Abstract
The role of nicotinic acetylcholine receptors (nAChR) in nicotine dependence (ND) is well established; CHRNA7, encoding the α7 subunit, has a still uncertain role in ND, although it is implicated in a wide range of neuropsychiatric conditions. CHRFAM7A, a hybrid gene containing a partial duplication of CHRNA7, is possibly involved in modulating α7 nAChR function. The aim of this study was to investigate the role of CHRNA7 and CHRFAM7A genetic variants in ND and to test the hypothesis that α7 nAChR variation may modulate the efficacy of varenicline treatment in smoking cessation. We assessed CHRNA7 and CHRFAM7A copy number, CHRFAM7A exon 6 ∆2 bp polymorphism, and sequence variants in the CHRNA7 proximal promoter in an Italian sample of 408 treatment-seeking smokers. We conducted case-control and quantitative association analyses using two smoking measures (cigarettes per day, CPD, and Fagerström Test for Nicotine Dependence, FTND). Next, driven by the hypothesis that varenicline may exert some of its therapeutic effects through activation of α7 nAChRs, we restricted the analysis to a subgroup of 142 smokers who received varenicline treatment. The CHRNA7 promoter variant rs28531779 showed association with both smoking quantitative measures (FNTD p = 0.026, β = 0.89, 95% CI 0.11–1.67; CPD p = 0.006, β = 4.82 95% CI 1.42–8.22). Moreover, in the varenicline-treated subgroup we observed association of CHRFAM7A copy number with 6 months smoking abstinence (p = 0.035, OR = 3.18, 95% CI = 1.09–9.30). Thus, our study points to a possible role of genetic variation in CHRNA7 and CHRFAM7A in tobacco addiction mechanisms and response to varenicline treatment.
Similar content being viewed by others
Introduction
Tobacco use is the largest preventable cause of death in industrialized countries. Nicotine, the main addictive substance present in tobacco smoke, binds and activates neuronal nicotinic acetylcholine receptors (nAChRs) and causes neuroplastic changes in central neural circuits, including the mesolimbic dopamine system, that lead to the development of tobacco dependence [1]. Nicotine dependence (ND) is influenced by genetic variation and environmental factors, with an estimated heritability of about 50% [2]. Genome-wide association studies (GWAS) analysis of smoking phenotypes and ND have identified common single nucleotide polymorphism (SNP) associations within genes encoding different nAChR subunits, including the clusters on chromosomes 15q25 (CHRNA5-CHRNA3-CHRNB4) and 8p11 (CHRNB3-CHRNA6) [2, 3] and the CHRNA4 gene [4]. Moreover, preclinical studies suggest that a diversity of nAChRs with different regional and cellular expression patterns and sensitivities to nicotine may contribute to tobacco addiction [1, 5, 6].
The α7 is the only neuronal nAChR subunit able to form homopentameric nAChRs, which are distinguished from heteropentameric nAChRs by a number of unique physiological and pharmacological properties. They are characterized by fast activation, rapid desensitization and high-Ca2+ permeability, and specific binding for selective ligands that include the antagonist α-bungarotoxin. Alpha7 nAChRs, together with ß2 containing nAChRs (ß2∗nAChRs), are widely expressed within the central nervous system. The activation of ß2∗nAChRs has an established role in promoting ND phenotypes [1, 5, 6], while the role of the homopentameric α7 nAChR in ND is still not entirely understood, though recent studies suggested that α7 nAChRs may be involved in addiction mechanisms [7, 8], possibly by modulating the activity of ß2∗nAChRs in the ventral tegmental area [9]. Alpha7 nAChRs are also expressed in peripheral systems, with a role in modulation of inflammatory response [10]. α7 nAChRs are thus likely to play multiple important roles in cognition and the immune system.
Altered expression and function of the α7 nAChR have been associated with many neuropsychiatric diseases including schizophrenia, Alzheimer’s disease, autism spectrum disorder (ASD), and epilepsy [11]. CHRNA7, the gene encoding for the α7 nAChR subunit, is located on human chromosome 15q13.3, which is amongst the most unstable regions of the human genome. Microdeletions of chromosome 15q13.3, including CHRNA7, have been established as pathogenic for a wide range of phenotypes and neuropsychiatric conditions (idiopathic generalized epilepsy, intellectual disability, ASD and schizophrenia) [12,13,14,15] and CHRNA7 has been implicated as the major candidate gene responsible for the clinical features expressed. Conversely, the pathogenicity of duplications of CHRNA7 is unclear and difficult to interpret, as they are detected across the same spectrum of neuropsychiatric disorders of the microdeletions, but without the same high penetrance, and they are also found in the general population with an estimated frequency of 0.6% [16].
