Abstract
Backgrounds
p53 is a tumor suppressor that prevents cancer onset and progression, and mutations in the p53 gene cause loss of the tumor suppressor function of the protein. The mutant p53 protein in tumor cells can form aggregates which contribute to the dominant-negative effect over the wild-type p53 protein, causing loss of p53 tumor suppression or gain of novel oncogenic functions. Mutations in p53 have been implicated in the pathogenesis of primary prostate cancer (PCa), and are often detected in recurrent and metastatic disease. Thus, targeting mutant p53 may constitute an alternative therapeutic strategy for advanced PCa for which there are no other viable options.
Methods
In this study, we used immunoprecipitation, immunofluorescence, clonogenic survival, and cell proliferation assays, flow cytometric analysis and in vivo xenograft to investigate the biological effects of ReACp53, a cell-permeable peptide inhibitor of p53 aggregation, on mutant p53-carrying PCa cells.
Results
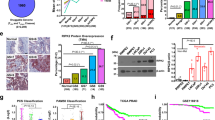
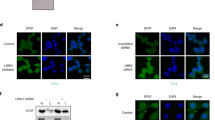
Our results show that ReACp53 targets amyloid aggregates of mutant p53 protein and restores the p53 nuclear function as transcriptional factor, induces mitochondrial cell death and reduces DNA synthesis of mutant p53-carrying PCa cells; ReACp53 also inhibits xenograft tumor growth in vivo.
Conclusions
The data presented here suggest a therapeutic potential of targeting mutant p53 protein in advanced PCa setting, which has a clinical impact for aggressive PCa with transforming how such tumors are managed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 4 print issues and online access
$259.00 per year
only $64.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016;66:7–30. https://doi.org/10.3322/caac.21332.
Barbieri CE, Baca SC, Lawrence MS, Demichelis F, Blattner M, Theurillat JP, et al. Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nat Genet. 2012;44:685–9. https://doi.org/10.1038/ng.2279.
Robinson D, Van Allen EM, Wu YM, Schultz N, Lonigro RJ, Mosquera JM, et al. Integrative clinical genomics of advanced prostate. Cancer Cell. 2015;162:454 https://doi.org/10.1016/j.cell.2015.06.053.
Vousden KH, Prives C. Blinded by the light: the growing complexity ofp53. Cell. 2009;137:413–31. https://doi.org/10.1016/j.cell.2009.04.037.
Leroy B, Fournier JL, Ishioka C, Monti P, Inga A, Fronza G, et al. The TP53 website: an integrative resource centre for the TP53 mutation database and TP53 mutant analysis. Nucleic Acids Res. 2013;41(Database issue):D962–9. https://doi.org/10.1093/nar/gks1033.
Robles AI, Harris CC. Clinical outcomes and correlates of TP53 mutations and cancer. Cold Spring Harb Perspect Biol. 2010;2:a001016 https://doi.org/10.1101/cshperspect.a001016.
He C, Li L, Guan X, Xiong L, Miao X. Mutant p53 gain of function and chemoresistance: the role of mutant p53 in response to clinical chemotherapy. Chemotherapy. 2017;62:43–53. https://doi.org/10.1159/000446361.
Okorokov AL, Sherman MB, Plisson C, Grinkevich V, Sigmundsson K, Selivanova G, et al. The structure of p53 tumour suppressor protein reveals the basis for its functional plasticity. EMBO J. 2006;25:5191–200. https://doi.org/10.1038/sj.emboj.7601382.
Silva JL, De Moura Gallo CV, Costa DC, Rangel LP. Prion-like aggregation of mutant p53 in cancer. Trends Biochem Sci. 2014;39:260–7. https://doi.org/10.1016/j.tibs.2014.04.001.
Joerger AC, Fersht AR. Structural biology of the tumor suppressor p53. Annu Rev Biochem. 2008;77:557–82. https://doi.org/10.1146/annurev.biochem.77.060806.091238.
Sabapathy K. The Contrived Mutant p53 Oncogene - Beyond Loss of Functions. Front Oncol. 2015;5:276 https://doi.org/10.3389/fonc.2015.00276.
