Abstract
Universal screening for Lynch syndrome in colorectal cancer is recommended, and immunohistochemistry for the mismatch repair proteins is commonly used. To reduce cost, some screen using only MSH6 and PMS2, with reflex to the partner stain if either are absent (two-stain method). An expression pattern revealing absent MSH2 and intact MSH6 is not expected, but could result in failed Lynch syndrome detection. We analyzed tumors with absent MSH2 but any degree of MSH6 expression to determine if the two-stain method could miss MSH2 mutations. One-thousand seven-hundred thirty colorectal cancer patients from the Ohio Colorectal Cancer Prevention Initiative underwent tumor screening using microsatellite instability and immunohistochemistry. The two-stain method was used for 1235 cases; staining for all four proteins was completed for 495 cases. The proportion of positive cells and staining intensity were reviewed for MSH6, as well as MSH2 when available. Patients with mismatch repair deficiency underwent next-generation sequencing of germline DNA for mismatch repair genes. If negative, tumor next-generation sequencing was performed to assess for somatic mutations. Overall, thirty-three (1.9%, 33/1730) MSH2-absent cases were identified. Of those, fourteen had no MSH6 expression but eight (0.5%, 8/1730) had ambiguous and eleven (0.6%, 11/1730) had convincing MSH6 expression that could have been interpreted as intact. Germline next-generation sequencing identified MSH2 mutations in 11/14 cases with absence of both stains, 7/8 cases with ambiguous MSH6 expression, and 9/11 cases with convincing MSH6 expression. All remaining cases, except one, had double somatic mutations. The two-stain method fails to detect some patients with Lynch syndrome: (1) significant staining weaker than the control may be incorrectly interpreted as intact MSH6, or (2) Weak or focal/patchy MSH6 can be retained with the absence of MSH2. Accordingly, we recommend the four-stain method be used for optimal Lynch syndrome screening detection.
Similar content being viewed by others
Introduction
Up to 15% of colorectal carcinomas have defective mismatch repair, characterized by microsatellite instability [1]. Defective mismatch repair is either acquired (the most common mechanism being MLH1 promoter methylation) or inherited (Lynch syndrome). Lynch syndrome accounts for at least 3% of all colorectal cancer cases and is caused by a germline mutation in one of the four mismatch repair genes (MLH1, MSH2, MSH6, PMS2) or EPCAM deletion [2, 3]. Individuals with Lynch syndrome have high lifetime risks for colorectal cancer, as well as increased risks for cancers of the endometrium, stomach, ovary and others [4]. Acquired somatic mutations in the mismatch repair genes mimic the features seen in defective mismatch repair colorectal cancer caused by Lynch syndrome, but are not thought to be associated with increased risk for metachronous cancers and typically cannot be transmitted from parent to offspring [5,6,7,8]. Regardless of whether the defective mismatch repair colorectal cancer is caused by inherited or acquired mutations, the patient may still benefit from immunotherapy, which has been proven effective in the treatment of microsatellite-unstable colorectal cancers [9].
Universal tumor screening of all colorectal cancers is recommended by many professional societies [10,11,12]. In addition to the presence of microsatellite instability, defective mismatch repair colorectal cancers typically have absent expression of one or more of the mismatch repair proteins, which corresponds to the dysfunctional gene. It is well known that the mismatch repair proteins dimerize to form heterodimer complexes. MSH2 and MSH6 proteins physiologically form a heterodimer MutSα, which recognizes mismatched DNA bases, initiating mismatch repair. If MSH2 is mutated, both MSH2 and MSH6 proteins should be absent on immunohistochemistry since MSH6 is not known to form a heterodimer complex with any other protein. However, MSH2 can form a heterodimer pairing with MSH3 when MSH6 is mutated, resulting in isolated absence of MSH6. MLH1 and PMS2 act in similar manner. Hence, screening algorithms for Lynch syndrome often utilize immunohistochemical staining to assess for the presence of the mismatch repair proteins, with absent expression patterns revealing defective mismatch repair and guiding germline testing [13].
