Abstract
IRF5 genetic variants leading to decreased IRF5 expression reduce risk for ulcerative colitis. However, how IRF5 regulates intestinal inflammation and contributes to the balance between defenses against intestinal pathogens and inflammation in vivo, and the cells mediating this balance, are not known. We found that deleting IRF5 in mice led to reduced intestinal inflammation in the T cell transfer colitis model, with reduced Th1 and Th17, and increased Th2 cytokines. However, with orally-administered invasive S. Typhimurium, IRF5−/− mice demonstrated an increased bacterial burden in the context of reduced Th1 and Th17 cytokines. IRF5 in macrophages was required for PDK1-dependent phagocytosis and for NFκB-dependent pathways mediating intracellular bacterial clearance. Despite reduced bacterial clearance pathways, in IRF5−/− mice exposed to high levels of resident intestinal bacteria after DSS-induced injury, the lower levels of inflammatory cytokines were associated with reduced intestinal permeability, and in turn, reduced bacterial translocation and intestinal inflammation. Consistent with the myeloid cell-intrinsic roles for IRF5 in vitro, mice with IRF5 deleted from myeloid cells demonstrated outcomes similar to those observed in IRF5−/− mice. While these data suggest that inhibition of IRF5 may be therapeutic in colitis, this needs to be balanced with the identified IRF5 role in protecting against intestinal pathogens.
Similar content being viewed by others
Introduction
Upon infectious challenges, the immune system must respond in a manner that clears microbes while minimizing excessive inflammation and tissue injury. The importance of this balance is particularly pertinent at mucosal surfaces, including the intestine, where the immune system is continuously exposed to microbes. Host pattern recognition receptors (PRRs) provide the initial recognition and response to microbes; this response includes induction of cytokine secretion and of microbial clearance pathways. Polymorphisms in genes that modulate PRR- or cytokine-initiated responses are frequently associated with immune-mediated diseases, including IBD.1 In these cases, reduced PRR responses may decrease IBD risk, but increase susceptibility to pathogens. However, immunological pathways can be more complex in intestinal tissues where there is ongoing microbial exposure, as a decreased ability to clear microbes can also result in uncontrolled inflammation (e.g. variants in NOD2 or in the NADPH complex2,3). Therefore, the manner in which a particular gene confers risk for IBD is not necessarily readily apparent. As such, a significant challenge has been to determine the mechanism(s) through which genes conferring altered IBD susceptibility contribute to intestinal immune homeostasis and disease pathogenesis.
IRF5 is required for optimal responses to PRRs.4,5,6,7 Genetic variants in IRF5 that decrease IRF5 mRNA8 and protein7 expression are associated with reduced susceptibility to ulcerative colitis, as well as to multiple other immune-mediated diseases, including SLE, rheumatoid arthritis, primary biliary cirrhosis, Sjogren’s disease, and systemic sclerosis.9 This broad range of immune-mediated diseases highlights a critical role for IRF5 in immunity and suggests that IRF5 may represent a nodal point for therapy of immune-mediated diseases.
Despite a paucity of information on intestinal IRF5 function, a few studies have examined IRF5 in bacterial clearance. In mice, IRF5 is required for optimal control of Leishmania donovani infection in the liver.10 In vitro, IRF5 in mouse myeloid cells is required for type I IFN induction following Mycobacterium tuberculosis11 or S. pyogenes12,13 co-culture. In these limited studies identifying contributions for IRF5 in bacterial clearance, the myeloid cell-intrinsic role for IRF5 in bacterial clearance in vivo was not examined. Further, the mechanisms through which IRF5 mediate this clearance were not broadly examined; myeloid cells, including macrophages, are critical in regulating bacterial infection and in IBD pathogenesis. Despite the IRF5 genetic associations with IBD,2,14 to our knowledge there are no reports clearly defining the role of IRF5 in intestinal inflammation or the cell subsets through which IRF5 might be mediating its contributions to intestinal responses in vivo. Such studies are critical given the distinct regulation and outcomes that can be observed between systemic versus intestinal immune responses (e.g. IL23).15
Here we asked if IRF5 contributes to the balance of intestinal inflammation and clearance of intestinal bacteria and also sought to define mechanisms and cell subsets through which IRF5 contributes to these functions. Using IRF5−/− mice, we show that IRF5 is required for PDK1-dependent phagocytosis of microbes and for the induction of antimicrobial pathways, including ROS (upregulation of NADPH complex members), RNS (upregulation of NOS2) and autophagy (induction of ATG5, ATG16L1 and IRGM), leading to clearance of intracellular bacteria in bone marrow-derived macrophages (BMMs). We also demonstrate that while IRF5 is important for regulating invasive enteric pathogens such as S. Typhimurium, during exposure to resident intestinal luminal bacteria in the setting of acute epithelial injury or chronic colitis, IRF5 contributes to the proinflammatory response to microbes and to the increased intestinal inflammation under these conditions. Finally, we show that myeloid cell-intrinsic IRF5 is required for regulation of invasive intestinal bacteria on the one hand, and for the inflammation observed upon exposure to high levels of resident intestinal luminal bacteria with epithelial injury on the other hand. These studies therefore define mechanisms by which the IBD-associated gene IRF5 regulates the balance between bacterial clearance and intestinal bacterial-induced inflammation.
Results
IRF5 contributes to chronic intestinal inflammation
To address the role of IRF5 in chronic colitis, we crossed IRF5−/− mice to RAG-1−/− mice and examined IRF5+/+, IRF5+/− and IRF5−/− x RAG-1−/− littermate control mice in the adoptive T cell transfer model of chronic colitis. Relative to IRF5+/+RAG-1−/− mice, IRF5−/−RAG-1−/− mice adoptively transferred with CD45RBhi CD4+ T cells demonstrated less weight loss, less severe histological inflammation, and lower levels of mucosal myeloperoxidase (MPO) activity (measure of myeloid cells) and stool lipocalin (measure of intestinal inflammation16) (Fig. 1a–e). Moreover, these mice demonstrated reduced levels of the Th1-associated cytokines IL12 and IFNγ, the proinflammatory cytokine TNF, and the Th17-associated cytokine IL17 in colon (Fig. 1f). In contrast, and consistent with select prior reports of IRF5−/− mice infected with L. donovani10 or in models of lupus and asthma,17,18,19 the Th2-associated cytokine IL13 and the anti-inflammatory cytokine IL10 in colon were increased (Fig. 1f). Serum of IRF5−/−RAG-1−/− recipients demonstrated similar cytokine patterns (Fig. 1g). IRF5+/−RAG-1−/− recipient mice generally demonstrated an intermediate phenotype (Fig. 1). Therefore, consistent with the reduced susceptibility to chronic ulcerative colitis in IRF5 genetic variant carriers with low IRF5 expression,2,6,7,14 IRF5−/− mice demonstrate a decreased severity of experimental chronic colitis with reduced Th1 and Th17 cytokines.