A partial duplication of CHRNA7, including exons 5 through 10, is present about 1.6 Mb centromeric to CHRNA7. The duplicated portion of CHRNA7 is fused to exons A–E of the FAM7A gene, resulting in a hybrid gene known as CHRFAM7A [17]. The formation of CHRFAM7A is human-specific and the duplication is evolutionary new: CHRNA7 and CHRFAM7A are highly homologous in their duplicated portion (99.9%). CHRFAM7A is present in variable number of copies; some individuals have only one copy of CHRFAM7A and rare subjects have no copies. The translation start site of CHRFAM7A is in exon B and it is in frame through the CHRNA7 sequence, leading to a peptide subunit, called dupα7, that is a truncated form of CHRNA7 missing the 5′ acetylcholine binding site [11, 18]. The dupα7 transcript has been identified in brain, immune cells, and the HL-60 cell line [19], although its translation product and function are still unknown. In vitro studies indicate that dupα7 assembles with α7 subunits, causing a decrease of acetylcholine-stimulated current [18, 19], thus dupα7 may act as a dominant negative regulator of α7 nAChR function.
CHRFAM7A also harbors a polymorphic 2 bp deletion within exon 6, rs67158670 -/TG (Δ2 bp), which is never observed in the homologous CHRNA7 sequence and is predicted to result in a truncated protein. However, since there are two methionine codons in exon 6, it is likely that translation could start from one of these methionines leading to a shorter dupα7 peptide [11, 18].
The CHRFAM7A Δ2 bp allele has been associated to schizophrenia [20], and to the P50 sensory gating deficit, a possible endophenotype for schizophrenia [21]. SNPs in the proximal promoter of CHRNA7 have also been associated to P50 inhibitory deficit [22], and to schizophrenia [23, 24]. Individuals with schizophrenia have a very high risk for tobacco dependence; although, the relationship between ND and schizophrenia is still unclear. Post-mortem studies have shown reduced levels of α7 nAChRs in multiple brain regions of these individuals [11]. Moreover, expression studies in post-mortem prefrontal cortex of patients with psychiatric disorders revealed an increased ratio of CHRFAM7A/CHRNA7 mRNAs in subjects with schizophrenia and bipolar disorder [25]. In vivo studies in rodents have demonstrated that reductions in α7 nAChR function promote nicotine use [7]; taken together, these finding suggests that altered function of α7 nAChRs could act as a potential mechanism of a shared vulnerability to tobacco use and schizophrenia [26]. Alternatively, there may be a causal effect of cigarette smoking on schizophrenia risk, or vice-versa, that schizophrenia might increase the risk of smoking behaviors, as a form of “self-medication” to alleviate symptoms and/or to reduce the side effects of antipsychotic drugs in schizophrenic patients [27]. Based on these lines of evidence, α7 nAChR has been considered a promising therapeutic target for the treatment of ND [9], as well as for improving cognition in complex disorders, such as schizophrenia [28, 29] and Alzheimer’s disease [30].
Varenicline, a drug approved by the US Food and Drug Administration for use as smoking cessation aid, is a α4ß2 nAChR partial agonist; although with lower affinity, varenicline is also a full agonist of α7 nAChRs [31]. The same drug has been shown to provide some cognitive improvement in people with schizophrenia [32, 33]. It is thus possible that activation of α7 nAChRs can contribute to the action of varenicline as a treatment for smoking cessation and schizophrenia.
We conducted a genetic study in order to investigate the possible role of α7 nAChR genetic variation in smoking phenotypes, as well as to test the hypothesis that α7 nAChR variation may modulate the efficacy of varenicline in smoking cessation. The study was conducted on a collection of 408 regular tobacco smokers, recruited at smoking cessation centers in northern Italy, including a subgroup of 142 individuals who were treated with varenicline. More specifically, the aims of this study were: (i) to investigate the association of genetic variants in the CHRNA7 and CHRFAM7A genes with smoking quantity and ND in the whole sample of 408 smokers; (ii) to test the effect of CHRFAM7A and CHRNA7 genetic variants on smoking cessation in varenicline-treated smokers, by contrasting individuals who successfully maintained smoking cessation at 6 months after treatment versus those who did not quit smoking.
In order to comprehensively characterize genetic variation at the CHRNA7 and CHRFAM7A loci in the sample of treatment-seeking smokers, we determined the copy number of both genes, as well as the genotypes of the CHRFAM7A exon 6 Δ2 bp polymorphism (rs67158670 -/TG); in addition, we resequenced the CHRNA7 proximal promoter region in order to test the effect of common and rare variants involved in CHRNA7 transcription regulation.