Immunohistochemical analysis for the four mismatch repair proteins has been shown to be cost-effective [2, 13]. However, in an effort to further reduce cost, some centers screen with an abbreviated “two-stain” method consisting of only the MSH6 and PMS2 proteins [14,15,16]. The underlying logic of the “two-stain” method is that, regardless of whether there is a mutation in the gene encoding for them or their heterodimer partners (MLH1 and MSH2), MSH6 and PMS2 should be lost. If MSH6 is absent on initial screening, reflex staining of MSH2 would be performed to clarify if both are absent (suggesting an underlying MSH2 gene mutation) or if only MSH6 is absent (suggesting an underlying MSH6 gene mutation). Likewise, if PMS2 is absent on initial screening, reflex staining of MLH1 would be performed.
Current literature suggests that there are three possible MSH2 and MSH6 protein expression patterns [17]: Intact expression of both proteins (implying wild-type MSH2 and MSH6 genes), isolated absence of MSH6 (implying germline or somatic MSH6 mutation[s]), or absence of both MSH2 and MSH6 (implying germline or somatic MSH2 mutation[s], or rarely MSH6 mutation). Any deviation from these three expression patterns can result in failed Lynch syndrome detection when using the two-stain method. An immunohistochemical expression pattern revealing absence of MSH2 with intact MSH6 is not expected, and to our knowledge, no published studies have addressed Lynch syndrome cases in which MSH6 staining could be interpreted as present despite a germline MSH2 mutation.
However, a fourth expression pattern was brought to our attention after the two-stain method failed to identify a patient with a germline MSH2 mutation at our institution. Despite the presence of MSH6 and PMS2 proteins on immunohistochemistry, given high clinical suspicion for Lynch syndrome, this patient received germline mismatch repair gene testing and was found to have a germline MSH2 mutation. After the germline mutation was identified, the tumor was retrospectively stained for MSH2 and it was unequivocally absent. Therefore, we hypothesized that the two-stain method could miss detecting patients with defective mismatch repair and MSH2-related Lynch syndrome since intact MSH6 expression would not trigger reflex MSH2 staining.
Materials and methods
Patients
A total of 3346 patients with newly diagnosed colorectal cancer enrolled from 51 participating Ohio hospitals into the Ohio Colorectal Cancer Prevention Initiative (ClinicalTrials.gov identifier: NCT01850654) between 01 January 2013 and 31 December 2016. Enrollment criteria for the Ohio Colorectal Cancer Prevention Initiative study has been previously described [18]. Institutional Review Board approval was obtained by the individual hospitals, Community Oncology Research Programs, or by ceding review to the Ohio State University Institutional Review Board. Written informed consent was obtained. For this analysis, we retrospectively reviewed all colorectal cancer cases received by June 2016 with available immunohistochemical slides (n = 1,730) for the staining intensity and proportion of stained cells of the MSH6 protein, and MSH2 was also reviewed when available.
Samples
Blood and a paraffin-embedded tumor block or unstained slides were submitted for each patient. Pathologists confirmed tumor histology and marked areas with at least 30% tumor and normal adjacent tissue. Blood and tissue (tumor and normal) underwent DNA extraction using standard methods [19].
Ohio colorectal cancer prevention initiative tumor screening strategy
Testing procedures for the Ohio Colorectal Cancer Prevention Initiative study have been previously described [18]. Briefly, all tumors were screened for defective mismatch repair by immunohistochemical analysis and/or microsatellite instability testing. Immunohistochemistry of the mismatch repair proteins was performed using the two-stain method as previously described; staining for all four mismatch repair proteins was done if microsatellite instability could not be performed due to insufficient tumor or if testing was already completed as part of the patient’s routine clinical care. Proteins with <5% staining were considered absent. Microsatellite instability testing was completed using tumor and normal DNA to detect a size change in microsatellites using the Promega MSI Analysis System (Version 1.2). This included fluorescently labeled primers for co-amplification of five repeat markers (BAT-25, BAT-26, NR-21, NR-24, MONO-27) [20, 21].
