout a host plant or to be picked up by phloem-feeding insects4,5,6. This makes the phloem a key tissue of interest for investigating host–pathogen interactions, as well as plant development. In recent years, the unique population of mRNAs found in the phloem have been at least partially identified in several plant species including Arabidopsis thaliana, melon, potato, grape, rice, and barley7,8,9,10,11,12,13,14,15. Identification of phloem-associated mRNAs has given greater insight into a range of plant physiological processes, such as sucrose loading and systemic acquired resistance (SAR) that occur within phloem tissues16,17,18. However, much information regarding the identity of phloem-associated genes and gene pathways remains to be investigated, especially within perennial plant species.
The phloem is comprised of companion cells (CCs) and sieve elements (SEs) surrounded by support cells including bundle sheath and phloem parenchyma3,19. At maturity, the anucleate SEs are dependent on adjacent CCs for genetic and metabolic capabilities. Together these cells form a pressurized system in which the phloem translocation stream flows from source to sink tissues. Isolating phloem contents from this pressurized system can be technically challenging. The majority of plants where phloem mRNAs have been identified are those that naturally exude sap upon wounding. For plant species that do not naturally exude sap, the addition of the chelator EDTA has been used to facilitate phloem bleeding20,21,22. However, this method has a risk of introducing contaminants to the phloem sap due to the damage of neighboring cells17,23,24,25. Other methods such as phloem collection from insect stylectomy or phloem microdissection can be technically difficult and costly to establish. In this study, we used a translating ribosome affinity purification (TRAP) approach to isolate mRNAs from phloem tissues26. In this method, a tagged ribosomal protein is expressed from phloem-specific promoters. This allows for the immunopurification of ribosome–mRNA complexes specifically from phloem tissues without disruption of the pressurized phloem system prior to mRNA harvesting. mRNAs eluted from tagged ribosomes (termed the translatome) are then identified by RNA-seq. A further advantage of this approach is that in contrast to total cellular mRNAs (termed the transcriptome) mRNAs associated with ribosomes are likely in the process of translation and thus more directly impact cell physiology.
In temperate crops such as fruit trees, phloem tissues are renewed annually in stems and newly developing leaves following a period of winter dormancy. Together with the xylem, phloem tissues establish sink to source movement of photoassimilates that drive carbohydrate partitioning. These fundamental functions are critical for numerous agronomic characteristics such as growth rate, fruit size, and fruit sweetness. For perennial tree crops, there exists little information regarding the identity of phloem-expressed gene products within the vascular tissues. Previous studies using phloem exudates derived by excision or aphid stylectomies have identified a number of proteins and RNAs present in the SEs of these woody perennials22,27. However, mature SEs are generally considered to be incapable of transcription and translation28,29,30,31. Thus, previously identified SE components are likely to be part of the translocation stream and in transit within the vasculature.
To better investigate phloem processes, specifically those that occur during leaf development and expansion, we have used a tissue-specific translatome approach to identify ribosome-associated mRNAs that are likely being actively translated within the leaf phloem of Prunus domestica L. Results from this study provide understanding into the changing cellular processes that occur during leaf phloem development, as well as the identity of specific genes associated with these processes. The utility of the TRAP system for studying phloem functions in perennial crops is also discussed.
Results
Isolation of translating ribosomes from plum
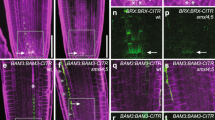
To identify phloem-specific mRNAs associated with ribosomes in plum, we generated transgenic Prunus domestica L. that express the Arabidopsis thaliana ribosomal protein L18 (RPL18) tagged with a His6-FLAG (HF) dual-epitope driven by either one of two phloem-specific promoters, pSUC2 or pSULTR2;2 that were acquired from A. thaliana, as well as the more ubiquitously expressed CaMV 35S promoter. A. thaliana RPL18 shares 87% amino-acid identity and 95% similarity with plum RPL18 (Fig. S1). A. thaliana pSUC2 has been previously shown to be expressed specifically in phloem vascular tissues in many plant species including pear, lime, and sweet orange trees32,33,34, whereas A. thaliana pSULTR2;2 has been shown to be expressed in phloem vascular tissues in A. thaliana and Nicotiana benthamiana15,35. pSUC2 and pSULTR2;2 have also been shown to have slightly different expression patterns with pSUC2 driving expression predominately in shoot CCs, whereas pSULTR2;2 expresses in both shoot CCs and bundle sheath cells15,35,36,37,38. To confirm the phloem-specific expression of pSUC2 and pSUTLR2;2 in Prunus domestica L., we created pSUC2::GUS and pSULTR2;2::GUS reporter lines. We found that GUS expression was observed in phloem tissues in plum leaves when driven by either pSUC2 or pSUTLR2;2 promoters but not in non-transgenic control plants (Fig. 1a). Consistent with previous reported results, we saw broader expression of GUS when driven by the pSULTR2;2 promoter compared with pSUC2.
