Abstract
DNA was obtained from matching micro-dissected, primary tumor cells, paired metastases, and peripheral blood mononuclear cells (germline) from patients with appendiceal mucinous neoplasms. We compared specimens from patient cohorts comprising low-grade adenomucinous neoplasm versus high-grade mucinous adenocarcinoma using a targeted, amplicon sequencing panel of 409 cancer related genes (Ion Torrent Comprehensive Cancer Panel, Thermo-Fisher, Waltham, MA). Copy number variants, single nucleotide variants and small insertions/deletions were identified using a multiplex algorithm pipeline (GATK, VarScan2, MuTect2, SIFT, SIFT-INDEL, PolyPhen-2, Provean). There were significantly more damaging variants in high-grade versus low-grade tumor cohorts. Both cohorts contained damaging, heterozygous germline variants (catenin β1; notch receptor 1 and 4) in pathways associated with cell-lineage specification (WNT, NOTCH). Damaging, somatic KRAS proto-oncogene, GTPase mutations were present in both cohorts, while somatic GNAS complex locus mutations were confined to low-grade neoplasms. Variants predominantly affected transcription factors, kinases, and stem cell signaling molecules in canonical pathways including epithelial to mesenchymal transition, stem cell pluripotency, p53, PTEN, and NF-қB signaling pathways. High-grade tumors demonstrated MYC proto-oncogene, bHLH transcription factor (MYC) and death domain associated protein (DAXX) amplification and damaging somatic variants in tumor protein p53 (TP53), likely to amplify an aggressive phenotype. Damaging APC, WNT signaling pathway regulator (APC) deletions were identified in metastatic tissue of both cohorts suggesting a role in invasive disease. Our data suggest that germline dysregulation of WNT and/or NOTCH pathways predisposes patients toward a secretory cell phenotype (i.e., goblet-like cells) upon acquisition of somatic KRAS mutations. Additional somatically acquired variants activating oncogenes MYC and DAXX and inhibiting the critical tumor suppressor, tumor protein TP53, were consistent with manifestation of a high-grade phenotype. These additional changes within the epithelial to mesenchymal transition signaling network (WNT, NOTCH, RAS/ERK/PI3K, PTEN, NF-қB), produce aggressive high-grade tumor characteristics by actively driving cells towards dedifferentiation, proliferation, and migration.
Similar content being viewed by others
Introduction
Appendiceal mucinous neoplasms are rare, primary epithelial malignancies of the appendix, with approximately 1000–2000 cases diagnosed each year in the United States [1, 2]. Peritoneal metastasis from appendiceal mucinous neoplasms is referred to as pseudomyxoma peritonei (PMP), a clinical entity that is characterized by peritoneal dissemination of mucinous ascites and mucinous tumor nodules [3, 4].
The cytohistologic features of neoplastic cells within these mucinous tumors are major determinants of prognosis and have been used to stratify patients into various proposed classification systems [3,4,5,6,7,8,9,10,11,12,13]. The eighth edition of the American Joint Committee on Cancer staging manual has adopted diagnostic terminology from a recent consensus statement of the Peritoneal Surface Oncology Group International [3, 14]. With respect to primary tumors, the American Joint Committee on Cancer defines (a) low-grade appendiceal mucinous neoplasms (LAMN) as having low-grade cytology without infiltrative invasion; (b) high-grade appendiceal mucinous neoplasms (HAMN) as having high-grade cytology without infiltrative invasion; and (c) mucinous adenocarcinomas (MACA) as having infiltrative invasion, i.e., destructive stromal invasion into the wall of the appendix. Primary mucinous adenocarcinomas can be either moderately differentiated (Grade 2: G2) or poorly differentiated (Grade 3: G3). Grade 2 cancers exhibit high-grade cytology and infiltrative invasion, while Grade 3 cancers demonstrate infiltrative invasion and most have signet ring cell differentiation. With respect to metastatic peritoneal disease, the American Joint Committee on Cancer advocates a three-tiered classification system for peritoneal disease based on the presence or absence of specific cytohistologic features, similar to those described by Davison et al. and Shetty et al. [7, 13]. They define well-differentiated neoplasms (low-grade, G1) as demonstrating low-grade cytology, absence of infiltrative invasion, low tumor cellularity, and no signet ring cells; moderately differentiated mucinous adenocarcinomas (high-grade, G2) as having high cytologic grade in the absence of signet ring cells; and poorly differentiated mucinous adenocarcinomas (high-grade, G3) as having signet ring cells.
The molecular alterations responsible for malignant transformation and the unique phenotype of mucinous appendiceal neoplasm/pseudomyxoma peritonei remain unclear. Recent genomic studies, including next-generation sequencing studies of targeted cancer genes and whole exomes, have identified some commonly mutated genes and altered signaling pathways affecting appendiceal mucinous neoplasia [7, 15,16,17,18,19,20,21,22,23,24]. In general, low-grade tumors are frequently associated with concurrent KRAS proto-oncogene, GTPase (KRAS) and GNAS complex locus (GNAS) mutations, while high-grade tumors have been characterized by KRAS and Tumor Protein p53 (TP53) mutations, with lower rates of GNAS mutations. Other less commonly identified mutated cancer genes include SMAD family members (SMAD2/3/4), phosphatidylinositol-4–5-bisphosphate 3-kinase-catalytic subunit α (PIK3CA), AKT serine/threonine kinase 1 (AKT1), and catenin β1 (CTNNβ1), which impact downstream TGFβ, PI3K/AKT, and WNT signaling pathways.
We performed high-resolution sequencing of low-grade and high-grade appendiceal mucinous neoplasms targeting 409 genes that exhibit tumor suppressor or oncogene activity associated with cancer. We sought to identify genomic aberrations that could provide dependable predictive biomarkers and/or effective targets for therapy. The overriding goal of the study was to identify “driver mutations” or a convergence of disparate mutations upon signaling pathways to help decipher the molecular mechanisms underlying these tumors.