IRF5−/− RAG-1−/− mice demonstrate less severe chronic colitis when transferred with WT CD45RBhi CD4+ T cells. CD45RBhi CD4+ T cells were adoptively transferred into IRF5+/+, IRF5+/− or IRF5−/− RAG-1−/− mice (n = 4–10; 3-4 months of age). a Body weights. b Representative histology of mice with no transfer (−) or with CD45RBhi CD4+ T cell transfer. Scale bar = 25 μm. c Histologic scores. d Colon myeloperoxidase. e Fecal lipocalin. f Colon cytokines. g Serum cytokines. Mean + SEM. *p < 0.05; **p < 0.01; ***p < 0.001; †p < 1 × 10−4; ††p < 1 × 10−5
IRF5−/− mice infected with oral S. Typhimurium demonstrate impaired immune responses and an increased bacterial burden
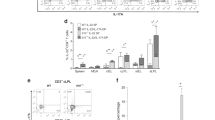
As IRF5 regulates PRR-induced outcomes, and PRR-initiated responses are critical in mediating bacterial clearance, we next assessed if IRF5 regulates responses to intestinal pathogens. IRF5+/+, IRF5+/− and IRF5−/− littermate control mice were orally infected with the enteric pathogen Salmonella enterica serovar Typhimurium and assessed 6 days later. IRF5−/− mice had a higher bacterial burden in the colon and feces, as well as in systemic organs including the liver and spleen relative to IRF5+/+ mice (Fig. 2a). Myeloid cells such as neutrophils play an important role in the clearance of acute enteric pathogens, including S. Typhimurium.20 Mucosal MPO levels were reduced in S. Typhimurium-infected IRF5−/− relative to IRF5+/+ mice (Fig. 2b). Immune cell-derived cytokines are also critical in regulating enteric pathogens, including S. Typhimurium.20 Colon and serum from S. Typhimurium-infected IRF5−/− mice demonstrated decreased levels of Th1 (IL12, IFNγ), Th17 (IL17), and proinflammatory (TNF) cytokines relative to infected IRF5+/+ mice (Fig. 2c, d). In contrast, and similar to what we observed with chronic colitis (Fig. 1f, g), the Th2-associated cytokine IL13 was elevated, as was the anti-inflammatory cytokine IL10 (Fig. 2c, d). IRF5+/− mice infected with S. Typhimurium generally demonstrated an intermediate phenotype (Fig. 2). Therefore, IRF5 is required for optimal production of Th1 and Th17 cytokines, reduction of Th2 cytokines, induction of colonic myeloid responses, and clearance of the enteric pathogen S. Typhimurium upon oral infection.
IRF5 is required for optimal clearance of S. Typhimurium upon oral infection in mice. IRF5+/+, IRF5+/− and IRF5−/− mice (n = 3–9; 3–3.5 months of age) were orally inoculated with S. Typhimurium. 6 days later mice were sacrificed. a S. Typhimurium CFU in the indicated tissues. b Colon myeloperoxidase. c Colon cytokines. d Serum Cytokines. Mean + SEM. Representative of 3 independent experiments. *p < 0.05; **p < 0.01; ***p < 0.001; †p < 1 × 10−4; ††p < 1 × 10−5
IRF5 promotes bacterial uptake in macrophages
Myeloid cells, including macrophages, are critical for the response to and clearance of enteric S. Typhimurium.20 We therefore first assessed the role of IRF5 in mediating phagocytosis and bacterial entry in macrophages in vitro. We confirmed reduced IRF5 expression in IRF5−/− compared to IRF5+/+ BMMs both at baseline and under lipid A (TLR4 ligand)-treated conditions where IRF5 expression is increased; IRF5+/− BMMs were intermediate in expression (Supplementary Fig. 1a). Myeloid cells in the intestinal lamina propria are continuously exposed to microbial ligands, and we21 and others22,23 have found that chronic PRR stimulation of macrophages increases anti-microbial mechanisms and efficacy of bacterial clearance. Chronic lipid A-treatment of MDMs similarly enhanced bacterial uptake (Fig. 3a). S. Typhimurium (GFP) entry at 20 min was decreased in IRF5−/− compared to IRF5+/+ BMMs in both untreated and chronic lipid A-treated conditions (Fig. 3a, Supplementary Fig. 1b); IRF5+/− BMMs generally demonstrated intermediate levels of bacterial entry (Fig. 3a). Similar results were observed with E. coli and S. aureus particles (Fig. 3a, Supplementary Fig. 1b). The PI3K pathway contributes to phagocytosis,24 such that we asked if IRF5 regulation of the PI3K pathway might be a mechanism through which IRF5 regulates bacterial phagocytosis. Lipid A treatment of both untreated and chronic lipid A-treated BMMs activated PDK1 (Fig. 3b), a contributor to PI3K pathway activation. This activation was reduced in IRF5−/− BMMs (Fig. 3b). Moreover, live bacterial and bacterial particle entry was reduced with either a PDK1 inhibitor or the PI3K pathway inhibitor wortmannin in chronic lipid A-treated BMMs (cell survival was intact with these inhibitors [Supplementary Fig. 1c]) (Fig. 3c). Therefore, IRF5 is required for optimal PDK1 activation and PI3K/PDK1-dependent bacterial entry into macrophages.
BMMs from IRF5−/− mice demonstrate reduced PRR-induced PDK1 activation and PI3K/PDK1-dependent phagocytosis. a, b BMMs from IRF5+/+, IRF5+/− or IRF5−/− mice were left untreated or treated for 48 h with 0.1 μg/ml lipid A (mature). a Cells were then co-cultured with S. Typhimurium-GFP, or FITC-labeled E. coli or S. aureus bioparticles for 20 min, and intracellular uptake was examined by flow cytometry (mean fluorescence intensity (MFI) + SEM) (3 replicates, representative of 2 independent experiments). b Cells were treated with lipid A for 20 min and examined for phosphorylated-PDK1 by Western blot. PDK1 and GAPDH served as loading controls. c BMMs were treated for 48 h with 0.1 μg/ml lipid A, then pre-treated with a PDK1 inhibitor (GSK) or wortmannin for 1 h and then co-cultured with bacteria or bacterial particles as per (a). MFI + SEM (3 replicates). Significance is compared to IRF5+/+ macrophages for each respective treatment condition or as indicated. Inh, inhibitors. *p < 0.05; **p < 0.01; ***p < 0.001; †p < 1 × 10−4; ††p < 1 × 10−5
IRF5 enhances intracellular bacterial clearance in macrophages
We next assessed if IRF5 was required for clearance of bacteria that have entered macrophages. We first confirmed enhanced bacterial clearance in chronic lipid A-treated macrophages (Fig. 4a), as per our prior studies.21 IRF5−/− BMMs demonstrated less effective clearance of intracellular S. Typhimurium both under untreated and chronic lipid A-treated conditions (Fig. 4a). This less effective bacterial clearance extended to additional bacteria, including adherent invasive E. coli (AIEC), a strain of E. coli enhanced in the ilea of Crohn’s disease patients,25 S. aureus, a resident bacteria (Fig. 4a), and E. faecalis, a resident intestinal luminal bacteria (Supplementary Fig. 2a). Therefore, despite a reduced level of bacterial entry, IRF5−/− BMMs demonstrate impaired clearance of those bacteria that do enter the cells. Consistent with the reduced cytokine secretion upon in vivo infection with S. Typhimurium in IRF5−/− mice (Fig. 2), IRF5−/− BMMs demonstrated reduced proinflammatory cytokine secretion upon co-culture with S. Typhimurium (Fig. 4b). Cytokine secretion was similarly reduced upon co-culture with AIEC and S. aureus (Fig. 4b). Anti-inflammatory cytokine secretion was also reduced (Fig. 4b). We further confirmed that IRF5 was required for cytokine secretion in BMMs upon exposure to a broad range of PRRs (Supplementary Fig. 2b). In contrast, cholera toxin-induced cytokines (Supplementary Fig 2c) and cholera toxin-induced bacterial clearance (Supplementary Fig 2d) were not regulated by IRF5 genotype. Importantly, IRF5 was required for optimal uptake and clearance of intracellular S. Typhimurium in colonic lamina propria macrophages (Fig. 4c). Moreover, IRF5−/− colonic macrophages demonstrated reduced S. Typhimurium-induced proinflammatory but increased anti-inflammatory IL10 cytokine secretion (Fig. 4d). Therefore, macrophage-intrinsic IRF5 promotes live bacterial-induced inflammatory cytokine secretion and clearance of intracellular bacteria.