Muller PA, Vousden KH. p53 mutations in cancer. Nat Cell Biol. 2013;15:2–8. https://doi.org/10.1038/ncb2641.
Lei J, Qi R, Wei G, Nussinov R, Ma B. Self-aggregation and coaggregation of the p53 core fragment with its aggregation gatekeeper variant. Phys Chem Chem Phys. 2016;18:8098–107. https://doi.org/10.1039/c5cp06538k.
Costa DC, de Oliveira GA, Cino EA, Soares IN, Rangel LP, Silva JL. Aggregation and prion-like properties of misfolded tumor suppressors: is cancer a prion disease? Cold Spring Harb Perspect Biol. 2016;8. https://doi.org/10.1101/cshperspect.a023614.
Stindt MH, Muller PA, Ludwig RL, Kehrloesser S, Dotsch V, Vousden KH. Functional interplay between MDM2, p63/p73 and mutant p53. Oncogene. 2015;34:4300–10. https://doi.org/10.1038/onc.2014.359. Epub 2014/11/25.
Rangel LP, Costa DC, Vieira TC, Silva JL. The aggregation of mutant p53 produces prion-like properties in cancer. Prion. 2014;8:75–84.
Stefani M. Protein misfolding and aggregation: new examples in medicine and biology of the dark side of the protein world. Biochim Biophys Acta. 2004;1739:5–25. https://doi.org/10.1016/j.bbadis.2004.08.004.
Brundin P, Melki R, Kopito R. Prion-like transmission of protein aggregates in neurodegenerative diseases. Nat Rev Mol Cell Biol. 2010;11:301–7. https://doi.org/10.1038/nrm2873.
Silva JL, Vieira TC, Gomes MP, Bom AP, Lima LM, Freitas MS, et al. Ligand binding and hydration in protein misfolding: insights from studies of prion and p53 tumor suppressor proteins. Acc Chem Res. 2010;43:271–9. https://doi.org/10.1021/ar900179t.
Ano Bom AP, Rangel LP, Costa DC, de Oliveira GA, Sanches D, Braga CA, et al. Mutant p53 aggregates into prion-like amyloid oligomers and fibrils: implications for cancer. J Biol Chem. 2012;287:28152–62. https://doi.org/10.1074/jbc.M112.340638.
Levy CB, Stumbo AC, Ano Bom AP, Portari EA, Cordeiro Y, Silva JL, et al. Co-localization of mutant p53 and amyloid-like protein aggregates in breast tumors. Int J Biochem Cell Biol. 2011;43:60–4. https://doi.org/10.1016/j.biocel.2010.10.017.
Higashimoto Y, Asanomi Y, Takakusagi S, Lewis MS, Uosaki K, Durell SR, et al. Unfolding, aggregation, and amyloid formation by the tetramerization domain from mutant p53 associated with lung cancer. Biochemistry. 2006;45:1608–19. https://doi.org/10.1021/bi051192j.
Rigacci S, Bucciantini M, Relini A, Pesce A, Gliozzi A, Berti A, et al. The (1-63) region of the p53 transactivation domain aggregates in vitro into cytotoxic amyloid assemblies. Biophys J. 2008;94:3635–46. https://doi.org/10.1529/biophysj.107.122283.
Ishimaru D, Andrade LR, Teixeira LS, Quesado PA, Maiolino LM, Lopez PM, et al. Fibrillar aggregates of the tumor suppressor p53 core domain. Biochemistry. 2003;42:9022–7. https://doi.org/10.1021/bi034218k. PubMed PMID: 12885235
Soragni A, Janzen DM, Johnson LM, Lindgren AG, Thai-Quynh Nguyen A, Tiourin E, et al. A designed inhibitor of p53 aggregation rescues p53 tumor suppression in ovarian carcinomas. Cancer Cell. 2016;29:90–103. https://doi.org/10.1016/j.ccell.2015.12.002.
Gregory CW, Johnson RT Jr., Mohler JL, French FS, Wilson EM. Androgen receptor stabilization in recurrent prostate cancer is associated with hypersensitivity to low androgen. Cancer Res. 2001;61:2892–8. Epub 2001/04/18.