Materials and methods
Samples
Thirty matched samples of peripheral blood mononuclear cells, primary tumor tissue and metastatic tissue were obtained from ten patients with peritoneal metastases from low-grade and high-grade appendiceal mucinous neoplasms. Low-grade tumors included five, primary low-grade appendiceal mucinous neoplasms and their corresponding low-grade (G1) metastatic tissue. High-grade primary tumors included 4 moderately differentiated mucinous adenocarcinomas (G2) and one poorly differentiated mucinous adenocarcinoma (G3) and their concordant high-grade (G2 and G3) metastatic tissue. The morphology of paired metastases closely mirrored that of their matching primary tumors, including signet ring cells in both primary tumor and metastases from the highest-grade (G3) specimen (Fig. 1). No goblet cell tumors were included in this study. All patients provided written consent for tissue banking and genetic testing. All experimental procedures were approved by the University of Pittsburgh Institutional Review Board (IRB # 02-077). Blood was obtained for germline DNA from peripheral blood mononuclear cells while tissue specimens underwent formalin-fixation and paraffin embedding.
Morphology of low-grade and high-grade appendiceal mucinous tumor and corresponding metastasis. Micrographs in upper and lower left panels were derived from a representative low-grade (LG: LAMN) mucinous tumor region (T: LG) and a paired metastatic specimen (M: LG) stained with hematoxylin and eosin. Images in right column were from a high-grade (HG: MACA-SRC) tumor specimen (T: HG) and corresponding metastatic tissue (M: HG). Arrows in these images point to cells with a “signet ring” morphology. All images were captured at ×40 magnification (Olympus BX-50 microscope)
DNA purification
Blood samples (1 × 106 to 5 × 106 cells) were extracted with the QIAamp DNA Micro kit (Qiagen, Valencia, CA). Briefly, peripheral blood mononuclear cells were centrifuged to pellet cells (800 rpm × 5 min) and all but 50 µl of supernatant was removed. The cells were lysed under denaturing conditions (56 °C) using Proteinase K and buffer ATL. Buffer AL was added to optimize DNA binding to the membrane (Min-elute column). Contaminants were washed away using ethanol-containing buffers AW1 and AW2 and DNA was eluted in 30 µl of Low Tris-EDTA (10: 0.1 mM Tris EDTA, pH 8.0).
Tumor dissection
Microdissection of formalin-fixed, paraffin-embedded tissue was performed manually using an inverted microscope (TE200 Eclipse, Nikon Instruments, Melville, NY) to obtain ≥95% tumor cells without inflammatory cells, necrotic tissue or normal cells. Cells were scraped from unstained slides (10 µm thickness) aligned in register with serial hematoxylin and eosin (H and E) stained specimens. DNA was purified using the QIAamp Formalin Fixed Paraffin Embedded DNA protocol (Qiagen). Briefly, dissected tissue was rinsed in xylene and 100% ethanol and air-dried. Deparaffinized tissue was suspended in lysis buffer (ATL + 2 mg Proteinase K × 24 h @ 56 °C, 60 rpm on Innova 4000 shaker, Eppendorf, Enfield, CT) and lysed tissues were heated (90 °C × 1 h) and bound to a Min-elute column (buffer AL in 50% ethanol). DNA was washed with increasing percentages of ethanol (AW1 and AW2), eluted in low tris-EDTA and concentration and quality determined by fluorometry and spectrophotometry (pico green: Qubit 2.0 fluorometer, Thermo Fisher, Waltham, MA; ND-1000 spectrophotometer, NanoDrop, Wilmington, DE). Molecular size (20–150 Kb) was determined on the Bioanalyzer 2100 (12000 DNA Chip, Agilent Technologies, Santa Clara, CA). Only samples (>100 ng/µl) with OD 260/280 >1.8 and fragments ≥300 base pairs were used for library construction.
Generation of amplicon libraries
Primers amplified DNA covering 7404 exons (AmpliSeq Comprehensive Cancer Panel Primer Pools, Thermo Fisher) in equimolar amounts per specimen distributed in multiplex PCR reaction mixes (10 ng each, 4000 primer pairs/reaction, Ion AmpliSeq HiFi Master Mix ver. 2.0, Thermo Fisher). PCR was performed (99 °C × 2 min, 13–17 cycles @ 99 °C × 15 s, 60 °C × 8 min, 10 °C × hold; Perkin Elmer 9700, Bridgeville, PA) followed by enzymatic digestion and phosphorylation of the multiplexed amplicons with FuPa reagent (50 °C × 10 min, 55 °C × 10 min, 60 °C × 20 min). Amplicons were then mixed with proprietary switch solution, barcodes (Ion Xpress Barcode Adapters 1-16 kit), Ion P1 adapters and ligase (ligation: 22 °C × 30 min; inactivation: 72 °C × 10 min, 10 °C hold × 30 min) for end modification. Adapter-ligated libraries were purified (Agencourt AMPure beads: XP System, Beckman Coulter, Danvers, MA) and pellets suspended in master mix to enrich for amplicons with P1 and barcode adapters (98 °C × 2 min, 5 cycles @ 98 °C × 15 s, 60 °C × 1 min, 10 °C × hold). The amplicons were purified and resuspended in 50uL Low Tris-EDTA (0.06–6.00 ng/µl).
Emulsion PCR and sequencing
Multiplex barcoded libraries were combined with ion spheres for clonal amplification during emulsion PCR (60 cycles, PGM OT2 200 bp kit, One Touch 2, Thermo-Fisher). Ion spheres were recovered and stained with Alexa Fluor 488 or 647 for Qubit assessment of beads containing template (20–36% polyclonal templates). Libraries were enriched using biotin labeled primers bound to streptavidin-coated, metallic beads (DynaBeads C1, Thermo Fisher). Ion spheres were stripped from the streptavidin beads (125 mM NaOH; 0.1% Tween 20) and sequencing was performed on the Ion Torrent Personal Genome Machine (PGM, Thermo-Fisher) using the 200 bp sequencing kit v2 (318v1 chip, pH 7.55, 500 flows). The torrent software suite (Torrent Suite v. 4.0) parsed barcoded reads, assessed read quality, performed trimming and aligned base calls to HG19 using the TMAP aligner (Torrent Mapping Alignment Program) to generate BAM (binary format) files.
