Abstract
Schilbach–Rott syndrome (SRS, OMIM%164220) is a disorder of unknown aetiology that is characterised by hypotelorism, epichantal folds, cleft palate, dysmorphic face, hypospadia in males and mild mental retardation in some patients. To date, 5 families and 17 patients have exhibited this phenotype, and recurrence in two of these families suggests an autosomal dominant inheritance. SRS overlaps with a mild form of holoprosencephaly (HPE), but array–CGH analysis and sequencing of some HPE-related genes (SEPT9, SHH and TWIST) did not reveal any variants in at least one family. Herein, we investigated by array–CGH analysis a 11-year-old female patient and her father, both exhibiting the typical SRS phenotype, disclosing in the daughter–father couple the same microduplication of chromosome 9q22.32q22.33 [arr[hg19]9q22.32(98,049,611_98,049,636)x3,9q22.33 (99,301,483_99,301,508)x3], involving eight genes, including PTCH1. The duplication segregated with the disease, since it was not found in the healthy paternal grandparents of the proband. The gain-of-function variants of the PTCH1 gene are responsible for a mild form of HPE. This is the first genetic variant found in SRS. This finding reinforces the hypothesis that SRS belongs to the HPE clinical spectrum and suggests to perform array–CGH in patients with SRS phenotype and, if negative, to consider a potential benefit from sequencing of HPE-related genes.
Similar content being viewed by others
Introduction
Schilbach and Rott [1] described ten relatives over five generations who exhibited a congenital disorder that is characterised by ocular hypotelorism, submucosal cleft palate and hypospadias in males. Other anomalies included blepharophimosis, upslant of palpebral fissures and a tendency for cutaneous syndactyly of the 3rd and 4th fingers and 2nd and 3rd toes. In 2002, Joss et al. [2] described a mother and two sons with cleft palates and facial appearances that closely resembled the Schilbach–Rott syndrome (SRS) (OMIM%164220). A father–son pair from Mexico who exhibited the typical features of SRS, such as ocular hypotelorism, cleft palate, hypospadias (only in the child) and microcephaly, further supports an autosomal inheritance [3]. De Carvalho et al. [4] described a 4-year-old girl with blepharophimosis, which is a typical facial gestalt and skeletal abnormalities seen in the blepharofacioskeletal syndrome (BFSS), and found a clinical overlap with SRS; the findings suggested that BFSS and SRS were of the same condition. Shkalim et al. [5] described another family with an autosomal dominant syndrome of hypotelorism, cleft palate/uvula, high-arched palate and mild mental retardation—symptoms resembling SRS—and noted a similarity in the facial appearance of patients with holoprosencephaly, even in the absence of holoprosencephaly on MRI. The results from genetic studies were normal in this case, including FISH for deletion of 22q11, karyotype analysis, fragile X testing, high-resolution comparative genomic hybridisation (CGH) and SEPT9, SHH and TWIST sequence analyses. These data suggest that SRS is an autosomal dominant developmental disorder of unknown aetiology.
Herein, we described a father–daughter couple exhibiting an SRS phenotype in whom we identified a 9q22.32q22.33 microduplication involving the PTCH1 gene.
Materials and methods
Informed consent for the genetic studies and the publication of images was obtained from all family members. Cognitive evaluations of patients were performed using the Wechsler Intelligence Scale for Children, revision III (WISC-III). Array–CGH analyses were performed on DNA extracted from peripheral blood of the proband and her parents using the Agilent Human Genome CGH Microarray Kit 60K (Agilent Technologies, Santa Clara, California, USA), as described previously [6, 7]. The array–CGH results were validated by a second array–CGH analysis. DNA extracted from peripheral blood was also obtained for paternal grandparents of the proband and a real-time PCR (QPCR) on the PTCH1 gene was performed, as previously described [8].
