Abstract
In our previous work, we proposed a genomic prediction method combing identical-by-state-based Haseman-Elston regression and best linear prediction with additive variance component only (HEBLP|A herein), the most essential component of genetic variation. Since the dominance effects contribute significantly in heterosis, it is desirable to incorporate the HEBLP with dominance variance component that is expected to enhance the predictive accuracy as we move to the further development: HEBLP|AD, a paralleled implementation of genomic prediction compared with genomic best linear unbiased prediction (GBLUP). The simulation results indicated that when the dominance effects contributed to a large proportion of genetic variation, HEBLP|AD and GBLUP|AD, having similar accuracy, both outperformed HEBLP|A; but when the dominance variation was none or little, HEBLP|A, HEBLP|AD, and GBLUP|AD had similar predictability. The analysis of real data from Arabidopsis thaliana F2 population also demonstrated the latter situation. In summary, HEBLP|AD performed stable whether a trait was controlled by dominance effects or not.
Similar content being viewed by others
Introduction
With the rapid development of high-throughput molecular marker techniques, such as single nucleotide polymorphisms (SNPs) and statistical approaches, genomic prediction first proposed by Meuwissen et al. (2001) has been successfully applied to genetic improvement of complex traits that are controlled by polygenic effects—numerous small-effect quantitative trait loci (QTL) (Schaeffer, 2006; Hayes et al. 2009; Jannink et al. 2010; Zhang et al. 2011; Riedelsheimer et al. 2012). Compared to the conventional marker-assisted selection (MAS), genomic prediction is far more accurate by utilizing all molecular marker information to estimate the breeding values of each individual in a candidate population (Heffner et al. 2009; Arruda et al. 2016).
In the early stage of genomic prediction methods, many models accounted only for additive effects (Meuwissen et al. 2001; Bernardo and Yu, 2007; Calus et al. 2008; VanRaden, 2008). However, dominance effects contribute to heterosis (Hua et al. 2003; Li et al. 2008), and therefore should be included in the models orienting hybrid breeding. Recent studies also show that genomic prediction models including dominance effects can improve the prediction accuracy (Denis and Bouvet, 2011; Su et al. 2012; Technow et al. 2012; Denis and Bouvet, 2013; Nishio and Satoh, 2014; de Almeida Filho et al. 2016; Wang et al. 2017; Liu et al. 2017; Resende et al. 2017).
In our previous study, we developed a fast genomic prediction approach (namely HEBLP, or HEBLP|A herein) combining identical-by-state (IBS)-based Haseman-Elston (HE) regression and best linear prediction (BLP). It can obtain the total additive genetic variance via a simple HE linear regression with reduced computation complexity, but only additive effects are included (Liu and Chen, 2017). The present study aims to develop the HEBLP with both the additive and dominance effects (HEBLP|AD) and to evaluate its predictive performance in the simulated and a real Arabidopsis thaliana F2 population.
Materials and methods
The Arabidopsis thaliana F2 population
We used the phenotype and genotype data of an Arabidopsis thaliana F2 population (namely P19) derived from a cross between Bay-0 and Lov-5 (Salomé et al. 2011). It consists of 384 individuals and 245 SNP markers. There are seven traits including days until visible flower buds in the center of the rosette (DTF1), days until inflorescence stem reached 1 cm in height (DTF2), days until first open flower (DTF3), rosette leaf number (RLN), cauline leaf number (CLN), total leaf number: sum of RLN and CLN (TLN), and leaf initiation rate (RLN/DTF1) (LIR1). For more details about the P19 population please refer to Salomé et al. 2011.
Statistical models
The linear model of a quantitative trait can be written as:
in which y is the n × 1 vector for the standardized phenotypic value of a quantitative trait measured from n individuals \(\left( {y_i = \frac{{y_i^\prime - \bar y}}{{\sigma _y}}} \right),\,y_i^\prime\) represents the raw phenotypic value; \(\bar y\) represents the mean value of the phenotypic values; and σy represents the standard error of the phenotypic values.); Za is the standardized genotype matrix of n rows and m columns for additive effects (m represents the number of markers.). Zd is the standardized genotype matrix n × m for dominance effects. In order to keep the additive and dominance variances orthogonal to each other, the coding schemes for additive and dominance effects should be tuned accordingly (Vitezica et al. 2017). For the ith individual at the kth locus, \(Z_{a,ik} = \frac{{x_{ik} - 2p_k}}{{\sqrt {2p_k(1 - p_k)} }},\) in which xik counts the number of reference alleles (2, 1, and 0 for AA, Aa, and aa, respectively) and pk the frequency of the reference allele A at the locus. \(Z_{d,ik} = \frac{{\delta _{ik} - 2pk}}{{2pk(1 - pk)}}\), in which δik is coded 0, 2pk, and (4pk−2), respectively for AA, Aa, and aa genotypes, respectively. F2 population the expected pk is 0.5, and the frequency for AA, Aa, and aa are 0.25, 0.5, and 0.25, respectively, under the Hardy-Weinberg equilibrium. The additive and dominance effects of the causal loci were represented by a and d, respectively; the additive effects follow \(N\left( {0,\sigma _d^2} \right)\); the dominance effects follow \(N\left( {0,\sigma _d^2} \right)\); and e is the residual error, following \(N\left( {0,\sigma _e^2} \right)\). Therefore, \({\it{{\rm var}}}{\mathrm{(}}{\it{y}}{\mathrm{) = }}{\it{\Omega }}_a\sigma _a^2 + {\it{\Omega }}_d\sigma _d^2 + I\sigma _e^2\), in which \({\it{\Omega }}_a = \frac{{z_az_a^\prime }}{m}\) is the additive genetic relationship matrix and \({\it{\Omega }}_d = \frac{{z_dz_d^\prime }}{m}\) is the dominance genetic relationship matrix.