a Histochemical analysis of Arabidopsis pSUC2 and pSULTR2;2 promoters in transgenic plums visualized by GUS staining in mid-vein cross sections. Phloroglucinol was used to stain xylem red. x xylem, p phloem. b Relative HF-RPL18 transgene expression in leaves. Quantitative RT-PCR analysis was performed with a primer set specific to HF-RPL18 and 18S rRNA was used as the internal control. Bars represent the mean of three biological replicates ± standard error. c Representative photographs of leaves collected at 2, 4, and 6 weeks post vernalization
To confirm expression of HF-RPL18, leaf tissue was collected from plum trees at 2, 4, and 6 weeks post vernalization. A vernalization treatment of 60 days was used to mimic the period of winter dormancy. This chilling period is required to initiate normal bud break and new leaf growth after trees are exposed to a period of favorable temperatures. Phloem tissues are renewed annually after dormancy in newly developing leaves. We chose to sample leaves every 2 weeks after dormancy to identify phloem-specific genes and pathways that contribute to this process. Quantitative RT-PCR (qRT-PCR) was used to monitor expression of the HF-RPL18 transcript at each time point. We observed the highest expression level from the p35S promoter, followed by pSUC2 then pSULTR2;2 (Fig. 1b). There was no significant difference in expression of HF-RPL18 between time points for p35S and pSUC2, but pSULTR2;2 had fourfold and twofold higher expression at 2 weeks than 4 weeks and 6 weeks (p-values = 0.003 and 0.024, respectively). To purify polysome–mRNA complexes, leaf extracts prepared with a polysome extraction buffer (PEB) were loaded onto a sucrose cushion and ultracentrifugation was used to pellet the polysome fraction. Any HF-RPL18 proteins that are not incorporated into ribosome complexes will remain in the supernatant and be removed. Anti-FLAG magnetic beads were added to the resuspended polysome fraction to capture tagged polysome–mRNA complexes. We could recover high-quality RNA from all plant lines expressing HF-RPL18 but not from non-transgenic controls (Fig. 2). qRT-PCR was used to determine HF-RPL18 expression in mRNAs isolated from polysomes compared with total RNAs. We found HF-RPL18 expression significantly increased 8.5 to 5.3-fold in pSULTR2;2 and pSUC2 translating mRNA compared with total RNA (p < 0.05) (Fig. 2c). In contrast, HF-RPL18 expression in p35S translating mRNA samples had a smaller 2.5-fold increase that was not statistically significant (p = 0.11). These data are consistent with phloem-specific expression of HF-RPL18 in pSULTR2;2 and pSUC2 transgenic plant lines and successful enrichment of tissue-specific transcripts. Together these results indicate that A. thaliana-derived HF-RPL18 is expressed in the predicted tissue-specific fashion in plum and can be used to capture mRNAs from translating ribosomes.
a Representative Agilent 2100 bioanalyzer gel-like image showing RNA isolated after incubation with Anti-FLAG magnetic beads taken from the unbound (blue labels) or bound (orange labels) fraction. High-quality RNA was successfully recovered in the bound fraction for all translatome HF-RPL18 plant lines, but not from non-transgenic controls. b Quantification of RNA recovered from the bound fraction. RNA was measured on a NanoDrop 2000 and is shown in nanograms. Bars represent the mean of four biological replicates ± standard error. c Relative HF-RPL18 transcript accumulation detected in total RNA (white bars) and translating mRNA samples (orange bars). Bars represent the mean of three biological replicates ± standard error. All samples are normalized to p35S total RNA
Phloem-associated translatome analysis
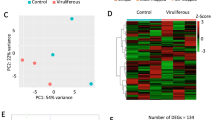
To characterize phloem-specific translatome profiles during post-dormancy leaf development, RNA was isolated from tagged polysome complexes of leaf extracts at 2, 4, and 6 weeks post vernalization (Fig. 1c). Three biological replicates, each comprised of leaf tissue collected from individual clonal trees of the same lines, were isolated for each promoter at each time point to generate a total of 27 libraries. Forty-three to 63 million paired-end reads were generated for each sample (Table S1). Paired-end reads were mapped with CLC Genomics Workbench v. 10.0.1 using the closely related Prunus persica genome version 2.0 as a reference39. Principal component analysis showed that samples were separated primarily on principal component 1 (29.7%) according to age and to a more limited extent on principal component 2 (17.5%) by construct (Fig. S2). As a control, we additionally sequenced ribosome-associated RNAs from non-transgenic leaves collected at 2 weeks. Ultracentrifugation was used to pellet the ribosome fraction and RNA was isolated from the resuspended pellet, but these non-transgenic samples were not incubated with anti-FLAG magnetic beads as they do not express a tagged protein. Non-transgenic samples separated on principal component 1 (37.4%) from the other 2-week samples (Fig. S3a). Differential gene expression analysis was done using CLC Genomics Workbench RNA-seq tool using a threshold of fold change > 2 and false discovery rate (FDR) p-value < 0.05. Non-transgenic samples had 6137, 6781, and 7297 differentially expressed genes (DEGs) when compared with p35S, pSUC2, and pSULTR2;2 samples respectively (Fig. S3b). We expected non-transgenic samples isolated by ultracentrifugation to be similar to p35S samples, as they both capture ribosomes for nearly all cell types. Although as expected, p35S did have the least number of DEGs when compared with non-transgenic controls, p35S samples were more similar to pSUC2 (700 DEGs) and pSULTR2;2 (686 DEGs) samples than non-transgenic controls (6137 DEGs). We concluded that p35S samples that had been through the immunopurification step were a more suitable control than non-transgenic ribosome-associated RNAs collected after ultracentrifugation.
After filtering and trimming, 72–90% of reads mapped to the reference genome, except for 4-week sample pSUC2-3, which had 43% of reads mapping to the genome. Of the reads that mapped, 66–94% mapped to exons, except for all three 4-week p35S samples and one 2-week p35S sample in which 31–53% of reads mapped to exons. Reads that did not map to exons mapped primarily to intergenic regions (Table S1). To our knowledge, Prunus domestica L. mapping rates to Prunus persica have not been previously published. However, percentages of mapped reads in this study are consistent with previous reports for other Prunus species. Peach-derived transcripts mapped to the Prunus persica reference genome have been reported at rates of 83–94%40,41. Although apricot-derived transcripts mapped to the Prunus persica reference genome at a lower rate of 64–72%42. Reads that did not map to exons could represent novel splice junctions or unannotated transcripts. In all samples, reads mapping to the Arabidopsis RPL18 His-FLAG tag sequence were present. AtRPL18 transcript levels were relatively consistent across biological replicates and there were no significant differences in reads mapping to the RPL18 His-FLAG tag sequence between time points (Table S1, Fig. S4).
