Abstract
Human papillomaviruses (HPVs) are the causative agents of carcinoma of the uterine cervix. A number of HPV genotypes have been associated with cervical cancer and almost all tumors associated with HPV show strong p16 expression. However, there is little information on the possible impact of the HPV genotype and p16 immunostaining on the clinicopathological features or their prognostic value in cervical carcinoma. We evaluated a series of 194 patients with HPV-positive cervical cancers treated at our institution, focusing on the clinicopathological features and the relationship of the HPV genotypes and p16 immunostaining with the prognosis. A single HPV type was identified in 149 (77%) tumors, multiple HPV infection was detected in 30 cases (15%), and undetermined HPV type/s were identified in 15 (8%) carcinomas. HPV 16 and/or 18 were detected in 156 (80%) tumors. p16 was positive in 186 (96%) carcinomas, but eight tumors (4%) were negative for p16 (seven squamous cell carcinomas, one adenocarcinoma); 5/8 caused by HPV 16 and/or 18. Patients with HPV 16 and/or 18 were younger (49 ± 15 vs. 57 ± 17 years, p < 0.01) and more frequently had nonsquamous tumors than patients with other HPV types (24% [37/156] vs. 0% [0/38]; p = 0.01). Neither the HPV type nor multiple infection showed any prognostic impact. Patients with p16-negative tumors showed a significantly worse overall survival than women with p16-positive carcinomas (45 vs. 156 months, p = 0.03), although no significant differences in disease-free survival were observed. In the multivariate analysis, negative p16 immunostaining was associated with a worse overall survival together with advanced FIGO stage and lymph node metastases. In conclusion, the HPV genotype has limited clinical utility and does not seem to have prognostic value in cervical cancer. In contrast, a negative p16 result in patients with HPV-positive tumors is a prognostic marker associated with a poor overall survival.
Similar content being viewed by others
Introduction
Human papillomaviruses (HPVs) have been identified as the causative agents for the development of cervical cancer [1]. HPVs are identified in almost all premalignant cervical lesions and in most cervical cancers [2,3,4,5]. It has been shown that different HPV genotypes have a different risk of progression to cervical cancer, and that the HPV 16 and 18 genotypes have a higher risk of progression to cancer than other HPV types [4, 6, 7]. It has been hypothesized that the HPV genotype may also influence the prognosis of the neoplasia once it has already developed [8,9,10]. However, few studies have analyzed the impact of this possible association, and thus, the possible clinicopathological and prognostic implications related to the different HPV genotypes are largely unknown.
p16 immunohistochemical overexpression is considered a good surrogate marker of HPV-associated tumors [11,12,13,14], independently of the HPV genotype involved in the development of cancer. HPV-associated and HPV-independent tumors can be found in vulvar, vagina, and head and neck cancer. p16 is commonly used to correctly classify these tumors, since cancers arising via the HPV-independent pathway are p16 negative and are more aggressive, whereas HPV-associated tumors show a strong overexpression of p16 and have better prognosis [15,16,17,18,19]. Interestingly, negative p16 immunostaining has been described in a small percentage of cases of the HPV-positive carcinomas of the vagina, vulva, and head and neck [15, 19, 20]. It is not clear whether the absence of p16 overexpression in HPV-positive tumors is due to a true absence of p16 expression or to a false positive result of the HPV test. Indeed, previous studies have suggested at least two ways of p16 suppression: allelic loss in the p16 region and/or promoter hypermethylation [21, 22], and it has been described that the inactivation of the tumor suppressor gene p16 due to genetic or epigenetic alterations can be associated with the absence of p16 immunostaining [23]. Interestingly, p16 inactivation due to either mutation or methylation is a relatively frequent event in different cancers and has been associated with tumor progression and dissemination [24] and with a more aggressive behavior [21, 25]. As almost all cervical cancers are considered to be HPV-associated, p16 is considered of little value in these tumors and consequently, there is little information on the frequency of and the clinical implications associated with the absence of p16 overexpression.
In the present study, we analyzed the HPV genotype in a large series of HPV-associated cervical cancers using a highly sensitive polymerase chain reaction (PCR) technique and p16 immunohistochemical staining and correlated the findings with the clinicopathological features and tumor behavior after a long-term follow-up.
Materials and methods
Study design
The study included all women with cervical cancer admitted to the Gynecological Oncology Unit of the Hospital Clinic of Barcelona from January 2012 to January 2016. The following inclusion criteria were required to be included in the study: (1) histologically confirmed diagnosis of cervical cancer; (2) a paraffin block with available tumor tissue for HPV DNA genotyping and p16 immunostaining; and (3) positive result of the HPV DNA test.