For HEBLP|A and HEBLP|AD methods, we estimated total additive \(\left( {\sigma _a^2} \right)\) and dominance \(\left( {\sigma _d^2} \right)\) genetic variance in the training population via Haseman-Elston regression (HE) as below
in which Y is a vector of \(\frac{{n\left( {n - 1} \right)}}{2}\) elements for the squared difference between a pair of individuals and Yij = (yi-yj)2; ωa is the additive genetic relatedness between a pair of individuals i and j, as found in the ith row and the jth column entry in Ωa; ωd is the dominance genetic relatedness between a pair of individuals i and j, similarly as found in the ith row and the jth column entry in Ωd. Alternative to HE, linear mixed model can be employed to estimate the additive and dominance variance components via restricted maximum likelihood (REML) algorithm. Of note, the difference between HE and linear mixed model are as below. HE is based on least squares, and it allows the analytical result for ba and bd, respectively. In contrast, REML is a model-based approach and the exact structure of the estimated variance, regardless of additive or dominance, remains elusive. Furthermore, as discussed in our previous study (Liu and Chen, 2017), the computational complex for HE is \({\cal O}(n^2)\), proportional to the square of sample size, but for REML \({\cal O}(n^3)\). The computational advantage of HE is important especially when the sample size is large.
Analytical results for the Haseman-Elston regression
The least-squares framework exists analytical results for the regression coefficient. Although, Eq 2 is a linear model of two regression coefficients, \(E\left( {b_a} \right) = \frac{{{\rm cov}(Y,\omega _a)}}{{{\rm var}(\omega _a)}}\) and \(E\left( {b_d} \right) = \frac{{{\rm cov}(Y,\omega _d)}}{{{\rm var}(\omega _d)}}\) because ωa and ωd are orthogonal for each locus. The general principal for deriving the analytical solution for E(ba) can be found in Chen’s study (Chen, 2014). For E(ba), \(cov\left( {Y,\omega _a} \right) = E\left( {Y\omega _a} \right) - E\left( Y \right)E\left( {\omega _a} \right) = E(Y\omega _a)\) because E(Y) = 0.
in which E(yi|xik) is the conditional probability of the phenotype given its genotype, ωa,ik as defined above. p(xik) takes value of 0.25, 0.5, and 0.25, respectively, given xik = AA, Aa, and aa. In quadric form
in which the general form of \({\boldsymbol{\beta }}^T = [\beta _1 + \left( {p_1 - q_1} \right)d_1,\beta _2 + \left( {p_2 - q_2} \right)d_2, \ldots ,\beta _m + \left( {p_m - q_m} \right)d_m]\) the vector for additive effects and IA an identity matrix with \({\boldsymbol{I}}_{A,kk} = \sqrt {2p_kq_k}\). For F2 populations, as pi = 0.5 the dominance effect di will be eliminated out from β. \({\cal M}_k = \left( {\begin{array}{*{20}{c}} {\begin{array}{*{20}{c}} {\rho _{1,k}^2} & {\rho _{2,k}\rho _{1,k}} \cr {\rho _{1,k}\rho _{2,k}} & {\rho _{2,k}^2} \end{array}} & {\begin{array}{*{20}{c}} \cdots & {\rho _{m,k}\rho _{1,k}} \cr \cdots & {\rho _{m,k}\rho _{2,k}} \end{array}} \cr {\begin{array}{*{20}{c}} \vdots & \vdots \cr {\rho _{1,k}\rho _{m,k}} & {\rho _{2,k}\rho _{m,k}} \end{array}} & {\begin{array}{*{20}{c}} \ddots & \vdots \cr \cdots & {\rho _{m,k}^2} \end{array}} \end{array}} \right)\), a symmetric matrix, indicating how the kth marker tags QTLs; for instance the entry at the ith row and the jth column ρi,k,ρj,k represents the joint LD of the ith and the jth QTLs tagged by the kth marker.