Methods
The sample
The study sample includes 408 Italian habitual smokers enrolled at three Smoking Cessation Centers (Modena, Parma and Imola) in northern Italy between 2012 and 2015. Exclusion criteria for participation in the study were age <18 or >70 years, or presence of severe hepatic and/or renal impairment. Each subject underwent 4 examinations (screening, baseline, 3- and 6-month follow-up). Data collected during the baseline examination included past medical, psychiatric and smoking history (smoking initiation age, number of cigarettes smoked per day, nicotine concentration per cigarette, family history of tobacco use, reasons for starting and quitting tobacco use, previous attempts to stop smoking), the Fagerström test for nicotine dependence (FTND) [34], the NEO Personality Inventory [35], and the Brief Wisconsin Inventory of Smoking Dependence Motives [36].
Following the baseline examination, all participants received group or individual cognitive-behavioral counseling with the support of a trained psychologist. In addition, 142 smokers received pharmacological treatment with varenicline (0.5 mg/die for 3 days, 0.5 mg twice a day for 4 days, 1 mg twice a day for 12 weeks). In all cases, the treatment was not interrupted before the planned 12 weeks period. Smokers who received varenicline set to quit smoking on the 8th day of therapy. The smokers who did not receive pharmacological treatment, set a quit date 10–15 day after starting the counseling sessions. All individuals were followed up at 3 and 6 months after treatment in order to assess maintenance of smoking cessation. A healthy population control sample was recruited among staff at Modena Policlinico University Hospital, including 139 never smokers and 55 light smokers (FTND scores = 0).
All participants provided a written informed consent to participate. This study was approved by the local Ethical Committee and took place in observation of the declaration of Helsinki (protocol number 2224/2013).
CHRFAM7A Genotyping
DNA for genotyping was extracted from blood (N = 380) or saliva (N = 28). Genotyping for the Δ2 bp polymorphism in exon 6 of CHRFAM7A was performed using a combined approach, as previously described [21, 37]. Specifically, to evaluate the presence of the 2 bp deletion, a region of 238 bp encompassing the polymorphism was amplified using 5′ fluorescent labeled PCR primers (forward: 5′-[6-FAM] GTTTCCATCACCCACACAGG-3′; reverse: 5′- AGCTTGCCCAGGAATAGGAA-3′). The Δ2 bp allele is specific for CHRFAM7A, while the wt (TG) allele is present on both CHRFAM7A and CHRNA7 genes. The PCR products were then analyzed by ABI3730 and Genemapper v3.0 software. The ratio between the 238 bp and the 236 bp fragment peak height allowed to determine the Δ2 bp genotype and relative copy number. The alleles were defined as follows: 0 = absence of the CHRFAM7A gene; 1 = wild-type (TG) allele of the exon 6 polymorphism (rs67158670); 2 = Δ2 bp allele. Each assay was performed twice. For all samples without the Δ2 bp allele, CHRFAM7A copy number was established by a quantitative Real Time PCR (qPCR) assay with Fast SYBR-green (Biorad) to amplify a region of 217 bp spanning the breakpoint between FAM7A exons D- A and CHRNA7 exons 5–10 (5′-TCCTTGCCAATCAACTTTATGA-3′, 5′-CACACCACCACACCTGGTTAAT-3′). The data were normalized to the reference gene FOXP2. Each assay was conducted in triplicate for the target region and for the control region. The relative copy number for the target region was determined using the ΔΔCt method with confidence interval as 2−(ΔΔCt+-SD).
CHRNA7 analysis
CHRNA7 copy number was established using SNP array data from the Illumina Infinium® PsychArray microarrays (Illumina, San Diego, California, USA) through three different CNV detection algorithms: PennCNV [38], QuantiSNP [39] and CNVPartition (Illumina). It was not possible to assess the CHRFAM7A copy number using genome-wide array SNP data as no SNPs uniquely mapping the CHRFAM7A gene are present in most commercial arrays.
The region encompassing 740 bp upstream the CHRNA7 start codon was sequenced in the sample of smokers, using two different PCR amplicons (5′-CATTAGGGTAACCACTGGGAAT-3′, 5′-AGGTGTGAGCGGGAGGTACT-3′; 5′-AGTACCTCCCGCTCACACCT-3′, 5′-GTGCAGCCCAGACAAGCA-3′). PCR products were then sequenced by Sanger method using the BigDye Terminator kit v1.1 (Life Technologies). Identified variants were submitted to the LOVD gene variant data base (https://databases.lovd.nl/shared/genes/chrna7; ID 00163820-00163832).