Chen X, Wong JY, Wong P, Radany EH. Low-dose valproic acid enhances radiosensitivity of prostate cancer through acetylated p53-dependent modulation of mitochondrial membrane potential and apoptosis. Mol cancer Res: MCR. 2011;9:448–61. https://doi.org/10.1158/1541-7786.MCR-10-0471. Epub 2011/02/10
http://p53.free.fr/Database/Cancer_cell_lines/cell_lines_1.0.pdf.
Thalmann GN, Anezinis PE, Chang SM, Zhau HE, Kim EE, Hopwood VL, et al. Androgen-independent cancer progression and bone metastasis in the LNCaP model of human prostate cancer. Cancer Res. 1994;54:2577–81.
Gurova KV, Rokhlin OW, Budanov AV, Burdelya LG, Chumakov PM, Cohen MB, et al. Cooperation of two mutant p53 alleles contributes to Fas resistance of prostate carcinoma cells. Cancer Res. 2003;63:2905–12.
van Bokhoven A, Varella-Garcia M, Korch C, Johannes WU, Smith EE, Miller HL, et al. Molecular characterization of human prostate carcinoma cell lines. Prostate. 2003;57:205–25. https://doi.org/10.1002/pros.10290.
Tai S, Sun Y, Squires JM, Zhang H, Oh WK, Liang CZ, et al. PC3 is a cell line characteristic of prostatic small cell carcinoma. Prostate. 2011;71:1668–79. https://doi.org/10.1002/pros.21383.
Picksley SM, Vojtesek B, Sparks A, Lane DP. Immunochemical analysis of the interaction of p53 with MDM2;–fine mapping of the MDM2 binding site on p53 using synthetic peptides. Oncogene. 1994;9:2523–9.
Burch LR, Midgley CA, Currie RA, Lane DP, Hupp TR. Mdm2 binding to a conformationally sensitive domain on p53 can be modulated by RNA. FEBS Lett. 2000;472:93–8.
Matsumoto M, Furihata M, Ohtsuki Y. Posttranslational phosphorylation of mutant p53 protein in tumor development. Med Mol Morphol. 2006;39:79–87. https://doi.org/10.1007/s00795-006-0320-0.
Chipuk JE, Kuwana T, Bouchier-Hayes L, Droin NM, Newmeyer DD, Schuler M, et al. Direct activation of Bax by p53 mediates mitochondrial membrane permeabilization and apoptosis. Science. 2004;303:1010–4. https://doi.org/10.1126/science.1092734.
Beckerman R, Prives C. Transcriptional regulation byp53. Cold Spring Harb Perspect Biol. 2010;2:a000935 https://doi.org/10.1101/cshperspect.a000935.
Lim S, Kaldis P. Cdks, cyclins and CKIs: roles beyond cell cycle regulation. Development. 2013;140:3079–93. https://doi.org/10.1242/dev.091744.
Alimirah F, Panchanathan R, Chen J, Zhang X, Ho SM, Choubey D. Expression of androgen receptor is negatively regulated byp53. Neoplasia. 2007;9:1152–9.
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr., Kinzler KW. Cancer genome landscapes. Science. 2013;339:1546–58. https://doi.org/10.1126/science.1235122.
Bielas JH, Loeb KR, Rubin BP, True LD, Loeb LA. Human cancers express a mutator phenotype. Proc Natl Acad Sci USA. 2006;103:18238–42. https://doi.org/10.1073/pnas.0607057103.
Hieronymus H, Schultz N, Gopalan A, Carver BS, Chang MT, Xiao Y, et al. Copy number alteration burden predicts prostate cancer relapse. Proc Natl Acad Sci USA. 2014;111:11139–44. https://doi.org/10.1073/pnas.1411446111.
Soundararajan R, Aparicio AM, Logothetis CJ, Mani SA, Maity SN. Function of Tumor Suppressors in Resistance to Antiandrogen Therapy and Luminal Epithelial Plasticity of Aggressive Variant Neuroendocrine Prostate Cancers. Front Oncol. 2018;8:69 https://doi.org/10.3389/fonc.2018.00069.