Additional immunohistochemical analysis for this study
Results of immunohistochemical expression were further assessed by determining the proportion and intensity of staining in tumor cells relative to internal tissue controls, which included stromal cells and lymphocytes [15]. Internal controls were used to verify appropriate staining and to serve as the basis for grading staining intensity of neoplastic cells; cases with control failure were discarded if repeat staining was not successful. Immunohistochemical analysis was performed by manual counting of the tumor nuclei in one whole section for each case. Staining intensity was graded as follows: (3) equal to or greater than control, (2) slightly less than control but with definitive nuclear staining, (1) less than control with ambiguous nuclear staining, or (0) no visible nuclear staining. Antibodies included MSH-2 Clone, Calbiochem FE11 (Mouse: NA27) and MSH-6 Clone, Epitomics EP49 (Rabbit: AC-0047).
Genetic testing
With regard to this analysis, patients with absence of MSH2 protein underwent germline genetic testing using next-generation sequencing of MSH2, MSH6 and EPCAM as part of a large multi-gene panel at the University of Washington. Genomic regions were captured using biotinylated RNA oligonucleotides (SureSelect) and sequenced on an Illumina HiSeq2000 instrument [22]. Large rearrangements were detected [23]. Patients without germline mutations underwent additional genetic testing of MSH2, MSH6 and EPCAM as part of a large multi-gene panel in tumor DNA at the University of Washington using methods previously described to assess for double somatic mismatch repair mutations [24].
Results
Of the 1730 cases reviewed, all had MSH6 immunohistochemistry completed; 1704 had confirmed intact MSH6 expression and 26 had absent MSH6 expression (1.5%, 26/1730). Of the cases reviewed, 495 selected cases had MSH2 immunohistochemistry completed as per the Ohio Colorectal Cancer Prevention Initiative protocol (reflex when MSH6 was absent, insufficient tumor for microsatellite instability, and/or all four proteins were stained as part of the patient’s clinical care). Of the 1730 cases, 462 had confirmed intact MSH2 expression, 1235 had implied intact expression of MSH2 as per the two-stain method, and 33 had absent MSH2 expression (1.9%, 33/1730).
Of the 33 cases with absence of MSH2, fourteen had no MSH6 expression as expected but eight (0.5%, 8/1730) had ambiguous (grade 1) MSH6 expression and eleven (0.6%, 11/1730) had convincing (grade 2–3) MSH6 expression. Germline next-generation sequencing identified a pathogenic or likely pathogenic germline MSH2 mutation in 11/14 cases with absence of both stains, 7/8 cases with ambiguous MSH6 expression, and 9/11 cases with convincing MSH6 expression. Of the remaining six cases without a germline mutation, five had confirmed double somatic MSH2 or MSH6 mutations, and one remains unexplained and could be due to either a germline or somatic MSH2 or MSH6 mutation that could not be detected with the current technology. The cases were divided into three groups based on MSH6 staining (Group 1, convincing MSH6 expression, Group 2 equivocal, and Group 3 none), (see Table 1). None of these cases showed diffuse MSH6 staining.
Group 1 (convincing MSH6 expression)
Eleven cases (0.6%) had absent MSH2 expression and convincing (grade 2 or 3) MSH6 expression in at least 5% of cells (Figs. 1, 2 and 3). Nine of the group 1 cases had a germline MSH2 mutation (4 had large deletions, 5 had splice site mutations). Of those with Lynch syndrome, two had grade 3 intensity with 18–44% of cells expressing MSH6 and seven had grade 2 intensity with 14–38% of cells expressing MSH6. One case had double somatic MSH2 mutations with grade 2 intensity and 30% staining. One case had double somatic MSH2 mutations with grade 3 intensity and 29% staining.