The demographic and clinicopathological features (age, smoking habit, clinical presentation, histological type, staging [International Federation of Gynecology and Obstetrics, FIGO 2009 classification], nodal status) and follow-up data were recorded. The study was approved by the Ethics Committee of the Hospital Clinic (registry HCB/2015/0517).
Study protocol
All women had been referred to the Gynecological Oncology Unit and were managed according to the standard clinical protocols of our center, which have been described elsewhere [26]. During the pretreatment evaluation, all patients underwent tumor biopsy, thorough clinical examination for FIGO 2009 staging, blood analysis, as well as abdominopelvic imaging evaluation (magnetic resonance imaging and/or computerized tomography scan). The primary location of the tumor in the uterine cervix was confirmed in all cases by the imaging and pathological evaluation, after careful review by a radiologist and a pathologist with experience in gynecological tumors [3]. Patients suspected of having endometrial or extra-uterine tumors were excluded.
Clinical management and treatment
Following the clinical guidelines [27] women with FIGO 2009 stage IA1 tumors without lymphovascular space invasion underwent a loop electrosurgical excision procedure or extra-fascial hysterectomy. Patients with FIGO 2009 stages IA1 with lymphovascular invasion, IA2, IB1, or IIA tumors underwent laparoscopy with intraoperative sentinel lymph node evaluation or pelvic lymphadenectomy [28]. Patients with negative nodes were treated either with radical vaginal hysterectomy assisted by laparoscopy or with radical trachelectomy, while patients with positive nodes underwent complete para-aortic and selective pelvic lymphadenectomy removing all suspicious or enlarged lymph nodes without hysterectomy. Patients with FIGO 2009 stage IB2, IIB, or III cervical cancer underwent a complete para-aortic lymphadenectomy with selective pelvic lymphadenectomy (in patients in whom enlarged lymph nodes were identified by imaging techniques or during the surgical procedure). According to the current guidelines for cervical cancer treatment, patients showing risk factors after radical surgery received adjuvant radiotherapy or chemoradiotherapy [27, 29]. Chemotherapy consisted of six single weekly doses of 40 mg/m2 cisplatin and was administered simultaneously with radiotherapy, which consisted of external beam radiation to the pelvic region followed by brachytherapy. A three-dimensional image-based treatment was planned, and radiotherapy was delivered 5 days per week (1.8 Gy/d) up to a dose of 45–50 Gy. Brachytherapy was administered using Iridium192 high-dose-rate source with a MicroSelectron projector (Nucletron Trading B.V., Leersum, Netherlands) (six fractions of 4–5 Gy, 2–3 fractions per week). A parametrial boost was performed in cases with neoplastic involvement of the parametria (mean dose of 12 Gy; range, 10–16 Gy). In patients with histologically confirmed metastatic para-aortic lymph nodes, in addition to pelvic radiotherapy, the radiation field was extended to the para-aortic area. Women with stage IV tumors were treated with chemotherapy or chemoradiotherapy, as described elsewhere [3].
Histological evaluation and tissue preparation
All samples had been routinely formalin-fixed and paraffin-embedded. The blocks were serially sectioned with a microtome using a sandwich method [30]. The first section (3-µm thick) was stained with hematoxylin and eosin and used for diagnosis; the second 3-µm thick section was used for p16 immunohistochemistry. The following two sets of sections (8-µm thick), were collected in RNAase-free reaction tubes for PCR analysis. A final 3 µm section was stained with hematoxylin and eosin and used for confirmation of the presence of the tumor. Paraffin blocks lacking tissue were cut in between the patient samples as controls to ensure the lack of contamination. Two gynecological pathologists evaluated all the specimens. The histological diagnosis was based on hematoxylin and eosin morphological criteria.
Nucleic-acid isolation, DNA evaluation, and HPV genotyping
DNA extraction and HPV genotyping were performed in formalin-fixed and paraffin-embedded tissue from either the pretreatment biopsy or the surgical specimen, as described previously [26]. Briefly, DNA was extracted by overnight incubation in 20 µL of proteinase K solution (1 mg/mL) at 56 °C. Subsequently, proteinase K was heat inactivated by incubation of the sections at 95 °C for 10 min, and samples were spun and cooled down at −20 °C for 1–2 min. DNA was isolated using a commercially available kit (QIAamp Tissue Kit; Qiagen, Hilden, Germany). DNA yields were quantified spectrophotometrically using the NanoDrop ND–1000 (Thermo Scientific NanoDrop, Wilmington, DE, USA).
To assess the quality of the DNA, β-globin PCR analysis was performed using the primers BGPCO3 and BGPCO5, as described elsewhere [3]. All samples were β-globin PCR-positive, indicating adequate DNA quality.