Statistical analysis
The distribution of CPD and FTND scores in our sample is shown in Supplementary Fig. S1. Although, the FTND is an ordinal variable ranging from 0 to 10, we modeled the FTND score as a continuous variable that satisfies the assumption of normality (Shapiro-Wilk test w = 0.99; p = 0.12). The distribution of CPD diverted from normality (Shapiro-Wilk test, w = 0.91; p = 5 × 10−15), possibly because most smokers tend to round off the number of CPD to multiples of 10, thus introducing some bias.
We used linear regression to test the association of CHRFAM7A copy number, the CHRFAM7A Δ2 bp variant, and CHRNA7 promoter variants with two quantitative measures of smoking: FTND and CPD; gender was included as covariate in the regression model. To test for the influence of the Δ2 bp polymorphism independently of CHRFAM7A copy number, regression analysis was conducted in the stratified sample of individuals with 1 or 2 copies of CHRFAM7A.
Logistic regression analysis was conducted to investigate the influence of genetic variants in CHRFAM7A and CHRNA7 on smoking cessation, using FTND score and gender as covariates. As above, the additive effect of the Δ2 bp allele was tested by logistic regression analysis of abstinent/non-abstinent status by stratification of the sample according to CHRFAM7A copy number. All the above analyses were performed using PLINK 1.9 [40] and STATA (version 9.0).
Rare CHRNA7 promoter variants were analysed using the Sequence Kernel Association Test (SKAT) [41], which aggregates individual score test statistics of a set of SNPs, as individually these SNPs are too rare for statistical analysis.
Results
Sample characterization
Table 1 shows the demographic and phenotypic characteristics of our cohort of 408 smokers. In the entire sample, the FNTD score is significantly correlated to CPD (r = 0.60; P < 10−5); we observed an association of gender with FTND (T-test p = 0.02) and CPD (T-test p < 0.0001), with males having a higher mean FTND (6.02, sd 2.27) and mean CPD (23.6, sd 10.78) compared to females (mean FTND: 5.52, sd 1.95; mean CPD: 19.37, sd 7.51). Therefore, to control the gender effect on smoking measure, all subsequent regression analyses were performed with adjustment for sex as a covariate. Age did not show significant association with either CPD (r2 = 0.008, r = −0.09, p = 0.08) or FNTD (r2 = 0.001, r = 0.035, p = 0.48), therefore this variable was not included as covariate in subsequent regression analyses.
Maintenance of abstinence at 6 months after treatment was investigated in the whole sample and in the subgroup of varenicline-treated subjects (Table 1). The abstinence rate was 28.5% in the untreated group and 37.3% in the varenicline-treated group (31.6% in the whole sample). The group of smokers who did not maintain smoking cessation had a significantly higher mean FTND score and CPD compared to the group who maintained abstinence, while there was no significant difference for gender and age (Table 1).
Analysis of genetic variation in CHRNA7 and CHRFAM7A with smoking phenotypes
Given that the promoter region of CHRNA7 is not adequately covered by SNP probes in the most commonly used genotyping arrays, we decided to comprehensively assess genetic variation in this region by resequencing. The region encompassing 740 bp upstream the CHRNA7 start codon, containing the CHRNA7 core promoter region [22], was sequenced in the sample of smokers. The screening led to the identification of thirteen variants: 11 rare variants and two common variants (MAF > 0.01) (Table 2). Focusing only on the two common variants rs28531779 and rs149637464 (MAF > 0.01), we performed a linear regression analysis to test for association of the SNPs with ND and smoking quantity, adjusting for sex. The rs28531779 SNP achieved a significant P-value for FTND (p = 0.026; β = 0.89; 95% CI 0.11–1.67) and CPD (p = 0.006; β = 4.82; 95% CI 1.42–8.22); rs28531779 minor allele (C) is associated to increased FTND score and CPD.
We also evaluated the cumulative effect of the 11 rare (MAF < 0.01) promoter variants on smoking measures (FTND, CPD) using the SKAT method [41], which produces a P-value indicating the degree of enrichment of rare variant associations within a genetic region. This burden analysis did not identify any statistically significant association.
Then, we investigated if the number of copies of the CHRNA7 and CHRFAM7A genes, as well as the CHRFAM7A exon 6 Δ2 bp polymorphism [37], may be associated to smoking status (case-control analysis) and/or to quantitative smoking phenotypes (CPD and FTND).
