Abstract
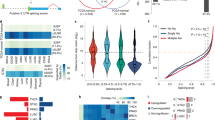
p53 is one of the most important tumor suppressor genes, and the exploration of p53-target genes is important for elucidation of its functional mechanisms. In this study, we identified Armadillo Repeat gene deleted in Velo-Cardio-Facial syndrome (ARVCF) as a direct target of p53 through ChIP-sequencing analysis. Activated p53 protein was found to bind to two distinct sites in the ARVCF gene, resulting in induction of ARVCF expression at both the mRNA and protein levels. We revealed that the knockdown of ARVCF inhibited p53-induced apoptosis. Interestingly, ARVCF interacted with hnRNPH2, which is involved in pre-mRNA splicing, and ARVCF knockdown induced dynamic changes in alternative splicing patterns. These results suggest that p53-induced ARVCF indirectly, but not directly, regulates p53 target selectivity through splicing alterations of specific genes. Thus, we demonstrated that the induction of ARVCF expression contributed to the tumor suppressive function of p53. Recently, it has been reported that many tumors have thousands of alternative splicing events that are not detectable in normal samples. ARVCF may play a role in alternative splicing events in cancer and may provide clues to explore novel approaches for cancer diagnosis and therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Vogelstein B, Lane D, Levine AJ. Surfing the p53 network. Nature. 2000;408:307–10.
Brosh R, Rotter V. When mutants gain new powers: news from the mutant p53 field. Nat Rev Cancer. 2009;9:701–13.
Idogawa M, Ohashi T, Sugisaka J, Sasaki Y, Suzuki H, Tokino T. Array-based genome-wide RNAi screening to identify shRNAs that enhance p53-related apoptosis in human cancer cells. Oncotarget. 2014;5:7540–8.
Ohashi T, Idogawa M, Sasaki Y, Tokino T. p53 mediates the suppression of cancer cell invasion by inducing LIMA1/EPLIN. Cancer Lett. 2017;390:58–66.
Ohashi T, Idogawa M, Sasaki Y, Suzuki H, Tokino T. AKR1B10, a transcriptional target of p53, is downregulated in colorectal cancers associated with poor prognosis. Mol Cancer Res. 2013;11:1554–63.
Idogawa M, Ohashi T, Sasaki Y, Maruyama R, Kashima L, Suzuki H, et al. Identification and analysis of large intergenic non-coding RNAs regulated by p53 family members through a genome-wide analysis of p53-binding sites. Hum Mol Genet. 2014;23:2847–57.
Idogawa M, Ohashi T, Sasaki Y, Nakase H, Tokino T. Long non-coding RNA NEAT1 is a transcriptional target of p53 and modulates p53-induced transactivation and tumor-suppressor function. Int J Cancer. 2017;140:2785–91.
McCrea PD, Park JI. Developmental functions of the P120-catenin sub-family. Biochim Biophys Acta. 2007;1773:17–33.
Chang GS, Chen XA, Park B, Rhee HS, Li P, Han KH, et al. A comprehensive and high-resolution genome-wide response of p53 to stress. Cell Rep. 2014;8:514–27.
Wang B, Niu D, Lam TH, Xiao Z, Ren EC. Mapping the p53 transcriptome universe using p53 natural polymorphs. Cell Death Differ. 2014;21:521–32.
Rappe U, Schlechter T, Aschoff M, Hotz-Wagenblatt A, Hofmann I. Nuclear ARVCF protein binds splicing factors and contributes to the regulation of alternative splicing. J Biol Chem. 2014;289:12421–34.
Olsson A, Manzl C, Strasser A, Villunger A. How important are post-translational modifications in p53 for selectivity in target-gene transcription and tumour suppression? Cell Death Differ. 2007;14:1561–75.
Katz Y, Wang ET, Airoldi EM, Burge CB. Analysis and design of RNA sequencing experiments for identifying isoform regulation. Nat Methods. 2010;7:1009–15.
Zhang D, Tang N, Liu Y, Wang EH. ARVCF expression is significantly correlated with the malignant phenotype of non-small cell lung cancer. Mol Carcinog. 2015;54:E185–191.
Idogawa M, Nakase H, Sasaki Y, Tokino T. Prognostic effect of long noncoding RNA NEAT1 expression depends on p53 mutation status in cancer. J Oncol. 2019;2019:4368068.
Kahles A, Lehmann KV, Toussaint NC, Huser M, Stark SG, Sachsenberg T, et al. Comprehensive analysis of alternative splicing across tumors from 8705 patients. Cancer Cell. 2018;34:211–224 e216.
Idogawa M, Sasaki Y, Suzuki H, Mita H, Imai K, Shinomura Y, et al. A single recombinant adenovirus expressing p53 and p21-targeting artificial microRNAs efficiently induces apoptosis in human cancer cells. Clin Cancer Res. 2009;15:3725–32.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60.
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28:511–5.
Fischer M. Census and evaluation of p53 target genes. Oncogene. 2017;36:3943–56.
Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Acknowledgements
The computations were partially performed on the NIG supercomputer at ROIS National Institute of Genetics.
Funding
This work was supported by JSPS KAKENHI (grant numbers: 19K08372, 19K07645, 16K09285, and 16K07122) and Takeda Science Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Suzuki, N., Idogawa, M., Tange, S. et al. p53-induced ARVCF modulates the splicing landscape and supports the tumor suppressive function of p53. Oncogene 39, 2202–2211 (2020). https://doi.org/10.1038/s41388-019-1133-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-1133-7
This article is cited by
-
MYEOV overexpression induced by demethylation of its promoter contributes to pancreatic cancer progression via activation of the folate cycle/c-Myc/mTORC1 pathway
BMC Cancer (2023)
-
Long noncoding RNA LINC01594 inhibits the CELF6-mediated splicing of oncogenic CD44 variants to promote colorectal cancer metastasis
Cell Death & Disease (2023)
-
Critical role of HOX transcript antisense intergenic RNA (HOTAIR) in gliomas
Journal of Molecular Medicine (2020)