Subjects
The patient, evaluated 2 years ago, was a 11-year-old female, the only child of a non-consanguineous couple. The personal and family history of her mother and family history of her father were unremarkable. The personal history of the father was characterised by the presence of anterior glandular hypospadias at birth, which was surgically treated at 18 months of age. He exhibited dysmorphisms of the face very similar to the daughter, including microbrachicephaly, hypotelorism, flat face, short philtrum, beaked and long nose with a notch on the tip, high-arched palate, short palpebral fissures, blepharophimosis and clinodactyly of 5th fingers (Fig. 1a, b). The paternal grandparents of the proband were healthy and did not show dysmorphism of the face. The proband’s pregnancy was complicated by maternal bleeding during the first trimester. Foetal ultrasound performed at 22 weeks of gestation revealed a foetal central cleft lip and a deceleration of foetal growth was observed during the last month of gestation. The parents decided to not perform invasive prenatal genetic testing. Spontaneous vaginal delivery occurred at 40 weeks of gestation, with a birth weight of 2300 g (<3rd percentile), a length of 47 cm (10th–25th percentile) and an OFC of 31 cm (3rd percentile). A central cleft of the lip and palate was immediately evident and surgically treated during the first few months of life. Motor development was achieved normally, but speech was delayed. Therefore, she received speech therapy. The patient did not experience seizures or other neurological disorders. Diagnostic workup included haematological and biochemical testing, urinalysis, ECG and echocardiography, ABR, abdominal ultrasounds, including urinary system evaluation and brain MRI, all of which were unremarkable. Her height was 140 cm (25th centile), her weight was 35.5 kg (50th centile) and her OFC was 48.5 cm (<<3rd centile) at the time of our evaluation (11 years old). Physical examination disclosed microbrachicephaly, short palpebral fissures, blepharophimosis, hypotelorism, long nose, short philtrum, flat malar region, asymmetric face with ptosis of the right eye, dental crowding, frontal angioma and clinodactyly of the 5th fingers (Fig. 1c, d).
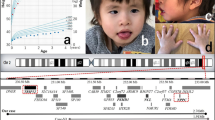
a, b Father of the proband, 40 years old. Note the hypotelorism, short philtrum, beaked, long nose with a notched tip, posteriorly rotated ears, short palpebral fissures and asymmetric face with ptosis of the left eye. c, d Proband, 11 years old. Note the similarity with the paternal phenotype: hypotelorism, short philtrum, long nose, small mouth with surgical scar of labiopalatoschisis, flat midface, posteriorly rotated ears, short palpebral fissures and asymmetric face with ptosis of the left eye
Results
The proband evidenced a full-scale IQ of 70, performance IQ of 75 and a verbal IQ of 65 on the WISC-III, which is in the borderline intellectual functioning range. The array–CGH analysis of the proband revealed a 1.2-Mb duplication of chromosome 9q22.32q22.33 [arr[hg19]9q22.32(98,049,611_98,049,636)x3,9q22.33 (99,301,483_99,301,508)x3]. These two patients have been registered in the Decipher database (https://decipher.sanger.ac.uk/) with the following identification codes: SAN376908 (proband) and SAN376909 (father) [9]. The array–CGH analyses were extended to the parents and disclosed the paternal origin of the duplication. These results were confirmed in the proband and in her father using a second array–CGH analysis (Fig. 2). We also performed a QPCR analysis of the PTCH1 gene in the healthy paternal grandparents who did not show the 9q22.32q22.33 microduplication (Fig. 3). The rearrangement involved the following OMIM genes: FANCC, PTCH1, HSD17B3, ERCC6L2, SLC35D2, HABP4, ZNF367 and CDC14B (Fig. 2).