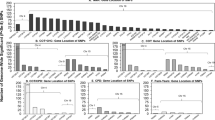
In order to test the hypothesis that genetic variation in CHRNA7 or CHRFAM7A contributes to risk to become nicotine dependent smokers, we determined the gene copy number of CHRNA7 and CHRFAM7A and the allelic status of the Δ2 bp variant (Methods) in the entire sample of treatment-seeking smokers and in a control population sample, consisting of 194 healthy subjects with no ND (FTND = 0) recruited from the same geographical region as the smoker sample. The majority of individuals carried two copies of each gene, while ~3% of individuals had three copies of CHRFAM7A, 15% of individuals had one copy, and 1% had no CHRFAM7A gene. CHRNA7 duplications were identified in five smokers and in one control. Not surprisingly, no CHRNA7 deletions were found in our sample, given the highly penetrant association of this rare microdeletion with different neuropsychiatric disorders [13,14,15]. The distribution of CHRNA7 and CHRFAM7A copy number genotypes did not significantly differ between smokers and controls, and was comparable to previously reported data in other European populations [11] (Suppl. Table 1).
Next, we assessed the allelic status of CHRFAM7A according to the deletion of the whole gene (“allele 0”), presence of either the wt (TG) allele for the exon 6 polymorphism[37] (“allele 1”), or the Δ2 bp allele (“allele 2”) (suppl. Table 2). There was no significant difference in allele frequency between the sample of smokers and controls.
Finally, we investigated the effect of CHRFAM7A genetic variants on quantitative measures of smoking quantity (CPD) and ND (assessed by FTND). Linear regression analysis, with adjustment for sex as covariate, did not reveal a significant effect for CHRFAM7A copy number on either smoking measure. Likewise, linear regression analysis stratified by CHRFAM7A copy number did not identify an effect of the Δ2 bp polymorphism (data not shown).
Association analysis for smoking abstinence in the sample of varenicline-treated smokers
A logistic regression analysis with adjustment for sex and FTND score as covariates was performed to test for an effect of CHRFAM7A copy number, the Δ2 bp allele and CHRNA7 promoter variants on smoking abstinence in the group of 142 smokers who received varenicline treatment. This analysis revealed an effect of CHRFAM7A copy number on abstinence (OR = 3.18, 95% CI = 1.09–9.30, p = 0.035) (Table 3). The logistic regression result is also supported by a higher cessation success rate in varenicline-treated smokers carrying 2 or 3 copies of the CHRFAM7A gene compared to those carrying 0 or 1 copies (1-sided Fisher exact test p = 0.048) (Table 4). The association of CHRFAM7A copy number and abstinence was only detected in the varenicline-treated smokers, while it was not present in the remaining sample of 266 smokers who did not receive varenicline treatment. These results suggest that variation in the number of copies of the CHRFAM7A gene may modulate the effectiveness of varenicline treatment as a smoking cessation aid.
Finally, logistic regression analysis in our sample of varenicline-treated or non-treated smokers did not reveal any significant influence on abstinence for the Δ2 bp polymorphism in CHRFAM7A and CHRNA7 promoter variants.
Discussion
The aim of this study was to examine the hypothesis that genetic variation affecting α7 nAChR function may influence smoking behaviors and the effectiveness of varenicline in smoking cessation. We investigated the CHRNA7 and CHRFAM7A loci, assessing copy number and sequence variants in these two genes, in a cohort of 408 treatment-seeking smokers, which included a subgroup of 142 individuals who were treated with varenicline. To our knowledge, this is the first study to comprehensively analyse genetic variation at these loci in relation to ND phenotypes.
Sequence analysis of the CHRNA7 promoter region led to the identification of an association of rs28531779 with ND and smoking quantity (FNTD and CPD). Our study is the first one to report an association of CHRNA7 variation with quantitative measures of smoking phenotypes. Interestingly, previous studies reported that variants in the CHRNA7 promoter are genetically associated to the P50 auditory evoked potential deficit [22, 42], a deficit frequently observed in schizophrenic patients and their relatives, and the same variants have been found to be more prevalent in schizophrenic subjects than controls; the specific rs28531779 variant has also been significantly associated to schizophrenia in a Danish case-control study [22, 23]. The mechanism of association between schizophrenia risk and cigarette smoking have been a subject of debate [27]. Functional analysis using the luciferase assay [22] demonstrated that the associated variants in CHRNA7 core promoter, including rs28531779, are linked to a reduced CHRNA7 promoter activity. Taken together, these data suggest that a decreased expression of α7 nAChR could be involved in susceptibility to both ND and schizophrenia with a pleiotropic effect, or that there might be a causal effect of cigarette smoking on schizophrenia, as recently suggested by Mendelian randomization analysis [43, 44, 45].
A second interesting finding of this study is the observation that CHRFAM7A copy number seems to influence the success of smoking cessation in subjects treated with varenicline. In particular, CHRFAM7A copy number positively correlates with smoking cessation success, with higher abstinence rates in subjects carrying two or three copies in comparison to subjects with 0–1 copies. Although, the biological bases for this effect is presently unknown, some evidence in reconstituted systems suggests that incorporation of dupα7 in α7 receptors may affect their sensitivity to varenicline [46].