Data analysis pipeline
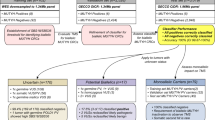
BAM files were realigned (GATK INDEL Realigner, Penn State University, State College, PA) and variant call files were generated using GATK (Genome Analysis Toolkit; UnifiedGenotyper v. 2.3) and VarScan 2 (http://dkoboldt.github.io/varscan/) in addition to the Torrent Suite variant caller (variant allele frequency cutoff ≥0.10) [25]. BAM files were also processed through an independent pipeline (Olympus Platform, GenomeNext, Columbus, OH) to identify germline and somatic mutations using MuTect 2 (https://www.broadinstitute.org/cancer/cga/mutect) [26]. Variant call files were filtered for overlapping calls using the GenomAnalytics application (GenomOncology Inc., Cleveland, OH) for non-synonymous, missense and nonsense mutations among matched samples including (1) single nucleotide substitutions and (2) small insertions or deletions <5 bases), (3) in somatic or germline samples, (4) in or out of the National Center for Biotechnology Information database of single nucleotide polymorphisms: https://www.ncbi.nlm.nih.gov/snp). Single nucleotide variants including substitutions and polymorphisms were filtered using PolyPhen-2, Provean and SIFT scores (Sorting Intolerant From Tolerant) for “star damaging” classification encompassing “possibly damaging”, “probably damaging,” and “damaging” variants [27, 28]. Insertion and deletion effects were analyzed using SIFT-INDEL and Provean algorithms to predict functional amino acid changes [29]. Variants identified by ≥2 of four callers underwent manual curation and validation using the Integrated Genomics Viewer (IGV, Broad Institute, Boston, MA) and annotation with multiple databases including ClinVar, Online Mendelian Inheritance In Man, Catalogue Of Somatic Mutations, National Cancer Institute Genome Data Commons [30].
Copy number analysis was performed using exon base counts for matched primary tumors, metastases and germline specimen base pileups. Briefly, total exon base calls were compiled after guanine-cytosine content correction and log2 normalization. Total base counts for matching primary tumors or metastases were compared for each position across identical bins (500 exons) and corrected by normalization using the mean ratio of primary tumor (or metastases) counts to matched germline values for each bin. Analysis of variance (ANOVA) was performed (Partek Genomics Suite, Partek Inc., St Louis, MO) for effects of tissue type on counts across germline, primary tumor, and metastatic samples. Post hoc comparisons (Student paired T-test) tested statistical significance corrected to a false discovery rate of 1% (q ≤ 0.01; p ≤ 0.005) where copy number variants exhibited a ≥1.5-fold change from germline values (−1.5≥ fold change ≥1.5). Copy number findings were confirmed by Nexus Copy Number software (v. 8.0, http://www.biodiscovery.com/nexus-copy-number/, Biodiscovery, Inc., El Segundo, CA), which performed normalization, segmentation, and identification of copy number events using a pooled “normal” reference file comprising germline BAM files for all patients. Two algorithms were used to define (1) statistically significant copy number alterations in individual samples (STAC: Significant Testing for Aberrant Copy number; p-value < 0.005) and (2) chromosomal regions with statistically significant gains and losses in cohorts (Q bound significance score = 0.05; GISTIC: Genomic Identification of Significant Targets in Cancer) [31].
Immunohistochemistry
Four-micron thickness, serial sections of formalin-fixed, paraffin-embedded tumors and metastases were stained with monoclonal antibodies for CTNNβ1 (mouse clone CTNNβ1, IgG1Κ, #M3539, Dako, Agilent Technologies) and c-MYC (rabbit clone Y69, IgG, #790–4628, Ventana Medical Systems, Tucson, Az). Deparaffinization through counterstaining was performed on the BenchMark Ultra Platform (Optiview DAB v3 kit; Ventana Medical Systems, Tucson, AZ). Slides were pretreated with Cell Conditioner 1 (CC1: # 950–124 @ 100 °C; [CTNNβ1: 24 min; c-MYC: 64 min] +OptiView Amplifier x 4 min) followed by incubation with primary antibody at 100 °C (CTNNβ1: 20 min, c-MYC: 28 min). Peroxidase blocking was performed with OptiView Peroxidase Inhibitor (3% H2O2) followed by the OptiView HQ Linker, Optiview HRP Multimer and OptiView DAB reagents according to the proprietary Ventana program. Counter staining was performed with Hematoxylin (8 min) and Bluing Reagent (buffered lithium carbonate: 4 min). Sections were assessed for the number of cells demonstrating positive staining by an anatomical pathologist blinded to the origin and classification stage of each sample.
Results
Patient characteristics
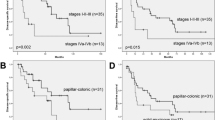
Disease progression and outcomes were markedly different between cohorts (Table 1). Patients with high-grade disease suffered recurrence with four of five succumbing to the disease (overall survival = 36.8 ± 12.7 months). In contrast, four of five patients with low-grade neoplasms were recurrence free at their 4-year follow-up appointment (53 ± 4.7 months) (Table 1).
Gains and losses (copy number variants)
Sequencing read depth averaged 184 ± 21 calls/base (Mean ± Std. dev) across the 7404 exons interrogated in this assay. ANOVA identified a significant effect of tissue type (peripheral blood mononuclear cells, primary tumors, metastases) on exon pileup counts, yielding more copy number events in high-grade tumors compared to low-grade tumors (p = 1.51 × 1021). This finding extended to metastatic specimens where copy number events were significantly higher in the metastases of high-grade tumors compared to metastatic specimens from the low-grade tumor cohort. Post-hoc STAC analysis identified significant gains in 96 genes and losses in 107 genes across low-grade primary tumors and significant gains in 177 genes and losses in 193 genes in high-grade tumors affecting at least 1 sample in each primary tumor cohort. The largest number of copy number variants was found in genes classified as transcriptional regulators, followed by kinases and enzymes. (IPA: version 28820210, Qiagen, Redwood City, CA) [32].
Shared gene amplifications in low-grade and high-grade tumors
Six amplifications were found in both tumor cohorts with incrementally higher magnitude and frequency in high-grade specimens (GISTIC 2.0: q < 0.05; Average Copy Number: Low grade = 2.7 ± 0.2; High grade = 3.1 ± 0.1) (Fig. 2). Four regions encoded cancer-related transcription factors, including nuclear factor, erythroid 2 like 2 (NFE2L2; chr 2: q31.2), cyclin E1 (CCNE1; chr 19: q12), Kruppel-like factor 6 (KLF6; chr 10, p15.1) and menin1 (MEN I; chr 11: q13.1). Two amplifications affected signal transduction molecules GNAS complex locus (GNAS; chr12: q13.32) and heat shock protein 90 alpha family class B member 1 (HSP90AB1; chr 14: q32.31). These amplifications were also identified in each of the metastatic specimens with the exception of KLF6 (Fig. 2).