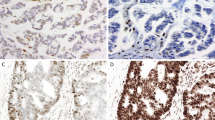
Convincing MSH6 staining (Grade 3 intensity) in the absence of MSH2 staining. The tumor cells demonstrate MSH6 nuclear staining as strong as the internal control cells (lymphocytes and stromal cells), (b) ×200 and (d) ×400. Notice MSH2 staining is absent/lost in the tumor cells, (a) ×200 and (c) ×400
Two examples of weak MSH6 staining (Grade 2 intensity) in the absence of MSH2 staining. The tumor cells demonstrate patchy MSH6 nuclear staining somewhat weaker than the internal control cells (lymphocytes and stromal cells), (b) ×200 and (d) ×400. Notice MSH2 staining is absent/lost in the tumor cells, while a few tumor infiltrating lymphocytes are present, (a) ×200 and (c) ×400
Same as for Fig. 2
Group 2 (ambiguous MSH6 expression)
Eight cases (0.5%) had absent MSH2 expression and ambiguous (grade 1) MSH6 expression in at least 5% of cells (Figs. 4 and 5) . Seven of the group 2 cases had a germline MSH2 mutation (2 had frameshift mutations, 3 had missense mutations, 1 had an in-frame deletion, 1 had a large deletion). Of those with Lynch syndrome, 6–19% of cells expressed MSH6. One case had double somatic MSH2 mutations and 6% of cells expressed MSH6.
Two examples of ambiguous MSH6 staining (Grade 1 intensity) in the absence of MSH2 staining. The tumor cells demonstrate faint patchy MSH6 nuclear staining significantly weaker than the internal control cells (lymphocytes and stromal cells), (b) 200× and (d) 400×. Notice MSH2 staining is absent/lost in the tumor cells, while a few tumor infiltrating lymphocytes are present, (a) 200× and (c) 400×
Same as for Fig. 4
Group 3 (no MSH6 expression)
Fourteen cases (0.8%) had absence of both MSH2 and MSH6 expressionas as expected. Eleven of the group 3 cases had a germline MSH2 mutation (7 had large deletions, 2 had splice site mutations, 2 had frameshift mutations), two cases had double somatic MSH2 mutations and one case remains unexplained (MSH2 germline negative, one somatic MSH2 mutation).
Ten of the patients with Lynch syndrome shared two common MSH2 mutations; five had the exon 1–6 deletion (3 in group 1, 2 in group 3) and five had the c.942 + 3A > T mutation (4 in group 1 and 1 in group 3). Clinical and molecular data for the cases in groups 1–3 are summarized in Table 1.
Discussion
Retrospective review of 1,730 colorectal cancer cases undergoing universal tumor screening for Lynch syndrome using a modified two-stain method revealed that 19 cases (16 of which had Lynch syndrome) with absent MSH2 had some degree of retained MSH6 expression (grade 1–3, ≥5% cells staining) and could have been missed if the two-stain method was the only screening test utilized.
The interpretation of immunohistochemical stains should always be done in the presence of appropriate positive controls. Stains are generally considered present when the intensity is at least equivalent to the positive control. At times, it can be challenging to determine comparative staining intensities between the tumor and control. In this study, we considered that 3+ staining intensity would be interpreted by all pathologists as present, while 2+ could be considered present and 1+ would likely be interpreted as lost by most. Therefore, three cases would have likely been missed (MSH6 3+ present), while eight cases may have been missed (MSH6 2+ present), for a total of eleven cases not reflexed to MSH2, and thus missed. Even though the eight cases with MSH6 1+ present would likely have been called absent, it is important to confirm that these cases should be considered absent.
Patchy mismatch repair protein staining is not uncommon, although most microsatellite stable cases show diffuse staining for all four proteins. At times, patchy staining can be attributed to technical issues, particularly when the control staining shows a similar pattern. All of our study cases showed appropriate staining in internal control tissue throughout the slides, and the mismatch repair staining was interpreted as compared to control staining in adjacent stroma and benign crypts. It is important to note that none of our study cases showed diffuse MSH6 staining, but rather, ranged from 6 to 44%. Unfortunately, there is no literature that supports the use of a higher cutoff for MSH6 than that chosen for the other mismatch repair proteins, and would lead to significantly overcalling microsatellite instability. Additional studies with all four MMR stains upfront may be helpful to determine if a higher cutoff is indicated. Regardless of the percent, weaker staining than control should be considered for additional testing. Unusual MSH6 expression patterns have been previously documented, such as under expression of MSH6 in post-chemoradiation specimens [25,26,27], heterogeneous MSH6 staining in MLH1/PMS2-deficient tumors [28, 29], and greater conservation of MSH6 in in situ lesions compared to invasive carcinomas [30, 31]. However, our study demonstrated unexpected presence of MSH6 rather than absence.