BMMs and intestinal myeloid cells from IRF5−/− mice are less effective at clearance of intracellular bacteria and production of bacterial-induced inflammatory cytokines in vitro. a, b BMMs from IRF5+/+, IRF5+/− or IRF5−/− mice were: (a) left untreated or treated for 48 h with 0.1 μg/ml lipid A and then co-cultured with S. Typhimurium, AIEC or S. aureus as per Methods for ‘Intracellular bacterial clearance’. Bacterial CFU (4 replicates, representative of 3 independent experiments). b Co-cultured with S. Typhimurium, AIEC or S. aureus and supernatants were assessed for cytokines 24 h later (4 replicates). Significance for A & B is compared to IRF5+/+ macrophages for each respective treatment condition or as indicated. c, d Colonic lamina propria macrophages were co-cultured with S. Typhimurium and gentamicin was added 20 min later. c Bacterial CFU (6 replicates) assessed at the indicated times after gentamicin. d Cytokines (4 replicates) at 20 h. Mean + SEM. *p < 0.05; **p < 0.01; ***p < 0.001; †p < 1 × 10−4; ††p < 1 × 10−5
IRF5 promotes induction of ROS, RNS and autophagy pathways in macrophages
We next assessed mechanisms through which IRF5 contributes to intracellular bacterial clearance in macrophages. Reactive oxygen species (ROS) can contribute to microbial clearance.1 Furthermore, polymorphisms in genes in the NADPH complex required for ROS production are associated with increased risk for both the common form of IBD2 and early-onset IBD.3 ROS production is induced with chronic PRR ligand exposure26; to our knowledge how IRF5 regulates ROS production has not been reported. With chronic lipid A treatment ROS was induced in IRF5+/+ BMMs, whereas IRF5−/− BMMs demonstrated a less effective increase and IRF5+/− BMM ROS induction levels were intermediate (Fig. 5a). Chronic lipid A treatment of BMMs induced expression of p40phox, p47phox, p67phox, p22phox and gp91phox transcripts, whereas expression of these NADPH oxidase complex members was less effectively induced in IRF5+/− and IRF5−/− BMMs (Fig. 5b). We confirmed that ROS pathways were required for the clearance of the bacteria examined using a ROS inhibitor (Fig. 5c, Supplementary Fig. 3a).
BMMs from IRF5−/− mice demonstrate impaired PRR-induced ROS, RNS and autophagy pathways. BMMs from IRF5+/+, IRF5+/− or IRF5−/− mice were left untreated or treated with 0.1 μg/ml lipid A for 48 h and assessed for: (a) ROS production. Representative flow cytometry plot with mean fluorescence intensity (MFI) values and a summary graph of MFI (5 replicates, representative of 3 independent experiments). b, e, h Fold mRNA expression normalized to Gapdh (4 replicates, representative of 2 independent experiments). c, f, i Cells were treated for 1 h with inhibitors blocking the (c) ROS (NAC), (f) RNS (L-NAME), or (i) autophagy (3-MA) pathways and then intracellular bacterial clearance was assessed as in Fig. 4a (4 replicates). d Nitrite (4 replicates, representative of 2 independent experiments), (g) LC3II by Western blot (representative of 2 independent experiments). Mean + SEM. *p < 0.05; **p < 0.01; ***p < 0.001; †p < 1 × 10−4; ††p < 1 × 10−5
Reactive nitrogen species (RNS) can also contribute to bacterial clearance, and a combination of ROS and RNS pathways is central in maintaining homeostasis in the intestinal mucosa.27 IRF5 plays a role in NOS2 upregulation in mice following L. donovani infection,10 such that we examined if IRF5 regulates nitric oxide production and NOS2 expression, the molecule regulating its production, directly in macrophages in vitro. With chronic lipid A treatment, nitrite, a product of nitric oxide, was induced in WT BMMs, whereas IRF5−/− BMMs demonstrated a less effective increase in nitrite (Fig. 5d) and Nos2 expression (Fig. 5e). We confirmed that RNS pathways were required for optimal bacterial clearance using a RNS inhibitor (Fig. 5f, Supplementary Fig. 3b). Cholera toxin-induced ROS and RNS pathways were unimpaired in IRF5−/− BMMs, demonstrating that these pathways remained functional in these cells (Supplementary Fig 4a, b).
Autophagy is another key bacterial clearance mechanism induced with PRR stimulation28 and polymorphisms in the autophagy-associated gene ATG16L1 confer altered susceptibility to Crohn’s disease.2 A prior report examined the relationship between IRF5 and autophagy; it found that IRF5 inhibits Hepatitis C virus-induced autophagy in MH-14 and C-5B cell lines.29 In contrast, with 48 h lipid A treatment, LC3II expression (a marker of autophagy) increased in WT BMMs, whereas IRF5−/− BMMs did not as effectively induce LC3II (Fig. 5g). To address the pathways leading to autophagy induction, we examined select autophagy-associated molecules. Atg5, Atg16l1 and Irgm transcripts were induced with lipid A treatment, and this induction was impaired in IRF5+/− and IRF5−/− BMMs (Fig. 5h). We confirmed that autophagy pathways were required for optimal bacterial clearance using the autophagy inhibitor 3-methyladenine (3-MA) (Fig. 5i, Supplementary Fig. 3c). Therefore, IRF5 is required for optimal PRR induction of multiple bacterial clearance pathways and for intracellular bacterial clearance in macrophages.
IRF5 promotes TLR4-induced NFκB activation and binds to the promoters of antimicrobial pathway transcripts
To address additional mechanisms through which IRF5 regulates the identified antimicrobial pathways, we considered that it might contribute through its ability to both regulate essential PRR-induced proximal signaling pathways (at earlier times) and function as a transcription factor (at later times).
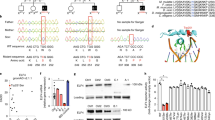
With respect to its ability to regulate PRR-induced proximal signaling pathways, IRF5 can form a complex with IRAK1 and TRAF6,4,30,31 and enhance PRR-induced NFκB signaling in human macrophages.6,7 An IRF5 requirement for PRR-induced signaling in mouse macrophages has not been reported. IRF5 phosphorylation was observed within 15 min of lipid A treatment (Fig. 6a). However, lipid A-induced NFκB activation at 15 min was reduced in IRF5−/− BMMs (Fig. 6b). Further, NFκB activation was required for optimal levels of baseline and TLR4-induced intracellular bacterial clearance and of the antimicrobial pathways we had defined to be IRF5-dependent (Fig. 6c–h). BMM viability remained intact under these conditions (Supplementary Fig 4c). Therefore, early PRR-induced signaling pathways regulated by IRF5 are, in turn, required for the induction of IRF5-dependent antimicrobial pathways.
IRF5-dependent NFκB activation regulates antimicrobial pathways in BMMs. a BMMs from IRF5+/+ mice were treated with 0.1 μg/ml lipid A for the indicated times. Phospho-IRF5 was assessed by Western blot. b BMMs from IRF5+/+, IRF5+/− or IRF5−/− mice were left untreated or treated with 0.1 μg/ml lipid A for 15 min and assessed for phospho-IκBα by Western blot. c–h BMMs were treated with 0.1 μg/ml lipid A for 48 h ± BAY 11-7082 (NFκB inhibitor). (c) Cells were co-cultured with S. Typhimurium, AIEC or S. aureus. Bacterial CFU (4 replicates). d ROS production (3 replicates). f Nitrite (3 replicates). e, g, h Fold mRNA expression normalized to Gapdh (4 replicates). i BMMs from IRF5+/+ mice were treated with 0.1 μg/ml lipid A for the indicated times. IRF5 binding to antimicrobial pathway gene promoters was assessed by ChIP (3 replicates). Mean + SEM. *p < 0.05; **p < 0.01; ***p < 0.001
We next considered the role of IRF5 as a transcription factor in the antimicrobial pathways identified. Consistent with its role as a transcription factor, IRF5 translocates to the nucleus 2–6 h after activation,32 and we had previously found that this translocation peaks in human macrophages 4 h after PRR stimulation.7 IRF5 binding to the promoters of the upregulated ROS, RNS and autophagy-associated genes was enhanced 4 h after lipid A treatment of BMMs, and further enhanced 4 h after treatment of 48 h lipid A pre-treated BMMs (Fig. 6i).
