Abstract
Background
Cancers are heterogeneous and contain various types of irregular structures that can go undetected when examining them with standard two-dimensional microscopes. Studies of intricate networks of vasculature systems, e.g., the tumour lymphatic microvessels, benefit largely from three-dimensional imaging data analysis.
Methods
The new DIPCO (Diagnosing Immunolabeled Paraffin-Embedded Cleared Organs) imaging platform uses three-dimensional light-sheet microscopy and whole-mount immunolabelling of cleared samples to study proteins and micro-anatomies deep inside of tumours.
Results
Here, we uncovered the whole three-dimensional lymphatic microvasculature of formalin-fixed paraffin-embedded (FFPE) tumours from a cohort of 30 patients with bladder cancer. Our results revealed more heterogeneous spatial deviations in more advanced bladder tumours. We also showed that three-dimensional imaging could determine tumour stage and identify vascular or lymphatic system invasion with higher accuracy than standard two-dimensional histological diagnostic methods. There was no association between sample storage times and outcomes, demonstrating that the DIPCO pipeline could be successfully applied on old FFPE samples.
Conclusions
Studying tumour samples with three-dimensional imaging could help us understand the pathological nature of cancers and provide essential information that might improve the accuracy of cancer staging.
Similar content being viewed by others
Introduction
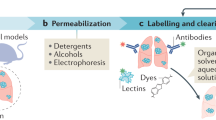
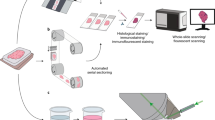
Cancers are heterogeneous, and there are various types of irregular structures that exist in three-dimensions (3D).1,2,3,4 To date, the lack of techniques and methods has limited researchers’ and physicians’ ability to spatially elucidate the entire cancer landscape. Studying solid tumours with traditional two-dimensional (2D) light microscopy restrict our findings to surface pictures.5 However, recent advances in tissue clearing techniques and light-sheet microscopy have enabled high-end 3D visualisation deep inside samples.6,7,8,9 Additionally, we recently optimised the use of formalin-fixed paraffin-embedded (FFPE) samples for whole-mount immunolabeling, clearing, and imaging with light-sheet microscopy, naming the approach DIPCO (Diagnosing Immunolabeled Paraffin-Embedded Cleared Organs, Fig. 1a).5 The time is ready for a new imaging platform to characterise cancers, which will fill the information gap in studying 3D objects, such as cancerous tumours, with 2D microscopy.
Application of the DIPCO pipeline for whole-mount immunostaining of LYVE-1. a Workflow of the DIPCO pipeline and conventional 2D histological methods. b Box-and-whisker plot of sample storage times of 30 FFPE bladder tumours. c Representative bright-field images of a deparaffinised FFPE bladder tumour pre-clearing and post-clearing. The grid lines are separated by 3 mm. d Volume rendering of light-sheet microscopy data from the LYVE-1 immunostained bladder cancer sample in c. Pseudo-colours indicate low (blue) and high (red) LYVE-1 expression. e Volumetric image of the lymphatic vasculature visualised by LYVE-1 immunostaining. Pseudo-colours indicate thin (blue) and thick (red) vessels. f Histograms of the LYVE-1 density kurtosis from 30 FFPE bladder samples of pTa-1 (blue, n = 16), pT2 (green, n = 6), or pT3-4 (red, n = 8) tumours, assessed with the DIPCO pipeline. The line within the box represents the median. The upper and lower quartiles are the bounds of the box, and the minimum and maximum values are the bars. *P < 0.05 by the two-tailed Mann-Whitney U-test. x,y,z- indicators, 500 μm
Studies of vasculature systems, e.g., the tumour lymphatic microvessels, benefit largely from using 3D imaging data analysis. Lymphatic dissemination is the major pathway for systemic tumour spread in patients with urinary bladder cancer.10 However, little knowledge exists about the spatial distribution of lymphatic microvessels within intact human bladder tumours. Herein, we applied the DIPCO pipeline to answer this question. Further, we demonstrated that cancer staging by 3D imaging data analysis provide more accuracy than standard 2D histological diagnostic methods.
