Abstract
CDK4/6 inhibitors are being used to treat a variety of human malignancies. In well-differentiated and dedifferentiated liposarcoma their clinical promise is associated with their ability to downregulate the MDM2 protein. The downregulation of MDM2 following treatment with CDK4/6 inhibitors also induces many cultured tumor cell lines derived from different types of malignancies to progress from quiescence into senescence. Here we used cultured human cell lines and defined a role for PDLIM7 and CDH18, regulating MDM2 protein in CDK4/6 inhibitor-treated cells. Materials from our previous phase II trials with palbociclib were then used to demonstrate that expression of CDH18 protein was associated with response, measured as both progression-free survival and overall survival. This supports the hypothesis that the biologic transition from quiescence to senescence has clinical relevance for this class of drugs.
Similar content being viewed by others
Introduction
The commitment to cell proliferation is initiated when extracellular signals converge at the cell cycle and induce the expression of D-type cyclins, their association with CDK4 and/or CDK6, and the activation of the holoenzyme complex [1,2,3]. The cyclin D-associated kinases are necessary for the proliferation of Rb-positive cells because they initiate the phosphorylation-dependent cascade that inactivates this tumor suppressor [2, 4]. Unchecked proliferation of Rb-positive tumor cells is commonly associated with mutations that dysregulate this pathway: including the overexpression of D-type cyclins, the mutation or overexpression of CDK4, or mutations in the INK4 family of CDK inhibitors [3, 5, 6]. The importance of cyclin D holoenzymes for inactivation of Rb and the development of cancer in mice prompted the development of CDK4/6 inhibitors to treat a variety of neoplasms [7, 8]. These inhibitors have had success, both as a monotherapy and in combination [9].
Multiple cellular mechanisms have been advanced to account for the clinical activity of CDK4/6 inhibitors (reviewed in Klein et al., Cancer Cell in press). Many Rb-positive cells exit the cell cycle after CDK4/6 inhibition [10,11,12,13,14,15,16]. Resistance to these drugs, either acquired or innate, has been suggested to be due to a failure of the tumor cell to exit in response to the drug, linked to a failure to mobilize cells of the tumor microenvironment, or associated with the inability of the tumor cell to progress from reversible quiescence into more permanent senescence.
The decision of a tumor cell to senesce after CDK4/6 inhibition is made after the cell has withdrawn from the cell cycle. This previously unrecognized transition, now called senescence after growth arrest or SAGA, is triggered in the CDK4/6 inhibitor-induced quiescent cell by the loss of MDM2 protein and increased focal localization of the chromatin-remodeling enzyme ATRX [17, 18]. Palbociclib (also known as PD0332991)-induced senescence is not due to increased p53 [13, 18], nor is it associated with increased DNA damage [17].
The PD0332991-induced downregulation of MDM2 and entry into senescence is observed in a number of different types of cancer cell lines, including those derived from well-differentiated and dedifferentiated liposarcoma (WD/DDLS), breast cancer, non-small cell lung cancer, and glioma [18]. In a small pilot study of seven patients with WD/DDLS treated with palbociclib, the downregulation of MDM2, but not the absolute amount of the protein, also associated with how patients respond to the drug [18]. Thus, to understand how palbociclib improves patient outcomes it is important to understand how MDM2 is regulated in PD0332991-treated cells.
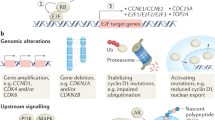
A number of cell type and signal-specific regulatory pathways can impact upon the accumulation of MDM2 protein (reviewed in ref. [19]). During SAGA, intrinsic E3 ligase activity is necessary for the downregulation of MDM2 [18]. HAUSP is a deubiquitinase that binds to MDM2 and removes ubiquitin from it, stabilizing the protein and allowing it to ubiquitinate other substrates [20, 21]. However, HAUSP dissociates from MDM2 as cells exit the cell cycle following palbociclib treatment, indicating that HAUSP does not play a role in whether quiescent cells downregulate MDM2 and proceed into senescence [18]. Thus, we set out to identify what stabilizes MDM2 protein in quiescent cells.
After attempting to knockdown five different genes whose proteins had been previously shown to inhibit MDM2 turnover [19], we show that one, PDLIM7, a PDZ and LIM domain-containing protein that binds to MDM2, is needed to stabilize MDM2 and prevent PD0332991-induced senescence. PDLIM7 was previously shown to inhibit MDM2 autoubiquitination and allow MDM2 to ubiquitinate p53 [22]. In cells that undergo senescence following PD0332991 treatment, we found that PDLIM7 was sequestered away from MDM2 by association with a type II cadherin, CDH18. In addition, both progression-free survival (PFS) and overall survival (OS) was significantly extended in patients with WD/DDLS tumors that are CDH18-positive and whom received palbociclib as a single agent in phase II clinical trials [23, 24]. This not only establishes that CDH18 and PDLIM7 impact on the regulation of MDM2 in PD0332991-induced quiescent cells, it reinforces the notion that SAGA contributes to the clinical outcome of patients treated with palbociclib.
Results
PDLIM7 knockdown allows PD0332991 to induce MDM2 turnover and senescence in cells that would otherwise quiesce
To gain insight into how MDM2 turnover was prevented in PD0332991-induced quiescent cells we carried out a small-targeted knockdown screen where we attempted to individually reduce expression of CREBBP, NEDD4-1, PDLIM7, PRKCD, or p300, all previously reported to regulate MDM2 auto-ubiqutination [19]. Parental WD/DDLS-derived LS8107 cells exit the cell cycle after PD0332991 treatment but MDM2 levels do not decrease and the cells do not undergo senescence. Throughout this manuscript we call such cells that when treated with PD0332991 do not undergo senescence as “non-responders”. In contrast, a ‘responder cell’ will exit the cell cycle after treatment with PD0332991 and progress into senescence. Knocking down MDM2 in PD0332991-induced quiescent non-responder cells will induce senescence [18].