Finally, we assessed the role of IRF5 in broader TLR4-induced transcriptional pathways, and the degree to which these might be shared with NFκB, given the IRF5-dependent role in NFκB activation. A previous study conducted TLR4-induced transcriptional analysis in IRF5+/+, IRF5+/− and IRF5−/− BMMs.33 We therefore selected the top 20 TLR4-induced transcripts and an additional 36 transcripts highlighted in this prior study33 (accession # E-MTAB-2032) for further analysis. We observed an IRF5-dependency for a broad range of TLR4-induced transcripts as per the prior study (Supplementary Fig 5a–c). Induction of the majority of these transcripts was also dependent on NFκB (Supplementary Fig 5a), such that IRF5 regulation of these transcriptional pathways might be through its role in regulating proximal signaling or its role as a transcription factor. However, in some cases NFκB regulation of the transcripts was reciprocal to that of IRF5 (Supplementary Fig 5b). Moreover, there were also transcripts that were IRF5-dependent and NFκB-independent (Supplementary Fig 5c) and those that were IRF5-independent, but NFκB-dependent (Supplementary Fig 5d). Taken together, these studies demonstrate that IRF5 regulates proximal, early signaling pathways required for antimicrobial clearance mechanisms, and at later times binds to the promoters of genes regulating these antimicrobial mechanisms.
IRF5 is required for optimal induction of anti-microbial pathways and intracellular bacterial clearance in LPS, IFNγ-differentiated (M1) macrophages
Macrophages can be polarized to a range of subtypes depending on environmental conditions, and one such subtype is the proinflammatory LPS, IFNγ-conditioned macrophage subset (also referred to as M1 macrophages). LPS, IFNγ-conditioned macrophages increase during infection with intestinal pathogens, including S. Typhimurium, and are particularly effective at clearing bacteria.20,34 We7 and others5 have found that IRF5 is required for optimal LPS, IFNγ-conditioned macrophage differentiation. Therefore, we assessed if IRF5 was required for optimal bacterial clearance and induction of antimicrobial pathways after culture in LPS, IFNγ-differentiating conditions. LPS, IFNγ-conditioned macrophages demonstrated appropriate induction of various M1 markers35 (Supplementary Fig. 6a) and of IRF5 expression (Supplementary Fig. 6b), as well as enhanced bacterial clearance (Supplementary Fig. 6c) and bacterial clearance pathways relative to non-polarized macrophages (Supplementary Fig. 6d–i). Moreover, we confirmed the requirement for IRF5 in the differentiation of LPS, IFNγ-conditioned macrophages (Supplementary Fig. 6a). In LPS, IFNγ-conditioned macrophages IRF5 was required for optimal intracellular bacterial clearance (Supplementary Fig. 6c) and for the specific antimicrobial pathways (Supplementary Fig. 6d–i) we had defined above. Therefore, in addition to the requirement for IRF5 in LPS, IFNγ-mediated macrophage differentiation, it is required for the optimal upregulation of antimicrobial pathways and bacterial clearance observed in these cells.
IRF5 promotes intracellular bacterial clearance and induction of antimicrobial pathways in additional myeloid cell subsets
As multiple myeloid cell subsets can participate in microbial clearance, we assessed if IRF5 was required for microbial clearance in neutrophils and in dendritic cells. IRF5 was required for optimal intracellular bacterial clearance and TLR4-induced ROS and RNS pathways in neutrophils (Supplementary Fig. 7a–c) and BMDCs (Supplementary Fig. 7d–f). Therefore, IRF5 contributes to microbial clearance pathways in multiple different myeloid cell subsets, which likely function in a combined manner to mediate bacterial clearance in vivo.
Myeloid cell-intrinsic IRF5 is required for optimal clearance of S. Typhimurium in vivo
Given the importance of IRF5 in regulating bacterial clearance in myeloid-derived cells in vitro, we asked if myeloid cell-intrinsic IRF5 contributes to the clearance of S. Typhimurium in vivo. To address this, we crossed IRF5fl/fl mice with LysM-Cre mice (IRF5Δmye) mice. We confirmed reduced IRF5 expression in BMMs from IRF5Δmye mice with and without lipid A treatment (Supplementary Fig. 8a). IRF5 expression in skin fibroblasts from IRF5Δmye mice was not decreased (Supplementary Fig. 8b), thereby demonstrating expected cell subset selectivity in IRF5 deletion in these mice. We further confirmed that BMMs from IRF5Δmye mice demonstrated decreased cytokine secretion to a range of PRR ligands (Supplementary Fig. 8c). BMMs from IRF5Δmye mice also demonstrated decreased efficacy in intracellular bacterial clearance, along with reduced induction of ROS, RNS and autophagy bacterial clearance pathways (Supplementary Fig. 9). Similar to IRF5−/− colonic myeloid cells (Fig. 4c–d), IRF5Δmye colonic myeloid cells demonstrated reduced S. Typhimurium entry and clearance, as well as reduced S. Typhimurium-induced proinflammatory and increased anti-inflammatory cytokines (Supplementary Fig. 10). Finally, IRF5Δmye mice demonstrated decreased LPS-induced serum cytokines compared to IRF5fl/fl controls (Supplementary Fig. 11). These studies verify that IRF5Δmye mice demonstrate effective IRF5 deletion in myeloid cells and functional consequences of this deletion in vitro and in vivo.
Similar to IRF5−/− mice, IRF5Δmye mice orally infected with S. Typhimurium demonstrated an increased burden of S. Typhimurium in colon and feces, as well as systemically compared to IRF5fl/fl controls (Fig. 7a). Also similar to IRF5−/− mice, S. Typhimurium-infected IRF5Δmye mice demonstrated decreased mucosal MPO activity (Fig. 7b) and decreased Th1-associated (IL12, IFNγ), Th17-associated (IL17) and additional proinflammatory (TNF) cytokines in both colon (Fig. 7c) and serum (Fig. 7d), whereas Th2-associated (IL13) and anti-inflammatory (IL10) cytokines were increased relative to IRF5fl/fl controls (Fig. 7c, d). Therefore, myeloid cell-intrinsic IRF5 is required for the induction of inflammatory responses and optimal regulation of oral S. Typhimurium infection in mice in vivo.
