Abstract
Purpose
Exome sequencing (ES) powerfully identifies the molecular bases of heterogeneous conditions such as intellectual disability and/or multiple congenital anomalies (ID/MCA). Current ES analysis, combining diagnosis analysis restricted to disease-causing genes reported in OMIM database and subsequent research investigation extended to other genes, indicated causal and candidate genes around 40% and 10%. Nonconclusive results are frequent in such ultrarare conditions that recurrence and genotype-phenotype correlations are limited. International data-sharing permits the gathering of additional patients carrying variants in the same gene to draw definitive conclusions on their implication as disease causing. Several web-based tools have been developed and grouped in Matchmaker Exchange. In this study, we report our current experience as a regional center that has implemented ES as a first-line diagnostic test since 2013, working with a research laboratory devoted to disease gene identification.
Methods
We used GeneMatcher over 2.5 years to share 71 novel candidate genes identified by ES.
Results
Matches occurred in 60/71 candidate genes allowing to confirm the implication of 39% of matched genes as causal and to rule out 6% of them.
Conclusion
The introduction of user-friendly gene-matching tools, such as GeneMatcher, appeared to be an essential step for the rapid identification of novel disease genes responsible for ID/MCA.
Similar content being viewed by others
INTRODUCTION
Developmental disorders are individually rare but together affect millions of people, significantly contributing to morbidity and remaining a major public health issue. Currently, the number of phenotypes with or suspected to have a Mendelian basis described in OMIM is 8634, with 200 novel phenotypes being described every year and many atypical and/or isolated phenotypes found in PubMed.1 Conditions characterized by intellectual disability and/or multiple congenital anomalies (ID/MCA) appeared particularly resistant to old approaches of gene identification likely because of the small number of cases with a specific phenotype and locus heterogeneity. During the past decade, exome sequencing (ES) has become highly powerful in identifying the molecular basis of ID/MCA.2,3,4 But because of the difficulty in recruiting an adequate number of affected cases with ultrarare conditions, the heterogeneous clinical spectrum, the uncertain functional impacts of variants, and the small number of known genes involved in diseases (~3500 genes), a large majority of results remain nonconclusive. Variants identified in a unique affected family need further supporting evidence to prove their pathogenicity.5 For that reason, further functional studies and/or recurrence of pathogenic variants in the same gene in unrelated affected cases are required.6,7 Gilissen et al.7 estimate that three unrelated cases with homozygous or compound heterozygous variants are sufficient to confirm the gene–disease link in autosomal recessive phenotypes, while five affected cases with heterozygous variants in the same gene are sufficient for an autosomal dominant phenotype. However, in clinical and/or molecular heterogeneous diseases, more additional cases and multiple functional studies are required to limit the possibility of finding patients with variants in the same gene just by chance.7 But, the restricted connections between scientists and clinicians limit the identification of other cases with similar phenotype and variants in the same gene, making the interpretation of ES data and the identification of novel disease genes harder.
During the past few years, several web-based tools have been developed to share phenotype and genotype data and to broaden the exchange between scientific and medical teams. In 2013, the Matchmaker Exchange project (http://www.matchmakerexchange.org/) was created to connect data-sharing tools and to facilitate the matching of cases using a common application-programming interface to accelerate the identification of novel disease genes in human diseases.8 Currently, seven projects developed by different institutions around the world are connected in Matchmaker Exchange: Australian Genomics Health Aliiance (AGHA) Patient Archive (https://mme.australiangenomics.org.au/#/home), DECIPHER (https://decipher.sanger.ac.uk/), GeneMatcher (https://www.genematcher.org/), matchbox (https://seqr.broadinstitute.org/), Monarch Initiative (https://monarchinitiative.org/), MyGene2 (https://mygene2.org/MyGene2/) and PhenomeCentral (https://www.phenomecentral.org/).8 The concept is to post de-identified phenotype or genotype of cases on one of these websites and be connected with other submitters with an interest in the same posted information. Each submitter receives an electronic message with contact information to exchange further clinical and molecular data that may confirm or reverse the involvement of the candidate gene in the disease being investigated. These interactions foster international collaboration and lead to functional studies to improve the knowledge on rare Mendelian phenotypes. For example, as of 1 August 2018, GeneMatcher has 5690 submitters from 76 countries and 9487 candidate genes submitted (https://genematcher.org/statistics). The number of users and candidate genes is clearly growing in data-sharing platforms8,9 and the connection between the platforms by Matchmaker Exchange improves the novel disease gene discovery.