Two independent lentiviral shRNA-targeting vectors efficiently reduced the level of PDLIM7, NEDD4-1, and p300 mRNAs (Fig. 1a). However, only in those in which PDLIM7 was reduced would PD0332991 treatment strongly induce the accumulation of senescence-associated beta-galactosidase (SA-β-gal), a marker of senescent cells (Fig. 1b). We confirmed that reducing PDLIM7 with the P2 shRNA also promoted the accumulation of SA-β-gal in another non-responder cell line, LS8313 (Supplemental Fig. 1). Furthermore, enforced expression of a his-biotin tagged mutant PDLIM7 in which the nucleic acid sequence was mutated to prevent interaction with the shRNA without affecting the protein-coding sequence prevented significant accumulation of SA-β-gal following treatment with PD0332991 (Supplemental Fig. 1).
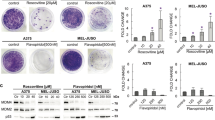
PDLIM7 knockdown allows PD0332991 to induce MDM2 turnover and senescence in LS8107 cells. a The non-responder cell line LS8107 was transduced with short hairpin expressing lentiviruses targeting either the indicated gene product or containing a non-specific sequence (scr), and selected in puromycin for 5 days. a Cells were harvested, RNA extracted, and qRT-PCR was performed to measure mRNA expression of the individual targets and quantitiated relative to the expression level in the LS8107 cells expressing the scr sequence. β-actin was used as a normalization control. The mean and standard deviation of three technical replicates is plotted. b Cells in a were treated with PD0332991 (PD) for 7 days and the number of cells staining for SA-β-gal quantified. c The cells transduced with the two different PDLIM7 knockdown lentiviral vectors (shP1 or shP2) or a non-specific vector (scr) were treated as in b, and MDM2 and PDLIM7 protein levels were detected using immunoblot. Tubulin served as a loading control. Top, a representative image is shown. Bottom, expression was quantified using densitometry and the mean and standard deviation of four independent experiments is shown. d LS8107scr and LS8107shP2 cells were treated as described in b and then exposed to 75 μg/mL cyclohexamide (CHX) for the time (min) indicated. MDM2 and tubulin were measured by immunoblot. A representative image is shown below and the relative amounts were quantified from two independent experiments (mean ± standard error of measurement) above. e The number of cells staining for SA-β-gal, HP1γ foci, and the number of ATRX foci per cell were quantified and the mean and standard deviation of four independent experiments is plotted
To confirm that reducing PDLIM7 altered the outcome of PD0332991-induced cell cycle exit we expanded our analysis to other markers of senescence. Seven days after the addition of PD0332991 to cell lines stably expressing either of the two different shRNAs targeting PDLIM7, LS8107shP1, or LS8107shP2, MDM2 levels decreased compared with a cell line stably expressing a scrambled shRNA, LS8107scr (Fig. 1c). Reduced levels of MDM2 were associated with the accelerated turnover of MDM2 in the LS8107shP2 cells treated with cyclohexamide (Fig. 1d). Similar results were observed in the LS8107shP1 cells. Furthermore, the accumulation of SA-β-gal-positive cells, senescence-associated HP1γ heterochromatic foci (SAHF)-positive cells, and the number of nuclear ATRX foci were increased after the addition of PD0332991 in both LS8107shP1 and LS8107shP2 cells but not LS8107scr cells (Fig. 1e). Thus, reducing PDLIM7 expression can alter the outcome of PD0332991-induced cell cycle exit, converting non-responder cells into responder cells.
The association of PDLIM7 with MDM2 is cell line and condition dependent
PDLIM7 is a PDZ and LIM domain-containing protein that can bind MDM2 and inhibits its autoubiquitination, but does not block E3 ligase activity, allowing MDM2/PDLIM7 complexes to catalyze the ubiquitination of other substrates [22]. PDLIM7 was readily detectable in extracts from both non-responder LS8107 and responder LS8817 cells, and the amount was not affected by treatment with PD0332991 (Fig. 2a).
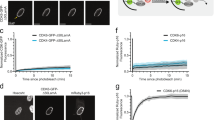
PDLIM7 is associated with MDM2 uniquely in LS8107 cells that quiesce and located in foci only in LS8817 cells that senesce. a The responder cell line LS8817 and non-responder cell line LS8107 were treated with 1 μM PD0332991 (PD) for 7 days. Cells were harvested for protein and MDM2 and PDLIM7 protein levels were detected using immunoblot. Tubulin served as a loading control. A representative image is shown. The mean expression value (PDLIM7/tubulin) quantified from four independent experiments is shown below each lane. b LS8817 and LS8107 cells were transduced with a vector expressing N-terminally tagged GFP-PDLIM7 and selected in puromycin. LS8817 cells were grown in serum-starved conditions with 0.5% serum for 4 days. MDM2 was immunoprecipitated and PDLIM7 immunoblotted (n = 3). IgG served as a control. c LS8817 and LS8107 cells were treated with either 1 μM PD0332991 (PD) for 7 days or grown in serum-starved conditions (0.5% serum) for seventy 2 h as shown. PDLIM7 was visualized by immunofluorescence (n = 3). d LS8817 and LS8107 cells were treated as shown. PDLIM7 and pan-cadherin were visualized by co-immunofluorescence. e LS8817 and LS8107 cells were fixed and incubated with antibodies against PDLIM7 and pan-cadherin followed by antibodies designed for the Sigma Duolink proximity ligation assay. Signal was visualized by immunofluorescence (n = 3)
LS8817 cells are derived from a different patient with WD/DDLS. The addition of PD0332991 induces these cells to exit the cell cycle and undergo senescence, with concomitant decreases in the level of MDM2, increases in the number of ATRX foci, and accumulation of SA-β-gal and SAHF-positive cells. On the other hand, serum starvation will induce these cells to exit the cell cycle, but the expression of MDM2 is not decreased, the number of ATRX foci does not increase, and SA-β-gal and SAHF-positive cells do not accumulate [17, 18].
