Abstract
Genetics has played an important role in risk stratification for plasma cell myeloma patients, providing therapeutic guidance. In this study, we investigated the correlation of bone marrow morphologic features and genetic aberrations, including gene expression profiles, translocations, and gene mutations. For the first time we show that high plasma cell volume, diffuse sheet growth pattern, immature cell morphology, high mitotic index, and increased reticulin fibrosis, significantly correlates with high risk disease determined by MyPRS gene expression profiles. Furthermore, we show the association between MyPRS risk stratification and chromosomal alterations and specific gene mutations. We also demonstrate the combinational effect of TP53 mutation and 17p loss on the histological changes in bone marrow.
Similar content being viewed by others
Introduction
Plasma cell myeloma is a malignant neoplastic proliferation of plasma cells with clinical and genetic heterogeneity. Bone marrow plasma cell histologic characteristics and staging in cohorts of multiple myeloma patients have been characterized and reported [1,2,3,4]. In addition to clinical features, genetics has provided an important independent prognostic indicator for myeloma patients. Many previous studies have demonstrated an important prognostic value of conventional cytogenetics and fluorescence in situ hybridization (FISH) based assessment of chromosomal alterations. Recent work and data propose molecular classification based on gene expression profiling has shown a promising role in determining prognostic risk status for myeloma patients [5].
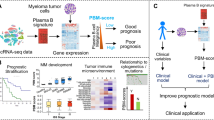
Seven different molecular subtypes were established by the University of Arkansas for Medical Sciences (UAMS) group through unsupervised hierarchic clustering of gene expression data, which later were included in the Myeloma Prognostic Risk Signature (MyPRS) [6]. The hyperdiploidy group is associated with trisomies of chromosomes. CD-1 and CD-2 groups are characterized by activating translocations of CCND1 and CCND3. MS group is associated with upregulation of FGFR3 or MMSET. MF group is defined by c-MAF or MAFB activation. LB group is associated with low bone lesions. The last subgroup, PR, is distinguished by increased expression of proliferation-associated genes. A GEP70 prognostic risk score was later developed based on expression levels of 70 genes. An individualized “Virtual Karyotype” was also created based on 813 gene expression, which makes up MyPRS, in conjunction with the molecular subtypes and risk score. It has been shown that the molecular subtypes and the 70-gene prognostic risk score significantly correlated with prognosis in myeloma patients [5,6,7,8]. The five year overall survival rate between high risk score cases and low risk score cases are dramatically different (28% vs 78%, p < 0.001) [9].
Plasma cell morphologic and molecular/genetic correlation has been rarely reported, such as lymphoplasmacytic morphology and the associated t(11:14) [10, 11]. We recently reported TP53 aberrations are associated with immature plasma cell morphology [12]. We were interested in further investigating the correlation of bone marrow histologic and morphologic features with MyPRS gene expression and gene mutation studies at a single academic center.
Methods
The study was comprised of 329 multiple myeloma bone marrow biopsies with available MyPRS risk scores, collected at a single large academic center, Mount Sinai Icahn School of Medicine Tisch Cancer Center from 01/2017 to 05/2018. Patients were consented with the Mount Sinai Institutional Review Board Tisch Cancer Institute Biorepository protocol [HS#: 10-00135; GCO#1: 06-0996 (0001)]. For patients with multiple biopsies, the biopsy with the highest plasma percentage was selected. The respective bone marrow morphology, FISH, and mutation profile data of 266 patients were collected.
Morphologic assessment
Bone marrow trephine biopsies were assessed for cellularity, percent volume plasma cells (Bartl stage), plasma cell infiltration patterns (interstitial, microclusters, nodules, diffuse sheets), mitotic figures per high power field (HPF) and fibrosis. Bartl stages were defined, based on the plasma cell burden in the biopsy, as stage I (<20 vol%), stage II (20–50 vol%) and stage III (>50 vol%) [1]. Biopsies were stained with CD138, kappa and lambda to characterize plasma cells. Reticulin stain was also performed to evaluate reticulin fiber content and graded as follows: none/0, mild/1, moderate/2, and severe/3. The aspirate smears were assessed for percentage of plasma cells based on a 200 cell differential count. Assessment of cytologic features was based on the modified Bartl grading system [13]; a three tiered grading scheme where grade 1 is assigned to cases with >70% of plasma cells showing morphologically mature plasma cells characterized by small-medium size, eccentric nucleus, coarse clumped chromatin and inconspicuous nucleolus. Grade 2 shows a mix of mature plasma cells and <50% plasma cells with nuclear atypia, open chromatin, lack of significant pleomorphism and small nucleoli. Grade 3 shows greater than 50% of plasma cells with marked cytologic atypia showing open chromatin, pleomorphism, and prominent nucleoli. The differences in these morphologic features were assessed between the MyPRS high risk cases and MyPRS low risk cases.