The denominator var(ωa) can be written as \(\frac{1}{{m^2}}\mathop {\sum}\nolimits_{k_1 = 1}^m {\mathop {\sum}\nolimits_{k_2 = 1}^m {\rho _{k_1k_2}^2} }\), understood as the averaged linkage disequilibrium between each pair of markers—including a marker with itself (see Appendix for the definition of effective number of markers me). Alternatively, var(ωa) can be expressed in quadric form
in which \(1^T = [1,1, \ldots 1]\) a vector for 1.
So, in quadric form
Similarly, for E(bd), we had
\(E\left( {Y\omega _d} \right) = \frac{1}{m}\mathop {\sum }\limits_{x_{ik}} \mathop {\sum }\limits_{x_{jk}} \omega _{d,ik}\omega _{d,jk}\left[ {E\left( {y_i{\mathrm{|}}x_{ik}} \right) - E\left( {y_j{\mathrm{|}}x_{jk}} \right)} \right]^2p\left( {x_{ik}} \right)p(x_{jk})\) and its quadric form
\(\frac{1}{m}{\boldsymbol{D}}^T{\boldsymbol{I}}_D\left\{ {\mathop {\sum }\limits_{k = 1}^m \left( {\begin{array}{*{20}{c}} {\begin{array}{*{20}{c}} {\rho _{1,k}^4} & {\rho _{2,k}^2\rho _{1,k}^2} \cr {\rho _{1,k}^2\rho _{2,k}^2} & {\rho _{2,k}^4} \end{array}} & {\begin{array}{*{20}{c}} \cdots & {\rho _{m,k}^2\rho _{1,k}^2} \cr \cdots & {\rho _{m,k}^2\rho _{2,k}^2} \end{array}} \cr {\begin{array}{*{20}{c}} \vdots & \vdots \cr {\rho _{1,k}^2\rho _{m,k}^2} & {\rho _{2,k}^2\rho _{m,k}^2} \end{array}} & {\begin{array}{*{20}{c}} \ddots & \vdots \cr \cdots & {\rho _{m,k}^4} \end{array}} \end{array}} \right)} \right\}{\boldsymbol{I}}_D{\boldsymbol{D}}\) in which \({\boldsymbol{D}} = [d_1,d_2, \ldots d_m]\) the vector for dominance effects and ID an identity matrix with ID,kk = 2pkqk.
The denominator is \({\rm var}\left( {\omega _d} \right) = \frac{1}{{m^2}}\mathop {\sum }\limits_{k_1 = 1}^m \mathop {\sum }\limits_{k_2 = 1}^m \rho _{k_1k_2}^4\), and in quadric form
So,
Although, E(ba) and E(bd) resemble each other, E(ba) has its kernel related to squared correlation ρ2, which is a term associated to the additive variance (Hill and Robertson, 1968), while E(bd) related to ρ4. In particular, the numerator involves the LD between a pair of markers and the denominator the LD between a pair of markers.
Of note, there are two kinds of F2 populations, the conventional F2 that is derived from F1 but not completely reproducible in term of genotypes, and in contrast there is “immortalized F2” (IF2), which can be reproduced accordingly. The IF2 can often be realized in two ways: via double haploid population (DH) (Liu et al. 2017) and from recombination inbred lines (RIL) (Hua et al. 2003). The LD differs upon F2/IF2 is used in practice. Between the \(k_1^{{\rm th}}\) and \(k_2^{{\rm th}}\) markers, for a conventional F2 and DH-derived F2 the squared correlation is \(\rho _{k_1,k_2}^2 = \left( {1 - 2c_{k_1,k_2}} \right)^2\) but \(\rho _{k_1,k_2}^2 = \left( {\frac{{1 - 2c_{k_1,k_2}}}{{1 + 2c_{k_1,k_2}}}} \right)^2\) for RIL-derived F2. For example, given the recombination of 0.1 between a pair of markers, their ρ2 = 0.64 for F2 and DH-derived F2 but 0.44 for RIL-derived F2. For dominance-associated terms, ρ4 = 0.41 for F2 and DH-derived IF2, and 0.2 for RIL-derived IF2.
For simplicity, we only consider the typical polygenic trait that the QTLs are randomly distributed along the genome, and, under this assumption, \(\sigma _a^2 = - \frac{{b_a}}{2}\) and \(\sigma _d^2 = - \frac{{b_d}}{2}\), respectively. A computer program that estimates additive and dominance heritability using Haseman-Elston regression is available from authors.