In the remaining sample of 266 smokers who did not receive varenicline treatment, we did not identify an effect of CHRFAM7A copy number, according to the hypothesis that the CHRFAM7A copy number influences smoking cessation by interacting with varenicline treatment. Our study thus supports the hypothesis of an involvement of α7 nAChRs activation in the varenicline mode of action for smoking cessation treatment.
In a similar fashion to variants in CHRNA7 promoter, the CHRFAM7A Δ2 bp polymorphism has been previously reported to be significantly associated to the P50 sensory gating deficit [21], as well as to schizophrenia [20] and bipolar disorder [47]; however, no evidence was found in our study for an effect of the CHRFAM7A Δ2 bp polymorphism on smoking phenotypes. These contradictory results could be explained by the small sample size of our study, thus lacking an adequate power to detect a significant association for this variant, or different ethnicity of investigated individuals between the studies; alternatively, the CHRFAM7A Δ2 bp may specifically influence psychiatric phenotypes.
It is worthy of note that no significant associations have been reported for CHRNA7 or CHRFAM7A in very large GWAS studies for smoking related traits [48, 49]. However, the lack of association signal at these loci could be explained by their complex architecture; indeed, investigation of this region by standard genotyping arrays or next generation sequencing is hampered by the presence of the duplication containing CHRFAM7A, and a very high C+G content. For instance, the most widely used commercial SNPs arrays (including Illumina 1M, Illumina Omni1-Quad and Affymetrix 6.0 array), do not contain SNP probes in the CHRNA7 promoter region. Likewise, the Δ2 bp CHRFAM7A variant is not represented on commercial SNPs arrays, and SNPs located in the duplicated portion of CHRNA7 and CHRFAM7A cannot be univocally mapped.
Finally, we acknowledge that the major limitation of the present study is the relatively small sample size that does not ensure sufficient power to draw definitive conclusions on the association results obtained. Replication of CHRNA7 and CHRFAM7A genetic analysis in larger and independent cohorts is therefore required to confirm our initial results.
In conclusion, this study is the first one to report an association of CHRNA7 promoter variants with smoking quantity and ND, suggesting that variation in CHRNA7 expression could have a critical role in tobacco addiction mechanisms. Furthermore, our work provides the first evidence that CHRFAM7A copy number variation could affect the response to varenicline treatment. Genetic factors are likely to play an important role in relapse after smoking cessation and in the limited efficacy of behavioral and pharmacologic approaches to smoking cessation. It has been estimated that 50% of the risk for a failed attempt at smoking cessation can be attributed to genetic factors [50]. Therefore, the identification of genetic factors that could improve the effectiveness of pharmacologic approaches to smoking cessation may represent a very interesting finding.
References
Changeux JP. Nicotine addiction and nicotinic receptors: lessons from genetically modified mice. Nat Rev Neurosci. 2010;11:389–401.
Greenbaum L, Lerer B. Differential contribution of genetic variation in multiple brain nicotinic cholinergic receptors to nicotine dependence: recent progress and emerging open questions. Mol Psychiatry. 2009;14:912–45.
Bierut LJ. Genetic vulnerability and susceptibility to substance dependence. Neuron. 2011;69:618–27.
Hancock DB, Reginsson GW, Gaddis NC, et al. Genome-wide meta-analysis reveals common splice site acceptor variant in CHRNA4 associated with nicotine dependence. Transl Psychiatry. 2015;5:e651.
Picciotto MR, Kenny PJ. Molecular mechanisms underlying behaviors related to nicotine addiction. Cold Spring Harb Perspect Med. 2013;3:a012112.
Gotti C, Guiducci S, Tedesco V, et al. Nicotinic acetylcholine receptors in the mesolimbic pathway: primary role of ventral tegmental area alpha6beta2* receptors in mediating systemic nicotine effects on dopamine release, locomotion, and reinforcement. J Neurosci. 2010;30:5311–25.
Brunzell DH, McIntosh JM. Alpha7 nicotinic acetylcholine receptors modulate motivation to self-administer nicotine: implications for smoking and schizophrenia. Neuropsychopharmacology. 2012;37:1134–43.
Harenza JL, Muldoon PP, De Biasi M, Damaj MI, Miles MF. Genetic variation within the Chrna7 gene modulates nicotine reward-like phenotypes in mice. Genes Brain Behav. 2014;13:213–25.