Ten microliters of isolated DNA were used for HPV detection and genotyping using the SPF10 PCR-LiPA25 system (Fujirebio, Gent, Belgium) as described elsewhere [8, 26]. This system allows the amplification and typing of high-risk HPV types (16, 18, 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, and 68), probably high-risk HPV types (26, 53, 66, 70, 73, and 82) and low-risk HPV types (6, 11, 40, 42, 43, 44, 54, 61, 62, 67, 81, 83, and 89). HPV DNA positive specimens not hybridizing with any of the 32 probes were classified as HPV type X (HPV X or undetermined type). Each run contained negative and positive controls to monitor the efficiency of DNA isolation, PCR amplification, hybridization, and genotyping procedures. None of the negative control samples were positive in the HPV PCR assay, indicating adequate absence of contamination.
Immunohistochemical detection of p16
All tumors were stained with p16 (CINtec Histology Kit, clone E6H4; Roche-Mtm-Laboratories, Heidelberg, Germany) following the manufacturer’s protocol. Immunohistochemistry was performed with the Autostainer Link 48 automated system (Dako Co, Carpinteria, CA, USA), using the EnVision system (Dako). Only cases with diffuse and strong nuclear and cytoplasmic staining (block staining) in all basal and suprabasal viable tumor cells were considered positive for p16, whereas cases with irregular or focal staining were considered negative, as were all cases with complete absence of p16 [26].
Data analyses
Data analyses were performed with the SPSS 23.0 statistical package (SPSS, Chicago, IL, USA). Categorical variables were expressed as absolute numbers and percentages. The chi-square exact test was used to compare qualitative variables. Quantitative variables were expressed as mean ± standard deviation, and the Student’s t or analysis of variance tests were used for comparisons.
Cases with a single HPV type were considered as single HPV infections, whereas cases with two or more HPV genotypes were classified as multiple HPV infections. Cases with HPV X were excluded from the analysis of single vs. multiple infection. HPV infections were grouped as: (a) HPV 16 infections (including single and multiple infections with HPV 16 and HPV genotypes other than HPV 18); (b) HPV 18 infections (which included single and multiple infections with HPV 18 and HPV genotypes other than HPV 16); (c) HPV 16 and 18 coinfections (HPV 16 and HPV 18 infection ± an infection by another HPV genotype); and (d) other HPV infections (including single or multiple infections containing other HPV genotypes different from HPV 16 or 18). HPV 16, HPV 18, and HPV 16 and 18 coinfections were grouped (HPV 16 and/or 18) for HPV analysis. The presence of other genotypes, including HPV X, were considered as other HPV types. FIGO staging 2009 [28], which was the staging system used at diagnosis and for the management of the patient, was used in the analyses. FIGO stages IA1 to IB1 were considered as early cervical cancers and FIGO stages IB2 to IV were deemed advanced tumors.
Disease-free survival was defined as the time from diagnosis to the first local recurrence or metastases. Overall survival was defined as the time from the date of diagnosis to the date of death or to the last follow-up. Deaths without documented progression were censored at the date of death. Survival data were analyzed with the Kaplan–Meier method, and survival curves were compared using the log-rank test. The level of significance was established at 0.05. Univariate and multivariate Cox models were used to analyze prognostic factors: histological type (squamous vs. nonsquamous cervical cancer), HPV genotype (HPV 16 and/or 18 vs. other HPV types), type of HPV infection (single vs. multiple infection), p16 staining (positive vs. negative), FIGO 2009 stage (early vs. advanced stage), and lymph node status (negative vs. positive nodes). The hazard ratio and the 95% confidence intervals were calculated for risk estimation. The multivariate Cox model included all variables showing statistical significance in the univariate models.
Results
From January 2012 to January 2016, 215 women with cervical cancer were referred to our center. Twenty-one cases (10%) were negative for HPV and were consequently excluded from the analysis. The clinicopathological characteristics of the HPV-negative tumors have been described previously [26]. The tumors from the remaining 194 patients (90%) were positive for HPV and were included in the study.
The mean age of the women included in the study was 50 years (standard deviation 15 years). The number of single and multiple infections and the distribution of the HPV genotype in the 194 HPV-associated cervical carcinomas are shown in Fig. 1. A single HPV infection was identified in 149 (77%) tumors, and multiple HPV infections were observed in 30 (15%) cases. Fifteen patients (8%) presented infection by an undetermined HPV type (HPV X). Overall (including single- and multiple-type infections), HPV 16 was the most frequent genotype identified (143/194; 74%), followed by HPV 18 (25/194; 13%), HPV 33 (15/194; 8%), HPV 45 (7/194; 4%), HPV 31 (4/194; 2%), HPV 52 (4/194; 2%), and HPV 56 (4/194; 2%). Of the 194 tumors, 156 (80%) had infection by HPV 16 and/or 18.