We report our experience on the use of GeneMatcher for sharing several genes with variants of uncertain significance to identify further evidence supporting their role as disease-causing.
MATERIAL AND METHODS
Patients
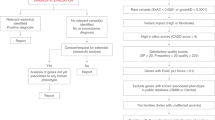
From June 2015 to January 2018, we identified 71 candidate genes (Table S1) by exome sequencing (ES) in 71 unrelated individuals with ID, including isolated ID/epileptic encephalopathy (EE) in 28/71 individuals and syndromic ID/MCA in 43/71 individuals (Supplemental Fig. 1A). The clinical data of all patients were shared in PhenomeCentral.
Data-sharing
We entered these 71 candidate genes in GeneMatcher: 66/71 genes were not associated with a human disease, 2/71 genes have previously been published in the literature but remained absent in OMIM, and 3/71 genes were reported in OMIM but with a different type of variant, such as large deletions versus missense (1/3) or associated to a more severe phenotype suggesting a novel gene–phenotype association or a phenotypic expansion (2/3) (Figure S1).
Data collection
After sharing in GeneMatcher, we gathered (1) the number of matches and collaborations initiated for each gene, (2) the number of definitive conclusions allowed by data-sharing, (3) the number of recruited cases to confirm the implication of the gene in the disease being investigated, (4) the time spent (in minutes, hours, or days) between the first submission and the first match, and between the first email contact and the first answer.
RESULTS
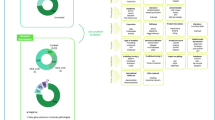
Among the 71 submitted candidate genes, 60 genes matched (84%), with an average of 4.2 matches per genes. The number of matches ranges from 1 to 34 matches with a median at 2 matches (Fig. 1a). In 23/60 genes (39%), the matching confirmed the gene’s pathogenicity whereas likely benign status was concluded in 6/60 genes (10%). Note that two candidate genes were confirmed likely pathogenic by other sources after the GeneMatcher submission (KLHL7 and UNC45A). The matches resulted in international collaborations with future publications for 28/60 matched genes (51%). We have also noted a significant increase in the number of matches since 2015 (Fig. 1c). At this time, 42/71 genes (58%) remain as candidate genes according to criteria determined by Gilissen et al.7 (Fig. 1b) because of insufficient evidence gathered by the matches so far (45%) or absent matching (13%).
Repartition of submitted genes in GeneMatcher. (a) Number of matches by submitted genes. (b) Repartition of the results after GeneMatcher submission. (c) Number of matches and submitted candidate genes between June 2015 and December 2017. (d) Waiting delay after first submission in 2015, 2016, and 2017.
In parallel, we measured the time interval between the submission and the first match (Figs. 1d and 2a). The waiting period varied from less than 5 minutes to a few months, with a median of 4 hours. Similarly, the response time after the first email sent to contact another submitter was very variable ranging from 1 minute to several months; the median was 31 hours. Nine of 38 submitters did not respond to our contacting emails. Among the 228 matches, 155 were within the United States and more than half of those were with GeneDx (Fig. 2b). We also often matched with submitters from Canada, the Netherlands, or France, and occasionally with other European countries, the Middle East, East Asia, and Australia, for a total of 19 individual countries.
DISCUSSION
This study demonstrates our positive experience with GeneMatcher, as 33% of the matched candidate genes were new genes implicated in rare diseases. Our matches resulted in a number of international collaborations with 8 scientific publications10,11,12,13,14,15,16 and 19 manuscripts in progress, 10 of them being led by our team (Figure S2).