Copy number variants in low-grade and high-grade appendiceal mucinous neoplasms. Six genes exhibited statistically significant amplifications (GISTIC 2.0: q < 0.05; ≥3 copies) in high-grade tumors (left panel, black bars). (See Results for gene names and chromosome location) The same genes from the low-grade cohort are included for comparison where the samples were significant by GISTIC analysis (q < 0.05) but did not exceed CN threshold for biological significance (≥3 copies) except MEN1. Metastases had copy number profiles similar to their primary tumors with statistically significant gains (GISTIC 2.0: q < 0.05; ≥3 copies) spanning the same gene domains in the high-grade tumors except KLF6 (right panel). Bars indicate copy number average of 5 individuals per cohort compared to 10 germline samples as the pooled normal reference value except where numbers 4 or 3 indicate a reduced number of samples were detected by GISTIC. Vertical error bars indicate the standard deviation for each gene copy number per cohort
Copy number variants unique to low-grade tumors
There were copy number alterations in only three genomic domains that affected low-grade neoplasms in three cases each but no high-grade tumors (Table 2). These included amplification of transcription factor 7-like 2 (TCF7L2: chr 10: q25.2) which plays a key role in the WNT/β-catenin signaling pathway [33]. There were copy number losses in two genes on chromosome X in three of the five low-grade samples comprising the androgen receptor (AR: chr X: q12) and SH2 domain-containing protein 1A (SH2D1A: chr X: q25) genes. These two losses and the amplification in TCF7L2 were also present in the three correlative metastatic specimens of this cohort.
Copy number variants unique to high-grade tumors
There were copy number alterations in five genomic domains affecting high-grade primary tumors (≥3 specimens) (Table 2). Two transcription factors, MYC proto-oncogene, bHLH transcription factor (Myc; chr 8: q24.21) and death domain associated protein (DAXX; chr 6: q21.32), exhibited amplifications in five and four patients respectively. The Pim1-proto-oncogene, serine/threonine kinase (PIM1; chr 6: p21.2) was significantly amplified in high-grade tumors. The matching metastatic specimens to these primary tumors of the high-grade cohort also contained these amplifications (MYC: n = 5, DAXX: n = 4; PIM1: n = 4). Two transcription factors, which regulate pluripotency versus specification of cell fate, POU class 5 homebox 1 (POU5F1; chr 6: p21.33) and GATA binding protein 2 (GATA2; chr 3: 21.3), exhibited significant copy number losses within high-grade tumors.
Somatic single nucleotide variants
Tumor-specific, damaging, non-synonymous single nucleotide variants were detected in KRAS and GNAS genes associated with cancer (COSMIC, ClinVar database) in low-grade tumors (≥3 specimens) (Table 3a). All five low-grade tumors contained a KRAS missense mutation (exon 2: rs121913529: G12A, G12D, G12V) while four patients in this cohort had concurrent damaging missense mutations in the GNAS gene (exon 1: S187A, exon 8: rs11554273: R844C, rs121913495: R844H). These KRAS mutations have been associated with increased RAS activity and pathogenic effects across a multitude of cancers, including mucinous appendix and colorectal cancer [34]. All five high-grade tumors exhibited activating missense mutations in the KRAS gene (rs121913529: G12D, G12V; rs121913530: G12S) but we detected no deleterious single nucleotide variants in the GNAS gene in the high-grade cohort. Three different sets of high-grade tumors contained damaging, missense substitutions in the MYB proto-oncogene, transcription factor (MYB) and tumor protein p53. MYB variants were damaging by SIFT, PolyPhen and Provean algorithms but were not reported in Cosmic or ClinVar databases. In contrast, each TP53 variant was “pathogenic” in both databases with mutations predicted to disrupt DNA binding in a variety of cancers (exon 4: rs28934578: R175H, exon 4: rs587780070, H179D, exon 6: rs11540652: R248W) [35].
Germline single nucleotide variants
There were shared, heterozygous germline variants in both low- and high-grade cohorts affecting two genes in corresponding germline, tumor and metastatic samples. (Table 3a) Three patients in the low-grade and four in the high-grade cohort exhibited an oncogenic mutation in catenin β1 (CTNNβ1) that blocks negative regulation of cytoplasmic CTNNβ1 levels through phosphorylation dependent degradation (exon 5: I198F). Cytoplasmic accumulation of CTNNβ1 comprised a gain of function activation of the WNT signaling pathway demonstrated in human colorectal cell lines [36]. All low-grade and two high-grade specimens contained a notch receptor 1 (NOTCH1) damaging, missense mutation responsible for degradation of the active notch intracellular domain (NICD) (exon 34: P2142H). These notch receptor 1 mutations increased NICD half-life resulting in constitutive notch receptor pathway activation [37].
Damaging insertions/deletions
Damaging insertions and deletions were infrequent. Four of five patients in each cohort exhibited the same germline deletions in notch receptor 4 (NOTCH4) resulting in potentially deleterious modifications of the signal peptide domain undetectable by nonsense mediated decay mechanisms (Table 3b). We also identified two different damaging frameshift deletions in the APC, WNT signaling pathway regulator (APC) gene exclusive to metastatic specimens of four low-grade (COSMIC: 18992) and four high-grade patients (COSMIC: 18730) resulting in truncations in nearby regions of the phosphoserine domain. These APC deletions have been previously identified in gastric cancer and were projected to escape nonsense mediated decay detection. (Supplement 1).
Pathway analysis
There was marked overlap of top ranked pathways in both tumor cohorts revealed through Ingenuity Pathways Analysis (IPA: version 28820210: Canonical Pathways, Qiagen, Hilden, Germany) when all significant copy number changes and all non-synonymous, germline and somatic single nucleotide variants and insertions/deletions found in more than one specimen were included. The top 10 canonical pathways for both groups comprised 14 distinct pathways with more molecules present (higher mutation load) in the high-grade cohort (Supplement Table 1). Including only damaging/pathogenic variants (germline and somatic) resulted in convergence into epithelial to mesenchymal transformation, embryonic stem cell pluripotency, PTEN and P53 signaling as the top canonical pathways for both cohorts. We dissected out variants in these pathways to account for phenotype similarities (shared variants) and differences (unique variants) between low and high-grade cohorts. For example, both cohorts exhibited activating mutations in CTNNβ1, NOTCH1 and KRAS genes as fundamental members of the WNT (cadherin E, frizzled class receptor 1 and PKA arms), NOTCH and KRAS-driven MAPK/ERK/PI3K signaling systems (Fig. 3). In contrast, damaging variants in AKT and TP53 combined with copy number alterations in MYC, DAXX and POU5F1 occurred predominantly in the high-grade cohort (Fig. 3). Immunohistochemical studies confirmed elevated expression of CTNNβ1 in both low and high-grade tumors and metastases (Fig. 4). CTNNβ1 accumulated both in the cytoplasm and nucleus associated with the exon 5, I198F mutation, which underlies this gain of function variant [36]. IHC expression studies confirmed that MYC was elevated predominantly in high-grade tumors and metastases consistent with the identification of copy number amplifications in this cohort (Fig. 4, Table 2).