Brunzell DH, McIntosh JM, Papke RL. Diverse strategies targeting alpha7 homomeric and alpha6beta2* heteromeric nicotinic acetylcholine receptors for smoking cessation. Ann N Y Acad Sci. 2014;1327:27–45.
Wang H, Yu M, Ochani M, et al. Nicotinic acetylcholine receptor alpha7 subunit is an essential regulator of inflammation. Nature. 2003;421:384–8.
Sinkus ML, Graw S, Freedman R, Ross RG, Lester HA, Leonard S. The human CHRNA7 and CHRFAM7A genes: a review of the genetics, regulation, and function. Neuropharmacology. 2015;96(Pt B):274–88.
Gillentine MA, Schaaf CP. The human clinical phenotypes of altered CHRNA7 copy number. Biochem Pharmacol. 2015;97:352–62.
Lowther C, Costain G, Stavropoulos DJ, et al. Delineating the 15q13.3 microdeletion phenotype: a case series and comprehensive review of the literature. Genet Med. 2015;17:149–57.
Shinawi M, Schaaf CP, Bhatt SS, et al. A small recurrent deletion within 15q13.3 is associated with a range of neurodevelopmental phenotypes. Nat Genet. 2009;41:1269–71.
Mikhail FM, Lose EJ, Robin NH, et al. Clinically relevant single gene or intragenic deletions encompassing critical neurodevelopmental genes in patients with developmental delay, mental retardation, and/or autism spectrum disorders. Am J Med Genet A. 2011;155a:2386–96.
Helbig I, Mefford HC, Sharp AJ, et al. 15q13.3 microdeletions increase risk of idiopathic generalized epilepsy. Nat Genet. 2009;41:160–2.
Gault J, Robinson M, Berger R, et al. Genomic organization and partial duplication of the human alpha7 neuronal nicotinic acetylcholine receptor gene (CHRNA7). Genomics. 1998;52:173–85.
Araud T, Graw S, Berger R, et al. The chimeric gene CHRFAM7A, a partial duplication of the CHRNA7 gene, is a dominant negative regulator ofalpha7*nAChR function. Biochem Pharmacol. 2011;82:904–14.
de Lucas-Cerrillo AM, Maldifassi MC, Arnalich F, et al. Function of partially duplicated human alpha77 nicotinic receptor subunit CHRFAM7A gene: potential implications for the cholinergic anti-inflammatory response. J Biol Chem. 2011;286:594–606.
Sinkus ML, Lee MJ, Gault J, et al. A 2-base pair deletion polymorphism in the partial duplication of the alpha7 nicotinic acetylcholine gene (CHRFAM7A) on chromosome 15q14 is associated with schizophrenia. Brain Res. 2009;1291:1–11.
Flomen RH, Shaikh M, Walshe M, et al. Association between the 2-bp deletion polymorphism in the duplicated version of the alpha7 nicotinic receptor gene and P50 sensory gating. Eur J Hum Genet. 2013;21:76–81.
Leonard S, Gault J, Hopkins J, et al. Association of promoter variants in the alpha7 nicotinic acetylcholine receptor subunit gene with an inhibitory deficit found in schizophrenia. Arch Gen Psychiatry. 2002;59:1085–96.
Bertelsen B, Oranje B, Melchior L, et al. Association Study of CHRNA7 promoter variants with sensory and sensorimotor gating in Schizophrenia patients and healthy controls: a danish case-control study. Neuromolecular Med. 2015;17:423–30.
Stephens SH, Logel J, Barton A, et al. Association of the 5’-upstream regulatory region of the alpha7 nicotinic acetylcholine receptor subunit gene (CHRNA7) with schizophrenia. Schizophr Res. 2009;109:102–12.
Kunii Y, Zhang W, Xu Q, et al. CHRNA7 and CHRFAM7A mRNAs: co-localized and their expression levels altered in the postmortem dorsolateral prefrontal cortex in major psychiatric disorders. Am J Psychiatry. 2015;172:1122–30.
Akbarian S, Kundakovic M. CHRNA7 and CHRFAM7A: psychosis and smoking? Blame the neighbors! Am J Psychiatry. 2015;172:1054–6.
Gage SH, Munafo MR. Smoking as a causal risk factor for schizophrenia. Lancet Psychiatry. 2015;2:778–9.
Freedman R. alpha7-nicotinic acetylcholine receptor agonists for cognitive enhancement in schizophrenia. Ann Rev Med. 2014;65:245–61.
Young JW, Geyer MA. Evaluating the role of the alpha-7 nicotinic acetylcholine receptor in the pathophysiology and treatment of schizophrenia. Biochem Pharmacol. 2013;86:1122–32.