Table 1 shows the main clinical and pathological characteristics of the patients with tumors caused by HPV 16 and/or 18 compared with the women with tumors by other HPV types. Patients with HPV 16 and/or 18 tumors were younger than those with tumors caused by other HPV types. Women with tumors with HPV 16 and 18 coinfections were younger than women with single HPV 16 or HPV 18 infections (39 ± 9 years for HPV 16 and 18 coinfections; 50 ± 15 years for single HPV 16 infections; and 47 ± 12 years for single HPV 18 infections; p = 0.05). A significant proportion of HPV 16 and/or 18 tumors were of the nonsquamous type (37/156; 24%). The proportion of adenocarcinomas was much higher in patients with HPV 18 or HPV 16 and 18 coinfections than with HPV 16 infection (36% vs. 42% vs. 18%, respectively, p < 0.01). In contrast, all cervical cancers caused by other HPV types were squamous cell carcinomas. No clinical or pathological differences were observed between women with HPV X vs. those with other HPV types (non HPV 16 and/or 18; data not shown). No clinical or pathological differences were observed between women with single vs. multiple HPV infections (data not shown).
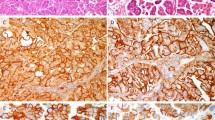
Of the 194 tumors included in the study, 186 (96%) were positive for p16 immunostaining, and only eight were negative (4%). In one of these p16-negative tumors, which showed a high-grade squamous intraepithelial lesion (HSIL) associated with invasive carcinoma, the premalignant lesion showed block p16 staining, whereas the invasive carcinoma was negative (Fig. 2).
Nonkeratinizing squamous cell carcinoma and the squamous lesion overlying the invasive tumor (case 3): a Hematoxylin and eosin, ×20; b Hematoxylin and eosin, ×100; c p16 immunostaining, ×20; d p16 immunostaining, ×100. The HSIL/CIN2–3 lesion shows strong, block type p16 staining in the intraepithelial lesion, whereas the invasive carcinoma is negative
Table 2 shows the clinical, histological, virological, and follow-up data of the eight cases with negative p16 immunostaining. There were no clinical or pathological differences between p16-positive and -negative cervical cancers: mean age 50 years (standard deviation 15) vs. 59 years (standard deviation 20), p = 0.08; smoking habit 44% vs. 25%, p = 0.40; nonsquamous cell type carcinoma 19% vs. 12%, p = 0.63; mean tumor size 36 mm (standard deviation 21) vs. 47 mm (standard deviation 18), p = 0.18; advanced FIGO 2009 staging 55% vs. 75%, p = 0.27; relapse 22% vs. 12%, p = 0.52; lymph node metastases 36% vs. 37%, p = 0.93.
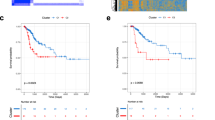
The median follow-up time was 63.6 months (95% confidence interval 7–154 months). No significant differences were found between the patients with tumors caused by HPV 16 and/or 18 compared with those with tumors caused by other HPV types in terms of disease-free survival (126 [95% confidence interval: 112–140] vs.116 [95% confidence interval: 86–147], respectively, p = 0.15) or overall survival (147 [95% confidence interval: 136–159] vs.134 [95% confidence interval: 104–164], respectively; p = 0.11). Figure 3 shows the disease-free survival (3A) and the overall survival (3B) of women with cervical cancer according to the HPV genotype. The disease-free survival of women with tumors caused by HPV 16, HPV 18, and HPV 16 and 18 coinfections was 133 [95% confidence interval: 118–147], 79 [95% confidence interval: 46–112], and 52 months [95% confidence interval: 35–69], respectively. However, these differences were not statistically significant (p = 0.23). The overall survival of patients with tumors caused by HPV 16, HPV 18, and HPV 16 and 18 coinfections was 151 [95% confidence interval: 139–163], 89 [95%confidence interval 60–117], and 59 months [95% confidence interval: 45–73], respectively, but neither did these differences reach statistical significance (p = 0.38). No differences in terms of survival were found between single vs. multiple HPV infections or between women with squamous cell carcinomas and nonsquamous cell tumors (data not shown).
In contrast, women with p16-negative tumors showed a worse overall survival compared to women with p16-positive carcinomas (45 months [95% confidence interval: 22–68] vs. 156 months [95% confidence interval: 145–168]; p = 0.03). However, no statistically significant differences were observed in terms of disease-free survival (44 months [95% confidence interval: 21–68] vs.134 months [95% confidence interval: 120–148], respectively; p = 0.21) (Fig. 4). The results of the univariate and multivariate analysis for progression and mortality are shown in Table 3. Advanced FIGO 2009 stage and lymph node metastases were associated with a higher risk of relapse and mortality. Negative p16 immunostaining was not related to the risk of relapse but was associated with increased mortality. No significant variation was found when FIGO 2018 [31] restaging compared to FIGO 2009 was considered (data nor shown).