Predominant Signaling and Transcriptional Pathways Associated with Pathogenic Genomic Alterations. Green indicates genes with germline or somatic variants (single nucleotide variants, insertions/deletions, copy number variants) enriched in both low- and high-grade specimens (Ingenuity Pathways Analysis, see Methods and Supplement Table 1). The blue gradient circle is specific to low-grade tumors (GNAS: GαS subunit) and red gradient circles are specific to high-grade tumors. Mutations in E-cadherin (CDH-E), Frizzled, APC and β-Catenin (CTNNβ) predict activation of cell survival through the WNT Pathway as part of the Epithelial to Mesenchymal Transition network. Nuclear CTNNβ binds TCF/LEF transcription factors recruiting multiple co-activators (BIRC5: baculoviral IAP repeat containing 5, CCND1, COX2: cyclo-oxygenase 2, cJUN: Jun proto-oncogene, AP-1 transcription factor subunit, MMP: matrix metalloproteinase). Copy number losses in POU5F1 and single nucleotide variants in TCF3/4 indicate aberrant regulation of pluripotency and cell survival driven by CTNNβ in high-grade tumors. Mutated NOTCH signaling proteins fail to cleave the Notch ligand (NICD: nuclear intracellular domains) activating the NOTCH pathway and promoting interactions with c-MYC which is amplified in high-grade tumors. Activating KRAS mutations increase cell motility through the MAP kinase (MAPK/ERK: KRAS-BRAF-MEK1/2-ERK1/2) pathway in all tumors and HRAS proto-oncogene mutations through the PI3K (phosphatidylinositol-4,5-bisphosphate 3-kinase) gene and pathway (HRAS-PI3K-AKT). KRAS, RALGDS (Ral guanine nucleotide dissociation stimulator), RHO and JNK (mitogen-activated protein kinase 8) mutations enhance cell survival by inhibition of apoptosis. Amplification of DAXX and BCL2 through the FAS (Fas cell surface death receptor) and p53 pathways respectively block apoptosis in high-grade tumors
Expression of CTNNβ1 and c-MYC in low-grade versus high-grade appendiceal mucinous neoplasms. Micrographs were obtained from a representative low-grade mucinous tumor (T: LG, LAMN) and a high-grade mucinous tumor (T: HG; MACA-SRC). All tumors and metastases (not shown) exhibited significant CTNNβ1 expression regardless of grade. MYC staining was rare and patchy (n = 1; <10% of cells) detected in only one low-grade tumor while three of four high grade tumors tested demonstrated substantial MYC expression (5 to 50% of cells) with dense staining in the correlative metastatic tissues of all four high grade specimens that underwent testing
Discussion
We interrogated DNA from matching germline cells, primary tumors and metastases of low- and high-grade appendiceal mucinous neoplasms. These tumors are characterized by a unique extracellular, mucin-secreting phenotype prone to peritoneal dissemination rather than distant visceral metastasis. Paired tumors and metastases in both low- and high-grade cohorts contained several overlapping germline and somatic gain of function variants in genes of the WNT, NOTCH and MAPK signaling pathways with a higher mutation load in high- versus low-grade tumors. Common, heterozygous germline variants (CTNNβ1, NOTCH1 and 4) in both patient cohorts indicated a potential hereditary predisposition towards development of these tumors. The variant profile of metastases mirrored their correlative primary tumors with the addition of a damaging mutation in the APC gene. Variants in both tumor cohorts clustered in canonical signaling pathways associated with epithelial to mesenchymal transition, embryonic stem cell pluripotency and cell-fate specification (WNT/CTNNβ1, NOTCH, RAS/MAPK/ERK, PTEN, P53, NF-ĸB), critical networks with oncogenic potential including dedifferentiation, proliferation and migration [32,33,34,35,36,37]. In contrast to low-grade specimens, the high-grade cohort demonstrated alterations in a set of oncogenes and tumor suppressors (copy number gain: MYC, DAXX, PIM1; copy number loss: POU5F1; single nucleotide variant: TP53, MYB) that may help discriminate between low- and high-grade tumor phenotypes and provide new therapeutic targets for patients confronting the poor prognosis associated with high-grade appendiceal mucinous tumors (Table 4).
These data corroborate and extend previous studies demonstrating concurrent GNAS and KRAS activating mutations in low-grade appendiceal mucinous neoplasms, with infrequent GNAS mutations in high-grade tumors [15, 17, 18, 23]. GNAS encodes the α-subunit of the central G protein signal transduction complex and activating mutations are implicated in mucin-producing pancreatic intraductal papillary mucinous neoplasms, mucinous colorectal and appendix cancers [9, 15, 17, 18]. We identified statistically significant copy number amplifications (q < 0.05) spanning the GNAS domain in both cohorts that just surpassed our arbitrary cutoff for biological relevance in the high-grade cohort (+1.5 fold). Consequently, we cannot rule out a role for GNAS activation in this cohort. We detected KRAS G12 mutations in both tumor cohorts corroborating previous reports implicating KRAS-driven MAPK/ERK/PI3K signaling in mucin production in mucinous appendix and colorectal cancer [18, 34, 38]. For example, the Cancer Genome Atlas data for mucinous colorectal cancers contained higher rates of KRAS mutations compared to non-mucinous cancers, suggesting distinct molecular pathogenesis of these phenotypes and indicating that aberrant activation of MAPK/ERK/PI3K signaling pathways may promote mucinous tumorigenesis [39, 40].