In the upper figure, the three partial array–CGH profiles of the mother, proband and father show the presence of microduplication in the last two subjects involving the chromosomal region 9q22.32q22.33. In the lower figure, the gene content is illustrated. Please note the presence of the entire PTCH1 gene
Results of the QPCR performed on exons (ex) 6, 14, 23 of the PTCH1 gene (genomic DNA) in two healthy controls CTRL_A (grey) and CTRL_B (black), paternal grandmother (blue), paternal grandfather (red), proband (green) and her father (purple). Note that duplication of PTCH1 is present only in the father–daughter couple
Discussion
SRS is a very rare dysmorphic condition of unknown aetiology, with less than 20 patients reported to date [1,2,3,4,5]. SRS patients share facial features with patients who have holoprosencephaly (HPE), even when their brain appears normal, as in the microform of HPE [5, 10]. Microforms are defined as HPE with facial dysmorphisms (ocular hypothelorism, midface hypoplasia, cleft lip and/or palate), minimal (corpus callosum defect, posterior fossa abnormalities) or no brain involvement, possible extra craniofacial congenital defects, mostly visceral and renal/urinary anomalies and intellectual disability of various degrees [5]. Shkalim et al. [5] analysed the SHH gene, which is the gene most frequently involved in HPE in SRS patients; however, no functionally relevant variants were found. However, HPE exhibits a wide genetic heterogeneity with at least 12 different loci and 7 identified genes (SHH, SIX3, PTCH1, GLI2, ZIC2, CDON and TGIF) [11]. SHH is also the gene most commonly responsible for HPE microform [10]. Notably, in contrast to the loss-of-function variants and genomic deletions of PTCH1 found in basal cell naevus syndrome (BCNS or Gorlin syndrome, OMIM #109400), at least five different missense PTCH1 variants, encoding for the SHH receptor, that normally acts to repress SHH signalling [11], were found in patients with HPE or holoprosencephaly-like features and normal brain (HPE7 or HPE microform) [12,13,14]. These variants likely affect the protein function through a gain-of-function manner [14]. Moreover, Derwinska et al. [15] identified an ~360-kb duplication in 9q22.32, involving the entire PTCH1 gene, in a 21-month-old boy and his mother, who both exhibited a mild intellectual disability and microcephaly. Izumi et al. [16] described a syndrome that was characterised by short stature, microcephaly, cognitive delay and facial features in three patients (mother and two children) carrying a microduplication of ~2.3 Mb completely involving the PTCH1 locus. Other larger duplications involving the 9q22q31 were described in at least 30 patients who shared the following clinical features: short stature, microcephaly, intellectual disability, dysmorphic facial features and congenital heart disease [17,18,19]. However, there were small duplications of 9q22.32, such as those described by Derwinska et al. [15]. Izumi et al. [17] in this report, restrict the number of genes that are potentially responsible for the primary clinical phenotype of this newly, clinically recognisable, duplication syndrome (Fig. 4). In Table 1, we compare the phenotypes of SRS with those of our patients, patients with a small 9q22.3 microduplication and patients with gain-of-function PTCH1 variants. There are many clinical features shared by these patients, such as microcephaly, short stature, high or cleft palate, hypotelorism, syndactyly or clinodactyly, and usually mild, intellectual disability; instead, other characteristics, such as brain anomalies and hypospadia, are more variable.
The role of PTCH1 in the pathogenesis of the main features (microcephaly, hypotelorism, high palate and intellectual disability) that characterise the phenotype associated with 9q22.3 microduplication appears likely. In fact, it is plausible that a duplication of the entire PTCH1 gene, which might lead to an overexpression of the protein, could reduce SHH signalling during embryogenesis in a dose-dependent manner, since PTCH1 physiologically suppresses SHH activity. The expected clinical effect of SHH signalling repression is HPE, which is similar to the phenotype of patients with loss-of-function SHH variants. We used RT-PCR to attempt to dose RNA levels of PTCH1 in the peripheral blood cells of our patients, in order to verify whether the PTCH1 gene duplication produces a higher PTCH1 transcript expression. However, this gene is physiologically expressed at extremely low levels in this tissue, and these experiments failed (data not shown). The proband and her father declined the request to perform a skin biopsy that would be used to obtain fibroblasts, cells where PTCH1 expression is more pronounced. If the role of PTCH1 appears plausible in contributing to the SRS phenotype, the role of the microduplication 9q22.32q22.33 appears more obvious, since it segregates with the disease, being present in the affected father–daughter pair but absent in the healthy paternal grandparents of the proband. We cannot exclude a possible clinical contribution of other duplicated genes and non-coding RNA in the 9q22.32q22.33 region, but, to date, none of these genes has known to be implicated in HPE and/or autosomal dominant conditions. Biallelic loss-of-function FANCC variants are responsible for classical Fanconi anaemia (OMIM #227645); biallelic loss-of-function HSD17B3 variants are responsible for 17-beta-hydroxysteroido dehydrogenase III deficiency, which is an autosomal recessive form of male pseudohermaphrodistism (OMIM #264300); biallelic loss-of-function ERCC6L2 variants are responsible for bone marrow failure syndrome 2 (OMIM #615715). Until now, the other duplicated genes (SLC35D2, ZNF367, HABP4 and CDC14B) are not known to be associated with human diseases.