Discussion
In the present study, 80% of the HPV DNA positive cervical cancers showed infection by the HPV 16 and/or 18 genotypes, which is in keeping with previously published data [32]. HPV 33, 45, and 31 were the other genotypes included among the five most common types. These results are similar to the distribution of HPV genotypes previously reported [4, 32, 33]. The proportion of tumors with HPV X infection in our series (8%) was higher than the percentage reported in a large international series (less than 1%) that used the same HPV genotyping technique but also analyzed the DNA sequences to further characterize specimens initially classified as HPV X (specimens positive for HPV that had not hybridized with any of the 32 probes) [4].
There were few clinicopathological differences between tumors caused by HPV 16 and/or 18 and tumors caused by other HPV types. Patients with tumors caused by HPV 16 and/or 18 were significantly younger than those with tumors caused by other HPV types, which is in keeping with the higher progression rate of HPV 16 and HPV 18 infections [34]. In addition, a significant proportion (24%) of HPV 16 and/or 18 tumors were of the nonsquamous type, mainly due to the relatively high proportion of adenocarcinomas in the subset of tumors caused by HPV 18 (40%). These data are similar to previous reports [35].
HPV 16 and/or 18 have been linked to an increased risk of developing (HSIL) or cervical cancer than other high-risk HPV genotypes [7]. However, it is controversial whether the HPV genotype has prognostic value once cervical cancer has developed [7, 9]. Some studies have suggested a favorable prognosis for HPV 16 tumors compared with cervical cancers caused by other high-risk HPV genotypes [9]. Nevertheless, these results have not been confirmed by other studies [36]. In our series, a trend for better prognosis for HPV 16 cervical cancers was identified, but the differences were not statistically significant. Patients with HPV 16 also showed better disease-free and overall survivals compared with HPV 18 infections, but these results did not reach statistical significance either. It is also controversial whether women with cervical cancers showing multiple HPV infection have a poorer survival than those with cervical cancer in whom a single HPV infection is detected [37, 38]. In the present study, no differences in survival were observed between patients with cervical cancers with single and multiple HPV type infection, and the histological type had no impact on prognosis, which is in keeping with the results of previous studies [3, 26, 39].
In our series, the main prognostic factors were FIGO staging and the presence of lymph node metastases. The prognostic relevance of lymph node status, consistently observed in other studies [3, 26, 40, 41], has led to the recent modification in the FIGO staging to include the involvement of lymph nodes [31].
One of the most relevant results of our study is that HPV-positive cervical cancers with negative p16 immunostaining had a worse overall survival. There is scant information on the impact of p16 immunohistochemical staining on the prognosis of cervical cancer. Previous studies have shown that abnormalities in the tumor suppressor gene p16 are associated with an aggressive tumor behavior [21, 24, 42]. Interestingly, a recent meta-analysis has reported a better prognosis (disease-free survival) of cervical cancers associated with overexpression of p16 [43]. p16-negative staining in an HPV-associated tumor can be caused by different mechanisms. Loss of heterozygosity is a frequent event in several tumors and has been previously described in HPV-associated carcinomas [44]. Point mutations of p16, and more frequently, silencing of the p16 gene by hypermethylation of the promoter, have been reported in many human cancers including HPV-associated cancers [21, 45]. p16-negative staining may also represent a false positive result of the HPV detection technique. In this regard, in other areas, such as the vulva or the head and neck, it has been proposed that HPV infection cannot be reliably diagnosed by the detection of HPV DNA alone, and that a second test, such as p16 or the detection of E6/7 mRNA, should be required in order to classify a tumor as conclusively associated with HPV [17,18,19, 46, 47]. However, the evidence of one p16-negative tumor, which showed p16-positive staining in the surrounding intraepithelial lesion (Fig. 2), strongly favors the loss of p16 overexpression due to mutation or silencing of the gene. In the present series, the proportion of cervical cancers with negative p16 immunostaining was very small (4%). This proportion is in keeping with the percentages observed in previous series [11]. Interestingly, seven out of the eight cervical cancers which were negative for p16 in our series were squamous cell carcinomas. This contrasts with other studies that have shown a higher proportion of p16-negative immunostaining mainly in nonsquamous tumors [48].