Along with KRAS activation of RAS signaling pathways, aberrant stimulation of highly conserved developmental pathways, WNT and NOTCH, played a critical role in the manifestation of these tumors. Specifically, we identified damaging mutations in CTNNβ1 and NOTCH1 and 4 protein domains critical for degradation of these transcriptional regulators that otherwise accumulate in the cytoplasm and induce cell proliferation. Patients in both cohorts shared a potentially disruptive CTNNβ1 mutation (exon 5: I198F) in the second armadillo repeat immediately upstream of the axin binding site. Failure of axin binding inhibits formation of the “destruction complex” that catalyzes phosphorylation and targeting of CTNNβ1 for proteasomal degradation [36]. Similarly, all specimens contained a damaging NOTCH1 mutation (exon 34: P2142H) in the cytoplasmic PEST domain (C-terminal rich in Proline [P], glutamic acid [E], serine [S] and threonine [T]) which compromises degradation of active NICD (notch intracellular domain) [37]. Increased CTNNβ1 levels have been identified in endometrial, hepatocellular, ovarian, prostate and colon cancers while elevated NOTCH1 was found in breast, gastric, pancreatic and colon cancers [41,42,43]. We also identified damaging somatic truncations in the APC gene (exon 14/16/17) of metastatic tissues. These truncations altered the c-terminal domain of the APC gene important to cell migration and chromosomal segregation in colorectal cancer patients [44].
MYC amplifications and somatic TP53 mutations were prevalent in high-grade tumors. These accompanied gain of function mutations in CTNNβ1, NOTCH, and KRAS known to induce constitutive activation of MYC which is otherwise under the stringent control of WNT, NOTCH and MAPK/ERK signaling mechanisms [22, 45, 46]. Disruption of this tight homeostatic regulation coupled with significant MYC amplification in high-grade tumors suggested that promiscuous MYC activation may exacerbate the pathogenic phenotype. Acute overexpression of MYC alone in normal cells triggers checkpoints including TP53 that lead to cell growth arrest and apoptosis [46,47,48]. However, several high-grade tumors also demonstrated TP53 mutations which have been linked previously to elevated CTNNβ1 driving MYC expression, cancer stem cell-like features and carcinogenesis [48]. Site directed mutagenesis studies have established that the high-grade tumor mutations of TP53 found in this study caused significantly diminished (H179D) or totally defective DNA binding transactivation (R175H and R248W) resulting in loss of function [49]. Consequently, the combination of MYC deregulation coupled with KRAS mutations and/or loss of TP53 function in high-grade tumors may drive the aggressive phenotype and poor outcome in this cohort. Ongoing therapeutic targeting studies have been directed at MYC inhibition (e.g., JQ1, GSK525762) or depleting mutant TP53 (e.g., PRIMA-1, APR-246) in a variety of cancers [50, 51]. These studies may yield a novel approach to treat high-grade tumors while multiple small molecules are being tested to counteract mutations which drive WNT, NOTCH, and RAS pathways underlying the general appendiceal mucinous tumor phenotype [52,53,54].
We employed special methodological precautions because of the rarity of these samples. We utilized a stringent statistical threshold for selection of genomic aberrations comprising reliance on multiple variant callers, high base call depth (≥50), high quality scores (≥Phred 30) and high variant allele frequency cutoffs (vaf: ≥0.10). Targeted amplicon sequencing required less substrate (DNA ≤ 10 ng) and cost significantly less than exome and whole genome sequencing while providing higher base call depths and reduced analysis burden. Tumor preparation by formalin fixation and paraffin embedding introduced additional technical challenges beyond tissue heterogeneity (normal, necrotic, inflammatory cells) and cellular clonality in the form of DNA crosslinking, oxidation, deamination and degradation. We optimized our dissection, extraction, library preparation, QC methods and analysis pipelines to obtain high fidelity results from formalin-fixed, paraffin-embedded tissue. Careful selection and microdissection are critical to minimize cellular heterogeneity in these tumors. The contrasting results obtained by PCR versus sequencing for KRAS in the signet ring cell tumor may reflect different tumor regions from which the DNA was obtained for clinical diagnosis as opposed to this research study. We also performed a 24-hour lysis reaction while spiking proteinase K (12-h intervals) to reduce cross-linking and maximize yields. We established minimal fragment size for this assay (DNA > 300 bp) to assure high fidelity priming and replication during initial amplicon library preparation by comparison to a National Institute of Standards and Technology reference sample (HG003 NA24149; Ashkenazi Father, Genome In A Bottle Consortium) prepared by formalin fixation followed by paraffin embedding [55]. We also limited high-temperature extremes to reduce cytosine deamination and minimize errors associated with cytosine hydrolysis. Finally, we confirmed NGS assay efficacy by validating all GNAS, KRAS, CTNNβ1, NOTCH4 mutations and MYC and PIM1 copy number amplifications using Sanger sequencing, immunohistochemistry and digital droplet PCR (QX200, Bio-Rad, Hercules, CA) (Supplement 2).
In summary, these results support the hypothesis that appendiceal mucinous tumors arise from activating mutations in CTNNβ1, NOTCH1, NOTCH4, and KRAS that are important components of constitutive WNT, NOTCH, and RAS signaling pathways. A higher mutation load including additional alterations specific to high-grade tumors was identified (MYC, DAXX, PIM1, POU5F1, TP53, and MYB) that represent potential diagnostic and therapeutic targets for this deadly disease. It is tempting to speculate based on these studies that the low-grade phenotype progresses to the high-grade tumor through the accumulation of oncogenic mutations and/or genomic amplifications that include a major oncogene such as MYC. However, direct manipulation of critical target genes and pathways using an animal or cell culture model will be required to determine if the low-grade neoplastic phenotype progresses to a high-grade form or is an independent molecular entity. We hope future studies in larger patient cohorts including fresh or frozen tissue will validate the variants we identified as characteristic of low-grade or high-grade appendiceal mucinous neoplasms with sufficient power for prognostic classification based on phenotype and outcome.
References
McCusker ME, Cote TR, Clegg LX, Sobin LH. Primary malignant neoplasms of the appendix: a population-based study from the surveillance, epidemiology and end-results program, 1973–1998. Cancer. 2002;94:3307–12.
Turaga KK, Pappas SG, Gamblin T. Importance of histologic subtype in the staging of appendiceal tumors. Ann Surg Oncol. 2012;19:1379–85.