IRF5Δmye mice demonstrate less effective clearance of oral S. Typhimurium infection. IRF5WT or IRF5Δmye mice (n = 7–11; 2.5–3 months of age) were orally inoculated with S. Typhimurium and assessed 5 days later. a S. Typhimurium CFU. b Colon myeloperoxidase. c Colon cytokines. d Serum cytokines. Representative of 3 independent experiments. Mean + SEM. *p < 0.05; **p < 0.01; ***p < 0.001; †p < 1 × 10−4; ††p < 1 × 10−5
The burden of S. Typhimurium measured in intestinal tissues reflects a combination of mechanisms, including dissemination (e.g. intestinal barrier permeability), bacterial uptake, and intracellular bacterial clearance (e.g., intracellular bacterial growth, intracellular bacterial killing). We therefore assessed if enhanced intestinal barrier permeability under these infected conditions might allow for increased S. Typhimurium dissemination. FITC-dextran in the serum was equivalent in S. Typhimurium-infected IRF5fl/fl and IRF5Δmye mice (Fig. 8a). Interestingly, the burden of intestinal luminal commensal bacteria in intestinal lymphoid structures during S. Typhimurium infection was reduced relative to that in uninfected mice, and this reduction was greater in IRF5Δmye mice where the S. Typhimurium burden is greater (Fig. 8b). This is consistent with prior reports which highlighted more effective competition by enteric pathogens for the environmental niche.36 IRF5Δmye mice demonstrated increased fecal S. Typhimurium compared to IRF5fl/fl mice as infection progressed (Fig. 8c). Finally, we examined the levels of S. Typhimurium in IRF5Δmye macrophages in vitro over time so as to integrate the consequences of bacterial uptake, bacterial growth and bacterial clearance in these cells. As expected based on Fig. 3a, the levels of S. Typhimurium in IRF5Δmye macrophages were lower 20 min after co-culture due to the reduced ability of the cells to take up bacteria (Fig. 8d). Over the next 2 h the rate of intracellular growth was similar in IRF5fl/fl and IRF5Δmye macrophages (Fig. 8d). However, after 2 h IRF5fl/fl macrophages began to reduce intracellular bacteria, whereas IRF5Δmye macrophages were unable to do so as effectively, such that the intracellular bacterial load in IRF5Δmye macrophages surpassed that in IRF5fl/fl macrophages at these later times (Fig. 8d). Therefore, the increased bacterial burden in tissues of S. Typhimurium-infected IRF5Δmye mice is likely a combination of reduced cellular bacterial uptake leading to increased bacterial exposure and reduced intracellular bacterial clearance mechanisms.
IRF5Δmye mice demonstrate less effective clearance of oral S. Typhimurium infection. a–c IRF5WT or IRF5Δmye mice (2–2.5 months of age) were orally inoculated with S. Typhimurium and assessed 5 days later. a Intestinal permeability by FITC-dextran (n = 5). b Commensal bacterial DNA in Peyer’s patches and MLN (n = 5). c S. Typhimurium CFU in feces (n = 5). d BMMs from IRF5WT or IRF5Δmye mice were co-cultured with S. Typhimurium-GFP, gentamycin was added 20 min later, and intracellular S. Typhimurium was then assessed over the next 4 h by flow cytometry (4 replicates; representative of 3 independent experiments). Mean + SEM. Significance is compared to WT mice for each respective condition. *p < 0.05; **p < 0.01
IRF5 contributes to the inflammation observed during colonic injury with influx of resident colonic bacteria
Given the importance of IRF5 in regulating inflammatory responses to the invasive intestinal pathogen S. Typhimurium, we assessed if IRF5 also regulates inflammatory responses to high levels of resident intestinal luminal bacteria in the context of acute intestinal injury. While IRF5-deficiency reduced inflammation in chronic colitis (Fig. 1), the role of a given pathway may be very different during acute colitis or with distinct triggers. If the ability to clear acutely high levels of resident intestinal bacteria is impaired, the increased bacterial tissue burden may result in compensatory immune responses leading to increased inflammation. Therefore, the balance in the inflammatory response to and clearance of translocated resident luminal bacteria is critical in determining the outcome of intestinal injury. We therefore selected an intestinal injury model that disrupts the intestinal epithelial barrier and results in increased influx of intestinal resident bacteria, the dextran sodium sulfate (DSS) model of intestinal injury. IRF5−/− mice administered DSS in drinking water demonstrated less weight loss, histologic injury, mucosal myeloperoxidase and fecal lipocalin compared to IRF5+/+ mice (Fig. 9a–e). IRF5+/− mice generally demonstrated an intermediate phenotype in these measures (Fig. 9a–e). Consistent with reduced histological intestinal inflammation in IRF5−/− mice, colonic (Fig. 9f) and serum (Supplementary Fig. 12) proinflammatory (TNF), Th1-associated (IL12, IFNγ) and Th17-associated (IL17) cytokines were decreased compared to IRF5+/+ mice administered DSS. In contrast, the Th2-associated cytokine IL13 was increased, and IL10 was not different compared to IRF5+/+ mice (Fig. 9f, Supplementary Fig. 12).
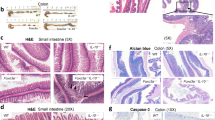
IRF5−/− mice develop less severe inflammation during DSS-induced injury. IRF5+/+, IRF5+/− and IRF5−/− mice were given 2.5% DSS in the drinking water. a Body weights. b Representative H&E of colon sections. Low magnification (LM) with scale bar = 1 mm. High magnification (HM) with scale bar = 100 μm. Dotted line on images represents ulcerated mucosa. c Histologic scores (n = 3–10). d Colon myeloperoxidase. e Fecal lipocalin. f Colon cytokines. g Colonic lamina propria was assessed for percentages of the indicated myeloid cell-derived populations by flow cytometry (n = 5–6 mice). h Bacterial CFU in MLN and spleen (n = 4–6; representative of 2 independent experiments). i Intestinal permeability as assessed by FITC-dextran (n = 6; representative of 2 independent experiments). Data are shown as mean + SEM. a, d–h: DSS for 8 days and then water for an additional 2 days (2-3 months of age). b, c, i: DSS for 8 days. Representative of 2 independent experiments for (d–f) or as indicated. DW, drinking water. *p < 0.05; **p < 0.01; ***p < 0.001
We next assessed how select myeloid cells in the colonic lamina propria were regulated. We focused on Gr1+ neutrophils and the proinflammatory P1 and P2 and anti-inflammatory P3/4 macrophage subsets as previously described.37 These lamina propria myeloid-derived cell subsets were not different between IRF5+/+, IRF5+/− and IRF5−/− mice at baseline (data not shown). Following oral DSS, the percent of Gr1+ neutrophils and proinflammatory macrophages (P1, P2) were reduced, whereas the anti-inflammatory P3/4 macrophage subset was increased in the lamina propria of IRF5−/− mice relative to IRF5+/+ mice (Supplementary Fig. 13a, Fig. 9g), consistent with the reduced inflammatory responses observed (Fig. 9a–f).
IRF5−/− mice demonstrated a reduced inflammatory immune response along with a decreased ability to clear the invasive pathogen S. Typhimurium (Fig. 2). However, resident luminal bacterial burdens can be more complex given the interplay between proinflammatory cytokines, the epithelial barrier, and bacterial translocation. In particular, inflammatory cytokines such as IFNγ and TNF can modulate epithelial tight junctions to increase intestinal permeability.38 Administration of DSS to WT mice increases intestinal permeability, which we confirmed by FITC-dextran (Supplementary Fig. 13b), and this was accompanied by increased bacterial translocation to mesenteric lymph nodes (MLNs), as well as systemically to spleen (Supplementary Fig. 13c). IRF5+/+, IRF5+/− and IRF5−/− mice did not differ in intestinal permeability (Supplementary Fig. 13d) or bacterial CFU in MLN or spleen (data not shown) under baseline conditions. However, with DSS-induced injury, bacterial levels in MLNs and spleen were reduced in IRF5−/− mice relative to IRF5+/+ mice (Fig. 9h). We therefore assessed if the reduced colonic proinflammatory cytokines observed under these conditions in IRF5−/− mice was associated with a decreased epithelial barrier defect. This could then lead to decreased bacterial translocation, and to the reduced bacterial burden observed in IRF5−/− mice. We found this to be the case (Fig. 9i). Therefore, despite the impaired bacterial clearance in IRF5−/− macrophages in vitro and in IRF5−/− mice in vivo upon infection with the invasive pathogen S. Typhimurium, the resident bacterial burden upon DSS exposure was actually reduced in the context of the reduced proinflammatory cytokines. As such, IRF5−/− mice demonstrate less DSS-induced intestinal injury and inflammation with decreased colon inflammatory cytokines, and this is associated with a reduced barrier defect, and in turn, a lower bacterial burden in intestinal lymphoid organs and systemically.