All images in the paper were taken with Nikon Plan Fluor under ×20, ×40 or ×100 high dry objective, Nikon Eclipse Ci microscope, Nikon DS-Ri2 camera, and NIS-Elements D 4.40.00 image acquisition software.
Molecular and genetic studies
Samples were assessed by metaphase karyotyping (conventional cytogenetics) as routinely performed in the clinical laboratory at Mayo Clinic in Rochester, MN [14]. FISH examination was carried out in conjunction with staining for cytoplasmic immunoglobulin, as routinely performed in the clinical laboratory at Mayo Clinic in Rochester, MN [14].
MyPRS prognostic risk score was calculated based on 70 gene expression profiling, and molecular subtype analysis through 700 gene expression profiling were performed at Signal Genetics. Virtual karyotype of 13 distinct chromosomes or sub-chromosomal regions was also created by MyPRS, based on the expression levels of 816 genes [5, 15, 16]. In this study, we treated both My PRS high risk score and high risk-border line as high risk and both low risk score and low risk-border line as low risk. Results from targeted next-generation sequencing studies (FoundationOne® Heme) were used for gene mutation analysis.
Statistics
Results are described as mean. T-tests, where p < 0.05 is significant, were used to compare age, bone marrow cellularity, mitoses/HPF, and genetic alteration rate per case between the MyPRS high risk and low risk cases. Fisher’s exact test or Chi square test were performed for analysis of different distribution in gender, race, molecular subtypes, morphologic features (including Bartl stage, growth pattern, nuclear morphology, fibrosis, mitotic index), light chain restriction, gene mutation and translocation between the two groups of patients. All statistical analyses were performed using GraphPad Prism, San Diego, CA, USA.
Results
Demographic and clinical characteristics of the cases
Biopsies from 266 myeloma patients were scored and analyzed (Table 1). There were 115 females (43%) and 151 males (57%). The average age of patients in the high risk group was 63.7 years (range = 34–90 years) and the average age in the low risk group was 64.8 years (range = 32–98 years). There was no significant difference in age (p = 0.3787), gender ratio (p = 0.0615), and racial distribution (p = 0.5065) between the two groups. The majority of biopsies in both the high and low risk group were not collected at diagnosis. The high risk group of patients were more likely to have active myeloma (84% vs 63%), higher International Staging System stage disease (stage III 27% vs 5%), and to be treated as well (62% vs 46%).
Different histologic features between high and low risk cases
First, we investigated whether high risk and low risk cases presented with different bone marrow histologic features (Table 2), which have not been previously reported. Compared to low risk, high risk cases showed overall higher marrow cellularity (53% vs 44%; p = 0.0026) and higher percentage of neoplastic plasma cells on bone marrow aspirate smears (30% vs 22%; p = 0.0084). Similarly, highest histological stage (Bartl stage 3) of tumor was more closely associated with high risk cases than low risk cases (p = 0.0001). This indicates that higher tumor burden exists in high risk diseases.
Histologically, neoplastic plasma cells can infiltrate the bone marrow with a spectrum of growth patterns, such as interstitial, microclusters, nodules, and sheets (Fig. 1). The diffuse sheet growth pattern was more commonly seen in the high risk cases (22% vs 10%, p = 0.0065) compared to the non-sheet growth patterns (interstitial, microclusters, nodules). According to the nuclear features, neoplastic plasma cells can be graded into three types (Fig. 2). Our results showed that the immature tumor more commonly occurred in the high risk cases (constituting 93% of all grade 3 morphology) (p = 0.0001). The average mitotic index approached statistically significance between the two risk group (2/HPF vs 1.4/HPF, p = 0.0871), which showed dramatic difference after excluding borderline high and low risk cases (3/HPF vs 1/HPF; p = 0.0267).