Carr NJ, Cecil TD, Mohamed F, Sobin LH, Sugarbaker PH, Gonzalez-Moreno S, et al. Peritoneal Surface Oncology Group I. A Consensus for Classification and Pathologic Reporting of Pseudomyxoma Peritonei and Associated Appendiceal Neoplasia: The Results of the Peritoneal Surface Oncology Group International (PSOGI) Modified Delphi Process. Am J Surg Pathol. 2016;40:14–26.
Carr NJ, Arends MJ, Deans GT, Sobin LH. Adenocarcinoma of the Appendix. In: Bosman FT CC, Hruban RH, Theise ND, editors. WHO classification of tumors of the digestive system. Lyon: IARC Press; 2010. pp. 95–8.
Bradley RF, Stewart JH, Russell GB, Levine EA, Geisinger KR. Pseudomyxoma peritonei of appendiceal origin: a clinicopathologic analysis of 101 patients uniformly treated at a single institution, with literature review. Am J Surg Pathol. 2006;30:551–9.
Carr NJ, McCarthy WF, Sobin LH. Epithelial noncarcinoid tumors and tumor-like lesions of the appendix. A clinicopathologic study of 184 patients with a multivariate analysis of prognostic factors. Cancer. 1995;75:757–68.
Davison JM, Choudry HA, Pingpank JF, Ahrendt SA, Holtzman MP, Zureikat AH, et al. Clinicopathologic and molecular analysis of disseminated appendiceal mucinous neoplasms: identification of factors predicting survival and proposed criteria for a three-tiered assessment of tumor grade. Mod Pathol. 2014;27:1521–39.
Misdraji J, Yantiss RK, Graeme-Cook FM, Balis UJ, Young RH. Appendiceal mucinous neoplasms: a clinicopathologic analysis of 107 cases. Am J Surg Pathol. 2003;27:1089–103.
Pai RK, Beck AH, Norton JA, Longacre TA. Appendiceal mucinous neoplasms: clinicopathologic study of 116 cases with analysis of factors predicting recurrence. Am J Surg Pathol. 2009;33:1425–39.
Pai RK, Longacre TA. Appendiceal mucinous tumors and pseudomyxoma peritonei: histologic features, diagnostic problems, and proposed classification. Adv Anat Pathol. 2005;12:291–311.
Ronnett BM, Yan H, Kurman RJ, Shmookler BM, Wu L, Sugarbaker PH. Patients with pseudomyxoma peritonei associated with disseminated peritoneal adenomucinosis have a significantly more favorable prognosis than patients with peritoneal mucinous carcinomatosis. Cancer. 2001;92:85–91.
Ronnett BM, Zahn CM, Kurman RJ, Kass ME, Sugarbaker PH, Shmookler BM. Disseminated peritoneal adenomucinosis and peritoneal mucinous carcinomatosis. A clinicopathologic analysis of 109 cases with emphasis on distinguishing pathologic features, site of origin, prognosis, and relationship to “pseudomyxoma peritonei”. Am J Surg Pathol. 1995;19:1390–408.
Shetty S, Natarajan B, Thomas P, Govindarajan V, Sharma P, Loggie B. Proposed classification of pseudomyxoma peritonei: influence of signet ring cells on survival. Am Surg. 2013;79:1171–6.
Overman MJ, Asare EA, Compton CC, Hanna NN, Kakar S, Kosinski LA, et al. Appendix Carcinoma. In: Amin MB, Edge S, Greene F, Byrd DR, Brookland RK, Washington MK, et al. editors. AJCC Cancer Staging Manual, 8th edn. Chicago, IL: Springer; 2017. pp. 237-250.
Alakus H, Babicky ML, Ghosh P, Yost S, Jepsen K, Dai Y, et al. Genome-wide mutational landscape of mucinous carcinomatosis peritonei of appendiceal origin. Genome Med. 2014;6:43.
Grady WM, Pritchard CC. Molecular alterations and biomarkers in colorectal cancer. Toxicol Pathol. 2014;42:124–39.
Liu X, Mody K, de Abreu FB, Pipas JM, Peterson JD, Gallagher TL, et al. Molecular profiling of appendiceal epithelial tumors using massively parallel sequencing to identify somatic mutations. Clin Chem. 2014;60:1004–11.
Nishikawa G, Sekine S, Ogawa R, Matsubara A, Mori T, Taniguchi H, et al. Frequent GNAS mutations in low-grade appendiceal mucinous neoplasms. Br J Cancer. 2013;108:951–8.
Noguchi R, Yano H, Gohda Y, Suda R, Igari T, Ohta Y, et al. Molecular profiles of high-grade and low-grade pseudomyxoma peritonei. Cancer Med. 2015;4:1809–16.
Nummela P, Saarinen L, Thiel A, Jarvinen P, Lehtonen R, Lepisto A, et al. Genomic profile of pseudomyxoma peritonei analyzed using next-generation sequencing and immunohistochemistry. Int J Cancer. 2015;136:E282–9.
Pietrantonio F, Perrone F, Mennitto A, Gleeson EM, Milione M, Tamborini E, et al. Toward the molecular dissection of peritoneal pseudomyxoma. Ann Oncol. 2016;27:2097–103.
Shetty S, Thomas P, Ramanan B, Sharma P, Govindarajan V, Loggie B. Kras mutations and p53 overexpression in pseudomyxoma peritonei: association with phenotype and prognosis. J Surg Res. 2013;180:97–103.
Singhi AD, Davison JM, Choudry HA, Pingpank JF, Ahrendt SA, Holtzman MP, et al. GNAS is frequently mutated in both low-grade and high-grade disseminated appendiceal mucinous neoplasms but does not affect survival. Hum Pathol. 2014;45:1737–43.
Szych C, Staebler A, Connolly DC, Wu R, Cho KR, Ronnett BM. Molecular genetic evidence supporting the clonality and appendiceal origin of Pseudomyxoma peritonei in women. Am J Pathol. 1999;154:1849–55.
Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD, Lin L, et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 2012;22:568–76.
Kelly BJ, Fitch JR, Hu Y, Corsmeier DJ, Zhong H, Wetzel A, et al. Churchill: an ultra-fast, deterministic, highly scalable and balanced parallelization strategy for the discovery of human genetic variation in clinical and population-scale genomics. Genome Biol. 2015;16:6.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
Ng PC, Henikoff S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003;31:3812–14.