We thus determined whether the binding of PDLIM7 to MDM2 was associated with outcome. To accomplish this, we transduced LS8107 and LS8817 cells with a N-terminally green fluorescent protein (GFP)-tagged PDLIM7, treated the cells as indicated and immunoprecipitated MDM2. Tagging PDLIM7 was necessary to facilitate its identification by immunoblot because the immunoglobulin heavy chain obscures the endogenous protein. GFP-PDLIM7 coprecipitated with MDM2 in both cycling LS8107 cells and serum-starved LS8817 cells, but not in the cycling LS8817 cells (Fig. 2b). Similar results were observed in both cell lines transduced with a C-terminally GFP-tagged PDLIM7. This suggested that the interaction of PDLIM7 with MDM2 was both cell line and condition dependent.
The cytosolic distribution of PDLIM7 is cell line and condition dependent
Although the expression of PDLIM7 could not account for the difference in MDM2 regulation following CDK4/6 inhibition in these two cell lines, the interaction of PDLIM7 with MDM2 was associated with the growth condition of LS8817 cells. PDZ domain proteins associate with actin and cytoskeletal structures [25,26,27], thus, we asked whether the localization of PDLIM7 might be associated with its ability to interact with MDM2. Immunofluorescence revealed that the localization of PDLIM7 was dynamically controlled. Focal depositions of cytosolic PDLIM7 were detected in the cycling LS8817 cells and in LS8817 cells that had undergone senescence following treatment with PD0332991, but not in quiescent serum-starved LS8817 cells (Fig. 2c). Focal depositions were also not detected in either cycling or PD0332991-treated quiescent LS8107 cells (Fig. 2c).
We next asked if other structural elements in the cell had a similar staining pattern in the two cell lines. There was no focal staining with antibodies raised against actin, γ-tubulin, or vimentin in either LS8817 or LS8107 cells (Supplemental Fig. 2). However, cytoplasmic foci were detected in LS8817 but not LS8107 cells stained with a pan-cadherin antibody (Supplemental Fig. 2).
Dual immunofluorescence indicated that the cytoplasmic cadherin foci overlapped with the PDLIM7 foci in LS8817 cells. No PDLIM7 or cadherin foci were identified in the LS8107 cells or serum-starved LS8817 cells (Fig. 2d). Confirming that these proteins were within 40 nm of each other we observed a positive signal in a proximity ligation assay (PLA) [28] in LS8817 cells (Fig. 2e). No such signal was observed when LS8817 cells were stained with individual antibodies or in LS8107 cells (Fig. 2e).
PDLIM7 is associated with CDH18 in foci
The surprising appearance of cadherin reactivity in the cytoplasm of the cell prompted us to investigate this further. The peptide immunogen used to generate the pan-cadherin antibody is found throughout the cadherin-C superfamily (Supplemental Fig. 3A) [29, 30]. To identify the specific cadherin responsible for this staining pattern we knocked down several individual family members using at least two independent lentiviral shRNA-targeting vectors in LS8817 cells and screened for pan-cadherin foci by immunofluorescence. Only those shRNAs targeting CDH18 consistently reduced the number of foci to less than five per cell and increased the number of cells with no foci to ~ 20% (Supplemental Fig. 3B).
To validate the effect of CDH18 loss on the focal staining of PDLIM7 we generated two different CDH18 mutant clones of LS8817 cells using CRISPR/Cas9. These independent knockout cell lines, LS8817CDH18KO1 and LS8817CDH18KO2, targeted CDH18 exons one and five, respectively. Sanger sequencing confirmed the presence of indels that lead to predicted frameshifts (Supplemental Fig. 3C), and CDH18 protein was reduced in both clones compared with parental cells (Supplemental Fig. 3D). In both cell lines, pan-cadherin foci were reduced or lost (Figs. 3a, b). PDLIM7 foci were also lost in both clones (Figs. 3a, b). Further we were unable to detect positive PLA signals (Fig. 3c). Consequently, PDLIM7 interacted with CDH18 in these cytoplasmic foci.
PDLIM7 is associated with CDH18 in foci in LS8817 cells. a The responder cell line LS8817 was unmanipulated (control), transduced with a PDLIM7 knockdown lentiviral vector (shP2), or a vector containing Cas9 and a vector containing a guide RNA against CDH18 (KO1 and KO2). PDLIM7 and pan-cadherin were visualized by co-immunofluorescence (n = 2). b The number of cells containing PDLIM7 and pan-cadherin foci were quantified. c Non-responder LS8107, responder LS8817, LS8817KO1, and LS8817shP2 cells were fixed and incubated with antibodies against PDLIM7 and CDH18 followed by antibodies designed for the Sigma Duolink proximity ligation assay. Signal was visualized by immunofluorescence (n = 2)
Knockout of CDH18 prevents PD0332991-induced MDM2 turnover and senescence
To determine whether differences in CDH18 expression might explain the differential response of cells to CDK4/6 inhibitors, we blotted CDH18 protein in a number of cell lines whose outcomes to CDK4/6 inhiibitors were previously characterized [18]. CDH18 protein was readily detected in LS8817, LS141, and LS0082 cells, and in the non-small cell lung cancer cell line H1975, all of which senesce following CDK4/6 inhibition (Supplemental Fig. 4). Expression of CDH18 was lower in LS8107 and LS8313 cells and in the non-small cell lung cancer cell line H358, all of which fail to senesce following CDK4/6 inhibition (Supplemental Fig. 4). All cells expressed PDLIM7, and although the amount varied, that did not correlate with the outcome of PD0332991 treatment, vis a vis quiescence or senescence (Supplemental Fig. 4).