Materials and methods
Sample collection
Thirty human FFPE samples from bladder cancers were included; namely, 2 from the Karolinska University Hospital in Sweden and 28 from the Medical University of Lublin in Poland. The tissues were fixed after surgery using formaldehyde and were then embedded in paraffin. One tissue block was randomly picked from each patient for further experiments. All tumours were histologically confirmed to be urothelial carcinomas. The tumours were staged according to the 2002 TNM staging system and graded according to the 2004 WHO classifications. All experiments were approved ethically (2011/421-31/1, 2015/1990-32/1, 2012/596-31/4, Karolinska University Hospital; KE-0254/222/2016, Medical University of Lublin).
Immunohistochemistry of paraffin-embedded sections (IHC-P)
FFPE sections (4–6 μm) were deparaffinised and rehydrated. Then, the antigen was removed, and endogenous peroxidase was quenched. After blocking, the sections were incubated overnight with the primary antibody for LYVE-1 (1:100, # ab33682, Abcam) with the appropriate species-specific secondary antibody. The specificity of the LYVE-1 immunosignal for detecting tumour lymphatic vessels was tested and confirmed using an alternative lymphatic marker Podoplanin (1:100, # ab10288, Abcam). Images were acquired with a fluorescence microscope (Cell Observer, Carl Zeiss, Jena, Germany).
Preparation and image processing for 3D analysis
Preparation and 3D imaging data processing of samples are described elsewhere.5 The lymphatic endothelial hyaluronan receptor LYVE-1 was targeted to label lymphatics within tumours.11,12 The primary and secondary antibody used was anti-LYVE-1 (1:100, # ab33682, Abcam) and Alexa 647-conjugated affinity purified F(ab’)2 fragment antibody (1:200, # 711-605-152, Jackson ImmunoResearch Laboratories), respectively. For tissue clearing, immunolabeled samples were incubated in methanol, dichloromethane, and finally dibenzyl ether.6 Cleared tumours were imaged using a custom-built light-sheet microscope.13
Amira (FEI) software was used for 3D volume rendering, vessel segmentation, and quantification.14 Images were processed and normalised using Amira and ImageJ (National Institutes of Health, Washington, DC) software. Lymphatics were segmented according to the LYVE-1 immunosignal level15 using an intensity-based threshold and spatial graph view algorithms of the Amira suite, which also calculated the vessel length and radius. Every vessel was automatically separated to the next branch as one segment and used for the analyses. The spatial heterogeneity feature of the LYVE-1 expression was examined by calculating the kurtosis, skewness, and variance of the LYVE-1 expression density for each 5-μm Z-section.5
Statistics
The values are given as the mean ± SE, median and interquartile range (IQR) for continuous variables, and frequency with percentage for categorical variables. Variables between groups were compared using the Mann-Whitney U-test. To assess the ability of the DIPCO pipeline, we carried out a receiver operating characteristic (ROC) curve analysis to distinguish cancers with advanced stages and vascular or lymphatic system invasion, i.e., lymphovascular invasion plus positive lymph node involvement. Finally, an area under the curve (AUC) value with a 95% confidential interval (CI) was determined for discrimination. Statistical significance was accepted for P values < 0.05. All analyses were performed using the SPSS version 22.0 statistical software package.
Results and discussion
Clinical FFPE samples from a cohort of 30 patients with bladder cancer, of which 20% had low-grade tumours and 80% had high-grade tumours, were assessed. The pathological T stage for Ta-1, T2, and T3-4 was in 53%, 20%, and 27%, respectively. Lymphovascular invasion was observed in five patients (17%) and positive lymph node involvement was observed in four patients (13%). The median storage time for the FFPE samples was 20 months (IQR, 13–71) (Fig. 1b) and no association between sample storage times and imaging quality was observed. The FFPE tumours were cleared (Fig. 1c), immunolabeled for LYVE-1 and studied applying the DIPCO pipeline.5 LYVE-1 is predominantly expressed on the initial lymphatic vessels and not on the collecting lymphatics.16, 17 The 3D imaging data analysis revealed heterogeneous lymphatic microvessels (Fig. 1d, Supplementary Video 1) with diversified vessel thicknesses (Fig. 1e, Supplementary Video 2) throughout the entire tumour. The specificity of the LYVE-1 antigen for detecting tumour lymphatics in bladder tumour tissue was verified using the alternative lymphatic marker Podoplanin (Supplementary Fig. 1).