Hu J, Ng PC. SIFT Indel: predictions for the functional effects of amino acid insertions/deletions in proteins. PLoS ONE. 2013;8:e77940.
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, et al. Integrative genomics viewer. Nat Biotechnol. 2011;29:24–6.
Mermel CH, Schumacher SE, Hill B, Meyerson ML, Beroukhim R, Getz G. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 2011;12:R41.
Krämer A, Green J, Pollard J Jr, Tugendreich S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics. 2014;30:523–30.
Polakis P Wnt signaling in cancer. Cold Spring Harb Perspect Biol. 2012; 4.
Mizutani N, Ito H, Hagiwara K, Kobayashi M, Hoshikawa A, Nishida Y, et al. Involvement of KRAS G12A mutation in the IL-2-independent growth of a human T-LGL leukemia cell line, PLT-2. Nagoya J Med Sci. 2012;74:261–71.
Rotter V, Prokocimer M. p53 and Human Malignancies. Adv Cancer Res. 1991;57:257–72.
Ilyas M, Tomlinson IPM, Rowan A, Pignatelli M, Bodmer WF. B-Catenin mutations in cell lines established from human colorectal cancers. Proc Natl Acad Sci USA. 1997;94:10330–4.
Wang K, Zhang Q, Li D, Ching K, Zhang C, Zheng X, et al. PEST domain mutations in Notch receptors comprise an oncogenic driver segment in triple-negative breast cancer sensitive to a γ-secretase inhibitor. Clin Cancer Res. 2015;21:1487–96.
Dilly AK, Song X, Zeh HJ, Guo ZS, Lee YJ, Bartlett DL, et al. Mitogen-activated protein kinase inhibition reduces mucin 2 production and mucinous tumor growth. Transl Res. 2015;166:344–54.
Furukawa T, Kuboki Y, Tanji E, Yoshida S, Hatori T, Yamamoto M, et al. Whole-exome sequencing uncovers frequent GNAS mutations in intraductal papillary mucinous neoplasms of the pancreas. Sci Rep. 2011;1:161.
Cancer Genome Atlas N. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–7.
Hugen N, Simons M, Halilovic A, van der Post RS, Bogers AJ, Marijnissen-van Zanten MA. et al. The molecular background of mucinous carcinoma beyond MUC2. J Pathol Clin Res2015;1:3–17.
Tai D, Wells K, Arcaroli J, Vanderbilt C, Aisner DL, Messersmith WA, et al. Review: Targeting the WNT Signaling Pathway in Cancer Therapeutics. Oncologist. 2015;10:1189–98.
Aster JC, Pear WS, Blacklow SC. The Varied Roles of Notch in Cancer. Annu Rev Pathol. 2017;12:245–75.
Hankey W, Frankel WL, Groden J. Review. Functions of the APC tumor suppressor protein dependent and independent of canonical WNT signaling: implications for therapeutic targeting. Cancer Metastasis Rev. 2018;1:159–72.
Dang CV. MYC on the path to cancer. Cell.. 2012;149:22–35.
Castell A, Larsson LG. Targeting MYC Translation in Colorectal Cancer. Cancer Discov. 2015;5:701–3.
Schmitt CA, McCurrach ME, de Stanchina E, Wallace-Brodeur RR, Lowe SW. INK4a/ARF mutations accelerate lymphomagenesis and promote chemoresistance by disabling p53. Genes Dev. 1999;13:2670–7.
Wood LD, Parsons DW, Jones S, Lin J, Sjoblom T, Leary RJ, et al. The genomic landscapes of human breast and colorectal cancers. Science. 2007;318:1108–13.
Kato S, Han SY, Liu W, Otsuka K, Shibata H, Kanamaru R, et al. Understanding the function–structure and function–mutation relationships of p53 tumor suppressor protein by high-resolution missense mutation analysis. PNAS. 2003;100:8424–9.
Chen H, Liu H, Qing G. Targeting oncogenic Myc as a strategy for cancer treatment. Signal Transduct Target Ther. 2018;3:5.
Duffy MJ, Synnott NC, Crown J. Mutant p53 as a target for cancer treatment. Eur J Cancer. 2017;83:258–65.
Krishnamurthy N, Kurzrock R. Targeting the Wnt/beta-catenin pathway in cancer: Update on effectors and inhibitors. Cancer Treat Rev. 2018;62:50–60.
Li L, Tang P, Li S, Qin X, Yang H, Wu C, et al. Notch signaling pathway networks in cancer metastasis: a new target for cancer therapy. Med Oncol. 2017;34:180.
Bahrami A, Hassanian SM, ShahidSales S, Farjami Z, Hasanzadeh M, Anvari K, et al. signaling pathway as a potential therapeutic target in the treatment of colorectal cancer. J Cell Physiol. 2018;233:2058–66.
Zook JM, Catoe D, McDaniel J, Vang L, Spies N, Sidow A, et al. Extensive sequencing of seven human genomes to characterize benchmark reference materials. Sci Data. 2016;3:160025.
Acknowledgements
We express our sincere gratitude to Clinton Miller from GenomOncology (Cleveland, OH) for providing access to the GenomAnalytics application which was instrumental in sorting and characterizing all variants in this study. Also, we appreciate the considerable efforts of James Hirmas, Christopher Dresher and Matthew Porter of GenomeNext (Columbus, OH) who performed an independent parallel variant analysis from the raw files using the Olympus platform to validate and extend the results of this study. We greatly appreciate the support of David Eaves and Cathy Valley from Bio-Rad Laboratories for their help in performing digital droplet PCR analysis of the copy number variant results obtained in this study. This project utilized the University of Pittsburgh Hillman Cancer Center shared resource facility (Cancer Genomics Facility) supported in part by award P30CA047904 (Dr. LaFramboise).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
LaFramboise, W.A., Pai, R.K., Petrosko, P. et al. Discrimination of low- and high-grade appendiceal mucinous neoplasms by targeted sequencing of cancer-related variants. Mod Pathol 32, 1197–1209 (2019). https://doi.org/10.1038/s41379-019-0256-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-019-0256-2
This article is cited by
-
The genetic profile and molecular subtypes of human pseudomyxoma peritonei and appendiceal mucinous neoplasms: a systematic review
Cancer and Metastasis Reviews (2023)
-
Molecular and clinicopathological features of appendiceal mucinous neoplasms
Virchows Archiv (2021)