The cause for the observed aberrant MSH2 absent, MSH6 patchy expression pattern is unclear. It is notable that six of the Lynch syndrome patients with some retained MSH6 expression shared two MSH2 mutations; two had the exon 1–6 deletion (both had grade 2 staining) and four had the c.942 + 3A > T mutation (one had grade 3 staining, three had grade 2 staining). These mutations are two of the most common MSH2 mutations found in individuals with Lynch syndrome [32, 33], so it is unclear if this finding is a coincidence or potentially related to the aberrant expression observed. It is known that some individuals with MSH2 deletion of exons 1–6 can have false-negative stability of BAT26 since that microsatellite is included in the deletion region of MSH2 [34]; however, this does not provide an explanation for the unusual immunohistochemical pattern we observed. It would be interesting to study the nature of aberrant splicing of the c.942 + 3A > T mutation to determine if it facilitates MSH6 presence with absence of MSH2.
A limitation of this study is that MSH2 immunohistochemistry was not done on all 1,730 patients. Therefore, this could be an underestimate of the number of defective mismatch repair-MSH2 cases missed by the two-stain method. However, we do not believe there would be many (if any) additional cases with MSH2 loss among those who only had the two-stain method because all such cases also had microsatellite instability analysis, and would have been flagged had there been discordant screening results (microsatellite instability and intact MSH6/PMS2). While this study focused on potentially missing MSH2-related Lynch syndrome with patchy MSH6 by the two-stained method, it will be of future interest to perform an analogous study with MLH1-related Lynch syndrome with intact/patchy PMS2.
We have shown that patchy MSH6 expression can be retained in colorectal cancer cases with absence of MSH2, and that the two-stain approach can miss a subset of MSH2-related Lynch syndrome. It is critical to identify these cases by universal tumor screening, since the majority were found to have Lynch syndrome due to germline mutations in the MSH2 gene. While the methodology of our study does not allow for direct epidemiological extrapolation of our findings to a general population, the identification of several cases with MSH2 mutations with ≥5% MSH6 nuclear expression suggests that retained MSH6 expression in MSH2 germline or double somatic mutations is not uncommon. Based on this work, we advocate for the use of all four MMR stains in universal tumor screening for Lynch syndrome. However, if the two-stain method is used, we recommend reflex MSH2 staining for all cases with focal/patchy or weak MSH6 staining, in addition to cases with absent MSH6 staining.
References
Ionov Y, Peinado MA, Malkhosyan S, Shibata D, Perucho M. Ubiquitous somatic mutations in simple repeated sequences reveal a new mechanism for colonic carcinogenesis. Nature. 1993;363:558–61.
Hampel H, Frankel WL, Martin E, et al. Feasibility of screening for Lynch syndrome among patients with colorectal cancer. J Clin Oncol. 2008;26:5783–8.
Hampel H, Frankel WL, Martin E, et al. Screening for the Lynch syndrome (hereditary nonpolyposis colorectal cancer). N Engl J Med. 2005;352:1851–60.
Lynch H, Lanspa S, Boman B, et al. Hereditary nonpolyposis colorectal cancer–Lynch syndromes I and II. Gastroenterol Clin North Am. 1988;17:679–712.
Mensenkamp AR,Vogelaar IP,van Zelst-Stams WA, et al. Somatic mutations in MLH1 and MSH2 are a frequent cause of mismatch-repair deficiency in Lynch syndrome-like tumors. Gastroenterology. 2014;146:643–6.