We then examined seven parameters: the 2D LYVE-1 density, 3D LYVE-1 density, architectural features of lymphatic vessel length and radius, spatial heterogeneity features of LYVE-1 density kurtosis, skewness, and variance (Supplementary Table 1). Of these parameters, only the 2D LYVE-1 density was acquired using 2D imaging of IHC-P. The 3D imaging data analysis demonstrated that the LYVE-1 density kurtosis was significantly higher in advanced tumour stages (Fig. 1f). There was no association between sample storage times and outcomes (Table 1), indicating that the DIPCO pipeline could be applied to study old FFPE samples.
To determine the clinical relevance of examining bladder tumours with 3D imaging, ROC curves were constructed thereafter to predict pathological features, such as advanced pT stage (Fig. 2a) and positive vascular or lymphatic system invasion (Fig. 2b). Measuring the 2D LYVE-1 density gave AUC values of 0.568 for detecting pT3 tumours or greater and 0.609 for detecting positive vascular or lymphatic system invasion (Fig. 2c). Both parameters were improved when they were transferred into 3D, especially to predict positive vascular or lymphatic system invasion (AUC = 0.658). Furthermore, assessing the LYVE-1 density kurtosis revealed the highest AUC values of 0.756 for detecting pT3 tumours or greater and 0.702 for detecting positive vascular or lymphatic system invasion (Fig. 2c). The remaining 3D imaging parameters, lymphatic vessel length and radius, were also better for predicting cancer stages compared to those of 2D imaging (Fig. 2c).
Predictive abilities and comparisons between 2D and 3D microscopy methods. ROC curve analysis of 30 FFPE bladder tumours for detecting pT3 tumours or greater (a) and positive vascular or lymphatic system invasion (b). c Table of AUC (95% confidence interval (CI)) values summarising the ROC-AUC analysis in (a) and (b). LVI lymphovascular invasion, LNI lymph node involvement
Light-sheet microscopy offers 3D imaging of cleared tissues deep inside of samples. Furthermore, this new technology can visualise immunolabeled samples with micro-scale resolution, possibly leading to diagnostic advantage in comparison with other 3D imaging alternatives, e.g., high-resolution magnetic resonance microscopy.18 Our results showed that light-sheet microscopy of cleared bladder tumours could uncover lymphatic microvasculatures running throughout entire tumours. Moreover, there was no association between the sample storage time and the quality of the 3D imaging analysis, as the DIPCO pipeline could be successfully applied to FFPE samples that were stored over 9 years.
Lymphangiogenesis is vital for various cancer types, participating in tumour development and metastasis.19, 20 We examined the clinical relevance of using 3D imaging to analyse the expression pattern of the lymphatic vessel marker LYVE-1, revealing more heterogeneous spatial deviations in samples from patients with advanced bladder cancers. The highest AUC values of LYVE-1 density kurtosis was presented for diagnosing pT3 tumours or greater (0.756) and positive vascular or lymphatic system invasion (0.702), exceeding values obtained from 2D imaging data analysis.
Precisely characterising and diagnosing tumours are essential for physicians to establish appropriate counselling and treatment plans for cancer patients. When treating patients with bladder cancer, accurate cancer staging prior cystectomies is important to select the correct candidates for neoadjuvant chemotherapy before surgeries. We believe that applying the DIPCO pipeline could enable pathologists to characterise typical cancer hallmarks and perform multi-region 3D analyses of tumours, resulting in more exact cancer diagnoses.
In summary, these results show the capacity of light-sheet microscopy to phenotypically characterise intact bladder tumours and to improve accuracy of cancer staging. The limitations of this study are its retrospective nature as well as the small cohort and the lack of survival data with treatment annotations, with the exception of three patients who were deceased.
References
Tabassum, D. P. & Polyak, K. Tumorigenesis: it takes a village. Nat. Rev. Cancer 15, 473–483 (2015).
Waclaw, B. et al. A spatial model predicts that dispersal and cell turnover limit intratumour heterogeneity. Nature 525, 261–264 (2015).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Junttila, M. R. & de Sauvage, F. J. Influence of tumour micro-environment heterogeneity on therapeutic response. Nature 501, 346–354 (2013).
Tanaka, N. et al. Whole-tissue biopsy phenotyping of three-dimensional tumours reveals patterns of cancer heterogeneity. Nat. Biomed. Eng. 1, 796–806 (2017).