Growth Pattern of neoplastic plasma cells. CD138 positive plasma cells infiltrate in interstitial (left upper, ×200 magnification), microcluster (right upper, ×200 magnification, less than 10 CD138 positive plasma cells per cluster), nodule (left lower, ×200 magnification, >10 CD138 positive plasma cells per cluster), and sheet pattern (right lower, ×200 magnification)
Fibrosis is often seen in bone marrow of the plasma cell myeloma patients (Fig. 3). We found that high risk cases tended to have more moderate to severe fibrosis than low risk ones (14% vs 4%, p = 0.0096). Consistent with our findings, it has been reported that fibrosis is significantly associated with inferior survival in myeloma patients [2, 17]. Interestingly, although the heavy chain types showed no dramatic difference between the two groups (Table S1), higher percentage of lambda light chain restriction was also found in the high risk cases (48% vs 31%, p = 0.0137).
Bone marrow fibrosis grading. Fibrosis in bone marrow, highlighted by reticulin staining, were graded as: MF0 = scattered linear reticulin with no intersections, MF1 = loose network of reticulin with many intersections, especially in perivascular areas, MF2 = diffuse and dense increase in reticulin with extensive intersections, and/or associated with focal osterosclerosis and MF3 = diffuse and dense increase in reticulin with extensive intersections and associated osterosclerosis (×400 magnification)
Additionally, the percentage of different cell types in the bone marrow aspirate based on 200 differential cell count (myeloid precursors, bands, segmented neutrophils, basophils, eosinophils, monocytes, erythroid and lymphocytes) were similar in high risk and low risk cases, except for blasts, which was lower in the high risk cases (0.5% vs 0.8%, p = 0.0041).
Different genetic alterations between high and low risk cases
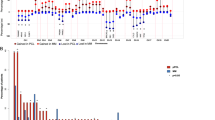
Next, we studied the association between MyPRS risk stratification and genetic alterations. Among the 7 molecular subtypes, poor prognostic types (MF, MS and PR) were closely associated with high risk group, whereas good prognostic types (CD1, CD2, HY, and LB) were closely associated with low risk group (p = 0.0001) (Table 3). 1q gain, 13q loss and IGH-FGFR3 were found to be associated with high risk diseases in both the MyPRS virtual karyotype and the Mayo FISH studies (Tables S2, 3).
In our targeted Next Generation Sequencing study, higher genetic alteration rate, including both mutations and translocations, was identified in the high risk cases, compared to low risk ones (2.7/per case vs 2.0/per case, p = 0.0012). In Table S4, we listed the top 10% most frequent translocations identified in the Next Generation Sequencing study and statistically significant difference was not identified between the high risk cases and low risk cases.
Among the top 10 most frequent mutations (Table 4) in our study, mutations of KRAS, NRAS, and BRAF were the recurrent ones. These genes were relatively equally distributed in both high risk and low risk cases. In addition, two genes involving the epigenetic modifiers, DNMT3A and TET2, were also frequently mutated in both high and low risk cases, consistent with the concept that epigenetic dysregulation is important for myeloma pathogenesis [18].
Furthermore, we also identified two mutated genes, TP53, and CD36, which were significantly associated with high risk cases (Table 4). TP53 is the most interesting one since it has been shown to contribute to poor survival in several reports [19,20,21,22]. We found 5 cases with both TP53 mutation and 17p loss (identified by Mayo FISH) and 4 of them were in high risk group. Compared to the wild-type cases and cases with only TP53 or 17p loss, these 5 cases were found to be more closely associated with sheet growth pattern, immature cell morphology, and high mitotic index (Table 5) (Fig. 4).
The second gene is CD36, with 4 mutations identified only in high risk cases. In this study, we did not see a significant correlation of mutant CD36 with the histological features, due to the low case numbers.