Tsai HK, Lehrer J, Alshalalfa M, Erho N, Davicioni E, Lotan TL. Gene expression signatures of neuroendocrine prostate cancer and primary small cell prostatic carcinoma. BMC Cancer. 2017;17:759 https://doi.org/10.1186/s12885-017-3729-z.
Khemlina G, Ikeda S, Kurzrock R. Molecular landscape of prostate cancer: implications for current clinical trials. Cancer Treat Rev. 2015;41:761–6. https://doi.org/10.1016/j.ctrv.2015.07.001.
Kluth M, Harasimowicz S, Burkhardt L, Grupp K, Krohn A, Prien K, et al. Clinical significance of different types of p53 gene alteration in surgically treated prostate cancer. Int J Cancer. 2014;135:1369–80. https://doi.org/10.1002/ijc.28784.
Chen H, Sun Y, Wu C, Magyar CE, Li X, Cheng L, et al. Pathogenesis of prostatic small cell carcinoma involves the inactivation of the P53 pathway. Endocr Relat Cancer. 2012;19:321–31. https://doi.org/10.1530/ERC-11-0368.
Guseva NV, Rokhlin OW, Bair TB, Glover RB, Cohen MB. Inhibition of p53 expression modifies the specificity of chromatin binding by the androgen receptor. Oncotarget. 2012;3:183–94. https://doi.org/10.18632/oncotarget.449.
Zhang Y, Hu Y, Wang JL, Yao H, Wang H, Liang L, et al. Proteomic identification of ERP29 as a key chemoresistant factor activated by the aggregating p53 mutant Arg282Trp. Oncogene. 2017;36:5473–83. https://doi.org/10.1038/onc.2017.152.
Wang G, Fersht AR. Multisite aggregation of p53 and implications for drug rescue. Proc Natl Acad Sci USA. 2017;114:E2634–E43. https://doi.org/10.1073/pnas.1700308114.
Acknowledgements
The authors thank Dr. David Eisenberg and Dr. Alice Soragni from David Geffen School of Medicine, UCLA for providing ReACp53 and its scramble control, and for their helpful comments and advice on experimental design.
Funding
This work is supported by Stand Up to Cancer–Prostate Cancer Foundation–Prostate Dream Team Translational Cancer Research Grant (SU2C-AACR-DT0812). This research grant was made possible by the generous support of the Movember Foundation. Stand Up To Cancer is a program of the Entertainment Industry Foundation administered by the American Association for Cancer Research. Jiaoti Huang is supported by grants from the National Institutes of Health (1R01CA181242, 1R01CA172603, 1R01CA205001, 1U54CA217297, 1R01CA212403, 1R01CA200853), Department of Defense Prostate Cancer Research Program (PC150382), and Prostate Cancer Foundation (Joyce and Larry Stupski Prostate Cancer Precision Oncology Special Challenge award). Lingfan Xu is supported by grants from National Natural Science Foundation (81902611), Youth Culturing Plan of National Natural Science Foundation (2018Kj16) and Anhui Natural Science Foundation (1908085QH337). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Zhang, Y., Xu, L., Chang, Y. et al. Therapeutic potential of ReACp53 targeting mutant p53 protein in CRPC. Prostate Cancer Prostatic Dis 23, 160–171 (2020). https://doi.org/10.1038/s41391-019-0172-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41391-019-0172-z
This article is cited by
-
Physiology and pharmacological targeting of phase separation
Journal of Biomedical Science (2024)
-
Amyloid aggregates induced by the p53-R280T mutation lead to loss of p53 function in nasopharyngeal carcinoma
Cell Death & Disease (2024)
-
Amyloid-like p53 as prognostic biomarker in serous ovarian cancer—a study of the OVCAD consortium
Oncogene (2023)
-
Targeting p53 pathways: mechanisms, structures, and advances in therapy
Signal Transduction and Targeted Therapy (2023)
-
Mutant p53 in cancer: from molecular mechanism to therapeutic modulation
Cell Death & Disease (2022)