To directly address whether CDH18 contributed to PD0332991-induced senescence and MDM2 turnover, we treated both LS8817 CDH18 knockout lines with PD0332991 and assayed a number of senescence markers seven days later. These lines were significantly impaired in their ability to undergo PD0332991-induced accumulation of SA-β-gal and SAHF (Fig. 4a). Furthermore, the LS8817CDH18KO1 and LS8817CDH18KO2 cells re-entered the cell cycle following removal of PD0332991 (Fig. 4b). MDM2 levels were not reduced in LS8817CDH18KO2 cells following treatment with PD0332991 (Fig. 4c), nor was the turnover of MDM2 accelerated to the same extent as seen in unmanipulated cells (Fig. 4d). Similar results were seen with LS8817CDH18KO1 cells. Collectively, this suggests that a CDH18 containing complex was sequestering PDLIM7 allowing for more rapid turnover of MDM2 and cellular senescence following treatment with PD0332991 (Fig. 4e).
Knockout of CDH18 prevents CDK4 inhibitor-induced MDM2 turnover and senescence in LS8817 cells. a LS8817 cells and LS8817 cells transduced with the CDH18 knockout guide RNAs (KO1 and KO2) were treated with 1 μM PD0332991 (PD) for 7 days and the number of cells (mean ± standard deviation) staining for SA-β-gal and HP1γ foci were quantified (n = 3). b LS8817, LS8817 KO1, and LS8817 KO2 cells were treated with 1 μM PD0332991 for 10 days and then plated in drug-free media and allowed to grow for 21 days to assess clonogenic growth. A representative image from three biologic replicates is shown. c MDM2 protein levels were detected using immunoblot on extracts from LS8817 and LS8817 KO2 cells. Tubulin served as a loading control. Top, a representative image is shown. Bottom, expression was quantified using densitometry and the mean and standard deviation of three independent experiments is plotted. d LS8817 and LS8817 KO2 cells were treated with 1 μM PD for 4 days and treated with 75 μg/mL cyclohexamide (CHX) for the time (min) indicated. A representative image is shown (bottom) and relative amounts of MDM2 and tubulin quantified using densitometry and plotted (upper). e Summary of MDM2 regulation after CDK4 inhibition
CDH18 expression in patients with WD/DDLS
In a small pilot study using seven pre- and post-treatment paired tumor sample biopsies from WD/DDLS patients who received palbociclib on clinical trial we had shown that the level of MDM2 was reduced in those who had a longer PFS [18]. This suggests that SAGA, induced by palbociclib, might be a mechanism that explains some of the clinical activity of this drug. To determine whether CDH18 might predict clinical benefit of the drug prior to therapy we measured its expression by immunohistochemistry in 49 surgical specimens obtained from patients who eventually enrolled in our palbociclib clinical trials and correlated this with PFS and OS.
The characteristics of these patients are shown in Supplemental Table 1. Details regarding the nature of the tissue samples and their staining with CDH18 antibodies are in the material and methods. The specificity of the antibody was validated in LS8817CDH18KO cell lines. Blinded pathology review was carried out and reactivity described as positive or negative. Infiltrating inflammatory cells in negative samples served as a positive control for staining, whereas endothelial cells in blood vessels were a negative control in the positive samples (Fig. 5a). Five samples were censored as positive and negative staining tumor cells were localized into specific domains within the mass. Such intratumoral heterogeneity is consistent with the variation reported for genetic markers in liposarcoma [31], but makes interpretation difficult. One patient was censored because control positive infiltrating inflammatory cells were not detected in the section. Information on the histology of the disease at the time of surgery, dose and scheduling of palbociclib, prior therapies, duration of survival after starting palbociclib, and CDH18 reactivity for the remaining 43 individual patients analyzed is shown in Supplemental Fig. 5.
CDH18 expression can stratify patient response to palbociclib monotherapy. a Patient tumor samples were obtained during surgery and formalin fixed and paraffin embedded. After antigen retrieval, IHC was performed using a CDH18-specific antibody. Hematoxylin and eosin was used as a counterstain. Samples were blinded during analysis. Representative images are shown from two patients with dedifferentiated histologies, a CDH18-negative tumor (left) and a CDH18-positive tumor (right). b PFS was plotted for patients with CDH18-positive and CDH18-negative tumor samples (p = 0.005). c Patients were grouped based on their PFS ( ≤ 12 weeks, n = 18; 12 < × < 24 weeks, n = 14; ≥ 24 weeks, n = 11). The percent of patients in each group that were CDH18-positive are shown in red and the percent of patients that were CDH18-negative shown in blue. d OS was plotted for patients with CDH18-positive and CDH18-negative tumor samples (p = 0.0007)
We plotted PFS (Figs. 5b, c) and OS (Fig. 5d) as a function of CDH18 expression in the remaining 43 patients. Patients whose tumors lacked CDH18 had worse outcomes than those who had CDH18 expression (PFS, p = 0.005; OS, p = 0.0007). Prior therapies and dosage schedule did not correlate with CDH18 staining and did not have a significant effect on either PFS or OS (Supplemental Table 2). However, all well-differentiated tumors were CDH18-positive, and because well-differentiated liposarcoma has a better prognosis than dedifferentiated liposarcoma [32], we reanalyzed the data after excluding well-differentiated samples. There was still a significant difference in outcome, with CDH18-positive patients having better survival, both by PFS (p = 0.05) and OS (p = 0.006) (Supplemental Figure 6).
For all CDH18-positive patients, the median PFS was 17.9 weeks (95% confidence interval, 17–25.9 weeks), and the median OS was 42.4 months (95% CI, 38.4 months to non-estimable (NE)). For CDH18-negative patients the median PFS was 9.0 weeks (95% CI, 6.1 weeks to NE) and OS was 19.8 months (95% CI, 8.9 months to NE). Although the power of the analysis is modest because the sample size is small, it does strengthen the hypothesis that SAGA is a clinically relevant biologic transition that underlies the efficacy of this class of drug.