Discussion
To achieve a better risk stratification for plasma cell myeloma patients, various systems, from the international staging system, revised international staging system, to Mayo stratification of myeloma and risk-adapted therapy (mSMART), chromosomal aberrations detected by cytogenetics and FISH have been commonly used [23, 24]. Recent studies have shown gene expression profiling-based molecular subtypes and prognostic risk score to have a superior role in the management of myeloma patients due to its high reproducibility [5, 15, 16]. Here we provide additional histologic and mutational genetic evidence for the prognostic value of this new method.
We report for the first time a systematic review of both bone marrow morphology and genetic aberrations in a large cohort of plasma cell myeloma patients seen at a single academic institution and cancer center. We correlated MyPRS risk stratification, defined by gene expression profiling, with bone marrow features including plasma cell morphologic grade, growth pattern, mitotic features and fibrosis. In addition, we also explored its correlation with genomic alterations identified by next generation sequencing.
First, we confirmed that the molecular subtypes and virtual karyotypes correlated with high risk and low risk gene expression, as reported in previous studies [5,6,7,8, 16, 25,26,27]. Next, we demonstrated that cases with MyPRS high risk score exhibited specific histologic features that correlate with poor clinical outcomes [1, 13]. In contrast to low-risk cases, the high risk cases were more commonly to present higher bone marrow cellularity, higher tumor burden, sheet like growth pattern, immature morphology, higher mitoses, and increased fibrosis. In our study, lambda light chain restriction was found to be more closely associated with the high-risk cases and its pathological role will be investigated in the future following studies. The cellular composition of bone marrow between high risk and low risk cases are similar, except for slightly lower percentage of blast in the high-risk cases.
In this study, the recurrent mutations involved the Ras/MAP kinase pathway (KRAS, BRAF, NRAS, and PTPN11) were identified equally in both high risk and low risk cases, suggestive of important role of this pathway in the tumorigenesis in myeloma patients. Consistent with our results, it has been reported that there is no significant survival difference in cases with these three mutations compared with those without [19]. The mutations involving the epigenetic modifiers, DNMT3A and TET2, were also identified in patients of both groups. It is noted that these are commonly mutated genes involved in clonal hematopoiesis of indeterminate potential [28, 29]. Given that CD138 plasma cell enrichment was not performed during the Next Generation Study study, we cannot exclude this possibility. Variant allele frequency may be helpful in clarifying this question. In addition, we also identified two mutated genes, TP53, and CD36, which were closely associated with the high risk cases. 17p loss identified by Mayo FISH did not show significant difference between high risk and low risk cases (16% vs 10%; p = 0.3015). Of note, inclusion of biopsies mixed at diagnosis and at relapse, treated and untreated might limit the 17p loss distribution difference between the low and high risk groups. Interestingly, cases with concurrent TP53 mutation and 17p loss appears to have more diffuse growth pattern, immature cell morphology, and higher mitotic index than wild-type cases and cases with only TP53 mutation or only 17p loss (Fig. 4). The potential bi-allelic dysfunction of TP53 might be the underlying mechanism. Consistent with our histological findings, it has been recently reported that myeloma patients with bi-allelic inactivation of TP53 is associated with much poor prognosis than the ones with wild-type or mono-allelic inactivation [30].
Although mutations of CD36 have been rarely reported, its gene expression and association with clinical outcome has been documented in both solid tumors and hematopoietic tumors including myeloma and B lymphoblastic leukemia [31,32,33]. However, the exact pathogenesis of CD36 is still elusive.
One limitation of the current study is that the biopsies at both diagnosis and relapse were included for the analysis. In our follow up project, we will focus on diagnostic biopsies and test the reproducibility. We show for the first time, after systemic investigation significant morphologic correlations between MyPRS and bone marrow morphology and gene mutations. Furthermore, we also studied the correlation between the genetic alterations associated with MyPRS high risk and the poor prognostic morphologic features. Our study provides morphologic insight associated with gene expression based risk stratification. Further analyses on a larger cohort is in process to establish criteria and a scoring system for screening and subsequent testing in clinical practice.
References
Bartl R, Frisch B, Fateh-Moghadam A, Kettner G, Jaeger K, Sommerfeld W. Histologic classification and staging of multiple myeloma. A retrospective and prospective study of 674 cases. Am J Clin Pathol. 1987;87:342–55.