To identify genes significantly enriched in phloem tissues, we compared pSUC2 or pSULTR2;2 samples to p35S samples for each time point. p35S is expressed in almost all tissue types including phloem, however, we would expect that phloem-derived mRNAs would be only a small fraction of all translating mRNAs isolated from p35S trees. Therefore, we would expect transcripts expressed predominately in phloem tissues to have significantly higher expression in pSUC2 and pSULTR2;2 samples when compared with p35S. Filtering transcripts identified in pSUC2 and pSULTR2;2 samples against p35S also compensates for any background non-phloem expression derived from the phloem promoters. Differential gene expression analysis was done using CLC Genomics Workbench RNA-seq tool using a threshold of fold change > 2 and FDR p-value < 0.05. A total of 1798 genes were identified as phloem enriched in the pSUC2 translatome for at least one time point, whereas 2149 genes were identified in the pSULTR2;2 translatome (Table S2).
Phloem-associated translatomes have only been previously characterized using TRAP in A. thaliana and N. benthamiana15,35. The number of plum phloem-associated genes identified in this study are consistent with results from N. benthamiana where 2308 genes were enriched in the pSUC2 translatome and 1236 genes were enriched in the pSULTR2;2 translatome15. Although in Arabidopsis, phloem translatomes included only 204–798 enriched genes15,35. To compare the plum and N. benthamaina phloem-enriched gene lists, we used the Arabidopsis best BLAST match for each gene. For plum, a total of 1740 unique genes were identified as associated with phloem pSUC2 or pSULTR2;2 translatomes, as many plum genes best BLAST match was the same Arabidopsis gene. A total of 368/1740 (21%) of these genes were previously identified as phloem enriched in N. benthamiana15. In contrast, 36/1740 (2%) of identified plum genes were also associated with phloem translatomes in Arabidopsis. However, there was greater overlap between plum phloem translatomes and the phloem transcriptome isolated from Arabidopsis phloem exudates with 174/1740 (10%) overlap9. Even when comparing the same species, phloem studies done by different groups have had only 8–22% overlap9,35,43. The low percent of overlap can be explained by differences in phloem isolation techniques and sampling times, which ranged from 1 week to 10 weeks. Thus, up to 21% overlap with previous identified phloem-associated transcripts is consistent with isolation of phloem tissues from plum.
To further validate that we had successfully isolated phloem-specific mRNAs, we first searched for known phloem-expressed genes. Eighty-eight Arabidopsis genes are annotated as associated with phloem tissues (PO:0005417). We found 16/88 (18%) of phloem-associated genes enriched in pSUC2 and/or pSULTR2;2 translatomes in our plum dataset (Table S3). In contrast, only 4/78 (5%) genes annotated as associated with xylem (PO:0005352) were enriched in our plum phloem dataset (Table S4). This is consistent with previous Arabidopsis phloem studies where 25% of phloem annotated genes were found in in phloem exudate or laser microdissection pressure catapulting derived phloem tissue compared with 13% of xylem annotated genes9. Taken together, the identification of known phloem-associated genes as significantly enriched in our phloem translatome dataset suggests the successful isolation of phloem-associated mRNAs from the plum translatome.
Identification of phloem-associated translatome alterations during leaf development
The highest number of phloem-enriched genes were found at the 6-week time point with a total of 1669 genes enriched in either the pSUC2 or pSULTR2;2 translatomes. Of these genes, 849 (51%) were significantly enriched in both translatomes. At 4 weeks, 1543 genes were identified as phloem enriched, with 322 (21%) significantly enriched in both phloem translatomes. While at 2 weeks, we found a total of 989 phloem-enriched genes with 397 (40%) significantly enriched in both phloem translatomes (Fig. 3a, Table S5). Twenty-one to 51% overlap between pSUC2 and pSULTR2;2 phloem translatomes is consistent with previous studies and is likely due to pSUC2 driving expression predominately in shoot CCs, whereas pSULTR2;2 expresses in both shoot CCs and bundle sheath cells15,35,36,37,38. Next, we chose to focus on the genes that were found to be significantly enriched in both pSUC2 and pSULTR2;2 translatomes for further analysis (Table S5). Of the 1100 genes enriched in both phloem translatomes, 128 (11.64%) genes were significantly enriched at all time points tested (Fig. 3b, Table S6). Among others, 144 were enriched only at the early 2-week time point, 69 were uniquely enriched at 4 weeks, and 547 genes were enriched only at 6 weeks (Fig. 3b). Gene ontology (GO) enrichment analysis using the best A. thaliana BLAST match for each gene, found that the plum phloem was enriched for genes associated with response to stress, defense responses, and protein phosphorylation at all time points tested (Fig. S5). At the 2-week time point, additional GO terms associated with defense and immune system responses were significantly enriched, while at 6-week genes associated with reproduction and transmembrane transport were uniquely enriched (Fig. S5).
a Total number of genes with > 2-fold expression that are unique or shared by pSUC2 or pSULTR2;2 phloem translatomes compared with p35S translatome at 2, 4, and 6 weeks post vernalization. b Number of unique and shared phloem-enriched genes across time points. Phloem-enriched genes were defined as > 2-fold expression in both pSUC2 and pSULTR2;2 phloem translatomes compared with p35S translatome
To further analyze how the phloem translatome is altered during development, we looked for those genes that were differentially expressed between leaf developmental time points. Differential gene expression analysis was done using CLC Genomics Workbench RNA-seq tool using a threshold of absolute value fold change > 2 and FDR p-value < 0.05. Of the 1100 genes enriched in both pSUC2 and pSULTR2;2 translatomes, 161/1100 (15%) were significantly downregulated over time in one or both phloem translatomes, 366/1100 (33%) were significantly upregulated, and the remaining 573/1100 (52%) did not significantly change expression over time (Table S5). We identified GO terms that were significantly overrepresented among the 527 phloem genes altered during leaf development and plotted the percentage of genes that were upregulated or downregulated for each GO term (Fig. 4, Table S7). DNA replication was the only GO term where the majority of genes were downregulated during leaf development. Of the 11 downregulated DNA replication genes identified, all displayed a reduction between the 2- and 4-week sampling times (Fig. 5a).