The presence of hypospadias in our patient and in SRS merits a final comment. SHH is one of the most important signalling molecules that contributes to genital tuberculum (GT) outgrowth and differentiation. SHH is expressed in the urethral plate epithelium (UE) during the hormone-independent phase of genital male embryogenesis, and it activates its pathway via the PTCH receptor, which is required for outgrowth, patterning and cell survival in the developing GT [20]. Mice with a targeted deletion of Shh exhibit initiation of the genital swellings, but outgrowth is not maintained, which indicates the absence of external genitalia [21]. Moreover, in a large Californian cohort of patients with hypospadias, several SNPs in SHH were associated with an increased risk [22]. Taken together, these observations suggest that hypospadias in our patient might result from the same SHH signalling dysfunction that leads to HPE microforms.
In summary, we identified a 9q22.32q22.33 duplication involving the PTCH1 gene in a father–daughter couple with SRS. This “copy number variation” is the first genetic variant found in SRS, and since array–CGH was negative in another SRS patient, genetic heterogeneity seems plausible. This finding supports the autosomal dominant inheritance of SRS and suggests that this condition belongs to the HPE-microform subgroup. These preliminary data suggest to perform array–CGH analysis and sequencing of HPE-related genes in the presence of an SRS phenotype. Further studies in a larger cohort of SRS patients are needed to reach a definitive conclusion.
References
Schilbach U, Rott HD. Ocular hypotelorism, submucosal cleft palate, and hypospadias: a new autosomal dominant syndrome. Am J Med Genet. 1988;31:863–70.
Joss SK, Paterson W, Donaldson MDC, Tolmie JL. Cleft palate, hypotelorism, and hypospadias: Schilbach-Rott syndrome. Am J Med Genet. 2002;113:105–7.
Becerra-Solano LE, Casillas-Avila MP, Diaz-Rodriguez M, Nastasi-Catanese JA, Toscano-Flores JJ, Ramirez-Duenas ML. Schilbach-Rott syndrome in a third family: further delineation of an autosomal dominant trait. Genet Couns. 2007;18:317–23.
de Carvalho DR, Rossi NF, Schellini S, Moretti-Ferreira D, Richieri-Costa A. Schilbach-Rott/blepharofacio skeletal syndrome in a Brazilian patient. Am J MedGenet. 2008;146A:2134–7.
Shkalim V, Baris HN, Gal G, Gleiss R, Calderon S, Wessels M, et al. Autosomal dominant syndrome of mental retardation, hypotelorism, and cleft palate resembling Schilbach-Rott syndrome. Am J Med Genet. 2009;149A:2700–5.
Prontera P, Bernardini L, Stangoni G, Capalbo A, Rogaia D, Romani R, et al. Deletion 2p15-16.1 syndrome: case report and review. Am J Med Genet. 2011;155A:2473–8.
Tokita MJ, Chow PM, Mirzaa G, Dikow N, Maas B, Isidor B, et al. Five children with deletions of 1p34.3 encompassing AGO1 and AGO3. Eur J Hum Genet. 2015;23:761–5.