Best linear prediction (BLP)
BLP method was used to predict the genotypic value of each line of the candidate population.
in which \(\hat g_2\) is the predicted genotypic values in the candidate population; \(y_1\) is the phenotypic values in the training population; Ωa21 and Ωd21 represent the additive and the dominance genetic relationship matrix between the candidate and the training population respectively; \(\hat \sigma _a^2\) and \(\hat \sigma _d^2\) represent the estimated additive and dominance variances respectively; the inverse of the V matrix is computed using \(V^{ - 1} = \left( {\hat \sigma _a^2\Omega _{a11} + \hat \sigma _d^2\Omega _{d11} + \hat \sigma _e^2I} \right)^{ - 1}\), in which Ωa11 and Ωd11 represent the additive and the dominance genetic relationship matrix for the training population respectively.
Results
Estimates of the heritability and predictability in the simulated F2 population
We simulated a quantitative trait from F2 experimental population. In the simulated F2 population, we assumed that 1001 equal-frequent biallelic markers were evenly distributed in one chromosome [the recombination rate was c between the ith and the (i + 1)th markers]. All markers were defined as QTLs whose additive and dominance effects follow a normal distribution. Each simulation scenario included 20 replications.
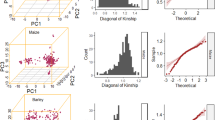
In order to assess the unbiasedness of estimating heritability via the three methods (HE|A, HE|AD, and REML|AD), we performed a Monte Carlo simulation experiment for a F2 population. When the simulated parameters were set as population size (n = 500), additive heritability (\(h_a^2 = 0.3\)), dominance heritability (\(h_d^2 = 0.2\)), and recombination rate (c = 0.01), the results showed that \(\hat h_a^2 = 0.271 \pm 0.075\) (via HE|A), \(\hat h_a^2 = 0.271 \pm 0.075\) and \(\hat h_d^2 = 0.193 \pm 0.039\) (via HE|AD), and \(\hat h_a^2 = 0.296 \pm 0.048\) and \(\hat h_d^2 = 0.226 \pm 0.052\) (via REML|AD) (Fig. 1). It indicated that all three methods could obtain unbiased estimates of parameters under the typical polygenic model.
The estimated heritability of additive and dominance based on a fixed population size (500) via HE|A, HE|AD, and REML|AD in 20 simulations when additive and dominance heritability was set at 0.3 and 0.2, respectively, and recombination rate (c) was set as 0.01. Here the HE|A only was used to estimate additive heritability. The vertical bar represents the standard deviation for 20 simulations
Moreover, we evaluated the prediction accuracy of HEBLP|AD, HEBLP|A, and GBLUP|AD under five environments in the simulated F2 population (Fig. 2). The size of both the training (nT) and the candidate population (nC) were 500 and 100 in all simulations. The squared correlation coefficient (r2) between the phenotypes and the predicted genotypic values was defined as the prediction accuracy.
Predictive ability based on a training population with a fixed population size (500), a candidate population with a fixed sample size (100), and a fixed recombination rate (c = 0.01) using HEBLP|A, HEBLP|AD, GBLUP|AD methods in 20 simulations. The value after capital letter A represents additive heritability and that after capital letter D represents dominance heritability (for example, A0.4_D0.0 represents \(h_a^2 = 0.4\) and \(h_d^2 = 0.0\)). The squared correlation coefficient (r2) between the phenotypes and the predicted genotypic values were defined as the prediction accuracy. The vertical bar represents the standard deviation for 20 simulations
In scenario 1 (\(h_a^2 = 0.4\), \(h_d^2 = 0\), and c = 0.01), the prediction accuracies were HEBLP|AD = 0.333 ± 0.066, GBLUP|AD = 0.314 ± 0.096, and HEBLP|A = 0.335 ± 0.067. In scenario 2 (\(h_a^2 = 0.40\), \(h_d^2 = 0.05\), and c = 0.01), the prediction accuracies were HEBLP|AD = 0.351 ± 0.065, GBLUP|AD = 0.355 ± 0.066, and HEBLP|A = 0.334 ± 0.065. The results of these two simulations indicated that the three methods had a similar predictive ability in the case of no or very small contribution of dominance effects to genetic variation. In scenario 3 (\(h_a^2 = 0.40\), \(h_d^2 = 0.1\), and c = 0.01), the prediction accuracies were HEBLP|AD = 0.388 ± 0.065, GBLUP|AD = 0.391 ± 0.066, and HEBLP|AD = 0.334 ± 0.067. In scenario 4 (\(h_a^2 = 0.4\), \(h_d^2 = 0.2\), and c = 0.01), the prediction accuracies were HEBLP|AD = 0.471 ± 0.063, GBLUP|AD = 0.475±0.064, and HEBLP|A = 0.335 ± 0.070. In scenario 5 (\(h_a^2 = 0.1\), \(h_d^2 = 0.6\), and c = 0.01), the prediction accuracies were HEBLP|AD = 0.553 ± 0.063, GBLUP|AD = 0.569 ± 0.067, and HEBLP|A = 0.079 ± 0.048. It indicated a similar predictability between HEBLP|AD and GBLUP|AD, and a significantly better performance than HEBLP|A in the case of a large contribution of dominance effects to genetic variation.