This study has some limitations. First, the use of formalin-fixed, paraffin-embedded samples could have resulted in worse sample quality, and consequently, misclassification of some of the tumors (e.g., multiple infections wrongly cataloged as simple infections, excess of cases classified as HPV X infections). However, we used a standardized and highly sensitive PCR technique and carried out the procedure under rigorous contamination controls as reported previously [26]. Another possible limitation is the small number of p16-negative cervical carcinomas. However, this is an uncommon situation in cervical cancer, and it is remarkable that, in spite of the small number of cases, the prognosis is clearly different.
In conclusion, our results suggest that the HPV genotype or multiple HPV infections do not have clinical or prognostic value in patients with cervical cancer. In contrast, women with HPV-positive cervical cancers negative for p16 have an impaired overall survival. Further studies including a larger number of cases are required to confirm these data and their potential use in clinical practice.
References
Darragh TM, Colgan TJ, Cox JT, Heller DS, Henry MR, Luff RD, et al. The lower anogenital squamous terminology standardization project for HPV-Associated Lesions: background and consensus recommendations from the College of American Pathologist and the American Society for Colposcopy and Cervical Pathology. Arch Pathol Lab Med. 2012;136:1266–97.
del Pino M, Rodríguez-Carunchio L, Alonso I, Torné A, Rodríguez A, Fusté P, et al. Clinical, colposcopic and pathological characteristics of cervical and vaginal high-grade lesions negative for HPV by Hybrid Capture 2. Gynecol Oncol. 2011;122:515–20.
Rodrıguez-Carunchio L, Soveral I, Steenbergen RDM, Torné A, Martinez S, Fusté P, et al. HPV-negative carcinoma of the uterine cervix: a distinct type of cervical cancer with poor prognosis. BJOG. 2015;122:119–27.
de Sanjose S, Quint WG, Alemany L, Geraets DT, Klaustermeier JE, Lloveras B, et al. Human papilomavirus genotype attribution in invasive cervical cancer: a retrospective cross-sectional worldwide study. Lancet Oncol. 2010;11:1048–56.
Bosch FX, Burchell AN, Schiffman M, Giuliano AR, de Sanjose S, Bruni L, et al. Epidemiology and natural history of human papilomavirus infections and type-specific implications in cervical neoplasia. Vaccine. 2008;26:1–16.
Alemany L, de Sanjosé S, Tous S, Quint W, Vallejos C, Shin HR, et al. Time trends of human papillomavirus types in invasive cervical cancer, from 1940 to 2007. Int J Cancer. 2014;135:88–95.
Khan MJ, Castle PE, LorinczAT, Wacholder S, Sherman M, Scott DR, et al. The elevated 10-year risk of cervical precancer and cancer in women with human papillomavirus (HPV) type 16 or 18 and the possible utility of type-specific HPV testing in clinical practice. J Natl Cancer Inst. 2005;97:1072–9.
Walboomers JM, Jacobs MV, Manos MM, Bosch FX, Kummer JA, Shah KV, et al. Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. J Pathol. 1999;189:12–9.
Onukia M, Matsumotoa K, Tenjimbayashia Y, Tasakab N, Akiyamab A, Sakuraib M, et al. Human papillomavirus genotype and prognosis of cervical cancer: favorable survival of patients with HPV16-positive tumors. Papillomavirus Res. 2018;6:41–5.
Barreto CL, Martins DB, De Lima Filho JL, Magalhães V. Detection of human papillomavirus in biopsies of patients with cervical cancer, and its association with prognosis. Arch Gynecol Obstet. 2013;288:643–8.
Kanthiya K, Khunnarong J, Tangjitgamol S, Puripat N, Tanvanich S. Expression of the p16 and Ki67 in Cervical Squamous Intraepithelial Lesions and Cancer. Asian Pac J Cancer Prev. 2016;17:3201–6.
Tringler B, Gup CJ, Singh M, Groshong S, Shroyer AL, Heinz DE, et al. Evaluation of p16INK4a and pRb expression in cervical squamous and glandular neoplasia. Hum Pathol. 2004;35:689–96.
Eleutério J Jr, Giraldo PC, Gonçalves AK, Cavalcante DI, de Almeida Ferreira F, Mesquita SM, et al. Prognostic markers of high-grade squamous intraepithelial lesions: the role of p16INK4a and high-risk human papillomavirus. Acta Obstet Gynecol Scand. 2007;86:94–8.
Carozzi F, Confortini M, Dalla Palma P, del Mistro A, Gillio-Tos A, de Marco L, et al. New Technologies for Cervical Cancer Screening (NTCC) Working Group. Use of p16-INK4a overexpression to increase the specificity of human papillomavirus testing: a nested substudy of the NTCC randomised controlled trial. Lancet Oncol. 2008;9:937–45.