Myeloid cell-intrinsic IRF5 contributes to inflammation during colonic injury and to the influx of resident colonic bacteria
We next assessed if myeloid cell-intrinsic IRF5 contributes to the inflammation observed with DSS-induced injury. Upon administration of DSS to IRF5Δmye mice, outcomes were similar to that observed in IRF5−/− mice, with decreased intestinal inflammation and inflammatory cytokines (e.g. TNF, Th1 and Th17 cytokines), increased anti-inflammatory colonic macrophages, reduced intestinal permeability, and reduced levels of intestinal bacteria in MLN and spleen (Fig. 10, Supplementary Fig. 14). Therefore, upon intestinal injury leading to increased intestinal luminal resident bacterial exposure, myeloid cell-intrinsic IRF5 contributes to the increased inflammation, barrier defect, and intestinal luminal bacterial burden observed.
IRF5Δmye mice develop less severe inflammation with DSS-induced injury. IRF5WT or IRF5Δmye mice were given 2.5% DSS in the drinking water. a Body weights. b Representative H&E of colon sections. Scale bar = 150 μm. c Histologic scores. d Colon myeloperoxidase. e Fecal lipocalin. f Colon cytokines (a–f n = 3–10; representative of 2 independent experiments). g Colonic lamina propria was assessed for percentages of the indicated myeloid cell-derived populations by flow cytometry (n = 8). h Bacteria in MLN and spleen (n = 5–7; representative of 2 independent experiments). i Intestinal permeability was assessed by FITC-dextran (n = 4–5; representative of 2 independent experiments). Mean + SEM. a, d–h DSS for 8 days and then water for an additional 2 days (2.5–3 months of age). b, c, i DSS for 8 days. *p < 0.05; **p < 0.01. DW drinking water
Discussion
This study defines contributions for the IBD-associated gene IRF5 to intestinal inflammation in mice in vivo. It finds that while IRF5 is required for optimal clearance of enteric invasive pathogens, it also contributes to the cytokines and inflammation observed during both acute and chronic intestinal inflammation in the context of resident intestinal microbes. Furthermore, myeloid cell-intrinsic IRF5 is critical in mediating this balance. Heterozygote IRF5 mice generally demonstrate an intermediate phenotype. These results are consistent with the increased risk for ulcerative colitis observed in carriers with genetic variants leading to increased IRF5 expression.2,7,8,14,39 However, these studies raise the possibility that while IRF5 genetic variants resulting in increased IRF5 expression confer a higher risk for UC, they may lead to improved clearance of invasive enteric pathogens; such patterns have been recently found in studies in vitro with these carrier cells.40 Conversely, similar to carriers of IRF5 genetic variants resulting in decreased IRF5 expression and decreased risk for UC, our studies highlight that reducing the IRF5 pathway may be a beneficial therapy in IBD, but that such individuals may be at higher risk for enteric infections.
We found that in the context of exposure to the invasive intestinal pathogen S. Typhimurium, the absence of IRF5 leads to decreased inflammatory (TNF), Th1 (IL12, IFNγ) and Th17 (IL17) cytokines, but also increased Th2 (IL13) and anti-inflammatory (IL10) cytokines in intestinal tissues (Fig. 2). Interestingly, whereas IRF5−/− BMMs demonstrated reduced IL10 secretion to live bacteria, IRF5−/− intestinal macrophages demonstrated increased IL10 secretion to live bacteria, thereby highlighting distinct regulation in colonic macrophages. Prior reports examining IRF5−/− mice found reciprocal cytokine regulation (decreased Th1 and Th17, and increased Th2 cytokines) with L. donovani10 and in models of lupus and asthma.17,18,19 In Th1- or Th17-mediated disease models this can result in decreased disease severity.17,18,41,42 In contrast, in the Th2-mediated models of asthma or Schistosoma japanicum this can lead to increased disease severity.19,43 That we now observe a similar regulation pattern of these cytokines in vivo with myeloid cell-intrinsic IRF5 deficient mice highlights the importance of IRF5 in myeloid cells as a mechanism for increased Th1- and Th17-, and decreased Th2-associated cytokines. Th2 pathway cytokines can contribute to intestinal inflammation.44 Therefore, the increased Th2 cytokines observed in intestinal tissues of IRF5−/− mice during both acute and chronic experimental colitis models (Figs. 1 and 9) may be contributing to some of the histological inflammation in the IRF5-/- mice, given that the inflammation is not necessarily reduced to the degree that might be expected relative to the reduction in Th1 and Th17 cytokines.
In this study we demonstrate that while the IBD-associated gene IRF5, in particular, myeloid cell-intrinsic IRF5, contributes to a robust induction of PRR-initiated proinflammatory and anti-microbial pathways which leads to more effective clearance of intestinal pathogens, it comes at the expense of disruption of the intestinal epithelial barrier, ongoing exposure to resident luminal microbes, and amplification of intestinal inflammation. Additional mechanisms through which IRF5 may be contributing to intestinal inflammation have yet to be determined, with considerations including roles in additional cell subsets and dysbiosis. The differential regulation of inflammatory versus infectious disease susceptibility and/or severity has been observed for SOCS1/LITAF/RMI2, IL12B, and FOXO3 region polymorphisms associated with IBD.2,45 Given the ability of IRF5 to regulate a broad range of inflammatory pathways across multiple immune-mediated diseases, it has promise as a nodal therapeutic target. As such, recent pre-clinical studies have demonstrated that delivery systems targeting IRF5 during inflammatory processes in vivo can improve outcomes. For example, nanoparticle-delivered IRF5 siRNA during acute myocardial infarction improved infarct healing46 and during spinal cord injury reduced adverse outcomes and promoted functional recovery.47 However, our studies indicate that when considering targeting of this pathway for immune-mediated diseases, including IBD, caution may be needed with respect to the threshold at which one attenuates the IRF5 pathway given the potential risk for microbial infections.
Materials and Methods
Mice
IRF5−/− mice (C57BL/6) were generously provided by Dr. Betsy J. Barnes and were crossed with C57BL/6 mice to generate littermate controls. IRF5−/− mice were crossed into RAG-1−/− mice. IRF5LoxP/LoxP mice (stock #017311, Jackson Laboratory) were crossed to LysM-Cre mice (stock #004781; Jackson Laboratory). Mice were maintained in a specific pathogen-free facility and cohoused, littermate mice were used between 2–5 months of age, with studies confirmed in both female and male mice. Experiments were performed according to the Yale University Institutional Animal Care and Use Committee and National Institutes of Health guidelines for animal use.
Abs and staining reagents
Antibodies to the following were used on a FACS Calibur or LSR II (BD Biosciences, San Jose, CA): fluorophore-labeled CD45, CD11c, Ly6C (Biolegend), CD19, F4/80, CD11b, Ly6G (eBiosciences), MHCII (BD Biosciences). The following were assessed by ELISA: IFNγ (BD Biosciences), IL6, IL12p40, IL13, IL17 (eBiosciences), IL10, TNF (Biolegend), MIP2 (Peprotech).