GeneMatcher is a powerful tool to recruit additional cases that previously were only recruited by communications in congresses or personal knowledge networks. These interactions are essential to accelerate the identification of novel disease genes.17 The number of pathogenic variants is higher in the ID/EE cohort (73%) than in the MCA cohort (26%), and this concerns mainly sporadic mode of inheritance (78%). These results are explained by predominantly submitted genes being implicated in sporadic ID at the beginning of data-sharing. While the vast majority of predicted loss-of-function variants were confirmed as pathogenic, only 25% of missense variants were confirmed as pathogenic (Figure S1C), illustrating the difficulties in interpreting the impact of missense variants in gene function.18,19,20
The response time to a match appears variable and depends on the time zones and submitters' constraints, including time constraints, competition, or the delay of internal communications. GeneMatcher proved to be time efficient because it took us an average of 1 month (median 10 days) to collect a minimum number of patients and to confirm the pathogenicity of 18/36 genes (50%). The median waiting time for the matching notification decreased from 36 hours in 2015 to 2 hours in 2017 (Fig. 1d). This decrease is likely explained by the growing number of genes submitted and submitters worldwide due to increased access to ES.9 The number of submitted genes has exponentially increased in GeneMatcher, from 500 genes in April 2014 to more than 8000 in January 2018, resulting in an important increased chance of matches.8 Most of the submitters in GeneMatcher are from North America and Europe, in particular countries that encourage the use of this platform and promote the international data-sharing, such as the United States, France, and the Netherlands (Figure S3).
The absence of matches or inconclusive matches (58% genes) do not exclude that the candidate genes submitted are the cause of the disease. Indeed, for some ultrarare diseases, the small number of clinicians or scientists in some disciplines sharing genotypic data makes it harder to identify additional cases using data-sharing platforms. This is particularly true for MCA without ID, such as the novel syndrome characterized by cholestasis, diarrhea, deafness, and bone fragility and caused by variants in UNC45A.13 The UNC45A gene did not match in GeneMatcher but was confirmed by the recruitment of three additional cases in specialized scientific meetings. The KLHL7 gene also did not match in GeneMatcher but was confirmed as the causal gene by the identification of additional consanguineous families in scientific meetings.14 To improve our chances of resolving rare Mendelian phenotypes, the GeneMatcher initiative needs to be better known and more used by medical specialists to share the candidate genes of their cases including cases with MCA without ID and recessive modes of inheritance. A longer-time will be necessary to get enough matches and information that will allow us to conclude that a candidate gene is the cause of a rare Mendelian phenotype. We identified the PACS2 gene as a candidate gene in November 2014 and only a few additional cases were identified by personal networks. It took more than a year to match for the first time in GeneMatcher and to collect a large enough cohort of patients confirming a hotspot missense variant as causal.15 GeneMatcher has progressively expanded with an increasing number of submitted genes and submitters, making it more efficient.8 After each match, the exchange of clinical and molecular data between submitters allowed additional strong arguments in favor, or not, of the involvement of the candidate genes. Following these discussions, the decisions to collaborate, to elaborate functional studies, and/or to recruit additional patients were made. The candidate genes could therefore be issued in the short or long term, depending on the submitter’s choice.
In conclusion, the identification of novel disease genes remains essential for clinical characterization, diagnosis, prognosis, and genetic counseling. GeneMatcher is a free, powerful, international data-sharing tool with the goal of connecting scientists/clinicians interested in the same candidate gene. We demonstrate its rapid efficiency to confirm (33%) or reverse (6%) the pathogenicity of a candidate gene and to develop collaborations in the rare Mendelian phenotype field. In the coming years, we would like GeneMatcher to expand the spectrum of phenotypes being entered into the system, including all cases of rare developmental disorders.
References
Boycott KM, Vanstone MR, Bulman DE, MacKenzie AE. Rare-disease genetics in the era of next-generation sequencing: discovery to translation. Nat Rev Genet. 2013;14:681–691.
Cooper DN, Chen JM, Ball EV, et al. Genes, mutations, and human inherited disease at the dawn of the age of personalized genomics. Hum Mutat. 2010;31:631–655.
Chong, JX Buckingham KJ, Jhangiani SN, et al. The genetic basis of Mendelian phenotypes: discoveries, challenges, and opportunities. Am J Hum Genet. 2015;97:199–215.