Comparison of computational time of HE|AD and REML|AD
We simulated F2 population based on 20 replications to evaluate the computational time of HE|AD and REML|AD. In this case, the parameters were set as population size (n = 500), additive heritability (\(h_a^2 = 0.2\)), dominance heritability (\(h_d^2 = 0.6\)), marker number (M = 3001), and recombination rate (c = 0.01). The result showed that \(\hat h_a^2 = 0.183 \pm 0.064\) and \(\hat h_d^2 = 0.568 \pm 0.068\) (via HE|AD), and \(\hat h_a^2 = 0.20 \pm 0.032\) and \(\hat h_d^2 = 0.636 \pm 0.084\) (via REML|AD), and that HE|AD and REML|AD took an average of 304 s and 3487 s in each simulation, respectively, demonstrating a significant computational advantage of HE|AD over REML|AD.
Comparison of heritability and predictability between F2 and IF2 derived from RIL using HEBLP|AD
We simulated F2 and IF2 derived from RIL population to evaluate HEBLP|AD. In this case, we simulated 1001 markers, among which 100 markers were sampled as QTLs. When we estimated the heritability and prediction accuracy, the markers representing QTLs were excluded. Each simulation scenario included 20 replications.
When the simulated parameters were set as training population size (nT = 500), candidate population size (nC = 100), additive heritability (\(h_a^2 = 0.5\)), dominance heritability (\(h_d^2 = 0.25\)), and recombination rate (c = 0.01), the results of the simulated F2 population showed \(\hat h_a^2 = 0.458 \pm 0.170\), \(\hat h_d^2 = 0.244 \pm 0.111\), and the predictability r2 = 0.614 ± 0.061 in the simulated F2 population; for the simulated IF2 populations, \(\hat h_a^2 = 0.480 \pm 0.129\), \(\hat h_d^2 = 0.230 \pm 0.075\), and the predictability r2 = 0.544 ± 0.071 in the simulated IF2 population derived from RIL population. As RIL-derived IF2 undergoing multi-generation selfing, its decayed LD resulted a much lower r2 than that of F2.
Approximation of prediction accuracy
To further understand the study, in the Appendix, we derived a formula of prediction accuracy including additive and dominance variance components. This derived result could be considered as an extension to those previously established by Daetwyler et al. (2008) and Goddard (2009).
The result showed that \(H^2\) was the upper bound of the prediction accuracy, and was further upon (1) the broad heritability (\(H^2 = h_a^2 + h_d^2\)), (2) the effective number of markers (me.a), (3) the effective number of markers of dominance heritability (me.d), and (4) the sample size of the training data. As me.a and me.d were determined by the recombination, when the markers were dense, the prediction accuracy could be further approximated as
in which l is the length of the chromosome (Morgan). Both Eq 4 and Eq 5 indicated that the upper bound of prediction accuracy was H2 when the sample size nT became infinite. Having evaluated the utility of the approximation, we found that the expected and observed prediction accuracy was consistent via HEBLP|AD under different recombination rates based on 10 simulations for F2 population (Table 1). Eq 4 and Eq 5 gave similar prediction for r2 when the markers were dense, and the accuracy of Eq 5 was reduced when the markers were sparse. The sample size of the candidate population could only influence the statistical power of the prediction accuracy because r2 followed \(\chi _1^2\) under the null hypotheses.
Genomic prediction of 7 traits in the Arabidopsis thaliana F2 population
The 7 traits, including DTF1, DTF2, DTF3, RLN, CLN, TLN, and LIR1 from a Arabidopsis thaliana F2 population were used to assess the prediction performance of HEBLP|A, HEBLP|AD, and GBLUP|AD.
We first analyzed the 7 traits via HE|A, HE|AD, and REML|AD, obtaining the estimated additive heritability varying from 0.080 to 0.582 (HE|A), 0.080 to 0.582 (HE|AD), and 0.158 to 0.731 (REML|AD), and the estimated dominance heritability varying from 0.009 to 0.052 (HE|AD), and 0.018 to 0.106 (REML|AD). The results demonstrated that dominance effects only accounted for a little proportion of genetic variation for these traits (Table 2).
Based on 100 replications, we found that the predictability of HEBLP|A, HEBLP|AD, and GBLUP|AD was similar for all traits (Table 3). For example, the prediction accuracies for DTF1 were 0.466 ± 0.028, 0.459 ± 0.032, and 0.440 ± 0.088 via HEBLP|A, HEBLP|AD, and GBLUP|AD, respectively. It indicated that, as is in the simulations, HEBLP|A, HEBLP|AD, and GBLUP|AD showed similar predictability in the case of a very small contribution of dominance effects to the genetic variation.