CD45RBhi CD4+ T cell transfer colitis
Spleen-derived CD45RBhi CD4+ T cells were sorted utilizing the FACSAria (BD Biosciences). Colitis was induced through the transfer of CD45RBhi CD4+ T cells (5 × 105) into IRF5+/+, IRF5+/− or IRF5−/− RAG-1−/− mice. Mice were monitored for weight loss and sacrificed after 6-7 weeks. H&E stained colon specimens were examined by a gastrointestinal surgical pathologist (JRT) blinded to the experimental treatment and activity was scored using a five-tier scale with a score of 0-2 for each measure (mucin depletion, proliferative response, surface damage, neutrophil infiltration, mononuclear cell infiltration) for a total possible score of 10.
Salmonella Typhimurium infection in vivo
Mice were orally inoculated with Streptomycin (20 mg/mouse) (VWR International, Bridgeport, NJ) and 24 h later orally inoculated with 1 × 106 colony forming units (CFU) of Salmonella enterica serovar Typhimurium (S. Typhimurium) grown under anaerobic conditions.
Dextran sodium sulfate colitis
Dextran Sodium Sulfate, MW ca 40,000 (Alfa Aesar, Tewksbury, MA) 2.5% (w/v) was added to the drinking water of mice with ad libitum water access. Upon harvest, some tissues were plated on brain heart infusion agar for bacterial colony counts. H&E stained colon specimens were examined by a gastrointestinal pathologist blinded to the experimental treatment (JRT), and activity was scored using a three-tiered scale with a 0–4 score for each measure (damage, regeneration and ulcer repair) for a total possible score of 12.
Generation and culture of BMMs
Bone marrow cell suspensions were cultured in complete DMEM containing 10% L929-conditioned medium. Cells were used at 6 to 8 days. In some cases BMMs were first treated with lipid A (Peptides International) for 48 h. In other cases BMMs were cultured with S. Typhimurium, Adherent Invasive Escherichia coli (AIEC) strain LF82 (generously from Dr. Emiko Mizoguchi), or Staphylococcus aureus at 5:1 MOI for 24 h, with 30 μg/ml gentamycin added after the first 20 min.
mRNA or DNA expression
RNA from cells was isolated (Trizol, Life Technologies, Carlsbad, CA), reverse transcribed, and quantitative PCR was performed as described.7 Each sample was run in duplicate and normalized to Gapdh. Bacterial DNA in tissues was assessed by quantitative PCR and normalized to Ptger. Primer sequences are available upon request.
Protein analysis
For protein analysis, tissue was suspended in a Triton lysis buffer and homogenized and ELISA for cytokines was performed. BMMs were lysed and protein detected by Western blot as per7 with antibodies to LC3II, phospho-PDK1, PDK1, phospho-IκBα, IRF5 (Cell Signaling Technology), IκBα (Santa Cruz Technology), or phospho-IRF5 (Invitrogen). Anti-GAPDH antibody (EMD Millipore, 6C5, dilution) was used as a loading control.
Bacterial entry/phagocytosis
FITC-labeled S. aureus or FITC-labeled E. coli bioparticles (1.5 × 106) (Molecular Probes), or 5:1 MOI S. Typhimurium-GFP (generously provided by Jorge E. Galan) were co-cultured with BMMs for 20 min, cell surface fluorescence was quenched with 0.25 mg/ml trypan blue, and cells were analyzed by flow cytometry. In some cases inhibitors were added: 3 μM GSK 2334470 (Tocris) (PDK1 inhibitor) or 100 nM wortmannin (MP Biomedicals).
Intracellular bacterial clearance
BMMs were infected with S. Typhimurium, AIEC, S. aureus at 5:1 MOI for 20 min, washed three times with PBS, incubated in HBSS medium containing 30 μg/ml gentamicin and then cultured for an additional 4 h. Cells were washed, lysed with 1% Triton X-100 (Sigma) and plated on MacConkey or LB agar. In some cases 20 mM N-acetylcysteine (NAC; Sigma-Aldrich), 10 mM Nω-nitro-L-arginine methyl ester hydrochloride (L-NAME), 10 mM 3-methyl adenine (3-MA) (Acros Organics) or 20 μM BAY 11-7082 (Calbiochem) were added 1 h prior to bacterial infection.
Myeloperoxidase assay
Colon was homogenized in hexadecyltrimethylammonium bromide (Sigma-Aldrich) buffer. MPO was assayed using o-dianisidine dihydrochloride (Sigma-Aldrich) and H2O2, and the change in optical density was measured at 450 nm.
FITC-dextran permeability assay
Mice were orally gavaged with FITC-dextran (40 mg/100 g body weight) 4 h before sacrifice. Serum concentration of FITC-dextran was measured by fluorometer at 488 nm.
Lipocalin
Frozen fecal samples were reconstituted in PBS/0.1% Tween 20, vortexed and supernatants were assessed for lipocalin concentrations by ELISA (R&D).
Intestinal lamina propria cell isolation
Colonic lamina propria cells were isolated as previously described.7 CD11b beads (Miltenyi Biotech) were used to isolate intestinal macrophages.
Intracellular ROS measurement
ROS was measured by flow cytometry using 1 μM of 2′, 7′-dichlorodihydrofluorescein diacetate (H2DCFDA) (Invitrogen).
Nitric oxide measurement
Griess reagent [0.1% N-(1-napthyl)ethylenediamine dihydrochloride + 1% sulfanilic acid in 5% phosphoric acid] (Sigma-Aldrich) was incubated with samples and absorbance measured at 548 nm.
Chromatin immunoprecipitation (ChIP)
ChIP was performed according to a modified protocol48 with anti-IRF5 (Abcam, Cambridge, MA) and an isotype control.
Statistical analyses
Statistical comparisons were assessed using a two-tailed Student’s t test. Bonferroni-Holm correction was used for multiple comparisons. Values of p < 0.05 were considered significant.
Change history
10 June 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Abraham, C. & Medzhitov, R. Interactions between the host innate immune system and microbes in inflammatory bowel disease. Gastroenterology 140, 1729–1737 (2011).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Dhillon, S. S. et al. Variants in nicotinamide adenine dinucleotide phosphate oxidase complex components determine susceptibility to very early onset inflammatory bowel disease. Gastroenterology 147, 680–689 e682 (2014).
Takaoka, A. et al. Integral role of IRF-5 in the gene induction programme activated by Toll-like receptors. Nature 434, 243–249 (2005).
Krausgruber, T. et al. IRF5 promotes inflammatory macrophage polarization and TH1-TH17 responses. Nat. Immunol. 12, 231–238 (2011).
Hedl, M. & Abraham, C. IRF5 risk polymorphisms contribute to inter-individual variance in pattern recognition receptor-mediated cytokine secretion in human monocyte-derived cells. J. Immunol. 188, 5348–5356 (2012).
Hedl, M., Yan, J. & Abraham, C. IRF5 and IRF5 disease-risk variants increase glycolysis and human m1 macrophage polarization by regulating proximal signaling and Akt2 activation. Cell Rep. 16, 2442–2455 (2016).
Graham, R. R. et al. A common haplotype of interferon regulatory factor 5 (IRF5) regulates splicing and expression and is associated with increased risk of systemic lupus erythematosus. Nat. Genet 38, 550–555 (2006).
Eames, H. L., Corbin, A. L. & Udalova, I. A. Interferon regulatory factor 5 in human autoimmunity and murine models of autoimmune disease. Transl. Res 167, 167–182 (2016).
Paun, A., Bankoti, R., Joshi, T., Pitha, P. M. & Stager, S. Critical role of IRF-5 in the development of T helper 1 responses to Leishmania donovani infection. PLoS Pathog. 7, e1001246 (2011).
Pandey, A. K. et al. NOD2, RIP2 and IRF5 play a critical role in the type I interferon response to Mycobacterium tuberculosis. PLoS Pathog. 5, e1000500 (2009).