A subset of gene ontology (GO) biological process terms overrepresented in phloem-enriched genes that are altered over the 2- to 6-week sampled time frame. Bars represent the percentage of genes in each category that are downregulated (blue) or upregulated (red). All significantly overrepresented GO biological process terms are listed in Table S7
Heatmaps showing changes to genes involved in a DNA replication, b systemic acquired resistance, c response to nutrients, and d reproduction in Prunus domestica L. leaves 2, 4, and 6 weeks post vernalization. Mean FPKM values from three biological replicates are shown on a log2 scale with z-scaling by row. Gene names from A. thaliana best BLAST matches are shown to the right of all heatmaps
Surprisingly, the defense response category was the most significantly overrepresented GO term (FDR p-value 1.40e–13) among the 527 phloem genes altered during leaf development (Table S7). Seventy-eight percent (92/118) of defense-related genes were upregulated over the course of leaf development (Fig. 4). Identified defense genes included 38 genes that are involved in SAR with 79% (30/38) showing the highest expression at 6 weeks in phloem translatomes (Fig. 5b). Additional GO categories where the majority of genes were upregulated over time included response to nutrient levels and reproductive processes (Figs. 4, 5). Response to nutrient levels had the highest percentage of upregulated genes over time at 93% (27/29 genes) (Fig. 4).
Validation of selected transcripts
To further confirm the results obtained by RNA-seq, 12 genes were selected for analysis using qRT-PCR. Eight of these genes were identified as phloem enriched in both phloem translatomes at all time points tested, two had higher expression in non-phloem tissues, and two had similar expression levels in all samples. Three additional translatome biological replicates were collected from a different set of trees at the 2-week post vernalization time point. qRT-PCR analysis mirrored the RNA-seq translatome study (Fig. S6). Combined these results indicate phloem-enriched mRNAs identified by TRAP-seq in this study are reproducible in additional biological replicates.
Discussion
Vascular phloem represents one of the most important and specialized tissues in higher plants. Yet our understanding of the phloem’s transcriptional and translational composition is generally limited, particularly for plant systems such as woody perennials. The goal of this study was to assess the phloem translatome responses during leaf development in plum trees. Here we used a new molecular approach to express a tagged ribosomal protein (AtRPL18) from two phloem-specific promoters, pSUC2 and pSULTR2;2, in order to capture mRNAs from phloem tissues. This approach allows for the isolation of phloem-specific mRNAs without having to physically separate phloem from surrounding tissues, which is extremely technically difficult. Another key advantage of this approach is that mRNAs that are associated with ribosomes are likely being translated and thus have a direct impact on cellular function.
In plum trees, phloem is renewed annually in developing leaves after a period of winter dormancy. Isolation of phloem-associated translating mRNAs during new leaf growth following dormancy allowed us to identify specific phloem-associated genes and their corresponding cellular processes that show differential regulation over a range of leaf developmental stages. Combined these findings provide a unique resource for understanding a range of phloem processes that include sink to source transition, plant defense, and the onset of reproduction. Knowledge into these processes also represents a first step toward future efforts to manipulate phloem processes for enhanced crop production and disease resistance.
Specific findings from these studies included a range of translational alterations, with many DEGs associated with specific leaf developmental and response processes. For example, genes involved in DNA replication represented the only category that was consistently downregulated over successive developmental time points. This was also the GO term that was most significantly overrepresented among all downregulated phloem-enriched genes. These genes included predicted plum homologs of the Arabidopsis MCM2, POK1, and UBC19, all of which are genes shown to be expressed primarily in actively dividing cells44,45,46,47,48. High expression of DNA replication genes during early leaf development in both phloem and non-phloem tissues is consistent with the transition from early rapid cellular division to later growth stages where cell division slows and growth predominantly occurs via cellular expansion. Interestingly, the downregulation of these genes at 4 and 6 weeks was significantly greater in p35S whole leaf translatome samples than in the pSUC2 and pSULTR2;2, phloem-associated translatomes, suggesting that phloem remains active for cell division even after the leaf reaches maturity.
The majority of phloem-enriched genes that were altered throughout the course of leaf development were upregulated. Among these genes, the GO term most significantly overrepresented was defense response. Among phloem altered defense genes, 38 genes were identified as involved in SAR. We found 34/38 of SAR genes were upregulated over the course of development with the majority (30/34) showing their highest levels of expression in the phloem at 6 weeks. Genes with the highest expression at 6 weeks included predicted homologs of EDS1, RLP33, and RLK1. An additional four genes had peak expression levels at the 4-week time point and included homologs of pathogenesis related 4 (PR4) and PR5. In contrast, a predicted homolog of WRKY70 was one of only four SAR genes that was downregulated over the course of leaf development in both phloem translatomes. WRKY70 showed high expression levels at 2 weeks and 4 weeks with reduced expression at 6 weeks. WRKY70 has been recently shown to directly repress SARD1, a positive regulator of salicylic acid synthesis, in an Arabidopsis protoplast system and has been reported as an important node for salicylic acid and jasmonic acid signaling in plant defense responses49,50. Thus, the downregulation of WRKY70 observed in the phloem could contribute to increased salicylic acid synthesis in mature plum leaves. Salicylic acid accumulation in mature leaves has been previously described and associated with age-related resistance to pathogens in Arabidopsis51,52,53. This is also consistent with the observed upregulation of the majority of SAR genes in plum phloem translatomes at 6 weeks, suggesting that the phloem within mature leaves contributes significantly to the activation of defense responses.
Within the phloem the GO category with the greatest percentage of upregulated genes at 93% (27/29) was response to nutrients. This category includes genes involved in the transport of essential elements involved in growth and development, such as homologs of nitrate and phosphate transport genes of NRT1/ PTR FAMILY 7.3, AGL44, and PHO1;H1. The upregulation of these genes within the leaf vasculature within mature leaf tissues is consistent with its transition to source tissue with significant nutrient import and export functions. Similarly, 65% (33/51) of genes categorized as involved in reproductive processes were upregulated during leaf development, even though the plum trees were still in a juvenile state and did not flower. This included homologs of Arabidopsis AGL6 and AGL16, which encode MADS box transcription factors involved in flowering and transition to a reproductive phase in Arabidopsis54,55. However, the predicted FT gene homolog in plum is not significantly enriched in phloem tissues at any time point. Thus, juvenile trees appear to be primed for flowering but lack the FT trigger needed to initiate the process. Collectively, these findings suggest there is a significant transition in the phloem translatome that occurs upon leaf maturation that closely represents the metabolic and developmental transitions associated with sink to source transition and reproduction status.