Discussion
The impact of the dominance heritability on predictive accuracy
The wide utilization of heterosis in the animals and plants, such as maize, rice, and cattle has significantly increased their productivity. In this study, we extended our previous method of HEBLP|A to HEBLP|AD. The simulation results demonstrated that (1) HEBLP|AD and GBLUP|AD are superior to HEBLP|A when the dominance effects can explain a significant proportion of genetic variation; (2) HEBLP|AD, GBLUP|AD, and HEBLP|A have a similar predictive ability when the dominance effects can only explain a small proportion of genetic variation. Furthermore, the real data from Arabidopsis thaliana F2 population was used to evaluate the three methods, and since the estimated heritability showed a small contribution of the dominance effects to genetic variation, the result was supportive to the second case in the simulation. de Almeida Filho et al. (2016) indicated that when the dominance effects consisted of only a small proportion in the total genetic variation, incorporating them into BayesA, BayesB, BL, and BRR would decrease the prediction accuracy. However, it is safe and stable to include dominance effects into HEBLP model under this circumstance. In addition, not limited to the F2 population as was demonstrated, HEBLP|AD is applicable as long as the populations promise the estimation of additive and dominance variance components (such as natural population of random mating).
In addition, we also provided an approximation of prediction accuracy for F2 population (Appendix). The genetic length of the chromosome, the density of markers, H2, and the sample size of the training population were key factors that would influence the prediction accuracy. The method presented in Appendix was general and could be applied to other populations. In this simulation, we simulated extremely long and single chromosome, which was unrealistic, and we will consider incorporating the real marker density into further study. We considered typical polygenic model only at present, but the interplay between genetic architecture will be included in our further studies.
Application of the genomic prediction in hybrid breeding of crops
The traditional strategy to cultivate hybrid crosses is to perform a large number of cross experiments between the inbred lines and furthermore select desirable hybrids. This process can be accelerated via combining genomic prediction approaches with immortalized F2 (IF2) population constructed by the doubled haploid (DH) population. Hua et al. (2003) first constructed IF2 population, which had the same genetic architecture as the conventional F2 population, can be generated via randomly permutated intermating of recombinant inbred lines (RILs) or DH population at present. In a hybrid breeding program, when sample size (n) of RIL or DH population is large and all crosses \(\left[ {\frac{{n(n - 1)}}{2}} \right]\) between inbred lines from the RIL or DH population need to be evaluated in the field trials, it will occupy large resources. To reduce the cost of genetic improvement, genomic prediction can be used to IF2 population to select hybrid crosses with high-hybrid performance. Guo et al. (2013) applied genomic prediction to an IF2 population derived from RIL population in maize, and Xu et al. (2014) did that in rice. Liu et al. (2017) has applied genomic prediction to IF2 population based on rapeseed DH population. However, construction of RIL population is time-consuming, and therefore the procedure of GP+IF2 (DH) will be a more efficient choice to pick out superior hybrids and potential lines with high-specific combining ability or general combining ability.
References
Arruda MP, Lipka AE, Brown PJ, Krill AM, Thurber C, Brown-Guedira G et al. (2016) Comparing genomic selection and marker-assisted selection for Fusarium head blight resistance in wheat (Triticum aestivum L.). Mol Breed 36:84
Bernardo R, Yu J (2007) Prospects for genomewide selection for quantitative traits in maize. Crop Sci 47:1082–1090
Calus MPL, Meuwissen THE, De Roos APW, Veerkamp RF (2008) Accuracy of genomic selection using different methods to define haplotypes. Genetics 178:553–561
Chen G-B (2014) Estimating heritability of complex traits from genome-wide association studies using IBS-based Haseman-Elston regression. Front Genet 5:107
Daetwyler HD, Villanueva B, Woolliams JA (2008) Accuracy of predicting the genetic risk of disease using a genome-wide approach. PLoS ONE 3:e3395
de Almeida Filho JE, Guimarães JFR, e Silva FF, de Resende MDV, Muñoz P, Kirst M et al. (2016) The contribution of dominance to phenotype prediction in a pine breeding and simulated population. Heredity 117:33–41