Alonso I, Felix A, Torne A, Fusté V, del Pino M, Castillo P, et al. Human papillomavirus as a favorable prognostic biomarker in squamous cell carcinomas of the vagina. Gynecol Oncol. 2012;125:194–9.
Nooij LS, terHaar NT, Ruano D, Rakislova N, van Wezel T, VTHBM Smit, et al. Genomic characterization of vulvar (pre) cancers identifies distinct molecular subtypes with prognostic significance. Clin Cancer Res. 2017;23:6781–9.
Alos L, Moyano S, Nadal A, Alobid I, Blanch JL, Ayala E, et al. Human papillomaviruses are identified in a subgroup of sinonasal squamous cell carcinomas with favorable outcome. Cancer. 2009;115:2701–9.
Larque AB, Hakim S, Ordi J, Nadal A, Diaz A, del Pino M, et al. High-risk human papillomavirus is transcriptionally active in a subset of sinonasal squamous cell carcinomas. Mod Pathol. 2014;27:343–51.
Rakislova, Clavero O, Alemany L, Saco A, Quirós B, Lloveras B, et al. Histological characteristics of HPV-associated and -independent squamous cell carcinomas of the vulva: a study of 1.594 cases. Int J Cancer. 2017;141:2517–27.
Smith EM, Rubenstein LM, Hoffman H, Haugen TH, Turek LP. Human papillomavirus, p16 and p53 expression associated with survival of head and neck cancer. Infect Agent Cancer. 2010;5:4.
Poetsch M, Hemmerich M, Kakies C, Kleist B, Wolf E, vom Dorp F, et al. Alterations in the tumor suppressor genep16(INK4A) are associated with aggressive behavior of penile carcinomas. Virchows Arch. 2011;458:221–9.
Yanagawa N, Osakabe M, Hayashi M, Tamura G, Motoyama T. Detection of HPV-DNA, p53 alterations and methylation in penile squamous cell carcinoma in Japanese men. Pathol Int. 2008;58:477–82.
Ferreux E, Lont AP, Horenblas S, Gallee MP, Raaphorst FM, von KnebelDoeberitz M, et al. Evidence for at least three alternative mechanisms targeting the p16INK4A/cyclin D/Rb pathway in penile carcinoma, one of which is mediated by high-risk human papillomavirus. J Pathol. 2003;201:109–18.
Karamitopoulou E, Zlober I, Koumarianou A, Patsouris ES, Peros G, Lugli A. Expression of p16 in lymph node metastases of adjuvantly treated stage III colorectal cancer patients identifies poor prognostic subgroups: a retrospective analysis of biomarkers in matched primary tumor and lymph node metastases. Cancer. 2010;116:4474–86.
Schulz P, Scholz A, Rexin A, Hauff P, Schirner M, Wiedenmann B, et al. Inducible re-expression of p16 in an orthotopic mouse model of pancreatic cancer inhibits lymphangiogenesis and lymphatic metastasis. Br J Cancer. 2008;99:110–7.
Nicolas I, Marimon L, Barnadas E, Saco A, Rodríguez-Carunchio L, Fuste P, et al. HPV-negative tumors of the uterine cervix. Mod Pathol. 2019;32:1189–96.
Marth C, Landoni F, Mahner S, McCormack M, Gonzalez-Martin A, Colombo N. Cervical cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2017;28:72–83.
FIGO Committee on Gynecologic Oncology. FIGO staging for carcinoma of the vulva, cervix, and corpus uteri. Int J Gynaecol Obstet. 2014;125:97–8.
del Pino M, Fusté P, Pahisa J, Rovirosa A, Martínez-Serrano MJ, Martínez-Román S, et al. Laparoscopic lymphadenectomy in advanced cervical cancer: prognostic and therapeutic value. Int J Gynecol Cancer. 2013;23:1675–83.
Van der Marel J, Berkhof J, Ordi J, Torné A, Del Pino M, van Baars R, et al. Attributing oncogenic human papillomavirus genotypes to high-grade cervical neoplasia: which type causes the lesion? Am J Surg Pathol. 2015;39:496–504.
Bhatla N, Aoki D, Sharma DN, Sankaranarayanan R. Cancer of the cervix uteri. Int J Gynaecol Obstet. 2018;2:22–36.
Clifford GM, Smith JS, Plummer M, Muñoz N, Franceschi S. Human papillomavirus types in invasive cervical cancer worldwide: a meta-analysis. Br J Cancer. 2003;88:63–73.
Alemany L, Pérez C, Tous S, Llombart-Bosch A, Lloveras B, Lerma E, et al. Human papillomavirus genotype distribution in cervical cancer cases in Spain. Implication for prevention. Gynecol Oncol. 2012;124:512–7.