The majority of phloem-enriched genes (53%) did not significantly change expression levels across the leaf developmental stages we tested. Seventy-four of these genes were significantly enriched in both phloem translatomes compared with p35S at all three time points. Included in this list were several kinases including a homolog of CRK10, which has been shown to be phloem mobile in Arabidopsis14. There were also genes associated with structural components of the phloem including homologs of Arabidopsis phloem protein 2-A1 and sieve-element occlusion-related 1. As expected, we found genes involved in the biosynthesis and transport of sugars were highly expressed in phloem tissues at all time points. Additionally, a predicted homolog of PHOSPHATE 2 was also identified in this gene set. In Arabidopsis, PHO2 is involved in the regulation of phosphate transport and has been shown to be expressed in the vasculature56,57. Thus, genes in this group that are similarly expressed and phloem enriched at all time points may represent genes involved in general phloem housekeeping functions or transport of nutrients needed throughout development.
In this study, TRAP was successfully used to capture phloem-specific mRNAs at three different developmental stages in plum leaves. The creation of transgenic translatome plant lines in plum and the adaptation of the TRAP method for use in fruit trees provides a powerful set of tools that can be used to identify changes to mRNAs in specific tissue types or at certain temporal stages. In A. thaliana, TRAP has been used to compare translatomes during developmental stages58 and in response to many abiotic and biotic stresses including low oxygen35,59, cold stress60, and pathogens15,61,62. Results from this study have identified specific expressed genes and processes that uniquely represent phloem responses during leaf development. These findings provide novel candidate genes and pathways that can be used in future studies aimed at improving fruit tree production.
Materials and methods
Plant material, transformations, and growth conditions
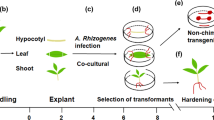
For the creation of transgenic plant lines, p35S::HF-RPL18, pSUC2::HF-RPL18, and pSULTR2;2::HF-RPL18 translatome constructs were kindly provided by Dr. J. Bailey-Serres, University of California, Riverside, CA, USA35. GUS expression constructs were created as described previously15. Briefly, upstream promoter sequences covering 2-kb upstream of the A. thaliana pSUC2 and pSULTR2;2 open reading frames were cloned into pBI101.1 (Clontech, Mountain View, CA, USA) in front of the GUS reporter open reading frame via primer-generated Sal1 and BamH1 restriction sites to create pSUC2::GUS and pSULTR2;2:GUS. Promoter::HF-RPL18 and promoter::GUS vectors were transformed into the Agrobacterium tumefaciens strain GV3101 and used for Agrobacterium-mediated transformation of plum (Prunus domestica L.) hypocotyl slices as previously described63. Hypocotyls were dissected from seeds of the open-pollinated cultivars “President” and “Stanley”. A minimum of seven independent transgenic lines for each construct were obtained and selected for using Kanamycin, propagated via tissue culture, rooted, and transplanted to soil. Transgene insertions were confirmed by quantitative RT-PCR (qRT-PCR) using primers 5ʹ-ATTTACAATTACCATGGGACATCAC-3ʹ and 5ʹ-CACCACCTCCCTTATCATCATC-3ʹ specific for the His6-FLAG-RPL18 transcript. All plum trees were maintained in a growth chamber at 25 °C, 75% humidity, with a 12-h photoperiod. After 12 weeks, plum leaves were stripped and plants were kept at 8 °C for a 60-day vernalization period before being moved back to the growth chamber.
GUS staining
Leaf and stem tissues were hand dissected (1–2 mm thick) with a razor blade then placed into 2 ml conical tubes filled with cold acetone on ice and incubated. The acetone from each sample was removed and the samples equilibrated with staining buffer64. Equilibrated samples were then developed in reaction buffer containing 2 mM of 5-bromo-4-chloro-3-indolyl-beta-d-glucuronic acid, overnight at 37 °C. Developed samples were rinsed in progressive concentrations of ethanol (30 min each at 20, 50, and 70%). Final treatments in 95% ethanol heated to 65 °C at 5- to 10-min increments was performed until the chlorophyll was fully removed. The samples were then observed and photographed in ethanol under a Zeiss Axiozoom microscope (Thornwood, NY, USA) fitted with a color camera.
Translating ribosomes affinity purification
Isolation of polysomes from non-transgenic or promoter::HF-RPL18 lines was done as previously described26,35,58. Briefly, frozen leaf tissue was homogenized in PEB (200 mM Tris-HCl, pH 9.0, 200 mM KCl, 25 mM ethylene glycol tetraacetic acid (EGTA) pH 8.0, 35 mM MgCl2, 1% (v/v) octylphenyl-polyethylene glycol (Igepal CA-630), 1% (v/v) polyoxyethylene 10 tridecyl ether, 1% (v/v) sodium deoxycholate, 5 mM dithiothereitol (DTT), 1 mM PMSF, 50 μg/mL cycloheximide, 50 μg/mL chloramphenicol, 0.5 mg/mL heparin) using 10 mL PEB per 5 g of tissue. The supernatant was loaded onto an 8 mL 1.6 M sucrose cushion. Samples were centrifuged at 170,000 g for 18 h at 4 °C to pellet the polysomes, which were then resuspended in 1 mL of PEB. For non-transgenic controls, RNA was isolated from the resuspended pellet using the Qiagen RNeasy kit (Qiagen, Valencia, CA, USA) according to the manufacturer's instructions.