Denis M, Bouvet J-M (2011) Genomic selection in tree breeding: testing accuracy of prediction models including dominance effect. BMC Proc 5:O13
Denis M, Bouvet JM (2013) Efficiency of genomic selection with models including dominance effect in the context of Eucalyptus breeding. Tree Genet Genomes 9:37–51
Goddard M (2009) Genomic selection: prediction of accuracy and maximisation of long term reponse. Genetica 136:245–257
Guo T, Li H, Yan J, Tang J, Li J, Zhang Z et al. (2013) Performance prediction of F1 hybrids between recombinant inbred lines derived from two elite maize inbred lines. Theor Appl Genet 126:189–201
Hayes BJ, Bowman PJ, Chamberlain AJ, Goddard ME (2009) Genomic selection in dairy cattle: progress and challenges. J Dairy Sci 92:433–443
Heffner EL, Sorrells ME, Jannink JL (2009) Genomic selection for crop improvement. Crop Sci 49:1–12
Hill WG, Robertson A (1968) Linkage disequilibrium in finite populations. Theor Appl Genet 38:226–231
Hua J, Xing Y, Wu W, Xu C, Sun X, Yu S et al. (2003) Single-locus heterotic effects and dominance by dominance interactions can adequately explain the genetic basis of heterosis in an elite rice hybrid. Proc Natl Acad Sci USA 100:2574–2579
Jannink J-L, Lorenz AJ, Iwata H (2010) Genomic selection in plant breeding: from theory to practice. Brief Funct Genom 9:166–177
Li L, Lu K, Chen Z, Mu T, Hu Z, Li X (2008) Dominance, overdominance and epistasis condition the heterosis in two heterotic rice hybrids. Genetics 180:1725–1742
Liu H, Chen G-B (2017) A fast genomic selection approach for large genomic data. Theor Appl Genet 130:1277–1284
Liu P, Zhao Y, Liu G, Wang M, Hu D, Hu J et al. (2017) Hybrid performance of an immortalized F2 rapeseed population is driven by additive, dominance, and epistatic effects. Front Plant Sci 8:815
Meuwissen TH, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829
Nishio M, Satoh M (2014) Including dominance effects in the genomic BLUP method for genomic evaluation. PLoS ONE 9:e85792
Resende RT, Resende MDV, Silva FF, Azevedo CF, Takahashi EK, Silva-Junior OB et al. (2017) Assessing the expected response to genomic selection of individuals and families in Eucalyptus breeding with an additive-dominant model. Heredity 119:245–255
Riedelsheimer C, Czedik-Eysenberg A, Grieder C, Lisec J, Technow F, Sulpice R et al. (2012) Genomic and metabolic prediction of complex heterotic traits in hybrid maize. Nat Genet 44:217–220
Salomé PA, Bomblies K, Laitinen RAE, Yant L, Mott R, Weigel D (2011) Genetic architecture of flowering-time variation in Arabidopsis thaliana. Genetics 188:421–433
Schaeffer LR (2006) Strategy for applying genome wide selection in dairy cattle. J Anim Breed Genet 123:218–223
Su G, Christensen OF, Ostersen T, Henryon M, Lund MS (2012) Estimating additive and non-additive genetic variances and predicting genetic merits using genome-wide dense single nucleotide polymorphism markers. PLoS ONE 7:e45293
Technow F, Riedelsheimer C, Schrag TA, Melchinger AE (2012) Genomic prediction of hybrid performance in maize with models incorporating dominance and population specific marker effects. Theor Appl Genet 125:1181–1194
VanRaden PM (2008) Efficient methods to compute genomic predictions. J Dairy Sci 91:4414–4423
Vitezica ZG, Legarra A, Toro MA, Varona L (2017) Orthogonal estimates of variances for additive, dominance, and epistatic effects in populations. Genetics 206:1297–1307
Wang X, Li L, Yang Z, Zheng X, Yu S, Xu C et al. (2017) Predicting rice hybrid performance using univariate and multivariate GBLUP models based on North Carolina mating design II. Heredity 118:302–310
Xu S, Zhu D, Zhang Q (2014) Predicting hybrid performance in rice using genomic best linear unbiased prediction. Proc Natl Acad Sci USA 111:12456–12461
Zhang Z, Zhang Q, Ding XD (2011) Advances in genomic selection in domestic animals. Chin Sci Bull 56:2655–2663
Acknowledgements
This study was supported by the National Natural Science Foundation of China (31771392 to G.-B.C.).
Author contributions
H.L. and G.-B.C. designed and performed the study as well as wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Appendix
Appendix
Factors influence prediction accuracy for F2 population
In this note, we try to outline the factors that influence the prediction accuracy for an F2 population.
For the training population, its phenotype can be expressed as
Here, we assume every marker is causal and has small effects, a typical polygenic trait. \(x_{a_j}\) and \(x_{d_j}\) are the orthogonal coding of the jth marker for the additive and dominance effect. var(y) = Vp the phenotypic variance, and \({\rm var}\left( {\mathop {\sum }\limits_{j = 1}^m b_{a_j}x_{a_j} + \mathop {\sum }\limits_{j = 1}^m b_{d_j}x_{d_j}} \right) = h_A^2 + h_D^2 = H^2\).