Discussion
MDM2 downregulation is necessary for CDK4/6 inhibitor therapy-induced senescence in a variety of cancer cell lines, and the loss of MDM2 is observed post palbociclib treatment in patients with WD/DDLS who have prolonged periods of PFS [18]. Given that senescence is a more desirable outcome of cytostatic therapy, we set out to define the mechanism controlling MDM2 turnover in response to PD0332991 in cell lines and then asked if this associates with the clinical outcome of WD/DDLS patients who were treated with palbociclib on a clinical trial. In this report, we show that the association of PDLIM7 with MDM2 can inhibit its turnover in cells that exit the cell cycle but fail to undergo senescence following exposure to PD0332991. In cells that do undergo senescence following drug treatment, PDLIM7 is sequestered in a complex with an intracellular cadherin, CDH18, in a serum-dependent manner. Furthermore, CDH18 expression correlates with the outcomes of patients with WD/DDLS who were treated with palbociclib, suggesting that SAGA is a bona fide mechanism of this drugs noted clinical activity.
There are two general outcomes of CDK4/6 inhibition-induced cell cycle exit in Rb-positive tumor cells. Cells can enter a quiescent state from which they are able to return to cycle following drug removal, or cells enter a senescent state from which they cannot return to cycle following drug removal [10,11,12]. Recently, it was shown that the decision of the cell to senesce follows upon its exit from the cell cycle, illuminating a new transition in cell cycle biology—senescence after growth arrest or SAGA.
The proteins required and the mechanisms by which they act during this transition in cells treated with CDK4/6 inhibitors are being actively pursued. MDM2 downregulation is necessary for this transition. Its E3 ligase activity is required to maintain the quiescent state. However, this is not because p53 is targeted by MDM2. Mutations affecting MDM2’s ability to bind to p53 do not affect its ability to suppress this transition, nor is p53 required for senescence induced by CDK4/6 inhibitors [13, 18].
Turnover of MDM2 during this transition is dependent on its own E3 ligase activity [18], and is prevented in cells that fail to senesce and remain in a quiescent state. HAUSP, a deubiquitinase, can bind to MDM2 [21]. HAUSP binding stabilizes MDM2 and allows it to ubiquitinate other substrates while removing the ubiquitins it places on itself. Nevertheless, HAUSP dissociates from MDM2 as the cells exit the cell cycle following treatment with PD0332991 regardless of the outcome [18], thus whatever prevents turnover in the quiescent cells must act downstream of this to stabilize MDM2. We show here that PDLIM7 binds to MDM2 inhibiting its turnover and preventing quiescent cells from progressing into senescence.
PDLIM7 had been previously reported to bind to MDM2, preventing its autoubiquitination and allowing MDM2 to trans-ubiquitinate p53 [19, 22]. Although PDLIM7 is ubiquitously expressed in the liposarcoma and non-small cell lung cancer cell lines we examined here, its association with MDM2 was cell type and condition specific. PDLIM7 could be sequestered in CDH18 complexes. Thus, sequestration of PDLIM7 in CDH18 complexes is a mechanism to control SAGA, after the dissociation of HAUSP from MDM2. Serum starvation can prevent this interaction in LS8817 cells allowing them to maintain themselves in quiescence rather than undergo senescence. Although the details of this regulatory event are not yet defined, it is tempting to speculate that this reflects context specific transformation events that sensitize cancer cells to CDK4/6 inhibitors. In normal untransformed cells, CDH18 and PDLIM7 interactions are not readily detected and such cells do not undergo CDK4/6 inhibitor-induced senescence.
In patients with WD/DDLS who received palbociclib, CDH18 is significantly associated with PFS and OS. As a predictive marker for PFS, it is quite clear that those patients who are negative for CDH18 are more likely to do poorly. When we model the successful outcome of PFS as ≥ 24 weeks, as defined by the primary endpoint of the clinical trials, the negative predictive value associated with the absence of CDH18 staining is 92.3% (95% CI, 64.0–99.8%). In contrast the positive predictive value of CDH18 staining is 33.3% (95% CI, 17.3–52.8%). Thus, IHC for CDH18 may identify those patients who would benefit from clinical strategies aimed at augmenting SAGA, potentially through inhibitor based combination strategies sequentially targeting CDK4/6 and the Ras signaling pathway, another regulator of SAGA [17].
Successful expansion of CDK4/6 therapies will require pre-treatment biomarkers that are predictive of response and are applicable in a large number of diverse tumor types [33]. WD/DDLS is characterized by amplification of 12q13–15 [32], which contains both the genes encoding MDM2 and CDK4. In WD/DDLS cell lines, the level of MDM2 is generally higher in asynchronously growing cells that will undergo senescence than in those that undergo quiescence following treatment with CDK4/6 inhibitors (Supplemental Fig. 4). Likewise, in the NSCLC cell lines we have looked at the level of MDM2 is generally higher in the growing cells that will undergo senescence than those that undergo quiescence (Supplemental Fig. 4). However, in neither cell type is it the level of MDM2 that determines the outcome—rather it is the reduction of that level from cycling to non-cycling that is associated with outcome. Indeed, a number of non-mesenchymal cell lines that do not have MDM2 amplification are induced to senesce by palbociclib treatment. Palbociclib can be used to successfully treat non-MDM2 amplified tumors, the most notable being HR+ Her2− breast cancers [34].
Remarkably, the twofold increase in the median PFS between CDH18-positive and CDH18-negative patients with WD/DDLS is comparable in magnitude to the twofold increase in the duration of response in HR + , Her2− breast cancer patients treated with letrozole and palbociclib. As of this writing, the only pre-treatment indication of response to one of the CDK4/6 inhibitor drugs, abemaciclib, is a signature of cyclin D activation, called DCAF [35, 36]. This is exclusively in cell lines. Our data suggest that CDH18 and a deeper understanding of SAGA might also predict if cell lines or a patient’s tumor would be innately resistant or clinically sensitive to CDK4/6 inhibitors.
Materials and Methods
Clinical Trials
The design and approvals of clinical trials with palbociclib have been previously described [23, 24]. Surgical resections were available from Memorial Hospital under IRB 10-094. PD0332991 was obtained from Selleckchem and used for the in vitro experiments.
Cell lines
The WD/DDLS cell lines used in this manuscript were generated by Samuel Singer’s laboratory and previously described [18, 37]. Culture conditions and the response of these cell lines to CDK4 inhibition were previously described [17, 18].