Immunopurification of polysomes from promoter::HF-RPL18 plants was done as described26,35 with minor modifications. Specifically, anti-FLAG magnetic beads, 50 μL (Sigma Chemical Company, St. Louis, MO, USA) were added to the resuspended pellet and incubated at 4 °C overnight with gentle rocking. The beads were recovered using a magnet and washed four times for 5 min with 1 mL of wash buffer (200 mM Tris-HCl, pH 9.0, 200 mM KCl, 25 mM EGTA, 35 mM MgCl2, 5 mM DTT, 50 μg/mL cycloheximide, 50 μg/mL chloramphenicol). The complexes were eluted by treatment of the magnetic beads with 100 μL of Elution Buffer (100 μL wash buffer, 10 μL of 5 mg/mL FLAG3 peptide—Sigma Chemical Company, St. Louis, MO USA, 0.5 μL of 2 U/mL RNAse OUT—Thermo Fisher Scientific Cleveland, OH, USA). RLT buffer plus 2-mercaptoethanol from the Qiagen RNeasy kit (Qiagen, Valencia, CA, USA) was added to the eluted complexes. A 0.5 × volume of 100% ethanol was then added and the sample transferred to Qiagen RNeasy columns. Washes and RNA elutions were performed according to the manufacturer’s instructions. Isolated RNA was measured on a NanoDrop 2000 and with the Qubit RNA Assay Kit (Thermo Fisher Scientific, Cleveland, OH, USA). Quality of the RNA was confirmed using a RNA ScreenTape assay (Agilent Technologies, Palo Alto, CA, USA).
RNA sequencing and analysis
Library preparation and paired-end sequencing on the Illumina HiSeq system was done by Genewiz (South Plainfield, NJ, USA). Reads that passed Illumina’s quality control filters were further processed. Reads were mapped to the Prunus persica v2.0 genome39. The International Peach Genome Initiative 2013, https://www.rosaceae.org/species/prunus_persica/genome_v2.0.a1 using the CLC Genomics Workbench v10.0.1 RNA-seq analysis tool and default parameters (Mismatch cost 2, Insertion cost 3, Deletion Cost 3, Length fraction 0.8, Similarity fraction 0.8, Max hits for a read 10—CLC Bio, Aarhus, Denmark) (Table S1). Total reads aligned to genes were used in all subsequent analyses. DEGs were identified using the CLC Genomics Workbench Differential Expression for RNA-Seq tool with a cutoff value of > 2-fold and FDR p-value < 0.05. DEGs are listed in Supporting Information in Tables S2-S6 and the dataset is available under GEO accession number GSE111738. GO enrichment analysis of DEGs was performed using the Arabidopsis best BLAST match and agriGO, an agriculturally focused web based GO analysis program65. The singular enrichment analysis tool was used to identify GO terms that were significantly enriched (FDR p-value < 0.05) using the TAIR10 genome as the background (Table S7). All heatmaps were generated in R using the heatmap.2 function in the gplots CRAN library (http://cran.r-project.org/web/packages/gplots/index.html). For GO enrichment heatmaps, the FDR p-value was used. For gene expression heatmaps, Log2 (FPKM) values exported from the CLC Genomics Workbench were used with z-scaling by row.
qRT-PCR validation of selected transcripts
RNA for qRT-PCR validation was isolated from immunopurified ribosomes of promoter::HF-RPL18 plants as previously described for three additional biological replicates collected from a different set of trees at the 2-week post vernalization time point. For these studies, 50 ng of isolated RNA was pretreated with RQ1 DNase (Promega, Madison, WI, USA), followed by reverse transcription using SuperScript III First-Strand Synthesis System and random hexamer primers (Invitrogen by Life Technologies, Carlsbad, CA, USA). SYBR green real-time qRT-PCR was performed in an ABI Prism 7100 (Applied Biosystems, Foster City, CA, USA). The 18S rRNA was chosen as an internal control for normalization. Primer sequences used for the amplification of all selected genes are provided in Table S8.
Data availability
The dataset is available under GEO accession number GSE111738.
References
Carella, P. et al. Comparative proteomics analysis of Arabidopsis phloem exudates collected during the induction of systemic acquired resistance. Plant Physiol. 171, 1495–1510 (2016).
Fu, Z. Q. & Dong, X. Systemic acquired resistance: turning local infection into global defense. Annu. Rev. Plant Biol. 64, 839–863 (2013).
Ham, B. K. & Lucas, W. J. The angiosperm phloem sieve tube system: a role in mediating traits important to modern agriculture. J. Exp. Bot. 65, 1799–1816 (2014).
Bendix, C. & Lewis, J. D. The enemy within: phloem‐limited pathogens. Mol. Plant Pathol. 19, 238–254 (2018).
Folimonova, S. Y. & Tilsner, J. Hitchhikers, highway tolls and roadworks: the interactions of plant viruses with the phloem. Curr. Opin. Plant Biol. 43, 82–88 (2018).
Hipper, C., Brault, V., Ziegler-Graff, V. & Revers, F. Viral and cellular factors involved in phloem transport of plant viruses. Front. Plant Sci. 4, 154 (2013).
Asano, T. et al. Construction of a specialized cDNA library from plant cells isolated by laser capture microdissection: toward comprehensive analysis of the genes expressed in the rice phloem. Plant J. 32, 401–408 (2002).
Omid, A., Keilin, T., Glass, A., Leshkowitz, D. & Wolf, S. Characterization of phloem-sap transcription profile in melon plants. J. Exp. Bot. 58, 3645–3656 (2007).
Deeken, R. et al. Identification of Arabidopsis thaliana phloem RNAs provides a search criterion for phloem‐based transcripts hidden in complex datasets of microarray experiments. Plant J. 55, 746–759 (2008).
Gaupels, F., Knauer, T. & van Bel, A. J. A combinatory approach for analysis of protein sets in barley sieve-tube samples using EDTA-facilitated exudation and aphid stylectomy. J. Plant Physiol. 165, 95–103 (2008).
Yu, Y., Lashbrook, C. & Hannapel, D. J. Tissue integrity and RNA quality of laser microdissected phloem of potato. Planta 226, 797–803 (2007).
Tetyuk, O., Benning, U. F. & Hoffmann-Benning, S. Collection and analysis of Arabidopsis phloem exudates using the EDTA-facilitated method. J. Vis. Exp. 80, 51111 (2013).