Pohanka M. Alpha7 nicotinic acetylcholine receptor is a target in pharmacology and toxicology. Int J Mol Sci. 2012;13:2219–38.
Mihalak KB, Carroll FI, Luetje CW. Varenicline is a partial agonist at alpha4beta2 and a full agonist at alpha7 neuronal nicotinic receptors. Mol Pharmacol. 2006;70:801–5.
Smith RC, Lindenmayer JP, Davis JM, et al. Cognitive and antismoking effects of varenicline in patients with schizophrenia or schizoaffective disorder. Schizophr Res. 2009;110:149–55.
Hong LE, Thaker GK, McMahon RP, et al. Effects of moderate-dose treatment with varenicline on neurobiological and cognitive biomarkers in smokers and nonsmokers with schizophrenia or schizoaffective disorder. Arch Gen Psychiatry. 2011;68:1195–206.
Fagerstrom KO. Measuring degree of physical dependence to tobacco smoking with reference to individualization of treatment. Addict Behav. 1978;3:235–41.
Costa PT Jr., McCrae RR. Stability and change in personality assessment: the revised NEO Personality Inventory in the year 2000. J Pers Assess. 1997;68:86–94.
Smith SS, Piper ME, Bolt DM, et al. Development of the brief wisconsin inventory of smoking dependence motives. Nicotine Tob Res. 2010;12:489–99.
Flomen RH, Collier DA, Osborne S, et al. Association study of CHRFAM7A copy number and 2 bp deletion polymorphisms with schizophrenia and bipolar affective disorder. Am J Med Genet B Neuropsychiatr Genet. 2006;141b:571–5.
Wang K, Li M, Hadley D, et al. PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 2007;17:1665–74.
Colella S, Yau C, Taylor JM, et al. QuantiSNP: an Objective Bayes Hidden-Markov Model to detect and accurately map copy number variation using SNP genotyping data. Nucleic Acids Res. 2007;35:2013–25.
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4:7.
Ionita-Laza I, Lee S, Makarov V, Buxbaum JD, Lin X. Sequence kernel association tests for the combined effect of rare and common variants. Am J Hum Genet. 2013;92:841–53.
Houy E, Raux G, Thibaut F, et al. The promoter -194 C polymorphism of the nicotinic alpha 7 receptor gene has a protective effect against the P50 sensory gating deficit. Mol Psychiatry. 2004;9:320–2.
Wium-Andersen MK, Orsted DD, Nordestgaard BG. Tobacco smoking is causally associated with antipsychotic medication use and schizophrenia, but not with antidepressant medication use or depression. Int J Epidemiol. 2015;44:566–77.
Gage SH, Munafò MR. Rethinking the association between smoking and schizo- phrenia. Lancet Psychiatry 2015; 2: 118–19.
Chen J, Bacanu SA, Yu H, et al. Genetic relationship between Schizophrenia and nicotine dependence. Sci Rep. 2016;6:25671.
Wang Y, Xiao C, Indersmitten T, Freedman R, Leonard S, Lester HA. The duplicated alpha7 subunits assemble and form functional nicotinic receptors with the full-length alpha7. J Biol Chem. 2014;289:26451–63.
Hong CJ, Lai IC, Liou LL, Tsai SJ. Association study of the human partially duplicated alpha7 nicotinic acetylcholine receptor genetic variant with bipolar disorder. Neurosci Lett. 2004;355:69–72.
Tobacco and Genetics Consortium. Genome-wide meta-analyses identify multiple loci associated with smoking behavior. Nat Genet. 2010;42:441–7.
Liu JZ, Tozzi F, Waterworth DM, et al. Meta-analysis and imputation refines the association of 15q25 with smoking quantity. Nat Genet. 2010;42:436–40.
Xian H, Scherrer JF, Madden PA, et al. The heritability of failed smoking cessation and nicotine withdrawal in twins who smoked and attempted to quit. Nicotine Tob Res. 2003;5:245–54.
Acknowledgements
We gratefully acknowledge all the subjects who have participated in the study. We thank Fondazione Sfameni for funding a PhD fellowship to C.C. We also thank Dr. Roberto D’Amico for statistical advice.
Funding:
This work was supported by the Italian Ministry of Health (RF2009-1549619) and by the University of Bologna (RFO2011-2014).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Cameli, C., Bacchelli, E., De Paola, M. et al. Genetic variation in CHRNA7 and CHRFAM7A is associated with nicotine dependence and response to varenicline treatment. Eur J Hum Genet 26, 1824–1831 (2018). https://doi.org/10.1038/s41431-018-0223-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-018-0223-2
This article is cited by
-
Translational implications of CHRFAM7A, an elusive human-restricted fusion gene
Molecular Psychiatry (2024)