Howald C, Merla G, Digilio MC, Amenta S, Lyle R, Deutsch S, et al. Two high throughput technologies to detect segmental aneuploidies identify new Williams-Beuren syndrome patients with atypical deletions. J Med Genet. 2006;43:266–73.
Firth HV, et al. DECIPHER: database of chromosomal imbalance and phenotype in humans using ensembl resources. Am J Hum Genet. 2009;84:524–33.
Mercier S, Dubourg C, Garcelon N, Campillo-Gimenez B, Gicquel I, Belleguic M, et al. New findings for phenotype-genotype correlations in a large European series of holoprosencephaly cases. J Med Genet. 2011;48:752–60.
Wallis D, Muenke M. Mutations in holoprosencephaly. Hum Mutat. 2000;16:99–108.
Marigo V, Davey RA, Zuo Y, Cunningham JM, Tabin CJ. Biochemical evidence that patched is the Hedgehog receptor. Nature. 1996;384:176–9.
Ming JE, Kaupas ME, Roessler E, Brunner HG, Golabi M, Tekin M, et al. Mutations in PATCHED-1, the receptor for SONIC HEDGEHOG, are associated with holoprosencephaly. Hum Genet. 2002;110:297–301.
Ribeiro LA, Murray JC, Richieri-Costa A. PTCH mutations in four Brazilian patients with holoprosencephaly and in one with holoprosencephaly-like features and normal MRI. Am J Med Genet A. 2006;140:2584–6.
Derwińska K, Smyk M, Cooper ML, Bader P, Cheung SW, Stankiewicz P. PTCH1 duplication in a family with microcephaly and mild developmental delay. Eur J Hum Genet. 2009;17:267–71.
Izumi K, Hahn A, Christ L, Curtis C, Neilson DE. Familial 9q22.3 microduplication spanning PTCH1 causes short stature syndrome with mild intellectual disability and dysmorphic features. Am J Med Genet A. 2011;155:1384–9.
Heller A, Seidel J, Hübler A, Starke H, Beensen V, Senger G, et al. Molecular cytogenetic characterisation of partial trisomy 9q in a case with pyloric stenosis and a review. J Med Genet. 2000;37:529–32.
Tiong K, Cotterill A, Falhammar H. Adult case of partial trisomy 9q. BMC Med Genet. 2010;11:26.
Hengstschläger M, Prusa AR, Repa C, Drahonsky R, Deutinger J, Pollak A, et al. Patient with partial trisomy 9q and learning disability but no pyloric stenosis. Dev Med Child Neurol. 2004;46:57–59.
Bouty A, Ayers KL, Pask A, Heloury Y, Sinclair AH. The genetic and environmental factors underlying hypospadias. Sex Dev. 2015;9:239–59.
Haraguchi R, Mo R, Hui C, Motoyama J, Makino S, Shiroishi T, et al. Unique functions of Sonic hedgehog signaling during external genitalia. Development. 2001;128:4241–50.
Carmichael SL, Ma C, Choudhry S, Lammer EJ, Witte JS, Shaw GM. Hypospadias and genes related to genital tubercle and early urethral development. J Urol. 2013;190:1884–92.
Acknowledgements
We gratefully acknowledge the family who participated in this study. This research was supported by the “Mauro Baschirotto Institute for Rare Diseases” Foundation. The funders played no role in the design of the study, the data collection, analysis and interpretation or writing of the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Dr. Sallicandro's work has been funded by the “Baschirotto Institute of Rare Disease” (BIRD) Foundation. She has received compensation for carrying out a study on possible genetic causes of Schilbach–Rott syndrome. All other authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Prontera, P., Rogaia, D., Sallicandro, E. et al. Schilbach–Rott syndrome associated with 9q22.32q22.33 duplication, involving the PTCH1 gene. Eur J Hum Genet 27, 1260–1266 (2019). https://doi.org/10.1038/s41431-019-0385-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-019-0385-6