Geurts-Giele WR, Leenen CH, Dubbink HJ, et al. Somatic aberrations of mismatch repair genes as a cause of microsatellite-unstable cancers. J Pathol. 2014;234:548–59.
Haraldsdottir S,Hampel H,Tomsic J, et al. Colon and endometrial cancers with mismatch repair deficiency can arise from somatic, rather than germline, mutations. Gastroenterology. 2014;147:1308–16.
Sourrouille I, Coulet F, Lefevre JH, et al. Somatic mosaicism and double somatic hits can lead to MSI colorectal tumors. Fam Cancer. 2013;12:27–33.
Le DT, Uram JN, Wang H, et al. PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med. 2015;372:2509–20.
Palomaki GE, McClain MR, Melillo S, Hampel HL, Thibodeau SN. EGAPP supplementary evidence review: DNA testing strategies aimed at reducing morbidity and mortality from Lynch syndrome. Genet Med. 2009;11:42–65.
Vasen HF, Blanco I, Aktan-Collan K, et al. Revised guidelines for the clinical management of Lynch syndrome (HNPCC): recommendations by a group of European experts. Gut. 2013;62:812–23.
Giardiello FM, Allen JI, Axilbund JE, et al. Guidelines on genetic evaluation and management of Lynch syndrome: a consensus statement by the US Multi-society Task Force on colorectal cancer. Am J Gastroenterol. 2014;109:1159–79.
Muller W, Burgart LJ, Krause-Paulus R, et al. The reliability of immunohistochemistry as a prescreening method for the diagnosis of hereditary nonpolyposis colorectal cancer (HNPCC)--results of an international collaborative study. Fam Cancer. 2001;1:87–92.
Hall G, Clarkson A, Shi A, et al. Immunohistochemistry for PMS2 and MSH6 alone can replace a four antibody panel for mismatch repair deficiency screening in colorectal adenocarcinoma. Pathology. 2010;42:409–13.
O’Regan T, Chau K, Tatton M, et al. Immunochemistry screening for Lynch syndrome in colorectal adenocarcinoma using an initial two-antibody panel can replace a four antibody panel. N Z Med J. 2013;126:70–7.
Mojtahed A, Schrijver I, Ford JM, Longacre TA, Pai RK. A two-antibody mismatch repair protein immunohistochemistry screening approach for colorectal carcinomas, skin sebaceous tumors, and gynecologic tract carcinomas. Mod Pathol. 2011;24:1004–14.
Kantelinen J, Kansikas M, Candelin S, et al. Mismatch repair analysis of inherited MSH2 and/or MSH6 variation pairs found in cancer patients. Hum Mutat. 2012;33:1294–301.
Pearlman R, Frankel WL, Swanson B, et al. Prevalence and spectrum of germline cancer susceptibility gene mutations among patients with early-onset colorectal cancer. JAMA Oncol. 2017;3:464–71.
Buckingham L, Flaws ML. Molecular diagnostics: fundamentals, methods and clinical applications. F.A. Davis Company; 2007.
Bacher JW, Flanagan LA, Smalley RL, et al. Development of a fluorescent multiplex assay for detection of MSI-high tumors. Dis Markers. 2004;20:237–50.
Bacher JW, Sievers CK, Albrecht DM, et al. Improved detection of microsatellite instability in early colorectal lesions. PLoS ONE. 2015;10:e0132727.
Pritchard CC, Smith C, Salipante SJ, et al. ColoSeq provides comprehensive lynch and polyposis syndrome mutational analysis using massively parallel sequencing. J Mol Diagn. 2012;14:357–66.
Nord AS, Lee M, King MC, Walsh T. Accurate and exact CNV identification from targeted high-throughput sequence data. BMC Genom. 2011;12:184.
Haraldsdottir S, Hampel H, Tomsic J, et al. Colon and endometrial cancers with mismatch repair deficiency can arise from somatic, rather than germline, mutations. Gastroenterology. 2014;147:1308–16.