Vissers LELM, Gilissen C, Veltman JA. Genetic studies in intellectual disability and related disorders. Nat Rev Genet. 2016;17:9.
Vissers LELM, van Nimwegen KJM, Schieving JH, et al. A clinical utility study of exome sequencing versus conventional genetic testing in pediatric neurology. Genet Med. 2017;19:1055–1063.
Cassa CA, Akle S, Jordan DM, Rosenfeld JA When “N of 2” is not enough: integrating statistical and functional data in gene discovery. Cold Spring Harb Mol Case Stud. 2017;3:a001099.
Gilissen C, Hoischen A, Brunner HG, Veltman JA. Disease gene identification strategies for exome sequencing. Eur J Hum Genet. 2012;20:490–497.
Sobreira N, Schiettecatte F, Valle D, Hamosh A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum Mutat. 2015;36:928–930.
Buske, OJ, Girdea M, Dumitriu S, et al. PhenomeCentral: a portal for phenotypic and genotypic matchmaking of patients with rare genetic diseases. Hum Mutat. (2015). https://doi.org/10.1002/humu.22851
Assoum M, Philippe C, Isidor B, et al. Autosomal-recessive mutations in AP3B2, adaptor-related protein complex 3 beta 2 subunit, cause an early-onset epileptic encephalopathy with optic atrophy. Am J Hum Genet. 2016;99:1368–1376.
Chiu, ATG, Pei SLC, Mak CCY, et al. Okur-Chung neurodevelopmental syndrome: Eight additional cases with implications on phenotype and genotype expansion. Clin Genet. 2018;93:880–890.
Lehalle D, Mosca-Boidron AL, Begtrup A, et al. STAG1 mutations cause a novel cohesinopathy characterised by unspecific syndromic intellectual disability. J Med Genet. 2017;54:479–488.
Esteve C, Francescatto L, Tan PL, et al. Loss-of-function mutations in UNC45A cause a syndrome associating cholestasis, diarrhea, impaired hearing, and bone fragility. Am J Hum Genet. 2018;102:364–374.
Bruel A-L, Bigoni S, Kennedy J, et al. Expanding the clinical spectrum of recessive truncating mutations of KLHL7 to a Bohring-Opitz-like phenotype. J Med Genet. 2017;54:830–835.
Olson HE, Jean-Marçais N, Yang E, et al. A recurrent de novo PACS2 heterozygous missense variant causes neonatal-onset developmental epileptic encephalopathy, facial dysmorphism, and cerebellar dysgenesis. Am J Hum Genet. 2018;102:995–1007.
Moutton S, Bruel AL, Assoum M, et al. Truncating variants of the DLG4 gene are responsible for intellectual disability with marfanoid features. Clin Genet. 2018;93:1172–1178.
Julkowska D, Austin CP, Cutillo CM, et al. The importance of international collaboration for rare diseases research: a European perspective. Gene Ther. 2017;24:562–571.
Richards S, Aziz N, Bale S, ACMG Laboratory Quality Assurance Committee, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–423.
Sivley RM, Dou X, Meiler J, Bush WS, Capra JA. Comprehensive analysis of constraint on the spatial distribution of missense variants in human protein structures. Am J Hum Genet. 2018;102:415–426.
Traynelis J, Silk M, Wang Q, et al. Optimizing genomic medicine in epilepsy through a gene-customized approach to missense variant interpretation. Genome Res. 2017;27:1715–1729.
Acknowledgements
This work was supported by grants from the Regional Council of Burgundy (to C.T.-R.), the FEDER, and PARI 2015. We thank GeneMatcher and collaborators.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Bruel, AL., Vitobello, A., Mau-Them, F.T. et al. 2.5 years’ experience of GeneMatcher data-sharing: a powerful tool for identifying new genes responsible for rare diseases. Genet Med 21, 1657–1661 (2019). https://doi.org/10.1038/s41436-018-0383-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-018-0383-z
Keywords
This article is cited by
-
PhenoDB, GeneMatcher and VariantMatcher, tools for analysis and sharing of sequence data
Orphanet Journal of Rare Diseases (2021)