Renier, N. et al. Mapping of brain activity by automated volume analysis of immediate early genes. Cell 165, 1789–1802 (2016).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Tainaka, K. et al. Whole-body imaging with single-cell resolution by tissue decolorization. Cell 159, 911–924 (2014).
Chung, K. & Deisseroth, K. CLARITY for mapping the nervous system. Nat. Methods 10, 508–513 (2013).
Fernandez, M. I. et al. Prognostic implications of lymphangiogenesis in muscle-invasive transitional cell carcinoma of the bladder. Eur. Urol. 53, 571–578 (2008).
Kato, T. et al. A quantitative analysis of lymphatic vessels in human breast cancer, based on LYVE-1 immunoreactivity. Br. J. Cancer 93, 1168–1174 (2005).
Padera, T. P. et al. Lymphatic metastasis in the absence of functional intratumor lymphatics. Science 296, 1883–1886 (2002).
Tomer, R., Ye, L., Hsueh, B. & Deisseroth, K. Advanced CLARITY for rapid and high-resolution imaging of intact tissues. Nat. Protoc. 9, 1682–1697 (2014).
Susaki, E. A. & Ueda, H. R. Whole-body and whole-organ clearing and imaging techniques with single-cell resolution: toward organism-level systems biology in mammals. Cell Chem. Biol. 23, 137–157 (2016).
Skobe, M. et al. Induction of tumor lymphangiogenesis by VEGF-C promotes breast cancer metastasis. Nat. Med. 7, 192–198 (2001).
Petrova, T. V. et al. Defective valves and abnormal mural cell recruitment underlie lymphatic vascular failure in lymphedema distichiasis. Nat. Med. 10, 974–981 (2004).
Makinen, T. et al. PDZ interaction site in ephrinB2 is required for the remodeling of lymphatic vasculature. Genes Dev. 19, 397–410 (2005).
Dashevsky, B. Z. et al. The potential of high resolution magnetic resonance microscopy in the pathologic analysis of resected breast and lymph tissue. Sci. Rep. 5, 17435 (2015).
Alitalo, K. The lymphatic vasculature in disease. Nat. Med. 17, 1371–1380 (2011).
Stacker, S. A. et al. Lymphangiogenesis and lymphatic vessel remodelling in cancer. Nat. Rev. Cancer 14, 159–172 (2014).
Acknowledgements
The authors would like to thank Prof. Justyna Szumiło, Department of Clinical Pathomorphology, Medical University of Lublin, Lublin, Poland for kindly providing human tissue samples. This study was supported by the Swedish Research Council (grants 2009-3364, 2013-3189, and 2017-00815 to P.U.), the Swedish Cancer Society (grant CAN 2013/802 and CAN 2016/801 to P.U.), the Linnaeus Center in Developmental Biology for Regenerative Medicine (DBRM) (P.U.), a Knut and Alice Wallenberg Foundation Grant to the Center for Live Imaging of Cells at the Karolinska (CLICK) Institutet (P.U.), the Royal Swedish Academy of Sciences (P.U.), the David and Astrid Hagelén Foundation (N.T.), and the Karolinska Institutet Research Foundation (N.T.).
Author information
Authors and Affiliations
Contributions
N.T. and P.U. designed the research, analysed the data, and wrote the manuscript. N.T., D.K., and S.K. performed the research. C.S., P.M., A.S., A.M., and P.W. provided conceptual advice.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. If you remix, transform, or build upon this article or a part thereof, you must distribute your contributions under the same license as the original. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/.
About this article
Cite this article
Tanaka, N., Kaczynska, D., Kanatani, S. et al. Mapping of the three-dimensional lymphatic microvasculature in bladder tumours using light-sheet microscopy. Br J Cancer 118, 995–999 (2018). https://doi.org/10.1038/s41416-018-0016-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-018-0016-y
This article is cited by
-
Consultation on UTUC Stockholm 2022
World Journal of Urology (2023)
-
Tissue clearing to examine tumour complexity in three dimensions
Nature Reviews Cancer (2021)
-
Harnessing non-destructive 3D pathology
Nature Biomedical Engineering (2021)
-
Tutorial: methods for three-dimensional visualization of archival tissue material
Nature Protocols (2021)
-
3D histopathology of human tumours by fast clearing and ultramicroscopy
Scientific Reports (2020)