Lentivirus constructs
As described previously [18], lentivirus vectors were generated in 293 T cells by triple transfection with the vector of interest, psPAX2, and pMD2.G. Infected cells were selected using puromycin (1 μg/mL) or blasticidin (3 μg/mL) as appropriate. shRNA were delivered in the pLKO.1 vector (Open Biosystems). shRNA sequences are as follows: PDLIM7 (GCGAGACTATGAGAAGATGTT and CGTCTGTGCGATATGTCAGAT), CREBBP (CCCGATAACTTTGTGATGTTT and GCTATCAGAATAGGTATCATT), NEDD4-1 (GCCTTTCTCTTGCCTGCATAT and CGGTTGGAGAATGTAGCAATA), PKCD (CGGCATGAATGTGCACCATAA and CAGAGCCTGTTGGGATATATC), and P300 (CCAGCCTCAAACTACAATAAA and CCCGGTGAACTCTCCTATAAT). To rescue knockdown, cells were infected with a lentivirus encoding His-Biotin tagged PDLIM7 with a mismatched sequence.
Real-time quantitative PCR
RNA was extracted from cells using the QIAGEN RNeasy kit per manufacturer’s instructions. Complementary DNA (cDNA) was synthesized from 1 μg of each RNA sample using the One Taq RT-PCR Kit and oligo-dT primers (New England BioLabs). cDNA was diluted 1:5 and 1 μL was used for qPCR per reaction. qPCR was performed by using 400 nM of each forward and reverse primer and SYBR Green PCR Master Mix (Life Technologies). qPCR was performed on Viia 7 Real-Time PCR System (Thermo Scientific). Primer sequences are as follows: PDLIM7 (Fw- CAGAGCCGCACCTCCATTG, Rev- TGGTGACACACGGGAGTCT), CREBBP (Fw- CCTGCCACGTCACAGACTG, Rev- GGCCAGAGTTACTATTGAGGAGG), p300 (Fw- GCTTCAGACAAGTCTTGGCAT, Rev- ACTACCAGATCGCAGCAATTC, PKCD (Fw- GTGCAGAAGAAGCCGACCAT, CCCGCATTAGCACAATCTGGA), NEDD4-1 (Fw- TCCAATGATCTAGGGCCTTTACC, Rev- TCCAACCGAGGATCTTCCCAT), β-actin (Fw- CATGTACGTTGCTATCCAGGC, Rev- CTCCTTAATGTACGCACGAT).
Immunoblots
Cells were lysed with a buffer containing 50 mM Tris-HCl, pH 7.4, 250 mM NaCl, 5 mM thylenediaminetetraacetic acid, 0.5% NP40, 2 mM phenylmethylsulfonyl fluoride, and protease inhibitors. Depending on the antibody used forty to eighty micrograms of protein were resolved by sodium dodecyl sulfate polyacrylamide gel electrophoresis and transferred to polyvinylidene difluoride membranes. Membranes were incubated overnight with primary antibodies.
The antibodies used for immunoblotting were MDM2 (Santa Cruz Biotechnology SMP14) 1:500, CDK4 (Santa Cruz Biotechnology C-22) 1:1000, PDLIM7 (Santa Cruz Biotechnology H-110) 1:2000, CDH18 (Abnova 6F7) 1:500, Tubulin (Santa Cruz Biotechnology C-11) 1:2000.
Protein expression was quantified by densitometry analysis using ImageJ software.
Immunoprecipitation
LS8817 and LS8107 cells were transduced with an LT3 lentivirus encoding GFP-tagged PDLIM7, and expression was induced by adding 10 μg/mL doxycycline to the media 2 days before cells were harvested. In LS8107 cells, expression of GFP-PDLIM7 was approximately equal to that of the endogenous protein. In LS8817 cells, expression was fourfold greater than the endogenous protein. Immunoprecipitation was performed by incubating 1.0–1.5 mg of protein lysate with 15–20 μL MDM2 SMP14 antibody or a mouse IgG control antibody overnight rotating at 4°C. Immune complexes were captured on either 30 μL of protein G sepharose or 20 μL of protein G dynabeads (catalog number 1003D) and subsequently eluted with 20–30 μL of 2 × sample buffer.
Senescence assays
All senescence assays were done 7–10 days after drug treatment and quantified as described previously [17, 18].
Immunofluorescence
Cells were plated in four-well glass chamber slides (Lab-Tek). Slides were fixed with 4% paraformaldehyde for 15 min, permeabilized with 0.1% triton for 5 min, blocked with phosphate-buffered saline (PBS) /0.5% Tween-20/1% bovine serum albumin for 20 min, and primary antibody was diluted in blocking buffer overnight. Dilutions for antibodies were as follows: HP1γ (Cell Signaling 05–690) 1:5000, ATRX (Santa Cruz Biotechnology H-300) 1:2000, pan-Cadherin (BD Biosciences 610181) 1:1000, PDLIM7 (Santa Cruz Biotechnology H-110) 1:2000. Fluorescent secondary antibodies (Alexa Flour Rabbit 488 and Mouse 546) were diluted at 1:500 and incubated for 1 h at room temperature and slides were counterstained with Hoechst for nuclear visualization. The Duolink Proximity Ligation Assay (Sigma) was performed according to manufacturer’s instructions.
CRISPR Cas9
lentiCas9-Blast and lentiGuide-Puro were a ‘gift’ from Feng Zhang (Addgene plasmid numbers 52962 and 52963). Guide sequences targeting CDH18 were designed using Benchling software. Guides were tested for successful genomic editing by the T7 endonuclease mismatch assay as described previously [38]. Guides were transduced into LS8817 cells and selected in puromycin for 6 weeks. Clones that demonstrated efficient cutting were sequenced and tested for the loss of CDH18 protein as measured by immunoblot. Cells were maintained in culture for up to eight weeks after sequencing while experiments were carried out.