Serra-Soriano, M., Navarro, J. A., Genoves, A. & Pallás, V. Comparative proteomic analysis of melon phloem exudates in response to viral infection. J. Proteom. 124, 11–24 (2015).
Thieme, C. J. et al. Endogenous Arabidopsis messenger RNAs transported to distant tissues. Nat. Plants 1, 15025 (2015).
Collum, T. D. & Culver, J. N. Tobacco mosaic virus infection disproportionately impacts phloem associated translatomes in Arabidopsis thaliana and Nicotiana benthamiana. Virology 510, 76–89 (2017).
Riesmeier, J. W., Hirner, B. & Frommer, W. B. Potato sucrose transporter expression in minor veins indicates a role in phloem loading. Plant Cell 5, 1591–1598 (1993).
Ruiz-Medrano, R., Xoconostle-Cázares, B. & Lucas, W. J. Phloem long-distance transport of CmNACP mRNA: implications for supracellular regulation in plants. Development 126, 4405–4419 (1999).
Ham, B.-K. & Lucas, W. J. Phloem-mobile RNAs as systemic signaling agents. Annu. Rev. Plant Biol. 68, 173–195 (2017).
Otero, S. & Helariutta, Y. Companion cells: a diamond in the rough. J. Exp. Botany, 68, 71–78 (2017).
King, R. & Zeevaart, J. Enhancement of phloem exudation from cut petioles by chelating agents. Plant Physiol. 53, 96–103 (1974).
Urquhart, A. A. & Joy, K. W. Use of phloem exudate technique in the study of amino acid transport in pea plants. Plant Physiol. 68, 750–754 (1981).
Atkins, C. A., Smith, P. M. & Rodriguez-Medina, C. Macromolecules in phloem exudates—a review. Protoplasma 248, 165–172 (2011).
Knoblauch, M. & van Bel, A. J. Sieve tubes in action. Plant Cell 10, 35–50 (1998).
Turgeon, R. & Wolf, S. Phloem transport: cellular pathways and molecular trafficking. Annu Rev. Plant Biol. 60, 207–221 (2009).
Dinant, S. & Kehr, J. Sampling and analysis of phloem sap. Plant Mineral Nutrients. Methods in Molecular Biology (Methods and Protocols) pp. 185–194 (Humana Press, Totowa, NJ, 2013).
Reynoso, M. A. et al. Translating Ribosome Affinity Purification (TRAP) followed by RNA sequencing technology (TRAP-SEQ) for quantitative assessment of plant translatomes. Methods Mol. Biol. 1284, 185–207 (2015).
Varkonyi-Gasic, E., Gould, N., Sandanayaka, M., Sutherland, P. & MacDiarmid, R. M. Characterisation of microRNAs from apple (Malus domestica’Royal Gala’) vascular tissue and phloem sap. BMC Plant Biol. 10, 159 (2010).
Oparka, K. J. & Turgeon, R. Sieve elements and companion cells—traffic control centers of the phloem. Plant Cell 11, 739–750 (1999).
Esau, K. The phloem. Encyclopedia of Plant Anatomy p. 505 (Gebrüder Borntraegger, Berlin-Stuttgart,1969).
Sjolund, R. & Shih, C. Freeze-fracture analysis of phloem structure in plant tissue cultures: I. The sieve element reticulum. J. Ultrastruct. Res. 82, 111–121 (1983).
Fisher, D. B., Wu, Y. & Ku, M. S. Turnover of soluble proteins in the wheat sieve tube. Plant Physiol. 100, 1433–1441 (1992).
Sun, Q. et al. High-efficiency and stable genetic transformation of pear (Pyrus communis L.) leaf segments and regeneration of transgenic plants. Acta Physiol. Plant. 33, 383–390 (2011).
Dutt, M., Ananthakrishnan, G., Jaromin, M., Brlansky, R. & Grosser, J. Evaluation of four phloem-specific promoters in vegetative tissues of transgenic citrus plants. Tree Physiol. 32, 83–93 (2012).
Miyata, L. Y. et al. GUS expression in sweet oranges (Citrus sinensis L. Osbeck) driven by three different phloem-specific promoters. Plant Cell Rep. 31, 2005–2013 (2012).
Mustroph, A. et al. Profiling translatomes of discrete cell populations resolves altered cellular priorities during hypoxia in Arabidopsis. Proc. Natl. Acad. Sci. USA 106, 18843–18848 (2009).
Stadler, R. & Sauer, N. The Arabidopsis thaliana AtSUC2 gene is specifically expressed in companion cells. Plant Biol. 109, 299–306 (1996).
Takahashi, H. et al. The roles of three functional sulphate transporters involved in uptake and translocation of sulphate in Arabidopsis thaliana. Plant J. 23, 171–182 (2000).
Truernit, E. & Sauer, N. The promoter of the Arabidopsis thaliana SUC2 sucrose-H+symporter gene directs expression of β-glucuronidase to the phloem: evidence for phloem loading and unloading by SUC2. Planta 196, 564–570 (1995).
Verde, I. et al. The Peach v2. 0 release: high-resolution linkage mapping and deep resequencing improve chromosome-scale assembly and contiguity. BMC Genomics 18, 225 (2017).
Rubio, M. et al. Analysis of gene expression changes in peach leaves in response to Plum pox virus infection using RNA‐Seq. Mol. Plant Pathol. 16, 164–176 (2015).
Wang, L. et al. Deep RNA-Seq uncovers the peach transcriptome landscape. Plant Mol. Biol. 83, 365–377 (2013).
Rubio, M. et al. Gene expression analysis of Plum pox virus (Sharka) susceptibility/resistance in apricot (Prunus armeniaca L.). PLoS ONE 10, e0144670 (2015).
Zhao, C., Craig, J. C., Petzold, H. E., Dickerman, A. W. & Beers, E. P. The xylem and phloem transcriptomes from secondary tissues of the Arabidopsis root-hypocotyl. Plant Physiol. 138, 803–818 (2005).