Jaisamrarn U, Castellsagué X, Garland SM, Naud P, Palmroth J, Del Rosario-Raymundo MR, et al. Natural history of progression of HPV infection to cervical lesion or clearance: analysis of the control arm of the large, randomised PATRICIA study. PLoS One. 2013;8:e79260.
Chen W, Sun H, Molijn A, Zeng L, Kang L, Jenkins D, et al. The variable characteristics of human papillomavirus in squamous cell carcinoma and adenocarcinoma of cervix in China. J Low Genit Trac Dis. 2018;22:355–61.
van Muyden RC, terHarmsel BW, Smedts FM, Hermans J, Kuijpers JC, Raikhlin NT, et al. Detection and typing of human papillomavirus in cervical carcinomas in Russian women: a prognostic study. Cancer. 1999;85:2011–6.
Nogueira-Dias-Genta ML, Martins TR, Mendoza-Lopez RV, Sadalla JC, de Carvalho JPM, Baracat EC, et al. Multiple HPV genotype infection impact on invasive cervical cancer presentation and survival. PLoS One. 2017;12:1–10.
Bosch FX, Lorincz A, Munoz N, Meijer CJ, Shah KV. The causal relation between human papillomavirus and cervical cancer. J Clin Pathol. 2002;55:244–65.
Xie X, Song K, Cui B, Jiang J, Yang X, Kong B. A comparison of the prognosis between adenocarcinoma and squamous cell carcinoma in stage IB–IIA cervical cancer. Int J Clin Oncol. 2018;23:522–31.
Quinn MA, Benedet JL, Odicino F, Maisonneuve P, Beller U, Creasman WT, et al. Carcinoma of the cervix uteri. Inter J Gynaecol Obstet. 2006;95:43–103.
Durowade KA, Osagbemi GK, Salaudeen AG, Musa OI, Akande TM, Babatunde OA, et al. Prevalence and risk factors of cervical cancer among women in an urban community of Kwara State, north central Nigeria. J Prev Med Hyg. 2012;53:213–9.
Saadallah-Kallel A, Abdelmaksoud-Dammak R, Triki M, Charfi S, Khabir A, Sallemi-Boudawara T, et al. Clinical and prognosis value of the CIMP status combined with MLH1 or p16 INK4a methylation in colorectal cancer. Med Oncol. 2017;3:147.
Huang K, Li LA, Meng YG, Fu XY. p16 expression in patients with cervical cancer and its prognostic significance: meta-analysis of published literature. Eur J Obstet Gynecol Reprod Biol. 2014;183:64–9.
Poetsch M, Schuart B-J, Schwesinger G, Kleist B, Protzel C. Screening of microsatellite markers in penile cancer reveals differences between metastatic and non-metastatic carcinomas. Mod Pathol. 2007;20:1069–77.
Esteller M, Corn PF, Baylin SB, Herman JG. A gene hypermethylation profile of human cancer. Cancer Res. 2001;61:3225–29.
de Sanjose S, Alemany L, Ordi J, Tous S, Alejo M, Bigby SM, et al. Worldwide human papillomavirus genotype attribution in over 2000 cases of intraepithelial and invasive lesions of the vulva. Eur J Cancer. 2013;49:3450–61.
Halec G, Alemany L, Quiros B, Clavero O, Höfler D, Alejo M, et al. Biological relevance of human papillomaviruses in vulvar cancer. Mod Pathol. 2017;30:549–62.
Stiasny A, Freier CP, Kuhn C, Schulze S, Mayr D, Alexiou C, et al. The involvement of E6, p53, p16, MDM2 and Gal-3 in the clinical outcome of patients with cervical cancer. Oncol Lett. 2017;14:4467–76.
Acknowledgements
We thank Donna Pringle for English revision of the article and Fran Calpe for image processing.
Funding
This research was funded in part by Instituto de Salud Carlos III (ICSIII)-Fondo de Investigación Sanitaria, and ERDF ‘One Way to Europe’ (PI17/00772).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nicolás, I., Saco, A., Barnadas, E. et al. Prognostic implications of genotyping and p16 immunostaining in HPV-positive tumors of the uterine cervix. Mod Pathol 33, 128–137 (2020). https://doi.org/10.1038/s41379-019-0360-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-019-0360-3
This article is cited by
-
Human papillomavirus-mediated carcinogenesis and tumor progression
Genome Instability & Disease (2021)
-
Aktuelle WHO-Klassifikation des weiblichen Genitale
Der Pathologe (2021)
-
CD274 (PD-L1), CDKN2A (p16), TP53, and EGFR immunohistochemical profile in primary, recurrent and metastatic vulvar cancer
Modern Pathology (2020)
-
Increased miR-20b Level in High Grade Cervical Intraepithelial Neoplasia
Pathology & Oncology Research (2020)