Vilkin A, Halpern M, Morgenstern S, et al. How reliable is immunohistochemical staining for DNA mismatch repair proteins performed after neoadjuvant chemoradiation? Hum Pathol. 2014;45:2029–36.
Bao F, Panarelli NC, Rennert H, Sherr DL, Yantiss RK. Neoadjuvant therapy induces loss of MSH6 expression in colorectal carcinoma. Am J Surg Pathol. 2010;34:1798–804.
Radu OM, Nikiforova MN, Farkas LM, Krasinskas AM. Challenging cases encountered in colorectal cancer screening for Lynch syndrome reveal novel findings: nucleolar MSH6 staining and impact of prior chemoradiation therapy. Hum Pathol. 2011;42:1247–58.
Graham RP, Kerr SE, Butz ML, et al. Heterogenous MSH6 loss is a result of microsatellite instability within MSH6 and occurs in sporadic and hereditary colorectal and endometrial carcinomas. Am J Surg Pathol. 2015;39:1370–6.
Shia J, Zhang L, Shike M, et al. Secondary mutation in a coding mononucleotide tract in MSH6 causes loss of immunoexpression of MSH6 in colorectal carcinomas with MLH1/PMS2 deficiency. Mod Pathol. 2013;26:131–8.
Walsh MD, Buchanan DD, Pearson SA, et al. Immunohistochemical testing of conventional adenomas for loss of expression of mismatch repair proteins in Lynch syndrome mutation carriers: a case series from the Australasian site of the colon cancer family registry. Mod Pathol. 2012;25:722–30.
Yurgelun MB, Goel A, Hornick JL, et al. Microsatellite instability and DNA mismatch repair protein deficiency in Lynch syndrome colorectal polyps. Cancer Prev Res). 2012;5:574–82.
van der Klift H, Wijnen J, Wagner A, et al. Molecular characterization of the spectrum of genomic deletions in the mismatch repair genes MSH2, MLH1, MSH6, and PMS2 responsible for hereditary nonpolyposis colorectal cancer (HNPCC). Genes Chromosomes Cancer. 2005;44:123–38.
Desai DC, Lockman JC, Chadwick RB, et al. Recurrent germline mutation in MSH2 arises frequently de novo. J Med Genet. 2000;37:646–52.
Pastrello C, Baglioni S, Tibiletti MG, et al. Stability of BAT26 in tumours of hereditary nonpolyposis colorectal cancer patients with MSH2 intragenic deletion. Eur J Hum Genet. 2006;14:63–8.
Acknowledgements
The Ohio Colorectal Cancer Prevention Initiative is supported by a grant from Pelotonia, an annual cycling event in Columbus, Ohio, that supports cancer research at The Ohio State University Comprehensive Cancer Center–James Cancer Hospital and Solove Research Institute. This study was also supported in part by the National Cancer Institute (grant P30CA16058). We thank Shawn Scully for his help with the figures.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
HH is on the scientific advisory board for Invitae Genetics and Genome Medical, has stocks in Genome Medical, has received an honorarium from the Beijing Genomics Institute and has received research study support (donated genetic testing) from Myriad Genetic Laboratories, Inc. The remaining authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Pearlman, R., Markow, M., Knight, D. et al. Two-stain immunohistochemical screening for Lynch syndrome in colorectal cancer may fail to detect mismatch repair deficiency. Mod Pathol 31, 1891–1900 (2018). https://doi.org/10.1038/s41379-018-0058-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-018-0058-y
This article is cited by
-
A 39 kb structural variant causing Lynch Syndrome detected by optical genome mapping and nanopore sequencing
European Journal of Human Genetics (2023)
-
Validity of a two-antibody testing algorithm for mismatch repair deficiency testing in cancer; a systematic literature review and meta-analysis
Modern Pathology (2022)
-
MSI-Testung
Der Pathologe (2021)
-
MSI testing
Der Pathologe (2021)
-
A practical guide to biomarkers for the evaluation of colorectal cancer
Modern Pathology (2019)