Immunohistochemistry
Formalin-fixed paraffin embedded blocks were sectioned at four microns. The well-differentiated or dedifferentiated status of the tumor and tumor cells were identified by visualization of a parallel section stained by Haemotoxylin and Eosin. Tissue sections ranged from 200 mm2 to 600 mm2 (with an average of 377 mm2) and were obtained from surgical resections carried out 100–2100 days prior to palbociclib therapy. Immunohistochemistry detection with CDH18 antibody was performed using the Discovery XT processor (Ventana Medical Systems). Tissue sections were blocked first for 30 min in MOM Block (Vector Labs, MKB-2213) in PBS. A mouse monoclonal antibody to CDH18 (Abnova, H00001016-M01) was used at a 3 μg/mL concentration and incubation was done for 4 h, followed by another 60 min incubation with biotinylated anti-mouse Secondary (Vector Labs, MOM Kit BMK-2202), at a 5.75 μg/mL concentration. Blocker D, Streptavidin-horseradish peroxidase and DAB detection kit (Ventana Medical Systems) were used according to the manufacturer instructions. Tumor cells were identified within the sample based on their nuclear morphology. Samples where > 80% of the tumor cells throughout contained cytoplasmic CDH18 staining were deemed positive.
Twenty-eight of the 49 patients received at least one prior systemic therapy sometime between this surgery and treatment with palbociclib. These included gemcitabine, doxil, docetaxel, brivanib, irinotecan, or an MDM2 inhibitor, either alone or in various combinations. The median number of therapies received was one (range from one to four). Two different dose schedules of palbociclib were used. Thirty-two patients received palbociclib at 125 mg daily for 21 days followed by a rest period of 7 days every 28 days, and 17 received 200 mg daily for 14 days with a rest period of 7 days every 21 days.
Statistical analysis
Disease progression and death were the endpoints of this study. Disease progression was defined as the time from the treatment start to the occurrence of disease progression; death was defined as the time from the treatment start to the occurrence of death owing to any cause or the date of last follow-up. The clinicopathologic variables examined were CDH18 staining (positive vs. negative), treatment dosage level (125 mg vs. 200 mg), disease subtype (well-differentiated vs. dedifferentiated), and prior therapy (yes vs. no). The probabilities of progression and death were each estimated using the Kaplan–Meier method. The associations of these outcomes with the clinicopathologic variables were examined using the log-rank test for categorical variables. P values ≤ 0.05 were considered significant. All analyses were performed using R version 3.2.0 (cran.r-project.org).
References
Ezhevsky SA, Nagahara H, Vocero-Akbani AM, Gius DR, Wei MC, Dowdy SF. Hypo-phosphorylation of the retinoblastoma protein (pRb) by cyclin D:Cdk4/6 complexes results in active pRb. Proc Natl Acad Sci USA. 1997;94:10699–704.
Harbour JW, Dean DC. The Rb/E2F pathway: expanding roles and emerging paradigms. Genes Dev. 2000;14:2393–409.
Sherr CJ, Roberts JM. CDK inhibitors: positive and negative regulators of G1-phase progression. Genes Dev. 1999;13:1501–12.
Lundberg AS, Weinberg RA. Functional inactivation of the retinoblastoma protein requires sequential modification by at least two distinct cyclin-cdk complexes. Mol Cell Biol. 1998;18:753–61.
Burkhart DL, Sage J. Cellular mechanisms of tumour suppression by the retinoblastoma gene. Nat Rev Cancer. 2008;8:671–82.
Knudsen ES, Knudsen KE. Tailoring to RB: tumour suppressor status and therapeutic response. Nat Rev Cancer. 2008;8:714–24.
Sherr CJ, Beach D, Shapiro GI. Targeting CDK4 and CDK6: from discovery to therapy. Cancer Discov. 2016;6:353–67.
Otto T, Sicinski P. Cell cycle proteins as promising targets in cancer therapy. Nat Rev Cancer. 2017;17:93–115.
Vora SR, Juric D, Kim N, Mino-Kenudson M, Huynh T, Costa C, et al. CDK 4/6 inhibitors sensitize PIK3CA mutant breast cancer to PI3K inhibitors. Cancer Cell. 2014;26:136–49.
Baughn LB, Di Liberto M, Wu K, Toogood PL, Louie T, Gottschalk R, et al. A novel orally active small molecule potently induces G1 arrest in primary myeloma cells and prevents tumor growth by specific inhibition of cyclin-dependent kinase 4/6. Cancer Res. 2006;66:7661–7.
Choi YJ, Li X, Hydbring P, Sanda T, Stefano J, Christie AL, et al. The requirement for cyclin D function in tumor maintenance. Cancer Cell. 2012;22:438–51.
Michaud K, Solomon DA, Oermann E, Kim JS, Zhong WZ, Prados MD, et al. Pharmacologic inhibition of cyclin-dependent kinases 4 and 6 arrests the growth of glioblastoma multiforme intracranial xenografts. Cancer Res. 2010;70:3228–38.
Yoshida A, Lee EK, Diehl JA. Induction of therapeutic senescence in vemurafenib-resistant melanoma by extended inhibition of CDK4/6. Cancer Res. 2016;76:2990–3002.
Bollard J, Miguela V, Ruiz de Galarreta M, Venkatesh A, Bian CB, Roberto MP, et al. Palbociclib (PD-0332991), a selective CDK4/6 inhibitor, restricts tumour growth in preclinical models of hepatocellular carcinoma. Gut. 2017;66:1286–96.
Witkiewicz AK, Borja NA, Franco J, Brody JR, Yeo CJ, Mansour J, et al. Selective impact of CDK4/6 suppression on patient-derived models of pancreatic cancer. Oncotarget. 2015;6:15788–801.
Yang C, Li Z, Bhatt T, Dickler M, Giri D, Scaltriti M, et al. Acquired CDK6 amplification promotes breast cancer resistance to CDK4/6 inhibitors and loss of ER signaling and dependence. Oncogene. 2017;36:2255–64.
Kovatcheva M, Liao W, Klein ME, Robine N, Geiger H, Crago AM, et al. ATRX is a regulator of therapy induced senescence in human cells. Nat Commun. 2017;8:386.