Gratz, N. et al. Type I interferon production induced by Streptococcus pyogenes-derived nucleic acids is required for host protection. PLoS Pathog. 7, e1001345 (2011).
Castiglia, V. et al. Type I interferon signaling prevents IL-1beta-driven lethal systemic hyperinflammation during invasive bacterial infection of soft tissue. Cell Host Microbe 19, 375–387 (2016).
Dideberg, V. et al. An insertion-deletion polymorphism in the interferon regulatory Factor 5 (IRF5) gene confers risk of inflammatory bowel diseases. Hum. Mol. Genet 16, 3008–3016 (2007).
Hue, S. et al. Interleukin-23 drives innate and T cell-mediated intestinal inflammation. J. Exp. Med 203, 2473–2483 (2006).
Chassaing, B. et al. Fecal lipocalin 2, a sensitive and broadly dynamic non-invasive biomarker for intestinal inflammation. PLoS ONE 7, e44328 (2012).
Feng, D. et al. Irf5-deficient mice are protected from pristane-induced lupus via increased Th2 cytokines and altered IgG class switching. Eur. J. Immunol. 42, 1477–1487 (2012).
Xu, Y. et al. Pleiotropic IFN-dependent and -independent effects of IRF5 on the pathogenesis of experimental lupus. J. Immunol. 188, 4113–4121 (2012).
Byrne, A. J. et al. A critical role for IRF5 in regulating allergic airway inflammation. Mucosal Immunol. 10, 716–726 (2016).
Patel, S. & McCormick, B. A. Mucosal inflammatory response to Salmonella typhimurium Infection. Front Immunol. 5, 311 (2014).
Lahiri, A. & Abraham, C. Activation of pattern recognition receptors up-regulates metallothioneins, thereby increasing intracellular accumulation of zinc, autophagy, and bacterial clearance by macrophages. Gastroenterology 147, 835–846 (2014).
Foster, S. L., Hargreaves, D. C. & Medzhitov, R. Gene-specific control of inflammation by TLR-induced chromatin modifications. Nature 447, 972–978 (2007).
West, A. P. et al. TLR signalling augments macrophage bactericidal activity through mitochondrial ROS. Nature 472, 476–480 (2011).
Freeman, S. A. & Grinstein, S. Phagocytosis: receptors, signal integration, and the cytoskeleton. Immunol. Rev. 262, 193–215 (2014).
Darfeuille-Michaud, A. et al. High prevalence of adherent-invasive Escherichia coli associated with ileal mucosa in Crohn’s disease. Gastroenterology 127, 412–421 (2004).
Lahiri, A., Hedl, M., Yan, J. & Abraham, C. Human LACC1 increases innate receptor-induced responses and a LACC1 disease-risk variant modulates these outcomes. Nat. Commun. 8, 15614 (2017).
Shiloh, M. U. et al. Phenotype of mice and macrophages deficient in both phagocyte oxidase and inducible nitric oxide synthase. Immunity 10, 29–38 (1999).
Sanjuan, M. A. et al. Toll-like receptor signalling in macrophages links the autophagy pathway to phagocytosis. Nature 450, 1253–1257 (2007).
Cevik, O. et al. Interferon regulatory factor 5 (IRF5) suppresses hepatitis C virus (HCV) replication and HCV-associated hepatocellular carcinoma. J. Biol. Chem. 292, 21676–21689 (2017).
Balkhi, M. Y., Fitzgerald, K. A. & Pitha, P. M. Functional regulation of MyD88-activated interferon regulatory factor 5 by K63-linked polyubiquitination. Mol. Cell Biol. 28, 7296–7308 (2008).
Inoue, M. et al. T cells down-regulate macrophage TNF production by IRAK1-mediated IL-10 expression and control innate hyperinflammation. Proc. Natl Acad. Sci. USA 111, 5295–5300 (2014).
Barnes, B. J., Kellum, M. J., Field, A. E. & Pitha, P. M. Multiple regulatory domains of IRF-5 control activation, cellular localization, and induction of chemokines that mediate recruitment of T lymphocytes. Mol. Cell Biol. 22, 5721–5740 (2002).
Saliba, D. G. et al. IRF5:RelA interaction targets inflammatory genes in macrophages. Cell Rep. 8, 1308–1317 (2014).
Wynn, T. A., Chawla, A. & Pollard, J. W. Macrophage biology in development, homeostasis and disease. Nature 496, 445–455 (2013).
Murray, P. J. Macrophage Polarization. Annu Rev. Physiol. 79, 541–566 (2017).
Lupp, C. et al. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe 2, 119–129 (2007).
Bain, C. C. et al. Resident and pro-inflammatory macrophages in the colon represent alternative context-dependent fates of the same Ly6Chi monocyte precursors. Mucosal Immunol. 6, 498–510 (2013).
Wang, F. et al. IFN-gamma-induced TNFR2 expression is required for TNF-dependent intestinal epithelial barrier dysfunction. Gastroenterology 131, 1153–1163 (2006).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet 47, 979–986 (2015).
Hedl, M., Yan, J., Witt, H. & Abraham, C. IRF5 is required for bacterial clearance in human m1-polarized macrophages, and irf5 immune-mediated disease risk variants modulate this outcome. J. Immunol. 202, 920–930 (2019).
Yang, L., Feng, D., Bi, X., Stone, R. C. & Barnes, B. J. Monocytes from Irf5-/- mice have an intrinsic defect in their response to pristane-induced lupus. J. Immunol. 189, 3741–3750 (2012).
Weiss, M. et al. IRF5 controls both acute and chronic inflammation. Proc. Natl Acad. Sci. USA 112, 11001–11006 (2015).
Xu, Z. et al. Innate scavenger receptor-A regulates adaptive T helper cell responses to pathogen infection. Nat. Commun. 8, 16035 (2017).
Bamias, G. & Cominelli, F. Role of type 2 immunity in intestinal inflammation. Curr. Opin. Gastroenterol. 31, 471–476 (2015).
Lee, J. C. et al. Human SNP links differential outcomes in inflammatory and infectious disease to a FOXO3-regulated pathway. Cell 155, 57–69 (2013).
Courties, G. et al. In vivo silencing of the transcription factor IRF5 reprograms the macrophage phenotype and improves infarct healing. J. Am. Coll. Cardiol. 63, 1556–1566 (2014).
Li, J., Liu, Y., Xu, H. & Fu, Q. Nanoparticle-delivered IRF5 siRNA facilitates M1 to M2 transition, reduces demyelination and neurofilament loss, and promotes functional recovery after spinal cord injury in mice. Inflammation 39, 1704–1717 (2016).
Zheng, S. & Abraham, C. NF-kappaB1 inhibits NOD2-induced cytokine secretion through ATF3-dependent mechanisms. Mol. Cell Biol. 33, 4857–4871 (2013).
Acknowledgements
This work was supported by R01AI120369, R01DK099097, and DKP30-34989, and the Crohn’s and Colitis Foundation. We thank Betsy Barnes for generously providing mice, Emiko Mizoguchi for reagents, and Matija Hedl for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
Project supervision and drafting of manuscript (C.A.); acquisition of data (S.P., J.Y., C.A.); data analysis (S.P., J.Y., J.R.T., C.A.); critical revision of manuscript (C.A., J.R.T.).
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Pandey, S.P., Yan, J., Turner, J.R. et al. Reducing IRF5 expression attenuates colitis in mice, but impairs the clearance of intestinal pathogens. Mucosal Immunol 12, 874–887 (2019). https://doi.org/10.1038/s41385-019-0165-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41385-019-0165-1
This article is cited by
-
Defactinib inhibits PYK2 phosphorylation of IRF5 and reduces intestinal inflammation
Nature Communications (2021)