Bartl R. Histologic classification and staging of multiple myeloma. Hematol Oncol. 1988;6:107–13.
Carter A, Hocherman I, Linn S, Cohen Y, Tatarsky I. Prognostic significance of plasma cell morphology in multiple myeloma. Cancer. 1987;60:1060–5.
Sailer M, Vykoupil KF, Peest D, Coldewey R, Deicher H, Georgii A. Prognostic relevance of a histologic classification system applied in bone marrow biopsies from patients with multiple myeloma: a histopathological evaluation of biopsies from 153 untreated patients. Eur J Haematol. 1995;54:137–46.
Van Laar RK, Borrelo I, Jabalayan D, Niesvizky R, Zielinski A, Leigh K, et al. MyPRS(R) molecular subtypes of multiple myeloma represent all high-risk FISH translocations included in the mSMART 2.0 and R-ISS guidelines. Blood. 2016;128:3264.
Zhan F, Huang Y, Colla S, Stewart JP, Hanamura L, Gupta S, et al. The molecular classification of multiple myeloma. Blood. 2006;108:2020–8.
Weinhold N, Heuck CJ, Rosenthal A, Thanendrarajan S, Stein CK, Van Rhee F, et al. Clinical value of molecular subtyping multiple myeloma using gene expression profiling. Leukemia. 2016;30:423–30.
Bergsagel PL, Chesi MV. Molecular classification and risk stratification of myeloma. Hematol Oncol. 2013;31(Suppl 1):38–41.
Shaughnessy JD Jr., Zhan F, Burington BE, Huang Y, Colla S, Hanamura I, et al. A validated gene expression model of high-risk multiple myeloma is defined by deregulated expression of genes mapping to chromosome 1. Blood 2007;109:2276–84.
Fonseca R, Blood EA, Oken MM, Kyle RA, Dewald GW, Bailey RJ, et al. Myeloma and the t(11;14)(q13; q32); evidence for a biologically defined unique subset of patients. Blood. 2002;99:3735–41.
Hoyer JD, Hanson CA, Fonseca R, Greipp PR, Dewald GW, Kurtin PJ. The (11;14)(q13; q32) translocation in multiple myeloma. A morphologic and immunohistochemical study. Am J Clin Pathol. 2000;113:831–7.
Taboada SE, Hussein S, Emmons F, El Jamal S, Houldsworth J, Teruya-Feldstein J. TP53Aberrations correlate with immature plasma cell morphology in multiple myeloma. Mod Pathol. 2018;31:558–9.
Goasguen JE, Zandecki M, Mathiot C, Scheiff JM, Bizet M, Ly-Sunnaram B, et al. Mature plasma cells as indicator of better prognosis in multiple myeloma. New methodology for the assessment of plasma cell morphology. Leuk Res. 1999;23:1133–40.
Rajan AM, Rajkumar SV. Interpretation of cytogenetic results in multiple myeloma for clinical practice. Blood Cancer J. 2015;5:e365.
van Laar R, Farmer P, Bender RA, Zielinski A, Leigh K, Brown N, et al. The 70- Gene MyPRSR prognostic risk score signature predicts increased risk of progression from MGUS to multiple myeloma requring treatment. Blood. 2016;128:3275.
van Laar R, Flinchum R, Brown N, Ramsey J, Riccitelli S, Heuck C, et al. Translating a gene expression signature for multiple myeloma prognosis into a robust high-throughput assay for clinical use. BMC Med Genom. 2014;7:25. https://doi.org/10.1186/1755-8794-7-25.
Hallgrimsdottir T, Porwit A, Bjorkholm M, Rossmann E, Steingrimsdottir H, Lund SH, et al. Bone marrow fibrosis in patients with multiple myeloma: a new prognostic factor for survival? Blood. 2013;122:1946.
Pawlyn C, Kaiser MF, Heuck C, Melchor L, Wardell CP, Murison A, et al. The spectrum and clinical impact of epigenetic modifier mutations in myeloma. Clin Cancer Res. 2016;22:5783–94.