Kovatcheva M, Liu DD, Dickson MA, Klein ME, O’Connor R, Wilder FO, et al. MDM2 turnover and expression of ATRX determine the choice between quiescence and senescence in response to CDK4 inhibition. Oncotarget. 2015;6:8226–43.
Riley MF, Lozano G. The many faces of MDM2 binding partners. Genes Cancer. 2012;3:226–39.
Brooks CL, Li M, Hu M, Shi Y, Gu W. The p53--Mdm2--HAUSP complex is involved in p53 stabilization by HAUSP. Oncogene. 2007;26:7262–6.
Li M, Brooks CL, Kon N, Gu W. A dynamic role of HAUSP in the p53-Mdm2 pathway. Mol Cell. 2004;13:879–86.
Jung CR, Lim JH, Choi Y, Kim DG, Kang KJ, Noh SM, et al. Enigma negatively regulates p53 through MDM2 and promotes tumor cell survival in mice. J Clin Invest. 2010;120:4493–506.
Dickson MA, Schwartz GK, Keohan ML, D’Angelo SP, Gounder MM, Chi P, et al. Progression-free survival among patients with well-differentiated or dedifferentiated liposarcoma treated with cdk4 inhibitor palbociclib: a phase 2 clinical trial. JAMA Oncol. 2016;2:937–40.
Dickson MA, Tap WD, Keohan ML, D’Angelo SP, Gounder MM, Antonescu CR, et al. Phase II trial of the CDK4 inhibitor PD0332991 in patients with advanced CDK4-amplified well-differentiated or dedifferentiated liposarcoma. J Clin Oncol. 2013;31:2024–8.
Fanning AS, Anderson JM. Protein-protein interactions: PDZ domain networks. Curr Biol. 1996;6:1385–8.
Krcmery J, Camarata T, Kulisz A, Simon HG. Nucleocytoplasmic functions of the PDZ-LIM protein family: new insights into organ development. Bioessays. 2010;32:100–8.
Urban AE, Quick EO, Miller KP, Krcmery J, Simon HG. Pdlim7 regulates Arf6-dependent actin dynamics and is required for platelet-mediated thrombosis in mice. PLoS ONE. 2016;11:e0164042.
Soderberg O, Gullberg M, Jarvius M, Ridderstrale K, Leuchowius KJ, Jarvius J, et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods. 2006;3:995–1000.
Halbleib JM, Nelson WJ. Cadherins in development: cell adhesion, sorting, and tissue morphogenesis. Genes Dev. 2006;20:3199–214.
van Roy F. Beyond E-cadherin: roles of other cadherin superfamily members in cancer. Nat Rev Cancer. 2014;14:121–34.
Kanojia D, Nagata Y, Garg M, Lee DH, Sato A, Yoshida K, et al. Genomic landscape of liposarcoma. Oncotarget. 2015;6:42429–44.
Tan MC, Brennan MF, Kuk D, Agaram NP, Antonescu CR, Qin LX, et al. Histology-based classification predicts pattern of recurrence and improves risk stratification in primary retroperitoneal sarcoma. Ann Surg. 2016;263:593–600.
O’Leary B, Finn RS, Turner NC. Treating cancer with selective CDK4/6 inhibitors. Nat Rev Clin Oncol. 2016;13:417–30.
Finn RS, Martin M, Rugo HS, Jones S, Im SA, Gelmon K, et al. Palbociclib and letrozole in advanced breast cancer. N Engl J Med. 2016;375:1925–36.
Gong X, Litchfield LM, Webster Y, Chio LC, Wong SS, Stewart TR, et al. Genomic aberrations that activate D-type cyclins are associated with enhanced sensitivity to the CDK4 and CDK6 inhibitor abemaciclib. Cancer Cell. 2017;32:761–76 e6.
Shapiro GI. Genomic biomarkers predicting response to selective CDK4/6 inhibition: progress in an elusive search. Cancer Cell. 2017;32:721–3.
Barretina J, Taylor BS, Banerji S, Ramos AH, Lagos-Quintana M, Decarolis PL, et al. Subtype-specific genomic alterations define new targets for soft-tissue sarcoma therapy. Nat Genet. 2010;42:715–21.
Guschin DY, Waite AJ, Katibah GE, Miller JC, Holmes MC, Rebar EJ. A rapid and general assay for monitoring endogenous gene modification. Methods Mol Biol. 2010;649:247–56.
Acknowledgements
We thank Annalee Tai for technical assistance, Maura Dickler for discussions on the clinical impact of palbociclib in breast cancer, and Marta Kovatcheva, Caroline Gleason, Andrew Schade (Eli Lilly and Company), Michael Overholtzer and others in our laboratories for discussions during the course of these studies. The research was funded in part through NIH/NCI Cancer Center Support Grant P30 CA008748 and the NCI Soft Tissue Sarcoma SPORE CA140146 (AK, WDT, MD, GKS). Additional funding was provided by the Maloris Foundation, The Linn Fund from Cycle For Survival, and a Heidi Connery Memorial Research Grant from the Sarcoma Foundation of America to AK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
These authors are co-communicating authors: Andrew Koff and William D. Tap
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Klein, M.E., Dickson, M.A., Antonescu, C. et al. PDLIM7 and CDH18 regulate the turnover of MDM2 during CDK4/6 inhibitor therapy-induced senescence. Oncogene 37, 5066–5078 (2018). https://doi.org/10.1038/s41388-018-0332-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0332-y
This article is cited by
-
Aging microenvironment and antitumor immunity for geriatric oncology: the landscape and future implications
Journal of Hematology & Oncology (2023)
-
CDH18 is a fetal epicardial biomarker regulating differentiation towards vascular smooth muscle cells
npj Regenerative Medicine (2022)
-
Exploiting senescence for the treatment of cancer
Nature Reviews Cancer (2022)
-
MDM2 inhibition in combination with endocrine therapy and CDK4/6 inhibition for the treatment of ER-positive breast cancer
Breast Cancer Research (2020)