Lipka, E. et al. The phragmoplast-orienting kinesin-12 class proteins translate the positional information of the preprophase band to establish the cortical division zone in Arabidopsis thaliana. Plant Cell 26, 2617–2632 (2014).
Müller, R., Morant, M., Jarmer, H., Nilsson, L. & Nielsen, T. H. Genome-wide analysis of the Arabidopsis leaf transcriptome reveals interaction of phosphate and sugar metabolism. Plant Physiol. 143, 156–171 (2007).
Ni, D. A. et al. The Arabidopsis MCM2 gene is essential to embryo development and its over‐expression alters root meristem function. New Phytol. 184, 311–322 (2009).
Shultz, R. W., Lee, T.-J., Allen, G. C., Thompson, W. F. & Hanley-Bowdoin, L. Dynamic localization of the DNA replication proteins MCM5 and MCM7 in plants. Plant Physiol. 150, 658–669 (2009).
Criqui, M. C. et al. Molecular characterization of plant ubiquitin-conjugating enzymes belonging to the UbcP4/E2-C/UBCx/UbcH10 gene family. Plant Physiol. 130, 1230–1240 (2002).
Zhou, M. et al. WRKY70 prevents axenic activation of plant immunity by direct repression of SARD1. New Phytol. 217, 700–712 (2018).
Li, H., Johnson, P., Stepanova, A., Alonso, J. M. & Ecker, J. R. Convergence of signaling pathways in the control of differential cell growth in Arabidopsis. Dev. Cell 7, 193–204 (2004).
Kus, J. V., Zaton, K., Sarkar, R. & Cameron, R. K. Age-related resistance in Arabidopsis is a developmentally regulated defense response to Pseudomonas syringae. Plant Cell 14, 479–490 (2002).
Carviel, J. L. et al. Investigation of intercellular salicylic acid accumulation during compatible and incompatible Arabidopsis-Pseudomonas syringae interactions using a fast neutron-generated mutant allele of EDS5 identified by genetic mapping and whole-genome sequencing. PLoS ONE 9, e88608 (2014).
Carella, P., Wilson, D. C. & Cameron, R. K. Some things get better with age: differences in salicylic acid accumulation and defense signaling in young and mature Arabidopsis. Front. Plant Sci. 5, 775 (2015).
Yoo, S. K., Wu, X., Lee, J. S. & Ahn, J. H. AGAMOUS‐LIKE 6 is a floral promoter that negatively regulates the FLC/MAF clade genes and positively regulates FT in Arabidopsis. Plant J. 65, 62–76 (2011).
Hu, J.-Y. et al. miR824-regulated AGAMOUS-LIKE16 contributes to flowering time repression in Arabidopsis. Plant Cell 26, 2024–2037 (2014).
Huang, T.-K. et al. Identification of downstream components of ubiquitin-conjugating enzyme PHOSPHATE2 by quantitative membrane proteomics in Arabidopsis roots. Plant Cell 25, 4044–4060 (2013).
Lin, S.-I. et al. Regulatory network of microRNA399 and PHO2 by systemic signaling. Plant Physiol. 147, 732–746 (2008).
Lin, S.-Y. et al. Profiling of translatomes of in vivo–grown pollen tubes reveals genes with roles in micropylar guidance during pollination in arabidopsis. Plant Cell 26, 602–618 (2014).
Branco‐Price, C., Kaiser, K. A., Jang, C. J., Larive, C. K. & Bailey‐Serres, J. Selective mRNA translation coordinates energetic and metabolic adjustments to cellular oxygen deprivation and reoxygenation in Arabidopsis thaliana. Plant J. 56, 743–755 (2008).
Juntawong, P., Sorenson, R. & Bailey‐Serres, J. Cold shock protein 1 chaperones mRNAs during translation in Arabidopsis thaliana. Plant J. 74, 1016–1028 (2013).
Moeller, J. R. et al. Differential accumulation of host mRNAs on polyribosomes during obligate pathogen-plant interactions. Mol. Biosyst. 8, 2153–2165 (2012).
Meteignier, L.-V. et al. Translatome analysis of an NB-LRR immune response identifies important contributors to plant immunity in Arabidopsis. J. Exp. Bot. 68, 2333–2344 (2017).
Petri, C., Scorza, R. & Srinivasan, C. Highly efficient transormation protcol for plum (Prunus domestica L.). Transgenic Plants pp. 191-199 (Humana Press, Totowa, NJ, 2012).
Bomblies, K. Whole mount GUS staining. Arabidopsis: A Laboratory Manual pp. 243-245 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 2000).
Du, Z., Zhou, X., Ling, Y., Zhang, Z. & Su, Z. agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res. 38, W64–W70 (2010).
Acknowledgements
We thank Professor Bailey-Serres, University of California Riverside for providing the translatome constructs. This work was supported by the USDA National Institute of Food and Agriculture, Plant-Associated Microbes and Plant-Microbe Interactions Program (2015–67013-23004) and the NSF Division of Integrative Organismal Systems (ISO-1644713).
Author information
Authors and Affiliations
Contributions
T.D.C, C.D., and J.N.C. planned and designed the research. T.D.C., E.L, and C.D.R. performed experiments and analyzed data. T.D.C., C.D., and J.N.C led the writing of the manuscript. All authors reviewed the manuscript for intellectual content and contributed to the final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Collum, T.D., Lutton, E., Raines, C.D. et al. Identification of phloem-associated translatome alterations during leaf development in Prunus domestica L.. Hortic Res 6, 16 (2019). https://doi.org/10.1038/s41438-018-0092-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-018-0092-4
This article is cited by
-
Defining the ‘HoneySweet’ insertion event utilizing NextGen sequencing and a de novo genome assembly of plum (Prunus domestica)
Horticulture Research (2021)
-
Genome-wide identification and transcript profiles of walnut heat stress transcription factor involved in abiotic stress
BMC Genomics (2020)
-
Red versus green leaves: transcriptomic comparison of foliar senescence between two Prunus cerasifera genotypes
Scientific Reports (2020)