According to linear regression theory, for the additive effect for the jth marker can be estimated as \(\hat b_{a_j} = \frac{{{\rm cov}(y,x_{a_j})}}{{{\rm var}(x_{a_j})}}\) and rewritten as \(b_{a_j} + \sigma _{\hat b_{a_j}}\), in which \(\sigma _{\hat b_{a_j}} = \frac{{\sigma _e^2}}{{N_T\sigma _{x_{a_j}}^2}}\) the sampling variance of the estimate; for the dominance effect, \(\hat b_{d_j} = \frac{{{\rm cov}(y,x_{d_j})}}{{{\rm var}(x_{d_j})}}\), and \(\sigma _{\hat b_{d_j}} = \frac{{\sigma _e^2}}{{N_T\sigma _{x_{d_j}}^2}}\). NT is the sample size of the training population, and m is the number of markers.
For the candidate population, the phenotype can be expressed as \(y_C = a + \mathop {\sum}\nolimits_{j = 1}^k {b_{a_j}\tilde x_{a_j}} + \mathop {\sum}\nolimits_{j = 1}^k {b_{d_j}\tilde x_{d_j} + \varepsilon _T}\), while the predicted genotypic values \(\hat y_C = \mathop {\sum}\nolimits_{j = 1}^k {\hat b_{a_j}\tilde x_{a_j}} + \mathop {\sum}\nolimits_{j = 1}^k {\hat b_{d_j}\tilde x_{d_j}}\). It is easy to derive the variance and covariance terms below.
The prediction accuracy is
For genetic value
The prediction accuracy between the true genotypic values and the predicted genotypic values can be written as squared Pearson’s correlation
This equation is an extension of the one as derived by Daetwyler et al. (2008), but here we include the dominance component. In practice, the prediction accuracy is more relevant to the effective number of loci, which can be understood as quasi-independent segment of the whole genome. So, the prediction accuracy is approximated as
in which me,a and me,d are the effective number of markers coded for additive and dominance effects.
As for markers not on the same chromosome, the LD is nearly zero, so \(m_{e.a} = \frac{{m^2}}{{m + \mathop {\sum }\nolimits_{i = 1}^k \mathop {\sum }\nolimits_{i \ne j}^k r_{ij}^2}}\)
If the recombination is based on Haldane map function, for F2 \(r_{ij}^2 = \exp \left( { - 4\left| {d_i - d_j} \right|} \right) = e^{ - 4d_{ij}}\), in which di,j = |di−dj| is the genetic distance (Morgan) between a pair of loci, and \(r_{ij}^2 = e^{ - 8d_{ij}}\). Obviously, when there is no LD between markers, \(r_{ij}^2 = 0\), and me,a = m, me,d = m. As \(r_{ij}^4 \le r_{ij}^2\), we have \(m_{e.a} \le m_{e.d} \le m\).
Further approximation for the prediction accuracy
For the additive component,
and for the dominance component,
in which \(c_{2l}\) is the recombination fraction given the genetic distance of 2l.
So, \(m_{e.a} = \left[ {\frac{1}{m} + \frac{{\left( {l_1 - \frac{{c_{2l}}}{2}} \right)}}{{2l^2}}} \right]^{ - 1}\), if the markers are dense, and m>>l1 (m is often greater than 10,000 along a single chromosome), \(m_{e.a} \approx 2l\); similarly, \(m_{e.a} \approx 4l\). So, the prediction accuracy can be further approximated as
when the density of markers is high.
So the expectation of the prediction accuracy is upon the training sample size, but the statistical significance of r2 depends on the sample size of the candidate sample size. Under the null distribution r2 follows \(\chi _1^2\), so the non-centrality parameter for the statistical test of r2 is \(\lambda = \frac{{n_Cr^2}}{{1 - r^2}}\), in which nC is the sample size of the candidate population.
In genomic prediction, the additive genomic relationship matrix can be used to estimate me.a. Given A, an nT × nT matrix, the additive genomic relationship matrix, if we estimate variance, \(\sigma _{A_o}^2\), of the \(\frac{{n_T\left( {n_T - 1} \right)}}{2}\) off-diagonal elements, and \(\hat m_{e.a} = \frac{1}{{\sigma _{A_o}^2}}\); similarly, we can have \(\hat m_{e.d} = \frac{1}{{\sigma _{D_o}^2}}\) for the dominance effective number of markers.
Rights and permissions
About this article
Cite this article
Liu, H., Chen, GB. A new genomic prediction method with additive-dominance effects in the least-squares framework. Heredity 121, 196–204 (2018). https://doi.org/10.1038/s41437-018-0099-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-018-0099-5
This article is cited by
-
A dimensionality-reduction genomic prediction method without direct inverse of the genomic relationship matrix for large genomic data
Plant Cell Reports (2023)
-
Including dominance effects in the prediction model through locus-specific weights on heterozygous genotypes can greatly improve genomic predictive abilities
Heredity (2022)
-
Transcriptome analysis reveals the molecular mechanisms of heterosis on thermal resistance in hybrid abalone
BMC Genomics (2021)