Bolli N, Avet-Loiseau H, Wedge DC, Van Loo P, Alexandrov LB, Martincorena I, et al. Heterogeneity of genomic evolution and mutational profiles in multiple myeloma. Nat Commun. 2014;5:2997.
Hideshima T, Cottini F, Nozawa Y, Seo HS, Ohguchi H, Samur MK, et al. p53- related protein kinase confers poor prognosis and represents a novel therapeutic target in multiple myeloma. Blood. 2017;129:1308–19.
Chang H, Yeung J, Qi C, Xu W. Aberrant nuclear p53 protein expression detected by immunohistochemistry is associated with hemizygous P53 deletion and poor survival for multiple myeloma. Brit J Haematol. 2007;138:324–9.
Deng SH, Xu Y, An G, Sui WW, Zou DH, Zhao YZ, et al. Features of extramedullary disease of multiple myeloma: high frequency of P53 deletion and poor survival: a retrospective single-center study of 834 cases. Cl Lymph Myelom Leuk. 2015;15:286–91.
Palumbo A, Avet-Loiseau H, Oliva S, Lokhorst HM, Goldschmidt H, Rosinol L, et al. Revised International Staging System for Multiple Myeloma: A Report From International Myeloma Working Group. J Clin Oncol. 2015;33:2863–9.
Mikhael JR, Dingli D, Roy V, Reeder CB, Buadi FK, Hayman SR, et al. Management of Newly Diagnosed Symptomatic Multiple Myeloma: Updated Mayo Stratification of Myeloma and Risk-Adapted Therapy (mSMART) Consensus Guidelines 2013. Mayo Clin Proc. 2013;88:360–76.
Shaughnessy J, Zhan FH, Barlogie B, Stewart AK. Gene expression profiling and multiple myeloma. Best Pr Res Cl Ha. 2005;18:537–52.
Chng WJ, Dispenzieri A, Chim CS, Fonseca R, Goldschmidt H, Lentzsch S, et al. IMWG consensus on risk stratification in multiple myeloma. Leukemia. 2014;28:269–77.
Johnson SK, Heuck CJ, Albino AP, Qu P, Zhang Q, Barlogie B, et al. The use of molecular-based risk stratification and pharmacogenomics for outcome prediction and personalized therapeutic management of multiple myeloma. Int J Hematol. 2011;94:321–33.
Boettcher S, Ebert BL. Clonal hematopoiesis of indeterminate potential. J Clin Oncol. 2019;37:419–22.
Steensma DP. Clinical consequences of clonal hematopoiesis of indeterminate potential. Blood Adv. 2018;2:3404–10.
Walker BA, Mavrommatis K, Wardell CP, Ashby TC, Bauer M, Davies F, et al. A high-risk, Double-Hit, group of newly diagnosed myeloma identified by genomic analysis. Leukemia. 2019;33:159–70.
Alaterre E, Raimbault S, Goldschmidt H, Bouhya S, Requirand G, Robert N, et al. CD24, CD27, CD36 and CD302 gene expression for outcome prediction in patients with multiple myeloma. Oncotarget. 2017;8:98931–44.
Li Z, Kang Y. Lipid metabolism fuels cancer’s spread. Cell Metab. 2017;25:228–30.
Newton JG, Horan JT, Newman S, Rossi MR, Ketterling RP, Park SI. CD36- positive B-lymphoblasts predict poor outcome in children with B-lymphoblastic leukemia. Pedia Dev Pathol. 2017;20:224–31.
Acknowledgements
We thank Dr. Michael Donovan, Director of Biorepository and Pathology CoRE, Saw (May) Li data collection, data management; Alexis De La Rosa, NERA summer student data collection; Allan Esperida, image editing; The Bone Marrow Laboratory Alexandra Yerzanskaya, Foxwell Emmons, Carolyn Nelson, our Myeloma Service at Mount Sinai Icahn School of Medicine, Bonnie Jenkins, Foundation Medicine and MyPRS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Hao, Y., Khaykin, D., Machado, L. et al. Bone marrow morphologic features, MyPRS, and gene mutation correlations in plasma cell myeloma. Mod Pathol 33, 188–195 (2020). https://doi.org/10.1038/s41379-019-0333-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-019-0333-6